Abstract
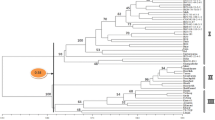
Velvetbean (Mucuna sp.) is a self-pollinated crop classified within the Leguminosae. Using AFLP markers, gene diversity and phenetic relationships were estimated in a collection of 40 velvetbean accessions from cultivated species and different eco-geographic regions. Eleven selective primer combinations generated a total of 508 amplification products. The average number of scorable fragments was 23 per primer combination. A total of 251 polymorphic markers was detected. The polymorphisms obtained ranged from 36% to 61% with an average of 49%. The final phenetic trees were constructed using Nei and Li’s coefficient of similarity with UPGMA. Other clustering algorithms were examined and all had high co-phenetic correlations, indicating the goodness of fit for the resulting phylogenetic trees. The phenetic tree as well as principal component analysis (PCA) separated the 40 velvetbean accessions into two main clusters. Bootstrap and Jackknife analyses were completed and their values indicated strong to moderate support for the two main clusters. This grouping confirmed the existing phenological difference with regard to maturity. The high values of the similarity coefficients observed (0.87 to 0.97) imply that the accessions used in this study are similar. The level of genetic variability detected within the velvetbean accessions with AFLP analysis suggests that it is a reliable, efficient, and effective marker technology for determining genetic relationships in velvetbean.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 19 June 2000 / Accepted: 1 March 2001
Rights and permissions
About this article
Cite this article
Capo-chichi, L., Weaver, D. & Morton, C. AFLP assessment of genetic variability among velvetbean (Mucuna sp.) accessions. Theor Appl Genet 103, 1180–1188 (2001). https://doi.org/10.1007/s001220100722
Issue Date:
DOI: https://doi.org/10.1007/s001220100722