Abstract
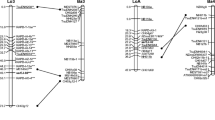
A map with 246 markers (11 isozymes and 235 RFLPs) was constructed using an interspecific F2 population between almond (cv Texas) and peach (cv Earlygold). RFLPs were obtained using 213 probes from the genomic and cDNA libraries of different species (almond, peach, P. ferganensis, cherry, plum and apple), including 16 almond probes which correspond to known genes. All markers were distributed in eight linkage groups, the same as the basic chromosome number of the genus, covering a total distance of 491 cM. The average map density was 2.0 cM/marker and only four gaps of 10 cM or more were found; the two largest gaps were 12cM each. This map was compared with one constructed previously with an intraspecific almond population sharing 67 anchor loci. Locus order was nearly identical and distances were not significantly different. A large proportion of the mapped loci (46%) had skewed segregations; in approximately half of them, the distortion was due to an excess of heterozygotes. One of the distorted regions could be associated with the position of the self-incompatibility gene of almond.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 6 November 1997 / Accepted: 26 May 1998
Rights and permissions
About this article
Cite this article
Joobeur, T., Viruel, M., de Vicente, M. et al. Construction of a saturated linkage map for Prunus using an almond×peach F2 progeny. Theor Appl Genet 97, 1034–1041 (1998). https://doi.org/10.1007/s001220050988
Issue Date:
DOI: https://doi.org/10.1007/s001220050988