Abstract
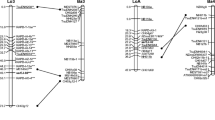
A genetic linkage map of Brassica juncea was constructed based on restriction fragment length polymorphism (RFLP) detected by anonymous cDNA markers from B. napus, using a segregating F1-derived doubled haploid (DH) progeny from a cross between a canola-quality mustard line (J90-4317) and a high-oil-content mustard line (J90-2733). The RFLP probes consisted of 229 cDNA probes from B. napus and a B. napus tandem repeat sequence, RDA2. The map consisted of 343 marker loci arranged in 18 major linkage groups plus five small segments with two to five marker loci, covering a total map distance of 2073 cM. Twenty-four percent of the markers were dominant in nature. Sixty-two percent of the marker loci were duplicated, and the majority were involved in inter-linkage group duplications, illustrating that complex duplications and subsequent rearrangements occurred after allopolyploidy. Deviation from the Mendelian segregation ratio for a DH population was observed for 27% of the markers. Two-thirds of these markers with a skewed segregation were clustered in 6 linkage groups and two unassigned segments. The overall average marker interval of the B. juncea map reported here was 6.6 cM, which would provide a marker density satisfactory for efficient use of the map in breeding applications, such as tagging of important agronomic traits and marker-assisted selection.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 14 May 1996 / Accepted: 11 October 1996
Rights and permissions
About this article
Cite this article
Cheung, W., Friesen, L., Rakow, G. et al. A RFLP-based linkage map of mustard [Brassica juncea (L.) Czern. and Coss.]. Theor Appl Genet 94, 841–851 (1997). https://doi.org/10.1007/s001220050485
Issue Date:
DOI: https://doi.org/10.1007/s001220050485