Abstract
Key message
A single dominant gene found in tetraploid and hexaploid wheat controls broad-spectrum race-nonspecific resistance to the foliar disease tan spot caused by Pyrenophora tritici-repentis.
Abstract
Tan spot is an important foliar disease of durum and common wheat caused by the necrotrophic fungal pathogen Pyrenophora tritici-repentis. Genetic studies in common wheat have shown that pathogen-produced necrotrophic effectors interact with host genes in an inverse gene-for-gene manner to cause disease, but quantitative trait loci (QTLs) with broad race-nonspecific resistance also exist. Less work has been done to understand the genetics of tan spot interactions in durum wheat. Here, we evaluated a set of Langdon durum—wild emmer (Triticum turgidum ssp. dicoccoides) disomic chromosome substitution lines for reaction to four P. tritici-repentis isolates representing races 1, 2, 3, and 5 to identify wild emmer chromosomes potentially containing tan spot resistance genes. Chromosome 3B from the wild emmer accession IsraelA rendered the tan spot-susceptible durum cultivar Langdon resistant to all four fungal isolates. Genetic analysis indicated that a single dominant gene, designated Tsr7, governed resistance. Detailed mapping experiments showed that the Tsr7 locus is likely the same as the race-nonspecific QTL previously identified in the hexaploid wheat cultivars BR34 and Penawawa. Four user-friendly SNP-based semi-thermal asymmetric reverse PCR (STARP) markers cosegregated with Tsr7 and should be useful for marker-assisted selection of resistance. In addition to 3B, other wild emmer chromosomes contributed moderate levels of tan spot resistance, and, as has been shown previously for tetraploid wheat, the Tsn1-Ptr ToxA interaction was not associated with susceptibility. This is the first report of a major dominant gene governing resistance to tan spot in tetraploid wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tan spot, also known as yellow leaf spot, is a foliar disease of common wheat (Triticum aestivum L., 2n = 6x = 42, AABBDD genomes), durum wheat (T. turgidum ssp. durum (Desf.) Husnot., 2n = 4x = 28, AABB genomes) and their relatives. Tan spot is caused by the necrotrophic pathogen Pyrenophora tritici-repentis (Died.) Drechs. and occurs in most wheat-growing regions of the world. The practice of no-till farming adopted in the 1970s to reduce soil erosion led to an increased incidence of tan spot because P. tritici-repentis overwinters in stubble residue. The primary symptoms of tan spot include large necrotic lesions surrounded by chlorotic borders and sometimes extensive chlorosis extending between necrotic lesions. Large areas of necrosis and/or chlorosis coalesce in highly susceptible genotypes leaving large areas of dead leaf tissue that result in a reduced capacity to perform photosynthetic functions, which ultimately leads to reductions in yield.
P. tritici-repentis is a member of the Pleosporales order of fungi and is known to produce necrotrophic effectors (NEs) (formerly referred to as host-selective toxins). At least three NEs are produced by P. tritici-repentis, including Ptr ToxA, Ptr ToxB, and Ptr ToxC (see Singh et al. 2010; Faris et al. 2013 for reviews). P. tritici-repentis race 1 produces Ptr ToxA and Ptr ToxC, race 2 produces Ptr ToxA, race 3 produces Ptr ToxC, race 4 is considered avirulent on wheat, and race 5 produces Ptr ToxB. Race 6 produces Ptr ToxB and Ptr ToxC, race 7 produces Ptr ToxA and Ptr ToxB, and race 8 produces all three NEs. Recognition of each of these NEs by single corresponding sensitivity genes in the host, namely Tsn1, Tsc2, and Tsc1, respectively, leads to a compatible interaction defined by the development of tan spot. Elimination or disruption of the dominant host allele negates recognition and leads to an incompatible (resistance) interaction. This system is referred to as inverse gene-for-gene where the pathogen essentially hijacks the plant’s own defense mechanism to cause disease (Faris et al. 2010).
In addition to the inverse gene-for-gene interactions, numerous tan spot resistance QTLs have been identified (see Faris et al. 2013 for review). Most notably, a resistance QTL on the long arm of chromosome 3B was identified in the Brazilian spring wheat cultivar BR34 (Faris and Friesen 2005), and more recently in the US spring wheat cultivar Penawawa (Kariyawasam et al. 2016). In both cases, the 3B QTL conferred good levels of resistance to all known races of P. tritici-repentis.
Compared to common wheat, the genetics of tan spot resistance in durum wheat has been less studied. The Tsn1-Ptr ToxA, Tsc2-Ptr ToxB, and Tsc1-Ptr ToxC interactions have all been shown to play important roles in the development of tan spot in common wheat (Faris et al. 2013; Kariyawasam et al. 2016; Liu et al. 2017). Virdi et al. (2016) recently showed that the Tsc2-Ptr ToxB interaction can also play a significant role in tan spot development in durum wheat. On the contrary, there is accumulating evidence indicating that a compatible Tsn1-Ptr ToxA interaction plays little to no role in tan spot development in durum wheat (Chu et al. 2010; Virdi et al. 2016). The effects of a compatible Tsc1-Ptr ToxC interaction in durum wheat have not been investigated.
A few studies on the genetics of tan spot resistance in durum wheat have revealed relevant loci other than the three NE sensitivity genes. For example, Singh et al. (2008a, b) used a tetraploid biparental population to show that two linked recessive genes on chromosome arm 3BL conferred resistance to different isolates of the fungus. Also, Chu et al. (2010) conducted a QTL study and reported regions associated with tan spot resistance on chromosomes 3A, 3B, 5A, and 7B. In addition to the Tsc2 locus, Virdi et al. (2016) reported resistance QTLs on chromosomes 4B and 6B using a biparental durum population. However, beyond these studies, little work has been done to dissect the genetics of tan spot resistance in durum wheat.
In polyploid wheat, disomic chromosome substitution lines consisting of a pair of chromosomes from a particular ‘donor’ accession substituted for a pair of homologous chromosomes in the background of a ‘recipient’ accession, are powerful tools for the genetic dissection of complex traits and identification of chromosomes harboring useful and/or novel genes. Dr. Leonard Joppa (USDA-ARS, Fargo, ND, retired) developed a set of disomic chromosome substitution lines in durum wheat using the wild emmer wheat [T. turgidum ssp. dicoccoides (Körn. Ex Asch. & Graebner) Aarons (2n = 4x = 28, AABB genomes)] accession IsraelA (syn. FA-15-3) as donor and North Dakota durum wheat variety Langdon (LDN) as the recipient (Cantrell and Joppa 1991). These substitution lines have been widely distributed and used for the investigation of various agronomic traits (Joppa and Cantrell 1990; Cantrell and Joppa 1991; Joppa et al. 1991; Joppa 1993; Stack et al. 2002; Singh et al. 2007).
Joppa (1993) crossed each of the chromosome substitution lines with LDN and developed segregating recombinant inbred chromosome line (RICL) populations. These populations serve as unique and useful tools because they segregate for individual recombinant chromosomes in a homozygous background allowing genetic analysis, mapping, and dissection of traits in isolation. The RICL populations have been used to evaluate traits such as grain protein content (Joppa et al. 1997), yield (Gonzalez-Hernadez et al. 2004), rachis brittleness (Nalam et al. 2006), segregation distortion (Kumar et al. 2007a, b), Fusarium head blight (FHB) resistance/susceptibility (Otto et al. 2002; Garvin et al. 2009), Septoria nodorum blotch and tan spot resistances (Faris et al. 2000; Gonzalez-Hernandez et al. 2009; Faris and Friesen 2009), and spike morphology (Faris et al. 2014b).
Here, we evaluated the LDN-IsraelA chromosome substitution lines for reaction to tan spot caused by P. tritici-repentis isolates representing races 1-3 and 5. Wild emmer chromosome 3B was found to harbor dominant race-nonspecific resistance and was therefore the subject of molecular and comparative mapping to determine whether the 3B resistance locus was the same as loci previously reported in hexaploid wheat. User-friendly PCR-based SNP markers developed for the locus should be useful for introgression of the gene into adapted durum backgrounds.
Materials and methods
Plant materials
The LDN-T. turgidum ssp. dicoccoides chromosome substitution lines were evaluated for reaction to tan spot using isolates Pti2 (race 1), 86-124 (race 2), 331-9 (race 3) and DW5 (race 5). Race 4 is considered an avirulent race and was not studied. All these races have been identified in North America (Friesen et al. 2005). For brevity, the chromosome substitution lines derived from T. turgidum ssp. dicoccoides accession IsraelA will be referred to as IsA-X, where ‘X’ refers to the T. turgidum ssp. dicoccoides chromosome being substituted for the homologous LDN chromosome. The set consists of substitution lines representing all tetraploid wheat chromosomes except for the line IsA-2B, which was not available because the IsraelA chromosome 2B poorly compensates for LDN 2B and plants are weak and mostly sterile (J.D. Faris and S.S. Xu, unpublished). In addition to LDN, IsraelA, and the substitution lines, the tan spot differential lines Salamouni, Glenlea, 6B365, and 6B662 (Lamari et al. 1995) were included in all inoculations.
The LDN × IsA-3B RICL population (hereafter referred to as the LD3B population) consisting of 91 homozygous recombinant lines was evaluated for reaction to the same isolates used to test the chromosome substitution lines. Also, a new cross between LDN and IsA-3B was made, and seven F1 plants from this cross were evaluated for reaction to tan spot using the race 2 isolate 86-124.
A panel of 21 wheat lines was used to evaluate STARP markers developed for SNP markers associated with the QTL on 3B (see below). The panel consisted of the tetraploid and hexaploid lines listed in Table 1.
Ptr ToxA infiltrations, fungal inoculations and statistical analysis
LDN, IsraelA, and the chromosome substitution lines were infiltrated with the necrotrophic effector Ptr ToxA as described in Liu et al. (2017). Reactions were scored 5 days after infiltration as either sensitive (presence of necrosis) or insensitive (no necrosis).
Plants were grown in super-cell containers (Stuewe & Sons Inc., Corvallis, OR) consisting of racks of 98 containers as described in Liu et al. (2017). The tan spot-susceptible North Dakota hard red winter wheat cultivar ‘Jerry’ was planted in the outside borders of each rack to reduce any edge effect. Inoculum preparation, plant inoculations, and disease evaluations were performed as described in Liu et al. (2017). The chromosome substitution sets, and later the LD3B population were grown in completely randomized designs with three seeds per container serving as an experimental unit, and two containers for each line per experiment. Each experiment was conducted at least three times; thus, the reaction of each line to each isolate consisted of 18 plants representing six replications. Disease reactions were scored seven days after inoculation using the 1–5 scale described by Lamari and Bernier (1989) where 1 is highly resistant and 5 is highly susceptible.
Bartlett’s Chi-squared test for homogeneity of error variances was conducted using PROC GLM in SAS program version 9.3 (SAS institute 2011) to determine whether inoculation experiments for each isolate were homogeneous among the substitution lines. Error variances were homogeneous among all experiments for each isolate (data not shown), and therefore the mean values were calculated and used for further analysis. Fisher’s Least Significant Difference (LSD) test was used to determine significant differences among means of the chromosome substitution lines at the 0.05 level of probability.
Molecular mapping
We constructed a genetic linkage map of chromosome 3B in the LD3B population using simple sequence repeat (SSR) markers known to detect loci on chromosome 3B. The SSR markers were selected from the BARC (Song et al. 2005), GWM (Röder et al. 1998), WMC (Somers et al. 2004), CFA (Sourdille et al. 2003), and HBG (Torada et al. 2006) primer sets. Polymerase chain reactions (PCR) were performed as described in Röder et al. (1998), and PCR-amplified products were visualized as described in Zhang et al. (2009). Linkage analysis was conducted using the MapDisto v1.7 program (Lorieux 2012) as described in Faris et al. (2014a). The RICLs of the LD3B population segregated into discrete resistant and susceptible classes for reaction to all the isolates and were therefore classified as either resistant or susceptible. Because resistance segregated as a qualitative trait, it was mapped as a single locus relative to the molecular markers on chromosome 3B.
Race-nonspecific tan spot resistance on chromosome arm 3BL in common wheat was previously identified using biparental populations derived from BR34 × Grandin (BG population) (Faris and Friesen 2005) and from Louise × Penawawa (LouPen population) (Kariyawasam et al. 2016). Linkage maps for these populations were constructed based on SSRs and target region amplified polymorphisms (TRAPs) for the BG population and SSRs and the Illumina iSelect 9 k array for the LouPen population. Recently, the Illumina iSelect 90 k array was used to genotype both the BG and the LouPen populations. To make comparisons among the chromosome 3B linkage maps of these two populations and the LD3B population, we selected all 90 k SNP markers on chromosome 3B in common between the BG and LouPen populations, and SSR markers in both populations that were in common either with each other, or with the LD3B population, to reconstruct the 3B linkage maps in both populations using MapDisto v1.8 (Lorieux 2012). The phenotypic data, which included data from all P. tritici-repentis races for both populations, were re-analyzed against the new 3B linkage maps using the software program QGene v4.3 (Joehanes and Nelson 2008) to determine precisely the relative position of the 3B QTL in each population.
Development and screening of semi-thermal asymmetric reverse PCR (STARP) markers
Four SNP markers that were closely associated with the 3B QTL in the BG and LouPen populations were converted to semi-thermal asymmetric reverse PCR (STARP) markers (Long et al. 2017). The primers used for STARP marker development are presented in Table 2. PCR was carried out using a total reaction volume of 10 μl consisting of 100 ng genomic DNA, 0.9 × NH4 + buffer, 1.5 mM MgCl2, 50 μM dNTPs, 0.8 M betaine, 0.04% (w/v) bovine serum albumin (BSA), 200 nM common reverse primer, 200 nM of each priming element-adjustable primer (PEA-primer 1 and PEA-primer 2), 40 nM of each asymmetrically modified allele-specific primer (AMAS forward primer 1 and AMAS forward primer 2), and 1.5 U of Taq DNA polymerase (without 3′ → 5′ exonuclease activity). PCR conditions were 94 °C for 5 min, followed by 6 cycles of 94 °C for 30 s and 56 °C for 2 min where the Ta/e decreased 1 °C each cycle. This touchdown cycle was followed by 40 cycles (45 cycles if using fluorescent PEA-primers) of 94 °C for 20 s, 62 °C for 30 s, 72 °C 1:30 min, followed by a final extension at 72 °C for 7 min. Amplicons were electrophoresed on 6% polyacrylamide gels, stained with GelRed™ nucleic acid stain (Biotium Inc., Fremont, CA), and visualized with a Typhoon™ FLA 9500 variable mode laser scanner (GE Healthcare Life Sciences, Marlborough, MA). For gel-free digital allele discrimination, PEA-primers 1 and 2 labeled with FAM and HEX fluorophores, respectively, were used in the STARP protocol. Fluorescence intensity of the amplified product was measured using a CFX384 Touch™ Real-Time PCR Detection System (Bio-Rad Laboratories, Inc., Hercules, CA) where STARP markers were used to genotype the three mapping populations (BG, LouPen, and LD3B) and the panel of 21 wheat lines mentioned above.
Results
Chromosome substitution line infiltrations with Ptr ToxA
Infiltrations with Ptr ToxA revealed that LDN was sensitive and IsraelA was insensitive as shown previously (Faris and Friesen 2009; Virdi et al. 2016). As expected, all of the substitution lines were sensitive to Ptr ToxA with the exception of IsA-5B (data not shown). In this case, the LDN 5B chromosome, which harbors Tsn1, was replaced by the non-Tsn1-containing 5B chromosomes of IsraelA, which rendered the plants insensitive to Ptr ToxA.
Inoculations of chromosome substitution lines
LDN was susceptible to tan spot caused by inoculations of all four isolates with average reaction types of 4.33, 4.38, 4.17, and 3.83 to Pti2 (race 1), 86-124 (race 2), 331-9 (race 3), and DW5 (race 5), respectively (Fig. 1, Fig. 2). In contrast, the wild emmer accession IsraelA was resistant to all four isolates with reaction types of 1.33, 1.63, 1.00, and 1.50, respectively. No chromosome substitution line was more susceptible than LDN (Fig. 2). IsraelA and the chromosome substitution line IsA-3B were significantly more resistant than LDN to all four P. tritici-repentis isolates (Fig. 2). Substitution lines IsA-3A and IsA-6A were each significantly less susceptible than LDN to three of the four isolates with IsA-3A being less susceptible to Pti2, 86-124, and DW5, and IsA-6A less susceptible to Pti-2, 86-124, and 331-9. The substitution line IsA-1A was less susceptible than LDN to isolates 86-124 and DW5, but it did not differ for Pti2 and 331-9.
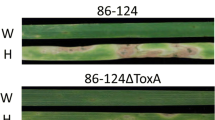
Leaves of Langdon (LDN), the LDN-Triticum turgidum ssp. dicoccoides accession IsraelA chromosome 3B disomic substitution line (IsA-3B), and the F1 derived from the cross between LDN and IsA-3B inoculated with Pyrenophora tritici-repentis isolates Pti2 (race 1), 86-124 (race 2), 331-9 (race 3), and DW5 (race 5)
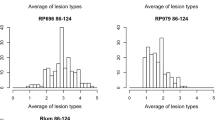
Bar graphs of average lesion type reactions of Langdon, IsraelA, and the chromosome substitution lines for Pyrenophora tritici-repentis isolates Pti2 (race 1), 86-124 (race 2), 331-9 (race 3), and DW5 (race 5). For each isolate, bar columns labeled with different lowercase letters are significantly different at P < 0.05. Bars colored in green indicate substitution lines that were significantly different from Langdon
Genetic analysis and mapping of the tan spot resistance in IsA-3B
Evaluation of the LD3B population for reaction to tan spot caused by all four P. tritici-repentis isolates revealed that responses of the RICLs could be clearly classified as either a reaction type less than 2.0 or a reaction type greater than 3.0, and therefore each RICL was classified in a qualitative manner, i.e., as either resistant (R) or susceptible (S) (Supplementary Table 1). Each RICL had identical scores for all isolates, indicating the same gene was likely conditioning race-nonspecific resistance. However, the segregation of the gene did not fit the expected 1:1 ratio, and instead, the population segregated 31 resistant:60 susceptible (χ2 = 9.24, P = 0.002).
To determine whether the resistance conferred by IsA-3B was dominant or recessive, we evaluated seven F1 plants derived from the LDN × IsA-3B cross along with eleven LDN plants and six IsA-3B plants with isolate 86-124. In this experiment, the F1 plants had an average reaction type of 1.14, whereas LDN and IsA-3B had average reaction types of 3.33 and 1.16, respectively, indicating that the resistance derived from IsA-3B was dominant (Fig. 1).
A total of 44 SSR markers were mapped in the LD3B RICL population resulting in a linkage map 177.1 cM in length (Fig. 3). Twenty-eight of the markers in the proximal region of the chromosome had segregation ratios that deviated significantly (P < 0.05) from the expected 1:1 ratio. The race-nonspecific tan spot resistance gene, which we designated as Tsr7, mapped within this region of distorted markers and cosegregated with SSR markers hbg218 and wmc69. Comparisons of common markers on the LD3B map and the deletion-based physical maps reported by Sourdille et al. (2004) indicated that gwm376 was located in the most proximal deletion bin on the long arm of chromosome 3B. Because Tsr7 mapped 1.2 cM distal to gwm376 on the LD3B map, Tsr7 is likely located in the proximal region of 3BL as well.
Genetic linkage maps of chromosome 3B developed in the BR34 × Grandin (BG) population (left), the Langdon × Langdon—Triticum turgidum ssp. dicoccoides disomic chromosome substitution line (LD3B) population (middle), and the Louise × Penawawa (LouPen) population (right). Markers are shown on the right of the linkage maps, and genetic distances are on the left. Composite interval mapping regression curves for reaction to tan spot caused by Pyrenophora tritici-repentis are shown to the right of the BG and LouPen maps with the LOD scale on the x axis. The boxed legend describes the different P. tritici-repentis isolates used. The phenotypic data for tan spot in the BG and LouPen populations were published in Faris and Friesen (2005) and Kariyawasam et al. (2016), respectively. The blue dashed lines indicate common SSR markers across maps. The orange and purple shaded regions on the BG and LouPen maps represent markers involved in an inversion between the two maps. Markers outlined in a red line indicate markers occupying the same locus. The green shaded markers represent the locus that defines the peak of the tan spot race-nonspecific resistance QTL on the BG and LouPen maps, and the Tsr7 locus on the LD3B map. The blue elliptical object indicates the centromere on the LD3B map
Re-mapping and analysis of tan spot resistance on chromosome 3B in the BG and LouPen populations
Genotyping of the BG and LouPen populations with the wheat Illumina 90 k iSelect array led to the identification of 934 and 340 SNP markers on chromosome 3B in each of the populations, respectively. Of these, a total of 148 SNP markers were common to both maps and were used to reconstruct the 3B linkage maps along with SSR markers that were in common between the two maps and/or with the LD3B map. The resulting maps consisted of 154 and 158 markers for the BG and LouPen populations, respectively (Fig. 3). These markers accounted for 50 loci spanning 174.95 cM on BG chromosome 3B, and 44 loci spanning 120.15 cM on LouPen chromosome 3B. The markers were perfectly collinear between the BG and LouPen 3B maps with one exception; an apparent inversion involving markers IWB5120, IWB36611, IWB56882, IWB11529, IWB8689, IWB63642, and barc218 occurred on the short arm (Fig. 3). Comparisons of all common SSR markers between the BG and LouPen chromosome 3B maps with the LD3B map indicated all were perfectly collinear.
Re-evaluation of the tan spot disease ratings in the BG and LouPen populations (Faris and Friesen 2005; Kariyawasam et al. 2016) revealed a locus at the seven cosegregating SNP markers (IWB7782, IWB370, IWB40103, IWB80300, IWB45103, IWB80906, and IWB80299) that defined the peak of the race-nonspecific tan spot resistance QTL (Fig. 3). In the BG population, the LOD values for tan spot reaction types ranged from 5.0 to 20.4 and explained from 17.5 to 55.0% of the variation in disease. LOD values ranged from 11.5 to 32.7 and the QTL explained from 24.4 to 55.1% of the variation in the LouPen population. SSR marker wmc69, which cosegregated with the Tsr7 locus in the LD3B population, also cosegregated with the seven SNP markers that defined the QTL peak in the LouPen population. Therefore, the tan spot resistance locus in all three populations was determined to be at the same genetic position on chromosome arm 3BL, providing strong evidence that the same gene likely conferred the observed race-nonspecific resistance in each population.
Development of STARP markers for Tsr7
STARP markers were developed based on the 90 k iSelect SNP markers IWB7782, IWB40103, IWB80300, and IWB80906, and the STARP markers were designated as fcp735, fcp736, fcp737, and fcp738, respectively. The four markers were used to genotype the BG and LouPen populations, and the allelic segregation of the markers was compared to the segregation of the corresponding 90 k iSelect SNP marker. The STARP marker segregation was identical to the corresponding SNP marker in each case (Fig. 3), indicating that the STARP markers detected the targeted loci. The STARP markers were then used to genotype the LD3B population, and linkage analysis indicated that all four markers cosegregated with the Tsr7 locus (Fig. 3).
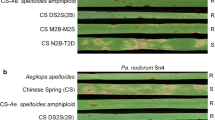
The four STARP markers were then used to genotype a small panel of wheat lines for which the status of the 3B resistance QTL was known from past research (Faris et al. 1997, 2012; Faris and Friesen 2005; Chu et al. 2008, 2010; Tadesse et al. 2010; Kariyawasam et al. 2016; JD Faris and TL Friesen unpublished data). Of the 21 lines, IsA-3B, LD3B-81, Penawawa, and BR34 were known to carry the 3B resistance QTL, whereas the remaining 17 lines were known to lack the QTL. Genotyping of the 21 lines indicated that the 3B QTL-containing lines shared the same marker allele for all four STARP markers. Among the lines that lacked the 3B QTL, 13 carried an allele different from the one harbored by the 3B QTL-containing lines, and four had the same allele as the resistant lines (Table 1, Fig. 4). This indicated that Opata 85, PI 41025, Chinese Spring, and ND495 have a recombination event(s) between the STARP marker loci and Tsr7, or the gene was deactivated by mutation without changing the marker haplotype.
Polyacrylamide gel electrophoresis assays of the four STARP markers fcp735, fcp736, fcp737, and fcp738 evaluated on the 21 wheat lines described in Table 1. The asterisks denote lines that are known to carry Tsr7, and lines without an asterisk are known to not carry Tsr7 based on previous genetic studies
To determine whether the STARP markers were suitable for genotyping using a digital assay, all four were used to genotype the BG population using a real-time detection system as described in Materials and Methods. Allele discrimination for all four was successful (Fig. 5) indicating that the four STARP markers are suitable for use in either gel or gel-free genotyping assays.
Discussion
Much less work has been done to understand the genetics of tan spot resistance in durum wheat than in common wheat. The LDN-T. turgidum ssp. dicoccoides chromosome substitution lines provide valuable tools for which to dissect and study the genetics of tan spot resistance in tetraploid wheat because they allow the effects of single chromosomes to be evaluated in isolation. Four chromosomes derived from IsraelA rendered LDN more resistant than euploid LDN and therefore likely possess tan spot resistance factors. Chromosome 3B from IsraelA conferred the highest levels of resistance to all four isolates.
Detailed comparative mapping of chromosome 3B in the BG, LouPen, and LD3B populations suggested that wild emmer accession IsraelA contains the same 3B resistance QTL as the hexaploid wheat cultivars BR34 and Penawawa (Faris and Friesen 2005; Kariyawasam et al. 2016). The gene underlying the QTL, which we designate as Tsr7, lies in a proximal region of 3BL. Tsr7 confers race-nonspecific resistance in a dominant fashion indicating the interaction with the pathogen is different from the canonical inverse gene-for-gene interactions involving dominant susceptibility genes with pathogen-produced NEs such as the Tsn1-Ptr ToxA interaction. Furthermore, Tsr7 resistance precludes susceptibility conferred by Tsn1 (Kariyawasam et al. 2016) making it a desirable target for cloning and deployment. The position of Tsr7 in the proximal region of the chromosome would likely pose difficulties for conventional map-based cloning strategies but make it an attractable candidate for alternate strategies such as MutChromSeq (Sánchez-Martin et al. 2016).
In addition to the wheat-P. tritici-repentis pathosystem, other systems involve inverse gene-for-gene interactions consisting of dominant host susceptibility genes and pathogen-produced NEs. The wheat-Parastagonospora nodorum system involves nine host gene-NE interactions that confer susceptibility, but no dominant resistance with resemblance to Tsr7 has been identified. However, the barley-Pyrenophora teres f. teres pathosystem consists of inverse gene-for-gene interactions involving dominant susceptibility genes and NEs as well as broad-spectrum dominant resistance genes, one of which resides on the long arm of chromosome 3H and could be homoeologous to Tsr7 (Koladia et al. 2017). We speculate that the resistance conferred by Tsr7 acts very early in the host–pathogen interaction and possibly prohibits penetration by the fungus or proliferation immediately after penetration, thereby precluding susceptibility conferred by the NE sensitivity genes such as Tsn1.
The finding that a wild emmer accession and two hexaploid bread wheat varieties carry the Tsr7 gene is intriguing, but it is difficult to discern the origin. It is possible that Tsr7 originated in the B-genome progenitor and passed through emmer (AB tetraploid) during the evolution of hexaploid wheat. However, gene flow between hexaploids and tetraploids subsequent to the formation of hexaploid wheat has occurred frequently (He et al. 2019), and it is possible that Tsr7 was transferred from a hexaploid to the wild emmer accession IsraelA. To support this notion, we found that IsraelA contains the domesticated form of the Q allele, which it acquired from domesticated tetraploid or hexaploid wheat subsequent to the domestication of wheat (Faris unpublished) indicating the possibility that IsraelA may have also acquired Tsr7 from a domesticated wheat form. Genome-wide association studies of large, diverse collections of tetraploid and hexaploid wheat would shed more light on the frequency of Tsr7, and the cloning of the gene would provide more insights regarding its origin.
The STARP markers developed in this research are closely linked to Tsr7 and provide user-friendly tools for marker-assisted selection. However, analysis of the panel of wheat lines showed that either recombination events or mutations have occurred in some lines resulting in reversal of the repulsion/coupling association of the markers and QTL. Therefore, it is important to know the marker allele/QTL status in initial material before using these markers for selection of resistance.
Chromosome 6A derived from IsraelA conferred moderate levels of resistance to Pti2, 86-124, and 331-9, and although not significant, it was slightly more resistant to DW5 than was LDN. Therefore, chromosome 6A from IsraelA appears to harbor race-nonspecific resistance gene(s) with minor effects. To our knowledge, no tan spot resistance genes have been identified on chromosome 6A in either hexaploid or tetraploid wheat. In evaluations done by Singh et al. (2006, 2007, 2008a), IsA-6A also showed less disease than other substitution lines for all the isolates they used.
Chromosome 3A from IsraelA conferred minor levels of resistance to Pti2, 86-124, and DW5. Tadesse et al. (2006) reported a single recessive gene conferring resistance to a race 1 isolate (ASC1) derived from the landrace Salamouni on chromosome 3A, and Chu et al. (2010) identified a QTL with minor effects on 3A for resistance to Pti2 and 86-124, with resistance effects derived from the durum cultivar Lebsock. In addition, Singh et al. (2008b) showed that a QTL on chromosome arm 3AS was associated with resistance to tan spot caused by a different race 1 isolate. The possibility that the chromosome 3A resistance in IsraelA might be the same as that identified in other sources will need further investigation.
The IsraelA set of chromosome substitution lines was previously screened by Singh et al. (2007) for reaction to tan spot caused by races 1, 2, 3 and 5. For that experiment, they used the same isolates for races 1-3 (Pti2, 86-124, and 331-9) as the current study and a different isolate (DW13) for race 5, and they infiltrated the entire set with Ptr ToxA. Their results showed that all the IsraelA substitution lines were sensitive to Ptr ToxA, with the exception of IsA-5B, which agreed with our findings that the recessive tsn1 allele on chromosome 5B is present in IsraelA.
However, there were some discrepancies between the results reported by Singh et al. (2007) and our results regarding the tan spot reactions obtained from fungal spore inoculation experiments. Singh et al. (2007) reported that the IsA-5B substitution line was resistant to tan spot caused by isolates Pti2 and 86-124 (races 1 and 2) and susceptible to isolates 331-9 and DW13 (races 3 and 5), whereas IsA-3B was resistant to isolates 331-9 and DW13, but not Pti2 and 86-124. Our results differ in that we found that IsA-5B was susceptible to all four tested isolates, and IsA-3B was resistant to all four. To confirm the results of our IsA-5B inoculations, we performed additional spore inoculations on plants derived from various seed sources, including seed from Dr. L.R. Joppa’s early increases of this line, and all results indicated that IsA-5B was susceptible. In further support of our findings, Virdi et al. (2016) reported that IsA-5B was susceptible to tan spot caused by 86-124 even though it was insensitive to Ptr ToxA. We do not have a plausible explanation for the discrepancy between the study by Singh et al. (2007) and the current study.
The finding that sensitivity to the NE Ptr ToxA in durum wheat is not associated with susceptibility to Ptr ToxA-producing isolates is consistent with previous research in tetraploid wheat. As summarized in Virdi et al. (2016), studies conducted to date indicate that the Tsn1-Ptr ToxA interaction is not associated with the development of tan spot in tetraploid wheat. However, in hexaploid wheat, the role of the Tsn1-Ptr ToxA interaction can range from nonsignificant to highly significant depending on the genetic background of the host. The reasons for this are not yet known, but the phenomenon is a subject of further study in our laboratories.
Although the Tsn1-Ptr ToxA interaction does not appear to be associated with tan spot susceptibility in tetraploid wheat, Virdi et al. (2016) showed that the Tsc2-Ptr ToxB interaction is important and significantly associated with susceptibility to chlorosis produced by race 5 isolates. The Tsc2 gene lies on the short arm of chromosome 2B (Friesen and Faris 2004; Abeysekara et al. 2010), and although the substitution line IsA-2B was not available for evaluation, effects of the Tsc2-Ptr ToxB interaction would not have been observed because neither LDN or IsraelA harbor Tsc2 (Fig. 2 and Virdi et al. 2016). Similarly, the Tsc1-Ptr ToxC interaction was not a factor in this research because LDN does not carry Tsc1 (Virdi et al. 2016), and IsraelA did not express chlorotic symptoms upon infection by Ptr ToxC-producing isolates (Pti-2 and 331-9) suggesting that it does not harbor Tsc1 either. Therefore, the Tsc1-Ptr ToxC interaction still has not been directly evaluated in tetraploid wheat.
The results of this work indicate that chromosome 3B from wild emmer accession IsraelA harbors Tsr7, a gene that confers high levels of race-nonspecific resistance to tan spot of durum wheat and it is likely the same as one that is known to exist in some lines of hexaploid wheat. The markers developed in this research should prove useful for deployment of the resistance and development of durum cultivars with improved tan spot resistance.
References
Abeysekara NS, Friesen TL, Liu ZH, McClean PE, Faris JD (2010) Marker development and saturation mapping of the tan spot Ptr ToxB sensitivity locus Tsc2 in hexaploid wheat. Plant Genome 3:179–189
Cantrell RG, Joppa LR (1991) Genetic analysis of quantitative traits in wild emmer (Triticum turgidum L. var. dicoccoides). Crop Sci 31:645–649
Chu CG, Friesen TL, Xu SS, Faris JD (2008) Identification of novel tan spot resistance loci beyond the known host-selective toxin insensitivity loci in wheat. Theor Appl Genet 117:873–881
Chu CG, Chao S, Friesen TL, Faris JD, Zhong S, Xu SS (2010) Identification of novel tan spot resistance QTLs using an SSR-based linkage map of tetraploid wheat. Mol Breeding 25:327–338
Faris JD, Friesen TL (2005) Identification of quantitative trait loci for race-nonspecific resistance to tan spot of wheat. Theor Appl Genet 111:386–392
Faris JD, Friesen TL (2009) Reevaluation of a tetraploid wheat population indicates that the Tsn1-ToxA interaction is the only factor governing susceptibility to Stagonospora nodorum blotch. Phytopathology 99:906–912
Faris JD, Anderson JA, Francl LJ, Jordahl JG (1997) RFLP mapping of resistance to chlorosis induction by Pyrenophora tritici-repentis in wheat. Theor Appl Genet 94:98–103
Faris JD, Haen KM, Gill BS (2000) Saturation mapping of a gene-rich recombination hot spot region in wheat. Genetics 154:823–835
Faris JD, Zhang Z, Lu HJ, Lu SW, Reddy L, Cloutier S, Fellers JP, Meinhardt SW, Rasmussen JB, Xu SS, Oliver RP, Simons KJ, Friesen TL (2010) A unique wheat disease resistance-like gene governs effector-triggered susceptibility to necrotrophic pathogens. Proc Natl Acad Sci USA 107:13544–13549
Faris JD, Abeysekara NS, McClean PE, Xu SS, Friesen TL (2012) Tan spot susceptibility governed by the Tsn1 locus and non race-specific resistance QTL in a population derived from wheat lines Salamouni and Katepwa. Mol Breeding 30:1669–1678
Faris JD, Liu ZH, Xu SS (2013) Genetics of tan spot resistance in wheat. Theor Appl Genet 126:2197–2217
Faris JD, Zhang QJ, Chao S, Zhang ZC, Xu SS (2014a) Analysis of agronomic and domestication traits in a durum x cultivated emmer wheat population using a high-density single nucleotide polymorphism-based linkage map. Theor Appl Genet 127:2333–2348
Faris JD, Zhang ZC, Garvin DF (2014b) Molecular and comparative mapping of genes governing spike compactness from wild emmer wheat. Mol Genet Genomics 289:641–651
Friesen TL, Faris JD (2004) Molecular mapping of resistance to Pyrenophora tritici-repentis race 5 and sensitivity to Ptr ToxB in wheat. Theor Appl Genet 109:464–471
Friesen TL, Ali S, Klein KK, Rasmussen JB (2005) Population genetic analysis of a global collection of Pyrenophora tritici-repentis, causal agent of tan spot of wheat. Phytopathology 95:1144–1150
Garvin DF, Stack RW, Hansen JM (2009) Quantitative trait locus mapping of increased Fusarium head blight susceptibility associated with a wild emmer wheat chromosome. Phytopathology 99:447–452
Gonzalez-Hernadez JL, Elias EM, Kianian SF (2004) Mapping genes for grain protein concentration and grain yield on chromosome 5B of Triticum turgidum (L.) var. dicoccoides. Euphytica 139:217–225
Gonzalez-Hernandez JL, Singh PK, Mergoum M, Adhikari TB, Kianian SF, Simsek S, Elias EM (2009) A quantitative trait locus on chromosome 5B controls resistance of Triticum turgidum (L.) var. dicoccoides to Stagonospora nodorum blotch. Euphytica 166:199–206
He F, Pasam R, Shi F, Kant S, Keeble-Gagnere G, Kay P, Forrest K, Fritz A, Hucl P, Wiebe K, Knox R, Cuthbert R, Pozniak C, Akhunova A, Morrell PL, Davies JP, Webb SR, Spangenberg G, Hayes B, Daetwyler H, Tibbits J, Hayden M, Akhunov E (2019) Exome sequencing highlights the role of wild-relative introgression in shaping the adaptive landscape of the wheat genome. Nat Genet 51:896–904
Joehanes R, Nelson JC (2008) QGene 4.0, an extensible Java QTL-analysis platform. Bioinformatics 24:2788–2789
Joppa LR (1993) Chromosome engineering in tetraploid wheat. Crop Sci 33:908–913
Joppa LR, Cantrell RG (1990) Chromosomal location of genes for grain protein content of wild tetraploid wheat. Crop Sci 30:1059–1064
Joppa LR, Hareland GA, Cantrell RG (1991) Quality characteristics of the Langdon durum-dicoccoides chromosome substitution lines. Crop Sci 31:1513–1517
Joppa LR, Du C, Hart GE, Hareland GA (1997) Mapping gene(s) for grain protein in tetraploid wheat (Triticum turgidum L.) using a population of recombinant inbred chromosome lines. Crop Sci 37:1586–1589
Kariyawasam GK, Carter AH, Rasmussen JB, Faris JD, Xu SS, Mergoum M, Liu ZH (2016) Genetic relationships between race-nonspecific and race specific interactions in the wheat-Pyrenophora tritici-repentis pathosystem. Theor Appl Genet 129:897–908
Koladia VM, Faris JD, Richards JK, Brueggeman RS, Chao S, Friesen TL (2017) Genetic analysis of net form net blotch resistance in barley lines CIho 5791 and Tifang against a global collection of P. teres f. teres isolates. Theor Appl Genet 130:163–173
Kumar S, Gill BS, Faris JD (2007a) Identification and characterization of segregation distortion loci along chromosome 5B in tetraploid wheat. Mol Genet Genomics 278:1870196
Kumar S, Stack RW, Friesen TL, Faris JD (2007b) Identification of a novel Fusarium head blight resistance quantitative trail locus on chromosome 7A in tetraploid wheat. Phytopathology 97:592–597
Lamari L, Bernier CC (1989) Evaluation of wheat lines and cultivars to tan spot Pyrenophora tritici-repentis based on lesion type. Can J Plant Pathol 11:49–56
Lamari L, Sayoud R, Boulif M, Bernier CC (1995) Identification of a new race in Pyrenophora tritici-repentis: implications for the current pathotype classification system. Can J Plant Pathol 17:312–318
Liu ZH, Zurn JD, Kariyawasam G, Faris JD, Shi GJ, Hansen J, Rasmussen JB, Acevedo M (2017) Inverse gene-for-gene interactions contribute additively to tan spot susceptibility in wheat. Theor Appl Genet 130:1267–1276
Long Y, Chao WS, Ma G, Xu SS, Qi L (2017) An innovative SNP genotyping method adapting to multiple platforms and throughputs. Theor Appl Genet 130:597–607. https://doi.org/10.1007/s00122-016-2838-4
Lorieux M (2012) MapDisto: fast and efficient computation of genetic linkage maps. Mol Breed 30:1231–1235
Nalam VJ, Vales MI, Watson CJW, Kianian SF, Riera-Lizarazu O (2006) Map-based analysis of genes affecting the brittle rachis character in tetraploid wheat (Triticum turgidum L.). Theor Appl Genet 112:373–381
Otto CD, Kianian SF, Elias EM, Stack RW, Joppa LR (2002) Genetic dissection of a major Fusarium head blight QTL in tetraploid wheat. Plant Mol Biol 48:625–632
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier M-H, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Sánchez-Martin J, Steuernagel B, Ghosh S, Herren G, Hurni S, Adamski N, Vrána J, Kubalákaová M, Krattinger SG, Wicker T, Doležel J, Keller B, Wulff BB (2016) Rapid gene isolation in barley and wheat by mutant chromosome sequencing. Genome Biol 17:221
Singh PK, Gonzalez-Hernandez JL, Mergoum M, Ali S, Adhikari TB, Kianian SF, Elias E, Hughes GR (2006) Identification and molecular mapping of a gene conferring resistance to Pyrenophora tritici-repentis race 3 in tetraploid wheat. Phytopathology 96:885–889
Singh PK, Mergoum M, Adhikari TB, Kianian SF, Elias EM (2007) Chromosomal location of genes for seeding resistance to tan spot and Stagonospora nodorum blotch in tetraploid wheat. Euphytica 155:27–34
Singh PK, Mergoum M, Gonzalez-Hernandez JL, Ali S, Adhikari TB, Kianian SF, Elias EM, Hughes GR (2008a) Genetics and molecular mapping of resistance to necrosis-inducing race 5 of Pyrenophora tritici-repentis in tetraploid wheat. Mol Breeding 21:293–304
Singh S, Bockus WW, Sharma I, Bowden RL (2008b) A novel source of resistance in wheat to Pyrenophora tritici-repentis race 1. Plant Dis 92:91–95
Singh PK, Singh RP, Duveiller E, Mergoum M, Adhikari TB, Elias EM (2010) Genetics of wheat-Pyrenophora tritici-repentis interactions. Euphytica 171:1–13
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Song QJ, Shi JR, Singh S, Fickus EW, Costa JM, Lewis J, Gill BS, Ward R, Cregan PB (2005) Development and mapping of microsatellite (SSR) markers in wheat. Theor Appl Genet 110:550–560
Sourdille P, Cadalen T, Guyomarc’h H, Snape JW, Perretant MR, Charmet G, Boeuf C, Bernard S, Bernard M (2003) An update of the Courtot x Chinese Spring intervarietal molecular marker linkage map for the QTL detection of agronomic traits in wheat. Theor Appl Genet 106:530–538
Stack RW, Elias EM, Mitchell Fetch J, Miller JD, Joppa LR (2002) Fusarium head blight reaction of Langdon durum-Triticum dicoccoides chromosome substitution lines. Crop Sci 42:637–642
Tadesse W, Hsam SLK, Zeller FJ (2006) Evaluation of common wheat cultivars for tan spot resistance and chromosomal location of a resistance gene in the cultivar ‘Salamouni’. Plant Breeding 125:318–322
Tadesse W, Reents HJ, Hsam SLK, Zeller FJ (2010) Monosomic analysis of tan spot resistance gene in the winter wheat cultivar ‘Arina’. Plant Breeding 129:477–479
Torada A, Koike M, Mochida K, Ogihara Y (2006) SSR-based linkage map with new markers using a population of common wheat. Theor Appl Genet 112:1042–1051
Virdi SK, Liu ZH, Overlander ME, Zhang ZC, Xu SS, Friesen TL, Faris JD (2016) New insights into the roles of host gene-necrotrophic effector interactions in governing susceptibility of durum wheat to tan spot and Septoria nodorum blotch. Genes Genomes Genetics 6:1. https://doi.org/10.1534/g3.116.036525
Zhang ZC, Friesen TL, Simons KJ, Xu SS, Faris JD (2009) Development, identification, and validation of markers for marker-assisted selection against Stagonospora nodorum toxin sensitivity genes Tsn1 and Snn2 in wheat. Mol Breed 23:35–49
Acknowledgements
The authors thank Shirley Zhong for technical assistance. This work was supported by USDA-Agricultural Research Service CRIS project 3060-21000-038-00D. This material is based upon work supported, in part, by the National Institute of Food and Agriculture, United States Department of Agriculture (USDA), under Hatch project number ND02224.
Author information
Authors and Affiliations
Contributions
JF and ZL initiated the study and designed the experiment. JF, SX and AC developed and/or maintained the mapping populations and genetic stocks. JF, MO, GK, and AC conducted mapping studies and/or contributed marker information and marker analysis. MO developed STARP markers. JF, MO, and GK conducted linkage analysis. ZH, MO, and GK conducted tan spot inoculations and analysis. JF and ZH wrote the manuscript and all authors contributed to the final version.
Corresponding authors
Ethics declarations
Conflict of interest
All authors have no conflict of interest.
Additional information
Communicated by Thomas Miedaner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Faris, J.D., Overlander, M.E., Kariyawasam, G.K. et al. Identification of a major dominant gene for race-nonspecific tan spot resistance in wild emmer wheat. Theor Appl Genet 133, 829–841 (2020). https://doi.org/10.1007/s00122-019-03509-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03509-8