Abstract
Key message
The RpsQ Phytophthora resistance locus was finely mapped to a 118-kb region on soybean chromosome 3. A best candidate gene was predicted and three co-segregating gene markers were developed.
Abstract
Phytophthora root rot (PRR), caused by Phytophthora sojae, is a major threat to sustainable soybean production. The use of genetically resistant cultivars is considered the most effective way to control this disease. The Chinese soybean cultivar Qichadou 1 exhibited a broad spectrum resistance, with a distinct resistance phenotype, following inoculation with 36 Chinese P. sojae isolates. Genetic analyses indicated that the disease resistance in Qichadou 1 is controlled by a single dominant gene. This gene locus was designated as RpsQ and mapped to a 118-kb region between BARCSOYSSR_03_0165 and InDel281 on soybean chromosome 3, and co-segregated with Insert11, Insert144 and SNP276. Within this region, there was only one gene Glyma.03g27200 encoding a protein with a typical serine/threonine protein kinase structure, and the expression pattern analysis showed that this gene induced by P. sojae infection, which was suggested as a best candidate gene of RpsQ. Candidate gene specific marker Insert144 was used to distinguish RpsQ from the other known Rps genes on chromosome 3. Identical polymerase chain reaction amplification products were produced for cultivars Qichadou 1 (RpsQ) and Ludou 4 (Rps9). All other cultivars carrying Rps genes on chromosome 3 produced different PCR products, which all lacked a 144-bp fragment present in Qichadou 1 and Ludou 4. The phenotypes of the analyzed cultivars combined with the physical position of the PRR resistance locus, candidate gene analyses, and the candidate gene marker test revealed RpsQ and Rps9 are likely the same gene, and confer resistance to P. sojae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phytophthora root rot (PRR), caused by Phytophthora sojae M. J. Kaufmann and J. W. Gerdemann, is one of the most economically destructive diseases of soybean (Kasuga et al. 1997; Schmitthenner 1999; Sugimoto et al. 2011). Phytophthora sojae is a soil-borne oomycete that is capable of infecting soybean plants at all developmental stages, resulting in symptoms including damping-off, root and stem rot, leaf yellowing, and wilting (Schmitthenner 1985; Tyler et al. 2007; Gunadi 2012; Lee et al. 2013). Phytophthora root rot was first observed in Indiana, USA in 1948, and has since been detected in all major soybean-producing areas worldwide (Kaufmann and Gerdemann 1958; Schmitthenner 1985; Anderson and Buzzell 1992; Jee et al. 1998; Grau et al. 2004; Dorrance and Grünwald 2009). In China, the disease was first reported in 1991 in Heilongjiang province, which is a major soybean-growing region (Shen and Su 1991). It subsequently spread to several other regions in Heilongjiang and Fujian provinces (Chen et al. 2004; Zhang et al. 2010). Globally, the annual economic losses resulting from PRR-infected soybean plants exceed US$1–2 billion (Wrather and Koenning 2006).
Generating Phytophthora-resistant soybean plants through the use of dominant Rps genes is the most economical and environmentally safe method to prevent this disease (Dorrance et al. 2003). To date, 27 Rps genes/alleles associated with 21 loci distributed on eight soybean chromosomes have been reported (i.e., Rps1a, Rps1b, Rps1c, Rps1d, Rps1k, Rps2, Rps3a, Rps3b, Rps3c, Rps4, Rps5, Rps6, Rps7, Rps8, Rps9, Rps10, Rps11, RpsYB30, RpsYD25, RpsSu, RpsZS18, RpsSN10, RpsYD29, RpsUN1, RpsUN2, RpsJS, and the Rps gene in the Waseshiroge soybean cultivar) (Zhu et al. 2007; Yao et al. 2010; Yu et al. 2010; Ortega 2011; Sugimoto et al. 2011; Sun et al. 2011; Wu et al. 2011; Lin et al. 2013; Zhang et al. 2013a, b; Ping et al. 2015).
The continuous use of Phytophthora-resistant cultivars leads to the breakdown of resistance over time because of the generation of new virulent P. sojae strains (Abney et al. 1997; Kaitany et al. 2001; Xue et al. 2015). Mutations and outcrossings between different strains increase the diversity of P. sojae populations (Förster et al. 1994). In China, the diversity among P. sojae populations is complex (Zhu et al. 2003; Cui et al. 2010; Zhang et al. 2010). Therefore, new Rps genes are constantly required to ensure sustainable disease control (Sugimoto et al. 2012).
The Rps genes follow a gene-for-gene model, and trigger a defense mechanism that elicits a hypersensitive response (Parlevliet 2002; Tyler 2007; Wang et al. 2011). Among the mapped Rps genes in the soybean genome, only three genes have been cloned, namely Rps1k, Rps2, and Rps4. These genes encode nucleotide-binding site, leucine-rich repeat (NBS-LRR) proteins, which are the most common R gene types in plants (Sandhu et al. 2004; Graham et al. 2002). Based on the sequence preceding the NBS domain, Rps1k and Rps4 are similar to a member of the coiled-coil NBS-LRR class, while Rps2 is considered to belong to the Toll/Interleukin1 receptor NBS-LRR class. Additionally, RpsYD29, RpsYD25, RpsUN1, and RpsUN2 are present in regions rich in NBS-LRR genes according to the information in the SoyBase database (Zhang et al. 2013b; Fan et al. 2009; Lin et al. 2013). Recently, Zhang et al. (2013a) mapped Rps10 in the genome of the Chinese soybean cultivar Wandou 15, and identified two candidate genes. The conserved sequences of the Rps10 candidate genes were determined to encode a serine/threonine kinase and an LRR domain.
There is an abundance of sources of Phytophthora-resistant cultivated soybeans and wild soybeans in China (Zhu et al. 2006; Xia et al. 2011a; Zhong et al. 2015). Soybean cultivar Qichadou 1 was bred at the Shandong Academy of Agricultural Sciences and released in 1995. Qichadou 1 is an elite cultivar with superior yield potential, and produces soybeans rich in fatty acids and proteins. It is also resistant to soybean cyst nematodes (races 1 and 3) and soybean mosaic virus. We previously found that Qichadou 1 plants exhibited a broad-spectrum resistance to P. sojae (data not shown).
The objectives of this study were to (1) characterize the inheritance of Phytophthora resistance in Qichadou 1 soybean, (2) finely map the Rps gene(s) with molecular markers and identify candidate gene(s), and (3) develop co-segregating candidate gene markers for P. sojae resistance in soybean, which would be relevant for molecular marker-assisted selection during breeding.
Materials and methods
Plant resources
A segregating population comprising 207 F2 individuals was generated by selfing a single F1 hybrid derived from a cross between Jikedou 2 (susceptible) and Qichadou 1 (resistant) soybean cultivars. The F2:3 families were developed through single seed descent of each F2 individual under field conditions in 2013. The seeds of each F2:3 family were harvested separately for genetic analyses and disease evaluations.
The following 19 P. sojae-resistant soybean differentials carrying unique Rps genes were tested to determine their reactions to the P. sojae isolates used in this study: Harlon (Rps1a), Harosoy 13XX (Rps1b), Williams 79 (Rps1c), PI 103091 (Rps1d), Williams 82 (Rps1k), L76–1988 (Rps2), L83–570 (Rps3a), PRX 146–36 (Rps3b), PRX 145–48 (Rps3c), L85–2352 (Rps4), L85–3059 (Rps5), Harosoy 62XX (Rps6), Harosoy (Rps7), PI 399073 (Rps8), Ludou 4 (Rps9), Wandou 15 (Rps10), Yudou 25 (RpsYD25), Yudou 29 (RpsYD29), and Youbian 30 (RpsYB30). Williams (rps) was used as a susceptible control. All soybean cultivars or lines were obtained from the China National Gene Bank at the Institute of Crop Sciences, Chinese Academy of Agricultural Sciences.
Phytophthora sojae isolates and phenotypic assessments
A total of 36 P. sojae isolates that differed in terms of virulence were used for phenotypic assessments of Qichadou 1, Jikedou 2, Williams, and the 19 soybean differentials. The soybean plants were inoculated using a hypocotyl inoculation technique (Haas and Buzzell 1976) (Supplementary Table 1). Briefly, all isolates were cultured on carrot agar medium and incubated at 25 °C in darkness for 7–10 days. A mycelial slurry was made by mixing the colonized agar and pushing it through a 10-ml syringe during the inoculation of soybean plants. The highly virulent Ps41-1 P. sojae isolate was used for phenotypic evaluations of the F2:3 population.
For disease evaluations, 15 soybean seeds of each differential were planted in vermiculite-filled paper cups (10-cm diameter) with bottom drainage and incubated in a greenhouse. For genetic analyses and gene mapping, 20 seeds of each F2:3 family were sown in paper cups. After 10 days, a mycelial slurry was used to inoculate hypocotyls that had been wounded with a 1-cm slit made by a syringe approximately 1 cm below the cotyledons. Inoculated seedlings were incubated in a misting room (100% relative humidity) at 25 °C for 2 days, and then placed in a greenhouse at 25 °C. Six days after inoculation, disease reactions were scored based on the percentage of dead seedlings.
Families, cultivars, or lines with less than 21% dead seedlings were considered resistant, while those with 21–79% or more than 79% dead seedlings were classified as segregating or susceptible, respectively (Gordon et al. 2006). Phenotypic assays were repeated three times.
DNA extraction and pooling for bulk segregant analysis
Genomic DNA was extracted from healthy soybean leaves sampled from the parents and F2 plants using the Plant Genomic DNA Kit (Tiangen Biotech, Beijing, China) according to the manufacturer’s protocols. For bulked segregant analysis, resistant and susceptible bulks were prepared by mixing equal amounts of DNA from either 10 homozygous-resistant or 10 homozygous-susceptible F2 individuals. Sample concentrations were adjusted to 20 ng/μl (Michelmore et al. 1991).
Molecular marker development and analysis
The bulked segregant analysis was employed for screening the polymorphisms between two contrasting parents using three types of PCR-based markers (Michelmore et al. 1991; Cregan et al. 1999). For SSR markers, the primer sets evenly distributed throughout the soybean genome were selected from the SoyBase database (http://www.soybase.org/) (Song et al. 2010). For further linkage analysis, the whole genome re-sequencing of the resistant (Qichadou 1) and susceptible (Jikedou 2) parents were performed on an Illumina HiSeqTM 4000 at Biomarker Technologies Corporation in Beijing, China. Thirty-five InDel (Insertion/Deletion polymorphisms) and twenty-eight single nucleotide polymorphism (SNP) markers were developed based on the polymorphisms of targeted genomic sequence between two parents using Primer Premier 5.0 software. Polymorphic markers were analyzed further to genotype the entire F2:3 mapping population.
For SSR and InDel markers, polymerase chain reaction (PCR) amplifications were completed in 10 μl reaction mixtures containing 20 ng genomic DNA, 5 μl 2 × PCR MasterMix (Tiangen Biotech), and 0.25 μl primers (10 μM each). The PCR program was as follows: 94 °C for 5 min; 35 cycles of 94 °C for 30 s, 53–61 °C (depending on primer pairs) for 30 s, and 72 °C for 30 s; 72 °C for 10 min. Samples were then cooled to 4 °C. The amplicons were resolved on 8% non-denaturing polyacrylamide gels and stained with ethidium bromide.
For SNP markers, PCRs were performed in a 10 μl volumes with 20 ng genomic DNA, 5 μl 2 × PCR MasterMix (Tiangen Biotech), 1 μl 10 × LC Green Plus (Biofire Diagnostics, Salt Lake City, UT, USA), 0.25 μl primers (10 μM each) and 10 μl mineral oil. The PCR reaction were performed as follows: 94 °C for 5 min; 50 cycles of 94 °C for 30 s, 61 °C for 30 s, and 72 °C for 30 s; 72 °C for 10 min. The melting-peak curves were acquired at temperatures ranging from 65 °C to 95 °C and analyzed using LightScanner software (version 2.0) (Idaho Technology, Salt Lake City, UT) according to the manufacturer’s instructions.
Data analysis and construction of genetic maps
Chi square (χ2) analyses were completed to determine the goodness-of-fit of the observed segregations with expected genetic ratios in the mapping population. A genetic linkage map for the target locus was then constructed using MapMaker 3.0 with a logarithm of odds threshold of 3.0 (Lincoln et al. 1993).
Quantitative real-time PCR analysis
The physical positions of the markers tightly linked to RpsQ were determined based on the soybean genome reference sequences using the BLAST program (http://soybase.org/SequenceIntro.php; http://www.Phytozome.net). The candidate genes present in the target regions were also analyzed using the BLASTN search program and the soybean database.
To determine the transcript abundance of the candidate gene in response to P. sojae, 10-day-old seedlings of Qichadou1 were inoculated with Ps41-1 using a standard hypocotyl inoculation method and a mock control. The stems were harvested at 0, 6, 12, 24, 36 and 48 h post inoculation and stored at −80 °C for further use. The roots, stems and leaves were also collected without inoculation for tissue-specific transcript abundance. Total RNA was isolated using RNAprep Pure Plant Kit (Tiangen Biotech, Beijing, China) and first-strand cDNA was synthesized using a PrimeScriptTMRT Reagent Kit (TaKaRa, Japan). Two candidate gene specific primers (F: CACACTTGCAGGCTTTTGTCT; R: GAACTCTCCTGGATGTCTTCCC) were used to determine its expression level. Real-time PCR was performed using the CFX96 Touch™ Real-Time PCR Detection System (Biobad, USA) with the SYBR® Premix Ex TaqII(TliRNaseH Plus) (TaKaRa, Japan) according to the manufacturer's instructions on the CFX96 Touch™ Real-Time PCR Detection System (Biobad, USA). The constitutive GmACT11 gene amplified with specific primers (F: ATCTTGACTGAGCGTGGTTATTCC; R: GCTGGTCCTGGCTGTCTCC) was used as an internal control. The gene expression levels were calculated using the 2−∆∆CT method. For each sample, three technological replications were conducted.
Isolation of the candidate gene
To generate genomic DNA fragments of the candidate genes for the parents, the 5′- and 3′-untranslated regions (UTRs) and gDNA were separately amplified using primers designed according to the candidate gene models. The PCR assay was completed using TransTaq ® HiFi DNA Polymerase (TransGen Biotech, Beijing, China) according to a standard protocol. The amplicons were purified from agarose gels using the Universal DNA Purification Kit (TianGen Biotech). The purified fragments were subcloned into the pEASY ®-T1 cloning vector (TransGen Biotech) and sequenced. Sequences were assembled and manually edited using DNAMAN 7.0 software. Syntenic sequences were compared between the resistant and susceptible genotypes using the MultAlin online tool (http://bioinfo.genotoul.fr/multalin/multalin.html). The primer sets are listed in Table 1.
Validation of the candidate gene marker
Based on the physical position of Insert144, Insert144 was in the cording region of the candidate gene and considered as candidate gene specific marker. To validate its utility, the marker was then used to check the allelism of RpsQ in the vicinity of its locus using the two parental lines and nine Rps gene isogenic lines (i.e., Rps1a, Rps1b, Rps1c, Rps1d, Rps1k, Rps7, Rps9, RpsYD25 and RpsYD29).
Results
Qichadou 1 is broadly resistant to Phytophthora sojae
Based on phenotypic reactions to P. sojae isolates, Qichadou 1 soybean plants were resistant to 30 of 36 isolates tested, indicating this cultivar is broadly resistant to P. sojae (Table 2). The reaction of Qichadou 1 to the 36 P. sojae isolates was not consistent with the reactions of the 19 differentials, although it was closest to that of Ludou 4 (Rps9), which was completely resistant to 26 isolates (Table 2). Qichadou 1 and Ludou 4 plants were affected differently by only four P. sojae isolates (i.e., PsAH5, PsJS10, PsJS6, and PsJMS2). Additionally, Qichadou 1 plants were more broadly resistant to P. sojae than the 19 differentials, except for Yudou 25 (RpsYD25) and Williams 82 (Rps1k), which were resistant to 33 and 30 isolates, respectively (Table 2).
Phenotypic analysis of the mapping population
Six days after inoculation, the reactions of the parents and the derived mapping population to P. sojae isolate Ps41-1 were as expected. All Jikedou 2 seedlings were dead, indicating susceptibility to isolate Ps41-1, whereas all Qichadou 1 seedlings were completely resistant, with only hypersensitive necrosis observed at infection sites.
The PRR phenotypes of the F2:3 mapping population comprising 207 families revealed there were 52 homozygous resistant, 101 heterozygous, and 54 susceptible families. A χ2 test indicated that this distribution matched a theoretical 1 RR: 2 Rr: 1 rr segregation ratio (χ2 = 0.16, p = 0.92) (Table 3). The results suggested the PRR resistance in Qichadou 1 plants was controlled by a single dominant gene, which was designated as RpsQ.
Genetic mapping of RpsQ
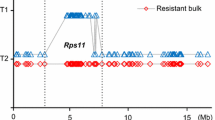
To map RpsQ in the Qichadou 1 genome, random SSR markers selected from the SoyBase database (http://www.soybase.org/) were first used to screen the parents and the bulks prepared for bulk segregant analysis. We focused on the chromosome harboring the Rps gene. Four markers (i.e., BARCSOYSSR_03_0012, Satt631, BARCSOYSSR_03_0204, and BARCSOYSSR_03_0250) on chromosome 3 were polymorphic between the parents and DNA bulks (Table 4; Fig. 1a). Linkage analyses revealed RpsQ was linked to these four polymorphic markers, and was flanked by Satt631 and BARCSOYSSR_03_0204. Thirty-four known SSR markers in the target region between Satt631 and BARCSOYSSR_03_0204 were screened. We found that three polymorphic SSR markers, BARCSOYSSR_03_0165, BARCSOYSSR_03_0176 and BARCSOYSSR_03_0184, were linked to RpsQ (Fig. 1a).
Genetic and physical map of RpsQ on chromosome 3. a Genetic linkage map of RpsQ. Molecular markers are on the left side of the map, while genetic distances (cM) are on the right side. Blue shading indicates the RpsQ target region. b Physical positions of molecular markers on chromosome 3. Loci names are on the right side of the map, while physical positions of molecular markers are provided on the left side. Blue shading indicates the RpsQ interval. c RpsQ region on the short arm of chromosome 3, with a red triangle indicating the RpsQ locus. (Color figure online)
To further delimit the target region, 35 InDel markers were secondly to screen the parents as well as two contrasting bulked and four polymorphic InDel markers (i.e., Insert11, Insert144, InDel281 and InDel286) were used to genotyping the mapping population. Thus, RpsQ was delimited to the BARCSOYSSR_03_0165-InDel281 interval of the short arm of the Chromosome 3, and co-segregated with Insert11and Insert144 (Fig. 1a). Lastly, 28 SNP markers were developed and polymorphic SNP marker SNP276 was also co-segregated with the phenotype of the mapping population (Fig. 1a). The final region containing RpsQ span approximately 118 kb (i.e., physical distance) according to the reference sequence for soybean cultivar Williams 82 (Fig. 1b, c).
Candidate gene analysis
Based on the gene annotations available in the Phytozome genomics resource (https://phytozome.jgi.doe.gov/pz/portal.html), 11 predicted genes were detected within the 118-kb target region (Supplementary Table 2) and two (i.e., Glyma.03g27000 and Glyma.03g27100) of them encode proteins with unknown function. By genomic-sequence comparison of the two parental lines, five predicted genes (i.e., Glyma.03g27200, Glyma.03g27400, Glyma.03g27500, Glyma.03g27600 and Glyma.03g27900) with non-synonymous variants in coding region were found, while the other four genes (i.e. Glyma.03g27300, Glyma.03g27700, Glyma.03g27800 and Glyma.03g28000) show synonymous variants in coding region. Among these five predicted genes, Glyma.03g027200 encoded a serine/threonine protein kinase that functioned as a receptor-like kinase (RLK) involved in signaling and plant defense activities (Afzal et al. 2008). Therefore, this gene was most likely the candidate gene of RpsQ.
To characterize the expression of the candidate gene to P. sojae in Qichadou1, a quantitative real-time PCR was performed. The examination of tissue-specific transcript abundance in Qichadou 1 showed that the candidate gene was constitutively and highly expressed in stems, followed by roots and leaves (Fig. 2a). The transcripts of the candidate gene rapidly increased in stem after P. sojae infection, reaching a maximum level at 24 h, followed by a rapid decline (Fig. 2b). According to the results, Glyma.03g027200 was suggested as a best candidate gene of RpsQ.
Expression patterns of the candidate gene in Qichadou 1. The infected samples were collected at 0, 6, 12, 24, 36, and 48 h post inoculated with isolate Ps41-1. a Expression patterns of the candidate gene mRNA level in various tissues of Qichadou 1. Roots, stems, and leaves were harvested from 10-day-old seedlings. b The candidate gene expression in stems of Qichadou 1 upon P. sojae infection. The infected samples were collected at 0, 6, 12, 24, 36 and 48 h post inoculated with isolate Ps41-1. The soybean GmACT11 gene was used as an internal reference. Values are mean ± SD of three biological replicates
Genomic sequence analysis
The genomic DNA sequences of the RpsQ candidate genes in the parents were determined based on the Glyma.03g027200 sequence, which was compared among the Qichadou 1, Jikedou 2, and Williams 82 cultivars (Supplementary Fig. 1). The RpsQ candidate gene in the Qichadou 1 genome was named STKQ, and its allele in the Jikedou 2 genome was referred to as stkq. The stkq sequence was identical to the corresponding sequence in the Williams 82 genome. The genomic STKQ sequence was 98% identical to stkq and Glyma.03g027200, which comprised a 173-bp 5′-UTR, a 3142-bp coding region, and a 260-bp 3′-UTR. There were 258 changes to the genomic STKQ sequence [i.e., single nucleotide polymorphisms (SNPs) and insertion/deletions (InDels)] (Supplementary Fig. 1). Moreover, the mean nucleotide diversity was higher at non-coding sites than at coding sites. Significant sequence differences included an 11-bp insertion in the 5′-UTR and a 144-bp insertion in the coding region of STKQ. The sequence of the 144-bp insertion was highly similar to a Glyma.03g027200 fragment (from position 734 bp to 878 bp), with the differences represented by a few SNPs (Supplementary Fig. 1). This suggested that the 144-bp insertion may have resulted from sequence duplications.
Analysis of candidate gene markers
Insert11 and Insert144 were codominant markers that co-segregates with the RpsQ (Figs. 1a, 3a, b), and were developed corresponding to the 11-bp (i.e., 5′-UTR) and 144-bp (i.e., STKQ coding region) insertions in Glyma.03g27200, respectively. The Insert11 InDel marker resulted in the PCR amplification of 221-bp and 232-bp fragments in Jikedou 2 and Qichadou 1 cultivars, respectively (Fig. 3a, b). Insert144 produced 437-bp and 581-bp fragments in Jikedou 2 and Qichadou 1, respectively (Fig. 3a, b). The two candidate gene markers were tested in the mapping population, and the results suggested that Insert11 and Insert144 can be used in marker-assisted selection (MAS) breeding. The RpsQ locus was on the short arm of chromosome 3, which contains 11 PRR resistance genes (i.e., Rps1a, Rps1b, Rps1c, Rps1d, Rps1k, Rps7, Rps9, RpsYD25, RpsYD29, RpsUN1, and the Waseshiroge Rps gene). To differentiate among these resistance genes, DNA was extracted from the differentials carrying all genes except RpsUN1 and the Waseshiroge Rps gene and analyzed using Insert144 (Fig. 4). The Qichadou 1 (RpsQ) and Ludou 4 (Rps9) 581-bp amplification products were identical. The amplicons for the other cultivars were similar to that for Jikedou 2, with all lacking the 144-bp fragment. These results suggested that RpsQ and Rps9 might represent the same gene.
Polymerase chain reaction analysis of progenies derived from the Jikedou 2 (susceptible) × Qichadou 1 (resistant) cross using the Insert11 and Insert144 candidate gene markers. a Co-segregation of Insert11 with the RpsQ locus in the parents and their 30 F2:3 progenies. b Co-segregation of Insert144 with the RpsQ locus in the parents and their 21 F2:3 progenies. M marker; P R Qichadou 1; Ps Jikedou 2; R resistant homozygote; H heterozygote; S susceptible homozygote
Discussion
In this study, we determined the Chinese soybean cultivar Qichadou 1 carried the RpsQ locus, which conferred broad spectrum Phytophthora resistance. This locus was mapped to a 118-kb region flanked by BARCSOYSSR_03_0165 and InDel281 on the short arm of chromosome 3 and co-segregated with Insert11 and Insert144 and SNP276. RpsQ was located in a region nearby 11 other known Rps genes, including five Rps1 alleles (Bernard et al. 1957; Mueller et al. 1978; Lin et al. 2013; Sun et al. 2014), Rps7 (Buzzell and Anderson 1992; Weng et al. 2001), Rps9 (Wu et al. 2011), RpsYD25 (Fan et al. 2009; Sun et al. 2011), RpsYD29 (Zhang et al. 2013b), RpsUN1 (Lin et al. 2013), and a Waseshiroge Rps gene (Sugimoto et al. 2011). To differentiate among these genes, the resistance spectrum of Qichadou 1 and 19 differentials known to carry Rps genes was investigated. Qichadou 1 exhibited a broad-spectrum resistance, which was most similar to the reaction of Ludou 4. Qichadou 1 was derived from a cross between the Peking and Ludou 4 cultivars. Peking is an important source of soybean cyst nematode resistance (races 1, 3, and 5) (Qiu et al. 1997), but it is susceptible to PRR (Xia et al. 2011b). In contrast, Ludou 4 is a P. sojae-resistant cultivar that carries Rps9 on chromosome 3 (Wu et al. 2011). Therefore, we hypothesized that RpsQ in the Qichadou 1 genome may have been inherited from the resistant parent, Ludou4. This suggests that RpsQ and Rps9 might be the same gene. However, the level of disease resistance in Qichadou 1 and Ludou 4 plants is not the same, likely because of differences in the genetic backgrounds between these two cultivars. A similar phenomenon was previously reported for other resistance genes, including Xa26 and Xa3 (Xiang et al. 2006). The level of resistance provided by Xa26 differs among lines and growth stages, and Xa26 symbolized xa9, causing dominance reversal at booting stages (Sidhu and Khush 1978).
A comparison between Rps genes and RpsQ regarding physical positions revealed that RpsQ was located at the Rps9 locus, which covers a 548-kb fragment, and was not close to the other Rps genes on chromosome 3. Furthermore, the candidate gene for RpsQ and Rps9 appears to be the same, because only Glyma.03g027200 encodes an LRR receptor kinase-like protein involved in disease resistance. The genomic sequence of the Rps9 candidate gene was determined based on the Ludou 4 genome, and a few SNPs were detected in the 3′- and 5′-UTRs during comparisons with STKQ in Qichadou 1 (Supplementary Fig. 1). Additionally, the Insert144 candidate gene marker generated the same amplicons for Ludou 4 and Qichadou 1, indicating RpsQ and Rps9 were likely the same gene.
According to deduced amino acid sequences, STKQ encoded a serine/threonine protein kinase, and belonged to the RLK class of resistance genes (Hulbert et al. 2001; Liu et al. 2007; Afzal et al. 2008). Plant RLK genes associated with diverse developmental pathways and pathogen recognition processes have been well studied (Li et al. 2006; Liu et al. 2007). Regarding plant–pathogen interactions, previous studies have concluded that Xa21 confers a high level of resistance to Xanthomonas oryzae in rice (Song et al. 1995), FLS2 regulates flagellin binding (Gómez-Gómez and Boller 2000), and SERK3/BAK1 modulates pathogen-associated molecular pattern-triggered immunity (Heese et al. 2007). Most RLKs contain extracellular repeats and an intracellular kinase, which enable the perception of elicitors and transmission of signals through phosphorylations, respectively (Morillo and Tax 2006). The RpsQ candidate gene consisted of an LRR extracellular domain and a serine/threonine kinase intracellular domain. Leucine-rich repeat domains vary considerably and evolve faster under diversifying selection pressures. This results in a versatile structural framework for the perception of extracellular signals, which influences resistance specificity (Braun and Walker 1996; Bai et al. 2002; Hulbert et al. 2001). In this study, segmental duplications occurred in the RpsQ candidate gene, thereby increasing the number of LRR-repeat units, which may have led to enhanced resistance to P. sojae (Dogimont et al. 2014; Ellis et al. 1999). The kinase domain affects protein phosphorylation-mediated signal transduction cascades, resulting in specific plant responses (Gachomo et al. 2003; Li et al. 2006). Three SNPs in the coding sequences resulted in three amino acid differences in the serine/threonine kinase motif between the RpsQ candidate gene in Qichadou 1 and its allele in Jikedou 2, which may be responsible for the differences in defense responses to P. sojae. A recent study regarding Rps genes in soybean concluded the Rps10 candidate gene encoded an LRR-RLK containing a serine/threonine kinase domain (Zhang et al. 2013a), with a similar structure to that of the protein encoded by the RpsQ candidate gene. Because of the complexity of disease resistance mechanisms, the activities of the RpsQ candidate gene that affect disease resistance require further characterization.
The use of Rps-containing soybean cultivars is the most effective and environmentally friendly strategy for managing PRR (Walters 2009). During the development of disease-resistant cultivars using marker-assisted selection, several genes with complementary disease resistance activities should be incorporated into an elite cultivar to delay the breakdown of resistance (Dorrance and Schmitthenner 2000; Pathan and Sleper 2008; Saghai Maroof et al. 2008). In this study, the Chinese soybean cultivar Qichadou 1 was resistant to 30 P. sojae isolates, and the presence of RpsQ made it a good resource for breeding new soybean cultivars. Moreover, three co-segregated markers specific for RpsQ were developed and validated in the mapping population. These results are relevant for the introgression of RpsQ into new commercial cultivars through marker-assisted selection or Rps gene pyramiding. Future studies will focus on the characterization of RpsQ activities during the interaction between soybean plants and P. sojae.
Author contribution statement
YL and ZZ conceived the study and designed the experiments. ZZ developed the genetic populations. YL, CZ and XW performed research. YL, SS, CZ, XW and ZZ analyzed data. YL, SS and ZZ organized the figures and prepared the manuscript.
References
Abney TS, Melgar JC, Richards TL, Scott DH, Grogan J, Young J (1997) New races of Phytophthora sojae with Rps1–d virulence. Plant Dis 81:653–655
Afzal AJ, Wood AJ, Lightfoot DA (2008) Plant receptor–like serine threonine kinases: roles in signaling and plant defense. Mol Plant Microbe In 21:507–517
Anderson TR, Buzzell RI (1992) Inheritance and linkage of the Rps7 gene for resistance to Phytophthora rot of soybean. Plant Dis 76:958–959
Bai JF et al (2002) Diversity in nucleotide binding site–leucine–rich repeat genes in cereals. Genome Res 12:1871–1884
Bernard RL, Smith PE, Kaufmann MJ, Schmitthenner AF (1957) Inheritance of resistance to Phytophthora root rot and stem rot in the soybean. Agron J 49:391
Braun DM, Walker JC (1996) Plant transmembrane receptors: new pieces in the signaling puzzle. Trends Biochem Sci 21:70–73
Buzzell RI, Anderson TR (1992) Inheritance and race reaction of a new soybean Rps1 allele. Plant Dis 76:600–601
Chen QH, Wang QY, Wang YC, Zheng XB (2004) Identification and sequencing of ribosomal DNA-ITS of Phytophthora sojae in Fujian. Acta Phytopathol Sin 34:112–116
Cregan PB et al (1999) An integrated genetic linkage map of the soybean genome. Crop Sci 39:1464–1490
Cui L, Yin W, Tang Q, Dong S, Zheng X, Zhang Z, Wang Y (2010) Distribution, pathotypes, and metalaxyl sensitivity of Phytophthora sojae from Heilongjiang and Fujian provinces in China. Plant Dis 94:881–884
Dogimont C, Chovelon V, Pauquet J, Boualem A, Bendahmane A (2014) The Vat locus encodes for a CC-NBS-LRR protein that confers resistance to Aphis gossypii infestation and A. gossypii-mediated virus resistance. Plant J 80:993–1004
Dorrance AE, Grünwald NJ (2009) Phytophthora sojae: Diversity among and within populations. In: Lamour K, Kamoun S (eds) Oomycete genetics and genomics: diversity, interactions, and research tools. J. Wiley & Sons, New Jersey, pp 197–212
Dorrance AE, Schmitthenner AF (2000) New sources of resistance to Phytophthora sojae in the soybean plant introductions. Plant Dis 84:1303–1308
Dorrance AE, McClure SA, DeSilva A (2003) Pathogenetic diversity of Phytophthora sojae in Ohio soybean fields. Plant Dis 87:139–146
Ellis JG, Lawrence GJ, Luck JE, Dodds PN (1999) Identification of regions in alleles of the flax rust resistance gene L that determine differences in gene-for-gene specificity. Plant Cell 11:495–506
Fan AY, Wang XM, Fang XP, Wu XF, Zhu ZD (2009) Molecular identification of Phytophthora resistance gene in soybean cultivar Yudou 25. Acta Agron Sin 35:1844–1850
Förster H, Tyler BM, Coffey MD (1994) Phytophthora sojae races have arisen by clonal evolution and by rare outcrosses. MPMI 7:780–791
Gachomo EW, Shonukan OO, Kotchoni SO (2003) The molecular initiation and subsequent acquisition of disease resistance in plants. Afr J Biotechnol 2:26–32
Gao H, Bhattacharyya MK (2008) The soybean-Phytophthora resistance locus Rps1-k encompasses coiled coil-nucleotide binding-leucine rich repeat-like genes and repetitive sequences. BMC Plant Biol 8:29. doi:10.1186/1471-2229-8-29
Gómez–Gómez L, Boller T (2000) FLS2: an LRR receptor–like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
Gordon SG, St Martin SK, Dorrance AE (2006) Rps8 maps to a resistance gene rich region on soybean molecular linkage group F. Crop Sci 46:168–173
Graham MA, Marek LF, Shoemaker RC (2002) Organization, expression and evolution of a disease resistance gene cluster in soybean. Genetics 162:1961–1977
Grau CR, Dorrance AE, Bond J, Russin J (2004) Fungal diseases. In: Boerma HR, Specht JE (eds) Soybeans: improvement, production and uses, 3rd edn. Agronomy Monogr. American Soc Agron Madison WI, pp 679–763
Gunadi A (2012) Characterization of Rps8 and Rps3 resistance genes to Phytophthora sojae through genetic fine mapping and physical mapping of soybean chromosome 13. Dissertation, The Ohio State University
Haas JH, Buzzell RI (1976) New races 5 and 6 of Phytophthora megasperma var. sojae and differential reactions of soybean cultivars for races 1 and 6. Phytopathology 66:1361–1362
Heese A et al (2007) The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Pro Natl Aca Sci USA 104:12217–12222
Hulbert SH, Webb CA, Smith SM, Sun Q (2001) Resistance gene complexes: evolution and utilization. Annu Rev Phytopathol 39:285–312
Jee HJ, Kim WG, Cho WD (1998) Occurrence of Phytophthora root rot on soybean (Glycine max) and identification of the causal fungus. RDA J Crop Prot Korea Repub
Kaitany RC, Hart LP, Safir GR (2001) Virulence composition of Phytophthora sojae in Michigan. Plant Dis 85:1103–1106
Kajava AV, Kobe B (2002) Assessment of the ability to model proteins with leucine-rich repeats in light of the latest structural information. Protein Sci 11:1082–1090
Kasuga T, Salimath SS, Shi J, Gijzen M, Buzzell RI, Bhattacharyya MK (1997) High resolution genetic and physical mapping of molecular markers linked to the Phytophthora resistance gene Rps1-k in soybean. MPMI 10:1035–1044
Kaufmann MJ, Gerdemann JW (1958) Root and stem rot of soybean caused by Phytophthora sojae n. sp. Phytopathology 48:201–208
Lee S, Mian R, McHale LK, Wang H, Wijeratne AJ, Sneller CH, Dorrance AE (2013) Novel quantitative trait loci for partial resistance to Phytophthora sojae in soybean PI 398841. Theor Appl Genet 126:1121–1132
Li XP et al (2006) Identification and functional characterization of a leucine-rich repeat receptor-like kinase gene that is involved in regulation of soybean leaf senescence. Plant Mol Biol 61:829–844
Lin F et al (2013) Molecular mapping of two genes conferring resistance to Phytophthora sojae in a soybean landrace PI 567139B. Theor Appl Genet 126:2177–2185
Lincoln SE, Daly MJ, Lander ES (1993) Construction of a genetic linkage map with MAPMAKER/EXP v3.0: a tutorial and reference manual. An whitehead institute technical report, Cambridge
Liu JL, Liu XL, Dai LY, Wang GL (2007) Recent progress in elucidating the structure, function and evolution of disease resistance genes in plants. J Genet Genom 34:765–776
Michelmore RW, Paran I, Kessell RV (1991) Identification of markers linked to disease-resistance genes by bulked segregate analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. P Natl Acad Sci USA 88:9828–9832
Morillo SA, Tax FE (2006) Functional analysis of receptor-like kinases in monocots and dicots. Curr Opin Plant Biol 9:460–469
Mueller EH, Athow KL, Laviolette FA (1978) Inheritance of resistance to four physiologic races of Phytophthora megasperma var. sojae. Phytopathology 68:1318–1322
Ortega MA, Dorrance AE (2011) Microsporogenesis of Rps8/rps8 heterozygous soybean lines. Euphytica 181:77–88
Parlevliet JE (2002) Durability of resistance against fungal, bacterial and viral pathogens; present situation. Euphytica 124:147–156
Pathan MS, Sleper DA (2008) Advances in soybean breeding. In: Stacey G (ed) Genetics and genomics of soybean. Springer, New York (NY), pp 113–134
Ping J et al (2015) Identification and molecular mapping of Rps11, a novel gene conferring resistance to Phytophthora sojae in soybean. Theor Appl Genet 1–7
Qiu BX, Sleper DA, Arelli AR (1997) Genetic and molecular characterization of resistance to Heterodera glycines race isolates 1, 3, and 5 in Peking. Euphytica 96:225–231
SaghaiMaroof MA, Tucker DM, Tolin SA (2008) Genomics of viral–soybean interactions. In: Stacey G (ed) Genetics and genomics of soybean. Springer Science and Business Media, New York, pp 293–319
Sandhu D, Gao H, Cianzio S, Bhattacharyya MK (2004) Deletion of a disease resistance nucleotide-binding-site leucine-rich-repeat-like sequence is associated with the loss of the Phytophthora resistance gene Rps4 in soybean. Genetics 168:2157–2167
Schmitthenner AF (1985) Problems and progress in control of Phytophthora root rot of soybean. Plant Dis 69:362–368
Schmitthenner AF (1999) Phytophthora rot of soybean. In: Hartman GL, Sinclair JB, Rupe JC (eds) Compendium of soybean diseases, 4th edn. The American Phytopathological Society Press (APS), St Paul, Minnesota, pp 39–42
Shen CY, Su YC (1991) Discovery and preliminary studies of Phytophthora megasperma on soybean in China. Acta Phytopathol Sin 21:298
Sidhu GS, Khush GS (1978) Dominance reversal of a bacterial blight resistance gene in some rice cultivars. Phytopathology 68:461–463
Song WY et al (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–1806
Song Q et al (2010) Abundance of SSR motifs and development of candidate polymorphic SSR markers (BArCSOYSSr_1.0) in soybean. Crop Sci 50:1950–1960
Sugimoto T et al (2011) Genetic analysis and identification of DNA markers linked to a novel Phytophthora sojae resistance gene in the Japanese soybean cultivar Waseshiroge. Euphytica 182:133–145
Sugimoto T et al (2012) Pathogenic diversity of Phytophthora sojae and breeding strategies to develop Phytophthora-resistant soybeans. Breed Sci 61:511–522
Sun S, Wu XL, Zhao JM, Wang YC, Tang QH, Yu DY, Gai JY, Xing H (2011) Characterization and mapping of RpsYu25, a novel resistance gene to Phytophthora sojae. Plant Breed 130:139–143
Sun J, Li L, Zhao J, Huang J, Yan Q, Xing H, Guo N (2014) Genetic analysis and fine mapping of RpsJS, a novel resistance gene to Phytophthora sojae in soybean [Glycine max (L.) Merr.]. Theor Appl Genet 127:913–919
Tyler BM (2007) Phytophthora sojae: root rot pathogen of soybean and model oomycete. Mol Plant Pathol 8:1–8
Walters D (2009) Disease control kin crops: biological and environmentally-friendly approaches. Blackwell Publishing, Chichester
Wang Q et al (2011) Transcriptional programming and functional interactions within the Phytophthora sojae RXLR effector repertoire. Plant Cell 23:2064–2086
Weng C, Yu K, Anderson TR, Poysa V (2001) Mapping genes conferring resistance to Phytophthora root rot of soybean, Rps1a and Rps7. J Hered 92:442–446
Wrather JA, Koenning SR (2006) Estimates of disease effects on soybean yields in the United States 2003 to 2005. J Nematol 38:173
Wu XL et al (2011) Identification, genetic analysis and mapping of resistance to Phytophthora sojae of Pm28 in soybean. Agr Sci China 10:1506–1511
Xia CJ, Zhang JQ, Wang XM, Wu XF, Liu ZX, Zhu ZD (2011a) Analyses of resistance genes to Phytophthora root rot in soybean germplasm. Chin J Oil Crop Sci 33:396–401
Xia CJ, Zhang JQ, Wang XM, Liu ZX, Zhu ZD (2011b) Analysis of genes resistance to Phytophthora root rot in soybean germplasm imported from America. Acta Agron Sin 37:1167–1174
Xiang Y, Cao Y, Xu C, Li X, Wang S (2006) Xa3, conferring resistance for rice bacterial blight and encoding a receptor kinase-like protein, is the same as Xa26. Theor Appl Genet 113(7):1347–1355
Xue AG, Marchand G, Chen Y, Zhang S, Cober ER, Tenuta A (2015) Races of Phytophthora sojae in Ontario, Canada, 2010–2012. Can J Plant Pathol 37:376–383
Yao HY, Wang XM, Wu XF, Xiao YN, Zhu ZD (2010) Molecular mapping of Phytophthora resistance gene in soybean cultivar Zaoshu18. J Plant Genet Resour 11:213–217
Yu AL et al (2010) Genetic analysis and SSR mapping of gene resistance to Phytophthora sojae race 1 in soybean cv Suinong 10. Chin J Oil Crop Sci 32:462–466
Zhang S et al (2010) Races of Phytophthora sojae and their virulences on soybean cultivars in Heilongjiang, China. Plant Dis 94:87–91
Zhang J, Xia C, Duan C, Sun S, Wang X, Wu X, Zhu Z (2013a) Identification and candidate gene analysis of a novel Phytophthora resistance gene Rps10 in a Chinese soybean cultivar. PloS One 8:e69799
Zhang J, Xia C, Wang X, Duan C, Sun S, Wu X, Zhu Z (2013b) Genetic characterization and fine mapping of the novel Phytophthora resistance gene in a Chinese soybean cultivar. Theor Appl Genet 126:1555–1561
Zhong C, Li YP, Sun SL, Liu ZX, Qiu LJ, Zhu ZD (2015) Identification of resistance and tolerance to Phytophthora sojae in wild soybean germplasm. J Plant Genet Resour 16:684–690
Zhu ZD, Wang HB, Wang XM, Chang RZ, Wu XF (2003) Distribution and virulence diversity of Phytophthora sojae in China. Sci Agric Sin 36:793–799
Zhu ZD, Huo YL, Wang XM, Huang JB, Wu XF (2006) Screening for resistance sources to Phytophthora root rot in soybean. J Plant Genet Resour 7:24–30
Zhu ZD, Huo YL, Wang XM, Huang JB, Wu XF (2007) Molecular identification of a novel Phytophthora resistance gene in soybean. Acta Agron Sin 33:154–157
Acknowledgements
The work was supported by the Special Fund for Agroscientific Research in the Public Interest (201303018), the Program of Protection of Crop Germplasm Resources (2015NWB030-14) from the Ministry of Agriculture of the People’s Republic of China, and the Scientific Innovation Program of Chinese Academy of Agricultural Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The experiments were performed in accordance with all relevant Chinese laws.
Additional information
Communicated by Henry T. Nguyen.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Y., Sun, S., Zhong, C. et al. Genetic mapping and development of co-segregating markers of RpsQ, which provides resistance to Phytophthora sojae in soybean. Theor Appl Genet 130, 1223–1233 (2017). https://doi.org/10.1007/s00122-017-2883-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2883-7