Abstract
Key message
Loci associated with variation in maize responses to two microbe-associated molecular patterns (MAMPs) were identified. MAMP responses were correlated. No relationship between MAMP responses and quantitative disease resistance was identified.
Abstract
Microbe-associated molecular patterns (MAMPs) are highly conserved molecules commonly found in microbes which can be recognized by plant pattern recognition receptors. Recognition triggers a suite of responses including production of reactive oxygen species (ROS) and nitric oxide (NO) and expression changes of defense-related genes. In this study, we used two well-studied MAMPs (flg22 and chitooctaose) to challenge different maize lines to determine whether there was variation in the level of responses to these MAMPs, to dissect the genetic basis underlying that variation and to understand the relationship between MAMP response and quantitative disease resistance (QDR). Naturally occurring quantitative variation in ROS, NO production, and defense genes expression levels triggered by MAMPs was observed. A major quantitative traits locus (QTL) associated with variation in the ROS production response to both flg22 and chitooctaose was identified on chromosome 2 in a recombinant inbred line (RIL) population derived from the maize inbred lines B73 and CML228. Minor QTL associated with variation in the flg22 ROS response was identified on chromosomes 1 and 4. Comparison of these results with data previously obtained for variation in QDR and the defense response in the same RIL population did not provide any evidence for a common genetic basis controlling variation in these traits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbe- or pathogen-associated molecular patterns (MAMPs or PAMPs) are highly conserved molecules, found in certain classes of microbes, that are usually essential for the microbe’s life cycle, fitness, and survival but are not specifically associated with pathogenicity (Segonzac and Zipfel 2011; Thomma et al. 2011; Choi and Klessing 2016). MAMPs are recognized by pattern recognition receptors (PRRs) at the plant cell surface, which then trigger a defense response that, in many cases, will confer a type of resistance known as MAMP- (or PAMP-) triggered immunity (MTI or PTI, Couto and Zipfel 2016). Well-studied MAMPs include epitopes of flagellin (flg22 and flgII-28), peptidoglycans (PGNs), lipopolysaccharides (LPSs), chitin, and translation elongation factor Tu (EF-Tu; Felix et al. 1993, 1999; Dow et al. 2000; Kunze et al. 2004; Gust et al. 2007; Cai et al. 2011; Clarke et al. 2013). A similar response is also activated by endogenous elicitors, called damage-associated molecular patterns (DAMPs), which are released into the extracellular space after a mechanical damage (Huffaker et al. 2011; Huffaker and Ryan 2007; Lotze et al. 2007; Pearce et al. 2010; Tanaka et al. 2014; Choi and Klessing 2016). MTI is considered to be a major cause of “non-host resistance in plants, the phenomenon whereby a plant–pathogen species is unable to cause disease on any member of a plant species” (Lee et al. 2016).
In cases in which a pathogen is adapted to a specific host, effector molecules secreted by the pathogen often allow the pathogen to suppress or overcome the host MTI response. A second level of disease resistance called effector-triggered immunity (ETI) often occurs in these cases in which resistance (R-) proteins detect specific effectors and trigger a response that is qualitatively similar to though quantitatively greater than PTI (Jones and Dangl 2006; Boller and Felix 2009; Dodds and Rathjen 2010). In many cases, a “hypersensitive response” (HR), in which rapid cell death occurs at the point of attempted pathogen penetration, is associated with ETI.
In plants, most known PRRs are plasma membrane-localized receptor-like kinases (RLKs) or receptor-like proteins (RLPs; Monaghan and Zipfel 2012). RLKs and RLPs both have extracellular and transmembrane domains. RLKs additionally possess a cytoplasmic kinase domain, while RLPs have a short cytoplasmic domain without obvious function. The Arabidopsis FLS2 (flagellin sensitive 2) gene encodes an RLK with an extracellular leucine-rich repeat (LRR) domain and a cytoplasmic serine/threonine kinase domain, which perceives of the flagellin epitope flg22 (Gómez-Gómez and Boller 2000; Chinchilla et al. 2006). The tomato FLS3 (flagellin-sensing 3) gene encodes another LRR-RLK that recognizes flgII-28, another epitope of flagellin. BAK1 (brassinosteroid insensitive 1-associated receptor kinase 1), another LRR-RLK, interacts with FLS2 and acts as a co-receptor (Sun et al. 2013). BAK1 also interacts with several other PRRs and is important for downstream signaling (Chinchilla et al. 2007; Heese et al. 2007). The rice OsCEBiP (chitin elicitor binding protein) and OsCERK1 genes encode a cell surface RLP and RLK respectively each of which carry extracellular lysin motifs (LysM). These two PRR proteins form a heterodimer capable of binding and triggering PTI in response to chitin (Shimizu et al. 2010). Chitin recognition in Arabidopsis appears to be somewhat different from the case in rice. The current model is that AtCERK1 interacts with AtLYK5, likely forming a heterotetramer, with AtLYK5 serving as the primary binding site for chitooligosaccharide but signaling through the active AtCERK1 kinase domain (Cao et al. 2014). Signaling downstream of the activated PRRs during the MAMP response is relatively well characterized and involves, in part, cascades of mitogen-activated protein kinase (MAP kinase) phosphorylation, leading to the phosphorylation and activation of sets of transcription factors (Bigeard et al. 2015).
The MAMP response includes changes in ion flux across the plasma membrane (Jabs et al. 1997; Pugin et al. 1997; Wendehenne et al. 2002; Mithöfer et al. 2005), production of reactive oxygen species (ROS; Felix et al. 1999; Segonzac et al. 2011; Lloyd et al. 2014), production of nitric oxide (NO; Lamotte et al. 2004; Foissner et al. 2000; Laxalt et al. 2007; Rasul et al. 2012), callose deposition (Brown et al. 1998; Millet et al. 2010; Luna et al. 2011), modification of phytohormone concentrations (Tsuda et al. 2008; Halim et al. 2009; Furch et al. 2014), and induction or repression of different plant defense-related genes (Nishizawa et al. 1999; Habib and Fazili 2007; Valdés-López et al. 2011; Szatmári et al. 2014). Quantitative measurements of one or more of these aspects of the response are usually used as a way of assessing the response strength. Growth inhibition due to prolonged activation of the MAMP response is also sometimes used as a proxy for the response (Gómez-Gómez et al. 1999). It seems clear that the response to different MAMPs can be qualitatively different (Vetter et al. 2016).
Several reports have demonstrated quantitative variation in the MAMP response across different lines within a species, and in some cases, quantitative trait loci (QTL) associated with this variation have been identified. Valdés-López et al. (2011) showed variation in the response to a mixture of flg22 and chitin among soybean genotypes and identified QTLs associated with total ROS production and expression (e)QTLs associated with expression variation of MAMP-responsive genes. Ahmad et al. (2011) identified two loci associated with flagellin-induced callose deposition in Arabidopsis thaliana. Vetter et al. (2012) reported extensive variation between different A. thaliana ecotypes in flg22 binding and in flg22-induced seedling growth inhibition. They also noted that these two traits were highly correlated. Vetter et al. (2016) found that growth inhibition induced by both flg22 and elf18 (an epitope of the MAMP EF-Tu) varied among genotypes and that the response to each was governed by a complex genetic architecture involving loci with generally small effects. They also found that the responses to the two MAMPs were largely uncorrelated and that the genetic architectures controlling them were likewise dissimilar. Veluchamy et al. (2014) investigated the responses of 13 heirloom tomato lines to three MAMPs: flg22, flgII-28, and csp22 (an active epitope from the bacterial cold shock protein). They showed that there was a significant variation among the lines for the response induced by each MAMP, though there was no significant correlation between the response induced by each MAMP and resistance to bacterial speck disease, caused by the bacterial pathogen Pseudomonas syringae. They also provided evidence for a single locus associated with variation in csp22 response.
The MAMP response in maize has not been characterized and no maize PRRs have been cloned. However, a maize DAMP, ZmPep1, has been shown to induce the defense response and to enhance disease resistance (Huffaker et al. 2011). The ZmPep1 receptor, a LRR-RLK called ZmPEPR1, has also been cloned (Lori et al. 2015).
Cultivating resistant varieties is one of the most important goals of most plant breeders. Quantitative disease resistance (QDR), in most cases the most durable form of resistance (Brown 2015), is usually conditioned by several genes with relatively small effects. Our understanding of the relationship between QDR and the MAMP response is very limited. In just a few cases, a link has been established or suggested. For example, two major QTLs associated with basal resistance to P. syringae pv. phaseolicola were reported in Arabidopsis, one of which encompasses FLS2 (Forsyth et al. 2010). In maize, we, and others, have identified several LRR-RLKS and LRR-RLPs as candidate genes for quantitative disease resistance (Kump et al. 2010, 2011; Poland et al. 2011). Probably, the best evidence for a link between QDR and the MAMP response comes from a study in which molecular variation in a PRR was shown to be associated with quantitative resistance to Fusarium oxysporum in Arabidopsis (Cole and Diener 2013).
As well as being among the most important crops on the planet, maize is a model organism for genetic studies. The goals of this study were to investigate intra-species variation in the MAMP response in maize, to identify QTL associated with his variation, and to determine whether there was any evidence for a connection between variation in the MAMP response and variation in QDR and other defense-related traits.
Materials and methods
Plant materials and growth condition
The maize nested association mapping (NAM) population is a meta-population of 5000 recombinant inbred lines made up of 25 sub-populations of 200 recombinant inbred lines (RILs) each. Each of these sub-populations is derived from a cross between the commonly-used maize line B73 and 25 different diverse lines (McMullen et al. 2009). The 26 NAM parental lines and 174 lines of the B73 × CML228 sub-population were used for investigating natural variation of responses to MAMPs.
For the initial studies on the transcriptional and NO response, all lines were grown in growth chambers at constant 27 °C on 16/8-h light/dark cycle. For all the ROS experiments, lines were grown in growth chambers at the NCSU Phytotron with a 16/8-h light/dark cycle at 25/18 °C. Six seeds were sown in each pot (diameter 15.2 cm, volume 1.65 L) containing standard substrates [33% Sunshine Redi-Earth Pro Growing Mix (Canadian Sphagnum peat moss 50–65%, vermiculite, dolomitic lime, 0.0001% silicon dioxide) and 66% pea gravel]. After germination, seedlings were removed until two seedlings remained in each pot.
MAMP elicitors
Two MAMPs used in this study were flg22 and chitooctaose. flg22 is a peptide corresponding to a conserved domain of bacterial flagellin and was ordered from Genscript (catalog# RP19986). Chitooctaose is a degradation product of chitin and soluble in water, and was ordered from Accurate Chemical and Scientific Corporation (Catalog # BCR57120010).
Gene expression assays using real-time quantitative reverse transcription PCR (qRT-PCR)
We assessed the expression of four established defense marker genes including endochitinase A (ECA), peroxidase 3 (PEX3), pathogenesis-related protein 1 (PR1), and endochitinase PR4 (ECPR4) genes (Huffaker et al. 2011; Table 1) following elicitation by MAMPs. For each line assessed, five leaf discs of 2 cm2 from five different 15-day-old plants were treated with H2O (mock), 1-µM flg22, or chitooctaose (treatment). 30 min later; leaf discs were immediately put into liquid nitrogen and stored at −80 °C until use.
Total RNA was extracted from mock, flg22- and chitooctaose-treated leaves using Trizol reagent (Invitrogen) according to the manufacturer’s specifications. Genomic DNA (gDNA) was removed from purified RNA using TURBO DNAse (Thermo Fisher Scientific Inc.) according to the manufacturer’s instructions. 2 µg of gDNA-free RNA were used to synthesize cDNA.
qRT-PCR was performed as described in Libault et al. (2008) in a Applied Biosystem qPCR machine (95 °C 10 min, and 45 cycles of 95 °C 10 s, 60 °C 1 min) using the housekeeping gene Actin to normalize the expression levels of target genes. Primer design was performed as described in Libault et al. (2010). Expression levels of the analyzed genes were calculated according to the equation \(E\text{ }=\text{ }{{P}_{\text{eff}}}^{(-\Delta {{C}_{\text{t}}})}\), where P eff is the primer set efficiency calculated using LinRegPCR (Ramakers et al. 2003) and ΔC t was calculated by subtracting the cycle threshold (C t) value of the housekeeping gene from the C t values of the gene analyzed. Fold changes were calculated between the ratios of the expression levels of MAMP-treated and mock samples, and expression levels were calculated for three biological replicates.
Nitric oxide (NO) assay
Five leaf discs (~1 cm each disc) from the same plants as those used in the gene expression assays were incubated in water for 12 h under dark conditions. Water was carefully removed and the leaf sections were incubated with shaking under dark conditions in 2.5-µM DAR-4M [cell-impermeable NO binding dye diaminorhodamine-4M (EMD Chemicals Inc. catalo # 251765)] with either H2O (mock), 1-µM flg22, or 1-µM chitooctaose (treatment). 100 µl of the solution from each mock, treatment, and cultivar were transferred into a 48-well plate after 2 h of incubation. Fluorescence of the supernatant was measured by excitation at 560 nm and emission at 575 nm. Fluorescence signal was corrected for the DAR-4M background. Determinations were in triplicate.
ROS assay
Two leaf discs (diameter 3 mm) from two plants of each line were taken from 10-day-old maize seedlings and floated on 50-µl H2O in a 96-well plate overnight. 50 µl of reaction solution carrying 1-µl 2-mg/ml L-012 (a sensitive chemiluminescence probe, which reacts with superoxide anion and produces luminescence at long wavelengths; Wako Pure Chemical Industries, Ltd., catalog # 120–04891) in Dimethyl Sulphoxide (DMSO), 1 µl of 2-mg/ml horseradish peroxidase (Sigma–Aldrich catalog # P6782), and 48-µl 2-µM MAMPs, was added just before measurement on a Synergy™ 2 multi-detection microplate reader (BioTek). All wells were read within a minute of the MAMP being added. In every case, the luminescence was recorded over a 60-min period 31 times at 2-min intervals, and ROS production was calculated as the sum of 31 photon counts over this period.
For the investigation of variation of responses to MAMPs across NAM parental lines, each line was assessed in four wells of each 96-well plate with one well of mock (without MAMPs) and three wells of treatment (with MAMPs). The common parent line B73 was always included as repeated check. There were four wells of blank in each plate. Three biological replicates were performed in every case. Each replication required two 96-well plates.
The RILs of the B73 × CML228 mapping population were assessed in sets of 20 in 96-well plates. Each line had one well of mock (without MAMPs) and three wells of treatment (with MAMPs). Four wells were left blank in each plate. Two sets of four wells were used for the CML228 line (positive control) and one set of four for B73. Two and three biological replicates of responses to flg22 and chitooctaose were measured in the population respectively. The method of ROS assay used in this part was the same as above except that the concentration of MAMPs was to be tested.
Data analysis and QTL mapping
Data was analyzed with the SAS software for Windows version 9.4 (SAS Institute Inc, 2014). Average of observations over wells corresponding to a particular line within each plate and repetition was calculated using SAS® MEANS procedure. Average value of the mock wells per plate was subtracted from the line mean response before analysis as a normalization technique. Data were analyzed using GLIMMIX procedure of SAS/STAT software for Windows version 13.2 (SAS Institute Inc, 2014), following the augmented design model with line and Group (checks vs. lines) as fixed-effect factors; repetition, plates within repetition, and repetition by line within group interaction were considered as random effects, assuming a residual normal distribution. Significance level was set up to 0.05. Line least-square means (LSMEAN) and their 95% confidence interval were estimated for each MAMP. Adjusted LSMEANS of the B73 × CML228 sub-population estimated using the augmented design was used in correlation analysis and QTL mapping. Pearson correlations were calculated with the CORR procedure.
Due to the limitations of the software, a subset of half of the available 7386 SNP markers were used in this study (Olukolu et al. 2014). Alternate markers were removed from the data set resulting in a set of 3693 SNP markers at uniform 0.4-cM intervals were used. The QTL analysis was performed using Windows QTL Cartographer software v2.5 based on composite interval mapping (CIM) with a walk speed of 1.0 cM (Silva et al. 2012). 1000 permutation tests were performed to determine likelihood of odds ratio (LOD) threshold value at a significance level of 0.05 for each trait.
Results
MAMPs-triggered responses in maize
Initially, to gauge variation among the NAM parents and to optimize the assay conditions, seven parents of the maize NAM population (B73, CML52, CML333, Ky21, Ki11, Ki3, and IL14H), were used to investigate MAMP-induced expression of defense-related genes and six parents (B73, CML52, CML322, Ki11, Ki3, and IL14H) were used to investigate NO production. Each of the four genes (ECA, PEX3, PR1, and ECPR4) investigated was induced by treatment with both MAMPs (flg22, chitooctaose) in each line (Fig. 1). In most cases, variability in the levels of induction was observed between the lines. Different genes displayed different patterns of induction across the lines and the patterns of variation differed somewhat between flg22 and chitooctaose for at least two genes (ECA and ECPR4; Fig. 1a, b). ECA expression was more highly induced by chitooctaose in B73 compared with other lines (Fig. 1a). Chitooctaose triggered stronger responses of ECPR4 than flg22 in all lines except for Ki3 (no data were available for IL14H; Fig. 1b). PEX3 showed stronger responses to MAMPs in IL14H, while PR1 was strongly induced by both MAMPs in Ky21 (Fig. 1c, d).
Expression assays of defense-related genes across different maize lines triggered by MAMPs. a–d Expression levels of ECA, ECPR4, PEX3, PR1; ND no data. Results shown are means of the ratio between MAMPs/mock from three biological replicates with error bars representing the standard error of three biological replicates
There were also differences in NO production across these lines after treatment with either flg22 or chitooctaose (Fig. 2). These data indicated that MAMPs can trigger responses in maize and that the responses are variable across lines and dependent on the MAMP used and the specific response being measured.
Natural variation of responses to MAMPs across different NAM parental lines
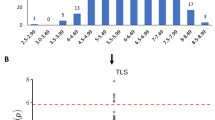
Total ROS production after treatment with flg22 and chitooctaose was assessed across the 26 NAM parental lines. Substantial variation was observed across these lines (Fig. 3). No response was detectable in several lines for each MAMP. The ROS response to flg22 and chitooctaose was significantly correlated across the NAM parental lines (Pearson correlation coefficient = 0.7, P < 0.0001), with CML228 having the strongest response to both flg22 and chitooctaose, while the common parent of the NAM population, B73, displayed a relatively low response to each MAMP. Figure 4 shows the detailed time kinetics of the MAMP-triggered oxidative burst over 60 min in line B73 and CML228. For CML228, the reaction peak of flg22 treatment appeared at about the 25 min time point, while that of chitooctaose treatment appeared at about the 10 min time point. Similar time kinetics was observed for the other lines in which responses were observed.
Responses of NAM parental lines to a flg22 and b chitooctaose. Data are expressed as total ROS production over 60 min; ROS production measured in relative light unit (RLU); results shown are means of three biological replicates with error bars representing the standard error; the treatment of each line was compared with that of B73; **P < 0.01; *P < 0.05
QTL analysis of MAMP responses
We performed QTL analysis of MAMP responses in the NAM RIL sub-population derived from the B73 × CML228 cross (population Z003, http://maizegdb.org/data_center/stock?id=9018763).
For our initial work, we used 2-μM concentrations of both MAMPs. However, before assessing the CML228 × B73 mapping population, we sought to determine the optimum concentrations of MAMPs to use for these lines. The flg22 response of CML228 was stronger than that of B73 at every concentration tested. The response increased with increasing concentration from 2 nM to 20 μM, but the difference between B73 and CML228 did not increase between 2 and 20 μM (Fig. 5). For chitooctaose, there was no significant difference in the response of CML228 between 2 and 20 μM. Therefore, we continued to use 2-μM concentrations of both MAMPs for subsequent work.
ROS production triggered by different concentrations of flg22 in line B73 and CML228. Data are expressed as total ROS production over 60 min; ROS production measured in relative light unit (RLU); results shown are means of three biological replicates with error bars representing the standard error of three biological replicates
We observed approximately normal distributions and substantial variation for the responses to each MAMP within the RIL population, including some transgressive segregation (Fig. 6). Correlations between replicates were moderate, especially for chitooctaose, but there were all highly significant (Table 2). ANOVA revealed significant differences (P < 0.0001) among lines and replicates, (Table 3). The responses to the 2 MAMPs were highly correlated within the CML228 × B73 RIL population (Pearson correlation coefficient = 0.76; Table 4).
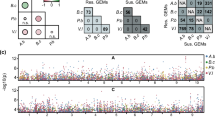
We used the adjusted LSMEAN values for each line to identify QTL for response to each MAMP. QTLs for flg22 and chitootaose responses were identified on chromosomes 1, 2, and 4 and on chromosome 2 respectively (Table 5). For the flg22 response, Qflg22-2 was a major QTL explaining 13.5% of the variation with CML228 contributing the allele for increased the response. The two other flg22-response QTL on chromosomes 1 and 4, Qflg22-1 and Qflg22-4, explained 6.5 and 4.9% of the variation, respectively. The allele for increased response also derived from CML228 for Qflg22-4 but from B73 for Qflg22-1. The two QTLs, the chitooctaose response, Qchitoo-2A, 2B, were close to each other on chromosome 2, and in each case, the CML228 allele increased the response (Table 5; Figure S1, S2). The two QTLs Qflg22-2 and Qchitoo-2A overlapped. Since we noted significant replication effects, we calculated QTL based on each replication separately (Figures S1, S2). We saw a consistent effect at the chromosome 2 QTL across reps for both flg22 and chitooctaose.
Correlation between MAMP responses and other disease and defense-related traits
We and others had previously assessed the CML228 × B73 population for resistance to the foliar diseases southern leaf blight, gray leaf spot, and northern leaf blight (SLB, GLS, and NLB respectively; Kump et al. 2011; Poland et al. 2011; Benson et al. 2015), for the strength of the HR-related traits (Olukolu et al. 2014) and for a leaf-flecking trait that was shown to be associated with disease resistance and ROS production (Olukolu et al. 2016). We compared the results of these previous studies with the data produced here to determine if there was any evidence for a shared genetic basis underlying variation in these responses. No significant correlations were observed between MAMP responses and any of the other traits except for weak correlations observed with SLB (Table 4). We also looked for overlap between QTL identified for these traits. Only one QTL, for HR severity, on chromosome 2 overlapped with any of the MAMP response QTL (Table 6).
Discussion
A variety of transcriptional and physiological responses that lead to MTI can be elicited by recognition of MAMPs by their corresponding PRRs. An interesting aspect is that regardless of the MAMP-PRR interaction, in general, a similar repertoire of genes involved in the defense response is induced (Wan et al. 2008). When flg22 was used to elicit Arabidopsis cell cultures, the differentially expressed genes were enriched for genes associated with signal transduction pathways such as transcription factors, and regulators of protein stability and phosphorylation (Navarro et al. 2004). Zipfel et al. (2004) reported that the expression of numerous defense-related genes was induced by flg22 in Arabidopsis. Transcripts induced in tomato by flgII-28 were similarly enriched for protein kinases and transcription factors (Rosli et al. 2013). Valdés-López et al. (2011) examined the transcriptional response to a mixture of chitin and flg22 in four soybean genotypes. Among the induced genes, six functional categories were overrepresented: regulation, protein modification, regulation of transcription, hormones, enzyme families, and transport. Considerable variation in the transcriptional profiles of MAMP-responsive genes across different soybean genotypes was also observed in this study.
Quantification of ROS and NO production has also been used as a method for measuring the strength of the MAMP response. Superoxide anions (O2 −), hydrogen peroxide (H2O2), and hydroxy radicals (.OH) are the main ROS, all of which can cause oxidative damage of DNA, proteins or lipids. ROS also play an integral role as signaling molecules in the regulation of numerous biological processes such as growth, development, and responses to biotic and/or abiotic stimuli in plants. NO is involved together with ROS in the activation of various stress responses in plants (Wojtaszek 1997; Ahlfors et al. 2009; Baxter et al. 2014; Schieber and Chandel 2014; Vidhyasekaran 2014, 2016).
In this study, we assessed the expression of four defense-related genes, NO and ROS production after treatment with two well-studied MAMPs, flg22 and chitooctaose, associated with bacteria and fungi, respectively. All of these different measurements identified differential responses to flg22 and chitooctaose across various maize genotypes. Differentials between lines can be exploited to identify genetic loci that underlie variation in these basal resistance responses. We chose to map QTL for the ROS response, simply because this was the most convenient metric to measure.
Several studies identified natural variation in the MAMP response in plants (Ahmad et al. 2011; Valdés-López et al. 2011; Vetter et al. 2012; Lloyd et al. 2014; Shi et al. 2015; Veluchamy et al. 2014) and in some cases mapped QTL associated with that variation (Ahmad et al. 2011; Valdés-López et al. 2011). Recently, genome-wide association (GWA) mapping has also been used in the dissection of the genetic basis underlying natural variation in MAMP-induced seedling growth inhibition (SGI) by the MAMPS flagellin and elf18 (Vetter et al. 2016). Variation in the MAMP response has not been examined previously in maize, though some aspects of both this response and the related DAMP response have been addressed in the previous studies (Huffaker et al. 2011; Oliveira-Garcia and Deising 2016).
We performed QTL analysis of ROS production triggered by flg22 and chitooctaose. One major QTL for flg22 response (Qflg22-2) was identified on chromosome 2, which overlapped with a QTL for the chitooctaose response (Qchitoo-2A). Two minor QTLs for flg22 responses dispersed on chromosomes 1 and 4. Genes located in the overlapping region of these two QTLs are listed in Table S1. Several of these genes are homologous to genes previous implicated in plant disease resistance or the defense response, including RLKs and toll/interleukin-1 receptor (TIR) domains-containing proteins (Nandety et al. 2013). The colocalization of these QTL together with the large and significant correlations between the flg22 and chitooctaose responses among the 26 NAM parental lines and within the CML228 × B73 RIL population suggest that, to a substantial extent, the genetic architectures controlling variation in these traits are shared. This finding contrasts with a recent study in Arabidopsis (Vetter et al. 2016) which identified “negligible correlation” in plant growth responses between the bacterial MAMPs EF-Tu and flagellin. As discussed above, five other metrics were used to measure the response to flg22 and chitoctaose, the expression of four genes, and NO production (Figs. 1, 2). As only six or seven lines were assessed for these responses, no firm conclusions can be drawn. However, it seems that, as with the ROS response, the responses to flg22 and chitooctaose were correlated within a metric, but that there does not seem to be much if any correlation across metrics. In other words, the apparent strength of the MAMP response may partly be a function of how exactly it is measured. Similar phenomena have been observed in Brassica napus (Lloyd et al. 2014).
We were further interested to examine whether variation in the MAMP response might be one of the causes of variation in QDR. QDR is usually controlled by multiple genes with small and sometimes inconsistent effects and tends to be more durable than major (R-) gene-mediated qualitative resistance (Poland et al. 2009). The NAM population, of which the CML228 × B73 RIL population used in this study is a part, has been used to dissect the genetic basis of variation in QDR to three maize diseases (SLB, NLB, GLS) and in the HR defense response (Kump et al. 2011; Poland et al. 2011; Benson et al. 2015; Olukolu et al. 2014). A leaf-flecking trait has also been mapped in the NAM population, and connections between leaf flecking, disease resistance, and ROS production were reported (Olukolu et al. 2016). There was no significant correlation between responses triggered by MAMPs and any of these other disease or defense-related traits, with the exception of a low correlations with SLB resistance (Table 4) and there was very little colocalization of QTL between these traits (Table 6). All the diseases for which we have data are caused by fungi. While chitooctase is a fungal MAMP, flg22 is an epitode of flagellin—a molecule found in bacteria. However, the response to neither is related substantially to variation in QDR. A study with various citrus genotypes suggested a link between variation in response to flg22 and resistance to citrus canker (Shi et al. 2015), while a study in tomato did not identify any link between variation in the MAMP response and resistance to bacterial speck (Veluchamy et al. 2014).
We can conclude, therefore, that maize is able to respond to the MAMPs flg22 and chitooctaose, that there is variation among different maize genotypes for this response and that the genetic basis of variation in these two responses, at least with respect to ROS production, is substantially shared in the populations we examined. However, our studies provide no evidence linking variation in the MAMP response to variation in QDR or other aspects of disease resistance.
Pathogens that are well adapted to their host have evolved multiple means by which to suppress MTI. Furthermore, as discussed above, the MAMP response is multifaceted and a study that measures only a single aspect may not provide a full reflection of what is occurring. These factors may explain the lack correspondence observed between the MAMP response and QDR and underline the fact that without substantially more work, these findings cannot be extrapolated beyond the population examined. Indeed, as noted above, other studies in other species have reached somewhat contrasting conclusions (Forsyth et al. 2010; Cole and Diener 2013; Shi et al. 2015). However, it is clear that the link between the MAMP response and quantitative disease resistance, as well as non-host resistance, is a topic deserving of further research.
Author contribution statement
PBK, GS, XZ and OVL planned the research, XZ and OVL performed the research and analyzed the data, CA performed statistical analyses, XZ and PBK wrote the manuscript, and GA, OVL, and CA edited the manuscript.
References
Ahlfors R, Brosché M, Kangasjärvi J (2009) Ozone and nitric oxide interaction in Arabidopsis thaliana: a role for ethylene? Plant Signal Behav 4:878–879
Ahmad S, Van Hulten M, Martin J, Pieterse CMJ, Van Wees SCM, Ton J (2011) Genetic dissection of basal defence responsiveness in accessions of Arabidopsis thaliana. Plant Cell Environ 34:1191–1206
Baxter A, Mittler R, Suzuki N (2014) ROS as key players in plant stress signaling. J Exp Bot 65:1229–1240
Benson JM, Poland JA, Benson BM, Stromberg EL, Nelson RJ (2015) Resistance to gray leaf spot of maize: genetic architecture and mechanisms elucidated through nested association mapping and near-isogenic line analysis. PLoS Genet 11:e1005045
Bigeard J, Colcombet J, Hirt H (2015) Signaling mechanisms in pattern-triggered immunity (PTI). Mol Plant 8:521–539
Boller T, Felix G (2009) A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu Rev Plant Biol 60:379–406
Brown JKM (2015) Durable resistance of crops to disease: a Darwinian perspective. Annu Rev Phytopathol 53:513–539
Brown I, Trethowan J, Kerry M, Mansfield J, Bolwell GP (1998) Localization of components of the oxidative cross-linking of glycoproteins and of callose synthesis in papillae formed during the interaction between non-pathogenic strains of Xanthomonas campestris and French bean mesophyll cells. Plant J 15:333–343
Cai R, Lewis J, Yan S, Liu H, Clarke CR, Campanile F, Almeida NF, Studholme DJ, Lindeberg M, Schneider D, Zaccardelli M, Setubal JC, Morales-Lizcano NP, Bernal A, Coaker G, Baker C, Bender CL, Leman S, Vinatzer BA (2011) The plant pathogen Pseudomonas syringae pv. tomato is genetically monomorphic and under strong selection to evade tomato immunity. PLoS Pathog 7:e1002130
Cao Y, Liang Y, Tanaka K, Nguyen CT, Jedrzejczak RP, Joachimiak A, Stacey G (2014) The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. eLife 3:e03766
Chinchilla D, Bauer Z, Regenass M, Boller T, Felix G (2006) The Arabidopsis receptor kinase FLS2 binds flg22 and determines the specificity of flagellin perception. Plant Cell 18:465–476
Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nürnberger T, Jones JDG, Felix G, Boller T (2007) A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448:497–500
Choi HW, Klessig DF (2016) DAMPs, MAMPs, and NAMPs in plant innate immunity. BMC Plant Biol 16:232
Clarke CR, Chinchilla D, Hind SR, Taguchi F, Miki R, Ichinose Y, Martin GB, Leman S, Felix G, Vinatzer BA (2013) Allelic variation in two distinct Pseudomonas syringae flagellin epitopes modulates the strength of plant immune responses but not bacterial motility. New Phytol 200:847–860
Cole SJ, Diener AC (2013) Diversity in receptor-like kinase genes is a major determinant of quantitative resistance to Fusarium oxysporum f.sp. matthioli. New Phytol 200:172–184
Couto D, Zipfel C (2016) Regulation of pattern recognition receptor signalling in plants. Nat Rev Immunol 16:537–552
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant-pathogen interactions. Nat Rev Genet 11:539–548
Dow M, Newman MA, von Roepenack E (2000) The induction and modulation of plant defense responses by bacterial lipopolysaccharides. Annu Rev Phytopathol 38:241–261
Felix G, Duran JD, Volko S, Boller T (1999) Plants have a sensitive perception system for the most conserved domain of bacterial flT (1999. Plant J 18:265–276
Felix G, Regenass M, Boller T (1993) Specific perception of subnanomolar concentrations of chitin fragments by tomato cells: induction of extracellular alkalinization, changes in protein phosphorylation, and establishment of a refractory state. Plant J 4:307–316
Foissner I, Wendehenne D, Langebartels C, Durner J (2000) In vivo imaging of an elicitor-induced nitric oxide burst in tobacco. Plant J 23:817–824
Forsyth A, Mansfield JW, Grabov N, de Torres M, Sinapidou E, Grant MR (2010) Genetic dissection of basal resistance to Pseudomonas syringae pv. phaseolicola in accessions of Arabidopsis. Mol Plant Microbe Interact 23:1545–1552
Furch ACU, Zimmermann MR, Kogel KH, Reichelt M, Mithöfer A (2014) Direct and individual analysis of stress-related phytohormone dispersion in the vascular system of Cucurbita maxima after flagellin22 treatment. New Phytol 201:1176–1182
Gust AA, Biswas R, Lenz HD, Rauhut T, Ranf S, Kemmerling B, Götz F, Glawischnig E, Lee J, Felix G, Nürnberger T (2007) Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J Biol Chem 282:32338–32348
Gómez-Gómez L, Boller T (2000) FLS2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
Gómez-Gómez L, Felix G, Boller T (1999) A single locus determines sensitivity to bacterial flagellin in Arabidopsis thaliana. Plant J 18:277–284
Habib H, Fazili KM (2007) Plant protease inhibitors: a defense strategy in plants. Biotechnol Mol Biol Rev 2:068–085
Halim VA, Altmann S, Ellinger D, Eschen-Lippold L, Miersch O, Scheel D, Rosahl S (2009) PAMP-induced defense responses in potato require both salicylic acid and jasmonic acid. Plant J 57:230–242
Heese A, Hann DR, Gimenez-Ibanez S, Jones AME, He K, Li J, Schroeder JI, Peck SC, Rathjen JP (2007) The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc Natl Acad Sci USA 104:12217–12222
Huffaker A, Dafoe NJ, Schmelz EA (2011) ZmPep1, an ortholog of Arabidopsis elicitor peptide 1, regulates maize innate immunity and enhances disease resistance. Plant Physiol 155:1325–1338
Huffaker A, Ryan CA (2007) Endogenous peptide defense signals in Arabidopsis differentially amplify signaling for the innate immune response. Proc Natl Acad Sci USA 104:10732–10736
Jabs T, Tschöpe M, Colling C, Hahlbrock K, Scheel D (1997) Elicitor-stimulated ion fluxes and O2 – from the oxidative burst are essential components in triggering defense gene activation and phytoalexin synthesis in parsley. Proc Natl Acad Sci USA 94:4800–4805
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Kump KL, Bradbury PJ, Wisser RJ, Buckler ES, Belcher AR, Oropeza-Rosas MA, Zwonitzer JC, Kresovich S, McMullen MD, Ware D, Balint-Kurti PJ, Holland JB (2011) Genome-wide association study of quantitative resistance to southern leaf blight in the maize nested association mapping population. Nat Genet 43:163–168
Kump KL, Holland JB, Jung MT, Wolters P, Balint-Kurti PJ (2010) Joint analysis of near-isogenic and recombinant inbred line populations yields precise positional estimates for quantitative trait loci. Plant Gen 3:142–153
Kunze G, Zipfel C, Robatzek S, Niehaus K, Boller T, Felix G (2004) The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 16:3496–3507
Lamotte O, Gould K, Lecourieux D, Sequeira-Legrand A, Lebrun-Garcia A, Durner J, Pugin A, Wendehenne D (2004) Analysis of nitric oxide signaling functions in tobacco cells challenged by the elicitor cryptogein. Plant Physiol 135:516–529
Laxalt AM, Raho N, Have AT, Lamattina L (2007) Nitric oxide is critical for inducing phosphatidic acid accumulation in Xylanase-elicited tomato cells. J Biol Chem 282:21160–21168
Lee H-A, Lee H-Y, Seo E, Lee J, Kim S-B, Oh S, Choi E, Choi E, Lee SE, Choi D (2016) Current understandings on plant nonhost resistance. Mol Plant Microbe Interact. doi:10.1094/MPMI-10-16-0213-CR
Libault M, Farmer A, Brechenmacher L, Drnevich J, Langley RJ, Bilgin DD, Radwan O, Neece DJ, Clough SJ, May GD, Stacey G (2010) Complete transcriptome of the soybean root hair cell, a single-cell model, and its alteration in response to Bradyrhizobium japonicum infection. Plant Physiol 152:541–552
Libault M, Thibivilliers S, Bilgin DD, Radwan O, Benitez M, Clough SJ, Stacey G (2008) Identification of four soybean reference genes for gene expression normalization. Plant Genome 1:44–54
Lloyd SR, Schoonbeek H, Trick M, Zipfel C, Ridout CJ (2014) Methods to study PAMP-triggered immunity in Brassica species. Mol Plant Microbe Interact 27:286–295
Lori M, van Verk MC, Hander T, Schatowitz H, Klauser D, Flury P, Gehring CA, Boller T, Bartels S (2015) Evolutionary divergence of the plant elicitor peptides (Peps) and their receptors: interfamily incompatibility of perception but compatibility of downstream signalling. J Exp Bot 66:5315–5325
Lotze MT, Zeh HJ, Rubartelli A, Sparvero LJ, Amoscato AA, Washburn NR, DeVera ME, Liang X, Tör M, Billiar T (2007) The grateful dead: damage-associated molecular pattern molecules and reduction/oxidation regulate immunity. Immunol Rev 220:60–81
Luna E, Pastor V, Robert J, Flors V, Mauch-Mani B, Ton J (2011) Callose deposition: a multifaceted plant defense response. Mol Plant Microbe Interact 24:183–193
McMullen MD, Kresovich S, Villeda HS, Bradbury P, Li H, Sun Q, Flint-Garcia S, Thornsberry J, Acharya C, Bottoms C, Brown P, Browne C, Eller M, Guill K, Harjes C, Kroon D, Lepak N, Mitchell SE, Peterson B, Pressoir G, Romero S, Oropeza Rosas M, Salvo S, Yates H, Hanson M, Jones E, Smith S, Glaubitz JC, Goodman M, Ware D, Holland JB, Buckler ES (2009) Genetic properties of the maize nested association mapping population. Science 325:737–740
Millet YA, Danna CH, Clay NK, Songnuan W, Simon MD, Werck-Reichhart D, Ausubel FM (2010) Innate immune responses activated in Arabidopsis roots by microbe-associated molecular patterns. Plant Cell 22:973–990
Mithöfer A, Ebel J, Felle HH (2005) Cation fluxes cause plasma membrane depolarization involved in β-glucan elicitor-signaling in soybean roots. Mol Plant Microbe Interact 18:983–990
Monaghan J, Zipfel C (2012) Plant pattern recognition receptor complexes at the plasma membrane. Curr Opin Plant Biol 15:349–357
Nandety RS, Caplan JL, Cavanaugh K, Perroud B, Wroblewski T, Michelmore RW, Meyers BC (2013) The role of TIR-NBS and TIR-X proteins in plant basal defense responses. Plant Physiol 162:1459–1472
Navarro L, Zipfel C, Rowland O, Keller I, Robatzek S, Boller T, Jones JDG (2004) The transcriptional innate immune response to flg22. interplay and overlap with Avr gene-dependent defense responses and bacterial pathogenesis. Plant Physiol 135:1113–1128
Nishizawa Y, Kawakami A, Hibi T, He DY, Shibuya N, Minami E (1999) Regulation of the chitinase gene expression in suspension-cultured rice cells by N-acetylchitooligosaccharides: differences in the signal transduction pathways leading to the activation of elicitor-responsive genes. Plant Mol Biol 39:907–914
Oliveira-Garcia E, Deising HB (2016) Attenuation of PAMP-triggered immunity in maize requires down-regulation of the key β-1,6-glucan synthesis genes KRE5 and KRE6 in biotrophic hyphae of Colletotrichum graminicola. Plant J 87:355–375
Olukolu BA, Bian Y, De Vries B, Tracy WF, Wisser RJ, Holland JB, Balint-Kurti PJ (2016) The genetics of leaf flecking in maize and its relationship to plant defense and disease resistance. Plant Physiol 172:1787–1803
Olukolu BA, Wang GF, Vontimitta V, Venkata BP, Marla S, Ji J, Gachomo E, Chu K, Negeri A, Benson J, Nelson R, Bradbury P, Nielsen D, Holland JB, Balint-Kurti PJ, Johal G (2014) A genome-wide association study of the maize hypersensitive defense response identifies genes that cluster in related pathways. PLoS Genet 10:e1004562
Pearce G, Yamaguchi Y, Barona G, Ryan CA (2010) A subtilisin-like protein from soybean contains an embedded, cryptic signal that activates defense-related genes. Proc Natl Acad Sci USA 107:14921–14925
Poland JA, Balint-Kurti PJ, Wisser RJ, Pratt RC, Nelson RJ (2009) Shades of gray: the world of quantitative disease resistance. Trends Plant Sci 14:21–29
Poland JA, Bradbury PJ, Buckler ES, Nelson RJ (2011) Genome-wide nested association mapping of quantitative resistance to northern leaf blight in maize. Proc Natl Acad Sci USA 108:6893–6898
Pugin A, Frachisse JM, Tavernier E, Bligny R, Gout E, Douce R, Guernb J (1997) Early events induced by the elicitor cryptogein in tobacco cells: involvement of a plasma membrane NADPH oxidase and activation of glycolysis and the pentose phosphate pathway. Plant Cell 9:2077–2091
Ramakers C, Ruijter JM, Lekanne Deprez RH, Moorman AFM (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339:62–66
Rasul S, Dubreuil-Maurizi C, Lamotte O, Koen E, Poinssot B, Alcaraz G, Wendehenne D, Jeandroz S (2012) Nitric oxide production mediates oligogalacturonide-triggered immunity and resistance to Botrytis cinerea in Arabidopsis thaliana. Plant Cell Environ 35:1483–1499
Rosli HG, Zheng Y, Pombo MA, Zhong S, Bombarely A, Fei Z, Collmer A, Martin GB (2013) Transcriptomics-based screen for genes induced by flagellin and repressed by pathogen effectors identifies a cell wall-associated kinase involved in plant immunity. Genome Biol 14:R139
Schieber M, Chandel NS (2014) ROS function in redox signaling and oxidative stress. Curr Biol 24:453–462
Segonzac C, Feike D, Gimenez-Ibanez S, Hann DR, Zipfel C, Rathjen JP (2011) Hierarchy and roles of pathogen-associated molecular pattern-induced responses in Nicotiana benthamiana. Plant Physiol 156:687–699
Segonzac C, Zipfel C (2011) Activation of plant pattern-recognition receptors by bacteria. Curr Opin Microbiol 14:54–61
Shi Q, Febres VJ, Jones JB, Moore GA (2015) Responsiveness of different citrus genotypes to the Xanthomonas citri ssp. citri-derived pathogen-associated molecular pattern (PAMP) flg22 correlates with resistance to citrus canker. Mol Plant Pathol 16:507–520
Shimizu T, Nakano T, Takamizawa D, Desaki Y, Ishii-Minami N, Nishizawa Y, Minami E, Okada K, Yamane H, Kaku H, Shibuya N (2010) Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice. Plant J 64:204–214
Silva LDCE, Wang S, Zeng ZB (2012) Composite interval mapping and multiple interval mapping: procedures and guidelines for using Windows QTL Cartographer. Methods Mol Biol 871:75–119
Sun Y, Li L, Macho AP, Han Z, Hu Z, Zipfel C, Zhou JM, Chai J (2013) Structural basis for flg22-induced activation of the Arabidopsis FLS2-BAK1 immune complex. Science 342:624–628
Szatmári Á, Zvara Á, Móricz ÁM, Besenyei E, Szabó E, Ott PG, Puskás LG, Bozsó Z (2014) Pattern triggered immunity (PTI) in tobacco: isolation of activated genes suggests role of the phenylpropanoid pathway in inhibition of bacterial pathogens. PLoS One 9:e102869
Tanaka K, Choi J, Cao Y, Stacey G (2014) Extracellular ATP acts as a damage-associated molecular pattern (DAMP) signal in plants. Front Plant Sci 5:446
Thomma BPHJ, Nürnberger T, Joosten MHAJ (2011) Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell 23:4–15
Tsuda K, Sato M, Glazebrook J, Cohen JD, Katagiri F (2008) Interplay between MAMP-triggered and SA-mediated defense responses. Plant J 53:763–775
Valdés-López O, Thibivilliers S, Qiu J, Xu WW, Nguyen THN, Libault M, Le BH, Goldberg RB, Hill CB, Hartman GL, Diers B, Stacey G (2011) Identification of quantitative trait loci controlling gene expression during the innate immunity response of soybean. Plant Physiol 157:1975–1986
Veluchamy S, Hind SR, Dunham DM, Martin GB, Panthee DR (2014) Natural variation for responsiveness to flg22, flgII-28, and csp22 and Pseudomonas syringae pv. tomato in heirloom tomatoes. PLoS One 9:e106119
Vetter M, Karasov TL, Bergelson J (2016) Differentiation between MAMP triggered defenses in Arabidopsis thaliana. PLoS Genet 12:e1006068
Vetter MM, Kronholm I, He F, Häweker H, Reymond M, Bergelson J, Robatzek S, de Meaux J (2012) Flagellin perception varies quantitatively in Arabidopsis thaliana and its relatives. Mol Biol Evol 29:1655–1667
Vidhyasekaran P (2014) PAMP signals in plant innate immunity: signal perception and transduction. Springer, Dordrecht, pp 85–87
Vidhyasekaran P (2016) Switching on plant innate immunity signaling systems: bioengineering and molecular manipulation of PAMP-PIMP-PRR signaling complex. Springer, Switzerland, pp 33–34
Wan J, Zhang XC, Neece D, Ramonell KM, Clough S, Kim SY, Stacey MG, Stacey G (2008) A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20:471–481
Wendehenne D, Lamotte O, Frachisse JM, Barbier-Brygoo H, Pugina A (2002) Nitrate efflux is an essential component of the cryptogein signaling pathway leading to defense responses and hypersensitive cell death in tobacco. Plant Cell 14:1937–1951
Wojtaszek P (1997) Oxidative burst: an early plant response to pathogen infection. Biochem J 322:681–692
Zipfel C, Robatzek S, Navarro L, Oakeley EJ, Jones JDG, Felix G, Boller T (2004) Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 428:764–767
Acknowledgements
This work was funded in part by DOE grant #DE-SC0014116, “Physiological and Molecular-Genetic Characterization of Basal Resistance in Sorghum”; XZ’s fellowship was funded by the China Scholarship Council Fellowship; we would like to thank Carole Saravitz, Joe Chiera, and the staff in NCSU Phytotron for their management of our plant materials, Jim Holland (USDA-ARS) and Sherry Flint-Garcia (USDA-ARS) for seed of the CML228 × B73 mapping population; we also would like to thank Jonathan Kressin (NCSU Department of Horticultural Science), Wall Crumpler, and John Godwin for access to and help with using the plate-reader, and Jenny Kimball, Yangrong Cao, Qin Yang, Yijian He, Guanfeng Wang, and Shannon Sermons for help, advice, and access to various materials.
Author information
Authors and Affiliations
Contributions
PBK, GS, XZ and OVL planned the research, XZ and OVL performed the research and analyzed the data, CA performed statistical analyses, XZ and PBK wrote the manuscript, and GA, OVL, and CA edited the manuscript.
Corresponding author
Ethics declarations
This work was funded in part by DOE Grant #DE-SC0014116, “Physiological and Molecular-Genetic Characterization of Basal Resistance in Sorghum”; XZ’s fellowship was funded by the China Scholarship Council Fellowship.
Conflict of interest
The authors declare that they have no conflict of interest.
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Jianbing Yan.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, X., Valdés-López, O., Arellano, C. et al. Genetic dissection of the maize (Zea mays L.) MAMP response. Theor Appl Genet 130, 1155–1168 (2017). https://doi.org/10.1007/s00122-017-2876-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2876-6