Abstract
Plant height and maturity are two critical traits in sorghum breeding. To develop molecular tools and to identify genes underlying the traits for molecular breeding, we developed 14,739 SNP markers used to genotype the complete sorghum [Sorghum bicolor (L.) Moench] mini core collection. The collection was evaluated in four rainy and three post-rainy season environments for plant height and maturity. Association analysis identified six marker loci linked to height and ten to maturity in at least two environments with at least two SNPs in each locus. Of these, 14 were in close proximity to previously mapped height/maturity QTL in sorghum. Candidate genes for maturity or plant height close to the marker loci include a sugar transporter (SbSUC9), an auxin response factor (SbARF3), an FLC and FT regulator (SbMED12), and a photoperiod response gene (SbPPR1) for maturity and peroxidase 53, and an auxin transporter (SbLAX4) for plant height. Linkage disequilibrium analysis showed that SbPPR1 and SbARF3 were in regions with reduced sequence variation among early-maturing accessions, suggestive of past purifying selection. We also found a linkage disequilibrium block that existed only among the accessions with short plant height in rainy season environments. The block contains a gene homologous to the Arabidopsis flowering time gene, LUMINIDEPENDENS (LD). Functional LD promotes early maturity while mutation delays maturity, affecting plant height. Previous studies also found reduced sequence variations within this gene. These newly-mapped SNP markers will facilitate further efforts to identify plant height or maturity genes in sorghum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant height and maturity are essential breeding objectives in crop plants. Reduced plant height in cereals was the basis for the Green Revolution (Asano et al. 2011; Sasaki et al. 2002; Spielmeyer et al. 2002). Maturity determines whether a variety can be grown in a particular cropping system or ecosystem. As a short-day tropical species with significant photoperiod sensitivity, sorghum [Sorghum bicolor (L.) Moench] flowers when the day length is shorter than its critical photoperiod. Photoperiod sensitive tropical sorghum varieties will not flower when grown in temperate regions because the day length during the summer never reaches the critical photoperiod, and by the time day length becomes short enough for flowering, temperatures are too low for optimal grain yield. Thus, much of the world collection of ~36,000 sorghum accessions flowers too late or is too tall for seed production in temperate zone environments. To overcome this limitation, the Sorghum Conversion Program was initiated in 1963 between the USDA and Texas A&M University to introgress day-neutral flowering alleles and dwarf height alleles into tall, photoperiod sensitive tropical sorghums to produce short photoperiod insensitive types suitable for growing and mechanical harvesting in temperate climates (Quinby 1974; Stephens et al. 1967).
A number of factors can affect the plant height. In sorghum, plant height is controlled by a few major genes based on mapping studies using molecular markers (reviewed by Salas Fernandez et al. 2009). The four major height loci in sorghum (Dw1, Dw2, Dw3 and Dw4) have been mapped with high resolution (Morris et al. 2013), although studies suggest that other loci also affect the plant height (Quinby and Karper 1954). Recessive dw alleles result in a form of dwarfism known as brachytic dwarfism. This type of dwarfism reduces the length of the internodes without affecting other agronomic traits including leaf number/size, flowering time/maturity, peduncle length or panicle size (Quinby 1974). Dw3 (SbPGP1) is the first sorghum height gene that has been cloned; it encodes an ATP-binding cassette type B (ABCB) auxin transporter (Multani et al. 2003). dw3 mutants exhibit reduced seedling auxin transport, reduced stalk height due to shortened lower internodes, thicker stems and altered stalk vasculature (Multani et al. 2003). The defect in auxin movement within dw3 mutants, may also affect other downstream processes, such as the regulation of cell wall extensibility and lignification by peroxidases. Schertz et al. (1971) demonstrated that in sorghum lines isogenic mainly for Dw3, short dw3 plants possess 40 % more peroxidase activity in their internodes than tall Dw3 plants. In addition to auxin, other plant hormones have been implicated in determining plant height. Gibberellins (GAs) are indispensible in stem elongation and flower induction. Association mapping has identified two markers, 44–2,080 (Wang et al. 2012) and pSB945 (Murray et al. 2009), and a height QTL (Sb-HT9.1/Dw1, Brown et al. 2008; Morris et al. 2013) physically close to a GA2-oxidase gene (Upadhyaya et al. 2012a; Wang et al. 2012). GA2-oxidase inactivates GAs, thus reducing the endogenous level which significantly decreases plant height (Biemelt et al. 2004).
Maturity is positively correlated with flowering time (Srinivas et al. 2009) and height (Kebede et al. 2001; Ali et al. 2008; Ritter et al. 2008; Upadhyaya et al. 2009). In sorghum, genetic studies have identified six maturity genes, Ma 1 , Ma 2 , Ma 3 , and Ma 4 (Quinby 1967) and Ma 5 and Ma 6 (Rooney and Aydin 1999). Both Ma 1 and Ma 3 have been cloned. Ma 3 encodes a phytochrome B (Childs et al. 1997) and Ma 1 suppresses flowering by activating the floral inhibitor CONSTANS and repressing the floral activators Early Heading Date 1, FLOWERING LOCUS T (FT), and Zea mays CENTRORADIALIS 8. Mutations in Ma 1 produce early-maturing grain sorghum plants (Murphy et al. 2011).
The first cloned flowering time gene was Arabidopsis LUMINIDEPENDENS (LD) (Lee et al. 1994) that regulates LEAFY (LFY) expression (Aukerman et al. 1999). LFY is critical in the transition from vegetative to flowering stage (Blazquez et al. 1998). Mutation in LD delays flowering in Arabidopsis (Lee et al. 1994). LD is the part of autonomous flowering pathway in which FLOWERING LOCUS C (FLC) is one of the central regulators of flowering in Arabidopsis; low FLC expression promotes early flowering. FLC expression is increased by FRIGIDA (FRI) but decreased by vernalization. A suppressor of FRIGIDA4 (SUF4) promotes early flowering as a result of low FLC expression. SUF4 by itself binds to the FLC promoter as a transcriptional activator, increasing FLC expression. A functional LD binds to SUF4 which suppresses SUF4. This decreases FLC expression and promotes early flowering (Kim et al. 2006). Mutation in LD delays flowering because without LD binding, SUF4 can now increase FLC expression to cause late flowering (Kim et al. 2006; Lee et al. 1994). Another player is the newly-discovered Cryptic Precocious (CRP)/MED12 in Arabidopsis. CRP promotes early flowering and mutation delays flowering under long-day conditions; it controls flowering time by downregulating FLC and upregulating FT (Imura et al. 2012).
The sorghum mini core collection was developed at the International Crops Research Institute for the Semi-Arid Tropics (ICRISAT) using 22,473 sorghum landraces from 76 countries around the world. Based primarily on height and maturity, a core collection of 2,247 landraces was selected (Grenier et al. 2001a). From the core collection, 242 landraces were selected to constitute the mini core collection, based on 11 qualitative and 10 quantitative traits (Upadhyaya et al. 2009). Both core and mini core collections represent the original 22,473 landraces in all phenotypic traits measured (Grenier et al. 2001b; Upadhyaya et al. 2009). The mini core has been used to map economically important traits in sorghum using SSR markers (Upadhyaya et al. 2012a, b; Wang et al. 2012). Furthermore, evaluation of the mini core with the SNP markers used in this study demonstrates that the collection is structured along both geographic origin and race (or subtypes) lines (Wang et al. 2013). In this study, we present the association mapping of height and maturity with 14,739 SNP markers using the mini core collection phenotyped in India’s tropic environments.
Materials and methods
Plant materials and growing conditions
A total of 242 accessions of the sorghum [Sorghum bicolor (L.) Moench] mini core collection (Upadhyaya et al. 2009) were evaluated in four rainy seasons (denoted as 2007R, 2008R, 2009R, and 2010R) and three post-rainy seasons (post-rainy seasons in 2004–2005, 2006–2007, and 2009–2010 are denoted as 2004PR, 2006PR and 2009PR) environments for height and maturity in India as described previously (Upadhyaya et al. 2012a, b). The 2004PR data were from Upadhyaya et al. (2009). Days are longer in rainy season than the post-rainy season except at the equator. Therefore, it takes longer for photosensitive sorghum varieties to flower in rainy season. Photosensitive varieties flower earlier during the post-rainy season when days are shorter, and consequently have a reduced plant stature due to shorter vegetative growth period (Grenier et al. 2001a).
Planting dates were June 18, 21, 26, and 23 for 2007R, 2008R, 2009R, and 2010R, respectively, and October 14, 26, and 27 for 2004PR, 2006PR and 2009PR, respectively. The plants were grown in an alpha design with two replications except 2010R when three replications were used. Each single-row plot was 4-m long with a row spacing of 75 cm, and plant spacing within a row of 10 cm. Ammonium phosphate was applied at the rate of 150 kg/ha before planting, and 100 kg/ha of urea was applied as top dressing 3 weeks after planting. Field plots were irrigated 2–3 times in the rainy and five at equal intervals in post-rainy seasons (each with 7 cm water). Plant height was measured of the main stalk at 50 % flowering and presented as a mean of five randomly selected plants. Days to 50 % flowering was used as a measure of maturity and was measured by the number of days from the 50 % seedling emergence to the date when 50 % of the plants had started flowering.
SNP development and genotyping
SNP development and genotyping has been described previously (Wang et al. 2013). Specifically, SNPs were generated using Illumina GAII× sequencing of DNA templates from specific sites targeted by restriction endonucleases as previously described (Wang et al. 2013). Each SNP name starts with its chromosome number followed by its physical location in bp (e.g., SNP chr1_46978664, resides on SBI-01 at position 46,978,664 bp). A total of 14,739 SNPs were used in this study.
Data analysis
Principal component analysis (PCA) was performed using the prcomp() of R (version 2.15.1, 64 bit; www.r-project.org/). Marker-trait associations were calculated using six models to evaluate the effects of population structure (Q, PC) and kinship (K): (1) naïve—without controlling for Q or K, (2) Q model—control for Q, (3) PCA model—control for PC, (4) K model—control for K, (5) QK model—control for both Q and K, and (6) the PK model—control for both PC and K. The naïve, Q, and PCA models were performed using GLM; the K, QK, and PK models were performed using mixed linear model (MLM) in TASSEL 3.0 (Casa et al. 2008; Pasam et al. 2012; Upadhyaya et al. 2012a, b; Yang et al. 2011; Yu et al. 2006). TASSEL 3.0 (Bradbury et al. 2007) was available from http://www.maizegenetics.net/. The Q matrix was calculated using STRUCTURE 2.3 (Pritchard et al. 2000) available from http://pritch.bsd.uchicago.edu/structure.html. The number of groups/subpopulations (k) was tested between 2 and 16 when running STRUCTURE. STRUCTURE was run with the admixture model, a burn-in period of 50,000 and 5,000 Markov Chain Monte Carlo repetitions as described by Wang et al. (2009), and genotypic data of the mini core collection from all SNP markers. The K matrix was generated in TASSEL with all SNP markers. Significance of associations between markers and traits was based on threshold p < 3.39 × 10−6, a stringent Bonferroni correction calculated by dividing 0.05 by 14,739 as in Murray et al. (2009). The measure of linkage disequilibrium, r 2, was calculated in TASSEL together with its p value.
Marker localization
SNP markers developed in this study are all identified by their chromosome locations. Markers previously identified as linked to height or maturity in sorghum were localized to the genome-based sequence information of relevant markers provided in Map Viewer in the NCBI website (http://www.ncbi.nlm.nih.gov/mapview/). For pSB RFLP markers, the marker sequences available from Map Viewer were searched against the sorghum genome presented in http://www.plantgdb.org/SbGDB/, http://www.phytozome.net/sorghum, or http://genome.jgi-psf.org/ to determine their physical locations. For RFLP markers with sequence from other grasses, the sequence was used to search the sorghum nucleotide and EST database in GenBank. Top-match sorghum sequence was then used as above to place that marker. For SSR markers, if their position was not given in Map Viewer or provided in Ramu et al. (2009), their primer sequences were then used to search the genome database in http://www.phytozome.net/sorghum for physical localization. Maps used were those in Map Viewer from Chittenden et al. (1994), Peng et al. (1999), Bhattramakki et al. (2000), Kong et al. (2000), and Bowers et al. (2003). Maps of chromosomes based on the physical distances in Mb were drawn using MapDraw 2.2 by Liu and Meng (2003). For localizing SNP markers to mapped height or maturity QTLs, only maps containing physically placeable markers are used.
Results
Evaluation of association mapping models
In total, 14,739 SNP markers were used in this study. To calculate the Q matrix, the number of subpopulations (k) in this mini core was first estimated by calculating the posterior probability ln(P(D)) for all possible k values between 2 and 16 in STRUCTURE 2.3. The “true” value of k was when ln(P(D)) increase plateaued or became minimal, as described by Casa et al. (2008). This k value has been determined to be 11 as described previously based mostly on ln(P(D)), but also on phylogenetic and principal component analyses (Wang et al. 2013). This was used to generate the Q matrix for both Q and QK models in this study.
To assess the utility of each model, ranked p values across all the environments from each model were plotted in a cumulative way for both height and maturity (Fig. 1) as described by Kang et al. (2008). A uniform p value distribution indicates an ideal model. Using this criterion, QK, PK, and K were the better models than naïve, Q or PCA in both height and maturity. The latter were characterized by the excess of low p values in both traits while QK, PK, and K models produced essentially similar results. This indicates that models controlling for either kinship alone or structure and kinship minimize the chance of spurious association.
Physical location of markers linked to height and maturity in sorghum. For each chromosome, on the left are physical positions of markers in mega base pairs (Mb). H or M in parenthesis: H marker linked to height, M marker linked to maturity. Open/closed bars to the left are mapped maturity/height QTLs from previous studies. Arrows indicate direction/range of the QTLs. Not drawn to scale relative to the whole chromosome but to scale for the physical positions shown
Association mapping
Because the results from K, QK, and PK models were similar, only results from the K model (Pasam et al. 2012) were presented in this paper. To identify true marker-trait associations, we used both p values and significant association of more than one SNP as is a case in a true association (Li et al. 2010; Morris et al. 2013). All associations in Table 1 had p values <3.39 × 10−6 in at least one environment. Furthermore, these associations had to involve at least two adjacent SNPs in each locus in at least two environments (p < 0.0001; Famoso et al. 2011). Consequently, association with lone SNPs, such as the one associated with SbPGP1/Dw3 (chr7_58616538; Table 1) were not selected. SNPs are considered as one locus if they are located within arbitrarily set 200 bp of each other. Using these criteria, we found eight marker loci associated with maturity and one with height (Table 1).
All nine marker loci showed strongest association only in two of the seven environments, 2007R and 2006PR (Table 1). No other environments produced significant association and only one marker locus was associated with height with p values lower than the threshold. The strongest association with maturity was the SNP locus of chr6_554248 in three of the four rainy season environments. There was only one strong association with height. The SNP locus of chr1_46978680 showed the strongest multiple consecutive SNP association with height with an average p value of 2.98 × 10−5 and included chr1_46978587, chr1_46978605, chr1_46978664, chr1_46978680, chr1_46978695, and chr1_46978697. This was the only association with height with a p value lower than the threshold. Chr4_62536438 was associated with height in 2007R with a p value of 4.99 × 10−5, but the association was less strong in the other six environment (data not shown).
Association of the traits with the two cloned known genes that control height/maturity was also tested. The height gene, SbPGP1/dw3 (Multani et al. 2003), was highly associated with height in 2009PR. Based on p values across environments, SbPGP1 was more likely associated with height in post-rainy, not in rainy season environments (Table 1). SbPGP1 (Sb07g023730) is located between 58,610,896 and 58,618,660 bp on chromosome 7 and our SNP for the gene was chr7_58616538. Ma1 (Sb06g014570—SbPRR37) is between 40,280,414 and 40,290,602 bp on chromosome 6 (Murphy et al. 2011) and a SNP marker in this region (chr6_40280035) was neither associated with maturity nor height. This is probably because the Ma genes control photoperiodic response in the temperature regions (Miller et al. 1968), while the phenotypic data used in this study were gathered in the tropics although an SSR marker close to SbPRR37 was associated with maturity with a p value above the Bonferroni-corrected threshold in a previous study (Upadhyaya et al. 2012a). Attempt was also made to analyze the phenotypic data with the newly-developed 265 K SNP markers (Morris et al. 2013). However, for unknown reasons, no known height or maturity loci were mapped by the markers. Therefore, these results were not presented here.
Colocalization with previously mapped QTLs
To validate the marker loci in Table 1, we colocalized the marker loci with previously mapped height or maturity QTLs to the sorghum chromosomes (Fig. 2). Of the nine marker loci, chr6_554248 was not clustered with previously mapped height or maturity QTLs. The remaining eight were localized in the vicinity of the QTLs. Four of the nine marker loci were also located toward chromosome ends (Fig. 2).
Physical location of markers linked to height and maturity in sorghum. For each chromosome, on the left are physical positions of markers in mega base pairs (Mb). H or M in parenthesis: H marker linked to height, M marker linked to maturity. Open/closed bars to the left are mapped maturity/height QTLs from previous studies. Arrows indicate direction/range of the QTLs. Not drawn to scale relative to the whole chromosome but to scale for the physical positions shown
On chromosome 1, of the two maturity QTLs mapped in Srinivas et al. (2009), one was anchored by Xtxp43 and Xtxp32 and the other was anchored by Xtxp75 and Xtxp284. Xtxp43 was 3.29 Mb from chr1_46978680. The height QTL mapped in Hart et al. (2001) covers a wide region spanning the centromere based on the anchor markers Xumc128 and Xcdo20. On chromosome 2, a maturity QTL was mapped next to Xtxp315 toward Xtxp56 by Srinivas et al. (2009), close to chr2_69639719 and close to the height marker pSB1224 identified by Murray et al. (2009). Chromosome 3 contained two SNP marker loci close to a maturity (flowering date) QTL flanked by Xtxp285 and Xtxp114 (Shiringani et al. 2010) (Fig. 2).
Two marker loci were associated with maturity on chromosome 6. Two overlapping QTLs (Srinivas et al. 2009), one for height and one for maturity were anchored by Xtxp145 (Fig. 2) and oriented by Xtxp317 (not shown in Fig. 2). Both are physically clustered with chr6_44980895 also in the region. Another maturity-associated SNP, chr6_554248, was located at the very end of the short arm of chromosome 6 and was not clustered with any known height/maturity QTLs. On chromosome 7, the height QTL by Klein et al. (2001) was anchored to Xtxp295. Another close height QTL by Murray et al. (2008a, b) could only be oriented and loosely localized by Xtxp295. The latter may be close to chr7_58616538 (SbPGP1) which was covered by the height QTL mapped in Srinivas et al. (2009). Toward the other end of chromosome 7, another height QTL anchored by Xtxp312 and oriented by Xtxp159 (Srinivas et al. 2009) was in the vicinity of chr7_3591025 and chr7_380316 (Fig. 2).
Candidate genes
To understand the causes of variation in height and maturity, we examined the genomic regions in each marker locus for the top four associations with maturity and the one with height for candidate genes. The locus associated with maturity with the lowest p value was that occupied by four SNPs: chr6_554233, chr6_554248, chr6_554251, and chr6_554279. They were all associated with maturity with identical p values. Since the flanking SNPs (chr6_338533 and chr6_616248) were not associated with maturity, the underlying gene might be close to the locus. Four of the five genes close to the locus were transporters and the closest gene encoded for a sugar transporter (Sb06g000520/SbSUC9: 548793–553157 bp). Similar to the locus of chr6_554233, the two flanking markers (chr3_58925778 and chr3_58992842) of locus of chr3_58977893, chr3_58977930, and chr3_58977944 were not associated with maturity either. Again, we looked for candidates close to the locus. There were two flanking genes, a serine/threonine protein kinase (Sb03g030730: 58968952–58971384 bp) and SbARF3 (Sb03g030740: 58980057–58985203 bp), but SbARF3 was closer (2.1 kb) to the locus than Sb03g030730 (8.9 kb). The locus of chr1_73244358 was 10.99 kb to Sb01g050280/SbMED12 (73255352–73267414 bp). The last of top four associations with maturity was with locus of chr6_44980895 which is 5 kb to an extensin gene (Sb06g016310: 44974245–44975602 bp) and 28.45 kb to a photoperiod response gene (Sb06g016300/SbPPR1: 44950503–44952442 bp).
The height SNP locus was inside a peroxidase gene (Sb01g027330/Prx53). All SNPs in the locus were in the last exon of the gene. Except for chr1_46978664 and chr1_46978697, all SNPs resulted in synonymous mutations (Table 2). The two non-synonymous SNPs showed stronger association with height in two rainy season environments (Table 3).
Linkage disequilibrium at selected loci
In an effort to find out if any of the loci significantly associated with maturity or height was under selection pressure, we calculated linkage disequilibrium using TASSEL. We separated the mini core based on maturity data in 2006PR and 2007R because these were the two environments in which significant associations were identified (Table 1). SNP genotypes of top 50 early- and top 50 late-maturing accessions for each of the nine loci were fed into TASSEL. In six loci, there was no difference in linkage disequilibrium between the early- and late-maturing accessions. In the remaining three loci, TASSEL could not calculate r 2, the measure of linkage disequilibrium, because in all three cases genotypes of the top 50 early-maturing accessions based on 2007R data were identical, while those of the top 50 late-maturing accessions were different. The results were similar based on 2006PR data for two of the three loci.
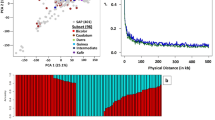
The most uniform locus was chr6_44980895 which is next to the SNP locus of chr6_44931382 with total 13 consecutive SNP markers (Fig. 3a). Genotypes consisted of the 13 SNPs from the top 50 early-maturing accessions based on the 2007R data were identical. In 2006PR data, only genotypes consisted of SNPs from chr6_44980895 locus were identical. Genotypes consisted of SNPs from chr6_44931382 locus were only shared by top 14 early-maturing accessions. Overall, 28 % of accessions were shared between top 50 early-maturing accessions in 2007R and 2006PR. Similarly, SNP genotypes in the locus of chr3_58977893 for the top 50 early-maturing accessions were also identical, while only 34 in 2006PR were identical (Fig. 3b). For SNP genotypes in the locus of chr6_554248, the top 50 early accessions were identical in 2007R, while the top 50 early accessions in 2006PR were different (Fig. 2c). Since the identical genotypes in the loci suggest that selection has occurred and mutations have been eliminated (Mather et al. 2007), these results indicate that both photoperiod response gene SbPPR1 and auxin response factor SbARF3 were under selection pressure, most likely as a result of artificial selection.
We also found two consecutive marker loci 99.61 kb apart from chromosome 3, chr3_72287401 and chr3_72387014, were in linkage disequilibrium. We calculated both r 2 and p values between the two SNPs in top 50 short/early and top 50 tall/late accessions in all environments. Based on the conventional r 2 threshold value of 0.1 (Nordborg et al. 2002; Palaisa et al. 2003; Remington et al. 2001) and p < 0.0001 (Caldwell et al. 2006), the pair was only in significant linkage disequilibrium in the top 50 short accessions in the four rainy season environments (Table 4; Fig. 4a). They were not in linkage disequilibrium in top 50 early or late accessions in any environment (data not shown). Fifty-eight percent of top 50 short accessions in the four rainy season environments were shared.
A haplotype block defined by chr3_72287401 and chr3_72387014 on sorghum chromosome 3. a Linkage disequilibrium between chr3_72287401 and chr3_72387014 and other SNPs (on the left) in the region as represented by r 2 (scale on the right). The left and right panels show linkage disequilibrium in 50 tallest and 50 shortest accessions in 2007R, respectively. For both panels, lower left are linkage disequilibrium p values and upper right are r 2 values. Arrow points to the r 2 value block between chr3_72287401 and chr3_72387014. b Gene and SNP distribution in the region; chr3_72287333 and chr3_72287335 are not shown in B. Scale based on screen shot of the region captured in www.phytozome.net
The above is clear evidence that there was a unifying selection for the genes in the block in short accessions, but not in tall accessions. Among the genes present in the region that could be related to height or maturity in sorghum was the homolog (Sb03g045030/SbLD1: 72338651–72347617 bp) of an Arabidopsis flowering time gene, LUMINIDEPENDENS (LD) (Fig. 3B).
Discussion
One shortcoming of association mapping is false-positive associations. MLM has been proposed to minimize such false-positives (Yu et al. 2006). Comparison of different association models including MLM indicates that QK, PK, and K were the better models than naïve, Q or PCA in both height and maturity. The more false positive-prone models of naïve, Q or PCA were characterized by the excess of low p values in both traits while QK, PK and K models produced essentially similar results. This indicates that models controlling for either kinship alone or structure and kinship minimize the chance of spurious association. This supports the conclusions reported in maize (Yang et al. 2011), barley (Pasam et al. 2012), and sorghum (Casa et al. 2008). We also compared QK models that use 168 well-spaced unlinked SNPs to generate both Q and K matrices and found that p value distribution was not as uniform as QK models that use 14,739 SNPs (data not shown).
We previously identified SSR markers linked to height or maturity also on chromosomes 6, 9, and 10 (Upadhyaya et al. 2012a) and 2, 6, and 9 (Wang et al. 2012). Many SNP markers associated with either height or maturity were also found in the regions on the three chromosomes, but were not presented because they did not meet our stringent selection criteria. On chromosome 6, chr6_42600524, chr6_56644978, and chr6_59912226 were associated with height with the respective p values of 7.75 × 10−4, 6.34 × 10−4, and 1.83 × 10−4. They were not considered because their adjacent SNPs were not associated with height and their association was only in one environment. Similarly, both chr6_48621460 and chr6_49571685 were associated with maturity with the p values of 2.85 × 10−7 and 3.48 × 10−7 which were lower than the threshold. Both were screened out because of the same reason. Adjacent SNPs of another height-associated marker, chr6_57689190, did show association also with height in both 2004PR and 2009PR. But because of high p values (>0.0001), chr6_57689190 was not included as a height marker. On chromosome 9, chr9_55288952, chr9_56249628, chr9_57984418, chr9_58349510, and chr9_58571457 were all associated with height, but just in one environment and for chr9_56249628, chr9_57984418, and chr9_58349510, their adjacent markers were not associated with height. Therefore, these were not listed as height markers on chromosome 9. On chromosome 10, chr10_55398654, chr10_57191748, and chr10_58701631 were rejected for the same reason. chr10_57423575 and chr10_57423587 were adjacent and were associated with height, but only in one environment. chr10_59454786 was significantly associated with maturity (p < 3.39 × 10−6), but its adjacent SNPs were not associated with maturity. The last marker, chr10_59519922, almost met our criteria. It was significantly associated with maturity in one environment and associated with p <0.0001 in two other environments. But its only adjacent SNP produced no data which made it a loner with no adjacent marker to validate its association. Otherwise, this marker would be very close to a height SSR marker (4–191) identified previously (Upadhyaya et al. 2012a). A summary comparing our previous results with the current study is provided in Table S1.
We have identified candidate genes linked to the associated loci, but not their flanking markers. SbSUC9 was identified close to the locus of chr6_554233 and was most homologous to (45 % identical and 66 % similar) AtSUC9 in Arabidopsis which was expressed in shoots and flowers. Functional AtSUC9 delay flowering and mutation causes early flowering in short-day; no other visible phenotype was observed in the mutant other than early flowering (Sivitz et al. 2007). The locus of chr3_58977893 was flanked by two genes: a serine/threonine protein kinase (Sb03g030730: 58968952–58971384 bp) and SbARF3 (Sb03g030740: 58980057–58985203 bp). SbARF3 (31–489 aa) was 42 % identical/59 % similar to the Arabidopsis auxin response factor 2 (AtARF2), a repressor of auxin signaling (Lim et al. 2010) but is most homologous (56 % identical/68 % similar in 4–430 aa) to AtARF3. AtARF2 mutants are taller (Okushima et al. 2005) and flower a week late with reduced fertility (Ellis et al. 2005; Schruff et al. 2006). AtARF3 is regulated by AG through GIK and is essential for flower development in Arabidopsis (Ng et al. 2009). SbMED12 close to the locus of chr1_73244358 is most homologous (45 % identical and 64 % similar) to Cryptic Precocious (CRP)/MED12 in Arabidopsis. CRP promotes early flowering and mutation in CRP as well as CRP overexpression delays flowering (doubles the time to flowering) under long-day conditions (no difference under short-day conditions) (Imura et al. 2012). This indicates that CRP is a negative regulator of FLC and positive regulator of FT. CRP also participates in auxin response as AtARF3 because both mutants produced similar gynoecium phenotype (Imura et al. 2012). SbPPR1 found in the locus of chr6_44980895 is 37 % identical/51 % similar to potato PHOR1 (CAC35703). PHOR1 acts in the GA signaling pathway and regulates photoperiodic tuber formation in potato. Suppression of PHOR1 expression results in early tuber formation, consequently higher tuber yield (Amador et al. 2001).
The height SNP locus was linked to a peroxidase gene Prx53. Peroxidase activity regulates plant height through its metabolism of auxin. Short sorghum lines isogenic mainly for dw3 have been shown to produce 40 % higher peroxidase activity in the internodes than the tall Dw3 isogenic lines (Schertz et al. 1971). Overproduction of peroxidase through overexpressing peroxidase genes produces shorter plants, while underproduction increases plant height in tobacco (Kristensen et al. 1997; Lagrimini et al. 1990, 1997): overproducers are 13–22 % taller whereas underproducers are 5–14 % shorter (Lagrimini et al. 1997).
The flowering time gene that may have been under selection, SbLD1, is homologous to Arabidopsis LD which is the first cloned flowering time gene in Arabidopsis and it encodes a homeobox protein (Lee et al. 1994). In the absence of FRI, functional LD interacts with SUF4 to reduce FLC expression which leads to early flowering (Kim et al. 2006). Mutation in LD therefore promotes late maturity (Lee et al. 1994). This correlates with higher level of linkage disequilibrium among short accessions observed in this study (Table 4). It is intriguing that the block is not in linkage disequilibrium in early- or late-maturing accessions. SbLD1 had 12 introns and 13 exons distributed in 8,966 bp. Zheng et al. (2011) sequenced two sweet sorghum (E-Tian and Keller) and one grain sorghum (Ji2731) varieties to identify SNPs. Sweet sorghums are usually tall and late maturing and grain sorghum are short and early maturing. No SNPs were discovered in the coding region of SbLD1 in the two sweet sorghums, but six were found in the grain sorghum Ji2731. Interestingly, five of the six SNPs were in the introns (one in 3′ UTR) and the only exonic SNP produced a synonymous mutation (Table 5). In addition, four different SNPs were found also in the coding region by Nelson et al. (2011) and three of those were in the introns and one in 3′ UTR (Table 5). The only indel found in the two studies was a single T insertion in poly-T tract in the third intron of SbLD1 in Ji2731 (Zheng et al. 2011). In our study of 243 sorghum accessions, the closest SNPs we found to SbLD1 were chr3_72287401 and chr3_72387014 (Fig. 4b). This is the further evidence that genetic variation is minimal in the gene which again suggests that SbLD1 may have been under selection.
Conclusion
Association mapping of 14,739 SNPs in the sorghum mini core collection evaluated in tropical environments identified nine SNP marker loci linked either to height or maturity. Of these, eight were in the close proximity of previously mapped height and maturity QTLs in sorghum. Candidate genes for maturity or height close to marker loci strongly associated with the traits include a sugar transporter (SbSUC9), an auxin response factor (SbARF3), an FLC and FT regulator (SbMED12), and a photoperiod response gene (SbPPR1) for maturity, and peroxidase 53 for height. Linkage disequilibrium analysis showed that SbPPR1 and SbARF3 were in regions with reduced genetic variation among early-maturing accessions, suggestive of past purifying selection. We also found a linkage disequilibrium block that existed only among short accessions in rainy season environments. The block contains a gene homologous to the Arabidopsis flowering time gene, LD. Functional LD promotes early maturity while mutation delays maturity, affecting plant height. Previous studies also found reduced genetic variations in this gene. These newly-mapped SNP markers will facilitate future efforts to identify height or maturity genes in sorghum.
References
Ali ML, Rajewski JF, Baenziger PS, Gill KS, Eskridge KM, Dweikat I (2008) Assessment of genetic diversity and relationship among a collection of US sweet sorghum germplasm by SSR markers. Mol Breed 21:497–509
Amador V, Monte E, García-Martínez JL, Prat S (2001) Gibberellins signal nuclear import of PHOR1, a photoperiod-responsive protein with homology to Drosophila armadillo. Cell 106:343–354
Asano K, Yamasaki M, Takuno S, Miura K, Katagiri S, Ito T, Doi K, Wu J, Ebana K, Matsumoto T, Innan H, Kitano H, Ashikari M, Matsuoka M (2011) Artificial selection for a green revolution gene during japonica rice domestication. Proc Natl Acad Sci USA 108:11034–11039
Aukerman MJ, Lee I, Weigel D, Amasino RM (1999) The Arabidopsis flowering-time gene LUMINIDEPENDENS is expressed primarily in regions of cell proliferation and encodes a nuclear protein that regulates LEAFY expression. Plant J 18:195–203
Bhattramakki D, Dong J, Chhabra K, Hart GE (2000) An integrated SSR and RFLP linkage map of Sorghum bicolor (L.) Moench. Genome 43:988–1002
Biemelt S, Tschiersch H, Sonnewald U (2004) Impact of altered gibberellin metabolism on biomass accumulation, lignin biosynthesis, and photosynthesis in transgenic tobacco plants. Plant Physiol 135:254–265
Blazquez MA, Green R, Nilsson O, Sussman MR, Weigel D (1998) Gibberellins promote flowering of Arabidopsis by activating the LEAFY promoter. Plant Cell 10:791–800
Bowers JE, Abbey C, Anderson S, Chang C, Draye X et al (2003) A high-density genetic recombination map of sequence-tagged sites for sorghum, as a framework for comparative structural and evolutionary genomics of tropical grains and grasses. Genetics 165:367–386
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Brown PJ, Rooney WL, Franks C, Kresovich S (2008) Efficient mapping of plant height quantitative trait loci in a sorghum association population with introgressed dwarfing genes. Genetics 180:629–637
Caldwell KS, Russell J, Langridge P, Powell W (2006) Extreme population-dependent linkage disequilibrium detected in an inbreeding plant species, Hordeum vulgare. Genetics 172:557–567
Casa AM, Pressoir GH, Brown PJ, Mitchell SE, Rooney WL, Tuinstra MR, Franks CD, Kresovich S (2008) Community resources and strategies for association mapping in sorghum. Crop Sci 48:30–40
Childs KL, Miller FR, Cordonnier-Pratt MM, Pratt LH, Morgan PW, Mullet JE (1997) The sorghum photoperiod sensitivity gene, Ma3, encodes a phytochrome B. Plant Physiol 113:611–619
Chittenden LM, Schertz KF, Lin YR, Wing RA, Paterson AH (1994) A detailed RFLP map of Sorghum bicolor × S. propinquum, suitable for high-density mapping, suggests ancestral duplication of sorghum chromosomes or chromosomal segments. Theor Appl Genet 87:925–933
Ellis CM, Nagpal P, Young JC, Hagen G, Guilfoyle TJ, Reed JW (2005) AUXIN RESPONSE FACTOR1 and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 132:4563–4574
Famoso AN, Zhao K, Clark RT, Tung CW, Wright MH, Bustamante C, Kochian LV, McCouch SR (2011) Genetic architecture of aluminum tolerance in rice (Oryza sativa) determined through genome-wide association analysis and QTL mapping. PLoS Genet 7:e1002221
Grenier C, Bramel-Cox PJ, Hamon P (2001a) Core collection of sorghum: I. Stratification based on eco-geographical data. Crop Sci 41:234–240
Grenier C, Hamon P, Bramel-Cox PJ (2001b) Core collection of sorghum: II. Comparison of three random sampling strategies. Crop Sci 41:241–246
Hart GE, Schertz KF, Peng Y, Syed NH (2001) Genetic mapping of Sorghum bicolor (L.) Moench QTLs that control variation in tillering and other morphological characters. Theor Appl Genet 103:1232–1242
Imura Y, Kobayashi Y, Yamamoto S, Furutani M, Tasaka M, Abe M, Araki T (2012) CRYPTIC PRECOCIOUS/MED12 is a novel flowering regulator with multiple target steps in Arabidopsis. Plant Cell Physiol 53:287–303
Kang HM, Zaitlen NA, Wade CM, Kirby A, Heckerman D, Daly MJ, Eskin E (2008) Efficient control of population structure in model organism association mapping. Genetics 178:1709–1723
Kebede H, Subadhi PK, Rosenow DT, Nguyen HT (2001) Quantitative trait loci influencing drought tolerance in grain sorghum (Sorghum bicolor L. Moench). Theor Appl Genet 103:266–276
Kim S, Choi K, Park C, Hwang HJ, Lee I (2006) SUPPRESSOR OF FRIGIDA, encoding a C2H2-Type zinc finger protein, represses flowering by transcriptional activation of Arabidopsis FLOWERING LOCUS C. Plant Cell 18:2985–2998
Klein RR, Rodriguez-Herrera R, Schlueter JA, Klein PE, Yu ZH, Rooney WL (2001) Identification of genomic regions that affect grain-mould incidence and other traits of agronomic importance in sorghum. Theor Appl Genet 102:307–319
Kong L, Dong L, Hart GE (2000) Characteristics, linkage-map positions, and allelic differentiation of Sorghum bicolor (L.) Moench DNA simple-sequence repeats (SSRs). Theor Appl Genet 101:438–448
Kristensen BK, Brandt J, Bojsen K, Thordal-Christensen H, Kerby KB, Collinge DB, Mikkelsen JD, Rasmussen SK (1997) Expression of a defence-related intercellular barley peroxidase in transgenic tobacco. Plant Sci 122:173–182
Lagrimini LM, Bradford S, Rothstein S (1990) Peroxidase-induced wilting in transgenic tobacco plants. Plant Cell 2:7–18
Lagrimini LM, Gingas V, Finger F, Rothstein S, Liu T (1997) Characterization of antisense transformed plants deficient in the tobacco anionic peroxidase. Plant Physiol 114:1187–1196
Lee I, Aukerman MJ, Gore SL, Lohman KN, Michaels SD, Weaver LM, John MC, Feldmann KA, Amasino RM (1994) Isolation of LUMINIDEPENDENS: a gene involved in the control of flowering time in Arabidopsis. Plant Cell 6:75–83
Li Y, Huang Y, Bergelson J, Nordborg M, Borevitz JO (2010) Association mapping of local climate-sensitive quantitative trait loci in Arabidopsis thaliana. Proc Natl Acad Sci USA 107:21199–21204
Lim PO, Lee IC, Kim J, Kim HJ, Ryu JS, Woo HR, Nam HG (2010) Auxin response factor 2 (ARF2) plays a major role in regulating auxin-mediated leaf longevity. J Exp Bot 61:1419–1430
Liu RH, Meng JL (2003) MapDraw: a Microsoft Excel macro for drawing genetic linkage maps based on given genetic linkage data. Heraditas (Beijing) 25:317–321
Mather KA, Caicedo AL, Polato NR, Olsen KM, McCouch S, Purugganan MD (2007) The extent of linkage disequilibrium in rice (Oryza sativa L.). Genetics 177:2223–2232
Miller FR, Quinby JR, Cruzado HJ (1968) Expression of known maturity genes of sorghum in temperate and tropical environments. Crop Sci 8:675–677
Morris GP, Ramu P, Deshpande SP, Hash CT, Shah T, Upadhyaya HD, Riera-Lizarazu O, Brown PJ, Acharya CB, Mitchell SE, Harriman J, Glaubitz JC, Buckler ES, Kresovich S (2013) Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc Natl Acad Sci USA 110:453–458
Multani DS, Briggs SP, Chamberlin MA, Blakeslee JJ, Murphy AS, Johal GS (2003) Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 302:81–84
Murphy RL, Klein RR, Morishige DT, Brady JA, Rooney WL, Miller FR, Dugas DV, Klein PE, Mullet JE (2011) Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum. Proc Natl Acad Sci USA 108:16469–16474
Murray SC, Sharma A, Rooney WL, Klein PE, Mullet JE, Mitchell SE, Kresovich S (2008a) Genetic improvement of sorghum as a biofuel feedstock: I. QTL for stem sugar and grain nonstructural carbohydrates. Crop Sci 48:2165–2179
Murray SC, Rooney WL, Mitchell SE, Sharma A, Klein PE, Mullet JE, Kresovich S (2008b) Genetic improvement of sorghum as a biofuel feedstock: II. QTL for stem and leaf structural carbohydrates. Crop Sci 48:2180–2193
Murray SC, Rooney WL, Mitchell SE, Kresovich S (2009) Sweet sorghum diversity and association mapping for Brix and height. Plant Genome 2:48–62
Nelson JC, Wang S, Wu Y, Li X, Antony G, White FF, Yu J (2011) Single-nucleotide polymorphism discovery by high-throughput sequencing in sorghum. BMC Genomics 12:352
Ng KH, Yu H, Ito T (2009) AGAMOUS controls GIANT KILLER, a multifunctional chromatin modifier in reproductive organ patterning and differentiation. PLoS Biol 7(11):e1000251
Nordborg M, Borevitz JO, Bergelson J, Berry CC, Chory J, Hagenblad J, Kreitman M, Maloof JN, Noyes T, Oefner PJ, Stahl EA, Weigel D (2002) The extent of linkage disequilibrium in Arabidopsis thaliana. Nat Genet 30:190–193
Okushima Y, Mitina I, Quach HL, Theologis A (2005) AUXIN RESPONSE FACTOR 2 (ARF2): a pleiotropic developmental regulator. Plant J 43:29–46
Palaisa KA, Morgante M, Williams M, Rafalski A (2003) Contrasting effects of selection on sequence diversity and linkage disequilibrium at two phytoene synthase loci. Plant Cell 15:1795–1806
Pasam RK, Sharma R, Malosetti M, van Eeuwijk FA, Haseneyer G, Kilian B, Graner A (2012) Genome-wide association studies for agronomical traits in a world wide spring barley collection. BMC Plant Biol 12:16
Peng Y, Schertz KF, Cartinhour S, Hart GE (1999) Comparative genome mapping of Sorghum bicolor (L.) Moench using an RFLP map constructed in a population of recombinant inbred lines. Plant Breed 118:225–235
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quinby JR (1967) The maturity genes of sorghum. Adv Agron 19:267–305
Quinby JR (1974) Sorghum improvement and the genetics of growth. A&M Univ Press, College Station
Quinby JR, Karper RE (1954) Inheritance of height in sorghum. Agron J 46:211–216
Ramu P, Kassahun B, Senthilvel S, Kumar CA, Jayashree B, Folkertsma RT, Reddy LA, Kuruvinashetti MS, Haussmann BIG, Hash CT (2009) Exploiting rice–sorghum synteny for targeted development of EST-SSRs to enrich the sorghum genetic linkage map. Theor Appl Genet 119:1193–1204
Remington DL, Thornsberry JM, Matsuoka Y, Wilson LM, Whitt SR, Doeblay J, Kresovich S, Goodman MM, Buckler ES (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc Natl Acad Sci USA 98:11479–11484
Ritter KB, Jordan DR, Chapman SC, Godwin ID, Mace ES, McIntyre CL (2008) Identification of QTL for sugar-related traits in a sweet × grain sorghum (Sorghum bicolor L. Moench) recombinant inbred population. Mol Breed 22:367–384
Rooney WL, Aydin S (1999) Genetic control of a photoperiod-sensitive response in Sorghum bicolor (L.) Moench. Crop Sci 39:397–400
Salas Fernandez MG, Becraft PW, Yin Y, Lübberstedt T (2009) From dwarves to giants? Plant height manipulation for biomass yield. Trends Plant Sci 14:454–461
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush GS, Kitano H, Matsuoka M (2002) Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 416:701–702
Schertz KF, Sumpter NA, Sarkissian IV, Hart GE (1971) Peroxidase regulation by the 3-dwarf height locus in sorghum. J Hered 62:235–238
Schruff MC, Spielman M, Tiwari S, Adams S, Fenby N, Scott RJ (2006) The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 133:251–261
Shiringani AL, Frisch M, Friedt W (2010) Genetic mapping of QTLs for sugar-related traits in a RIL population of Sorghum bicolor L. Moench. Theor Appl Genet 121:323–336
Sivitz AB, Reinders A, Johnson ME, Krentz AD, Grof CP, Perroux JM, Ward JM (2007) Arabidopsis sucrose transporter AtSUC9. High-affinity transport activity, intragenic control of expression, and early flowering mutant phenotype. Plant Physiol 143:188–198
Spielmeyer W, Ellis MH, Chandler PM (2002) Semidwarf (sd-1), “green revolution” rice, contains a defective gibberellin 20-oxidase gene. Proc Natl Acad Sci USA 99:9043–9048
Srinivas G, Satish K, Madhusudhana R, Reddy RN, Mohan SM, Seetharama N (2009) Identification of quantitative trait loci for agronomically important traits and their association with genic-microsatellite markers in sorghum. Theor Appl Genet 118:1439–1454
Stephens JC, Miller FR, Rosenow DT (1967) Conversion of alien sorghum to early combine types. Crop Sci 7:396
Upadhyaya HD, Pundir RPS, Dwivedi SL, Gowda CLL, Reddy VG, Singh S (2009) Developing a mini core collection of sorghum for diversified utilization of germplasm. Crop Sci 49:1769–1780
Upadhyaya HD, Wang Y-H, Sharma S, Singh S (2012a) Association mapping of height and maturity across five environments using the sorghum mini core collection. Genome 55:471–479
Upadhyaya HD, Wang Y-H, Sharma S, Singh S, Hasenstein KH (2012b) SSR markers linked to kernel weight and tiller number in sorghum identified by association mapping. Euphytica 187:401–410
Wang ML, Zhu C, Barkley NL, Chen Z, Erpelding JE, Murray S, Tesso T, Pederson GA, Yu J (2009) Assessment of genetic diversity and population structure of accessions in the US sweet sorghum germplasm collection. Theor Appl Genet 120:13–23
Wang Y-H, Bible P, Loganantharaj R, Upadhyaya HD (2012) Identification of SSR markers associated with height using pool-based genome-wide association mapping in sorghum. Mol Breed 30:281–292
Wang Y-H, Upadhyaya HD, Burrell AM, Sahraeian SM, Klein RR, Klein PE (2013) Genetic structure and linkage disequilibrium in a diverse, representative collection of the C4 model plant, Sorghum bicolor. Genes, Genomes, Genetics (G3) doi: 10.1534/g3.112.004861
Yang X, Gao S, Xu S, Zhang Z, Prasanna BM, Li L, Li J, Yan J (2011) Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol Breed 28:511–526
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zheng LY, Guo XS, He B, Sun LJ, Peng Y, Dong SS, Liu TF, Jiang S, Ramachandran S, Liu CM, Jing HC (2011) Genome-wide patterns of genetic variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol 12:R114
Acknowledgments
We thank Patricia E. Klein, Robert R. Klein, and A. Millie Burrell for reviewing and the two reviewers for their suggestions to improve the manuscript. The study was supported in part by the International Crops Research Institute for the Semi-Arid Tropics and the University of Louisiana at Lafayette.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
Experiments conducted for this study comply with the current laws of India and the United States.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. E. Melchinger.
H. D. Upadhyaya and Y.-H. Wang have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Upadhyaya, H.D., Wang, YH., Gowda, C.L.L. et al. Association mapping of maturity and plant height using SNP markers with the sorghum mini core collection. Theor Appl Genet 126, 2003–2015 (2013). https://doi.org/10.1007/s00122-013-2113-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-013-2113-x