Abstract
Anthracnose in sorghum caused by Colletotrichum sublineolum is one of the most destructive diseases affecting sorghum production under warm and humid conditions. Markers and genes linked to resistance to the disease are important for plant breeding. Using 14,739 SNP markers, we have mapped eight loci linked to resistance in sorghum through association analysis of a sorghum mini-core collection consisting of 242 diverse accessions evaluated for anthracnose resistance for 2 years in the field. The mini-core was representative of the International Crops Research Institute for the Semi-Arid Tropics’ world-wide sorghum landrace collection. Eight marker loci were associated with anthracnose resistance in both years. Except locus 8, disease resistance-related genes were found in all loci based on their physical distance from linked SNP markers. These include two NB-ARC class of R genes on chromosome 10 that were partially homologous to the rice blast resistance gene Pib, two hypersensitive response-related genes: autophagy-related protein 3 on chromosome 1 and 4 harpin-induced 1 (Hin1) homologs on chromosome 8, a RAV transcription factor that is also part of R gene pathway, an oxysterol-binding protein that functions in the non-specific host resistance, and homologs of menthone:neomenthol reductase (MNR) that catalyzes a menthone reduction to produce the antimicrobial neomenthol. These genes and markers may be developed into molecular tools for genetic improvement of anthracnose resistance in sorghum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sorghum (Sorghum bicolor (L.) Moench) is a versatile crop as its grains can be used as food or feed and its stalks can be used to produce juice, as animal feed, and building materials. Anthracnose in sorghum is caused by the fungus Colletotrichum sublineolum. It was previously thought that anthracnose in maize and sorghum was both caused by C. graminicola. But now molecular evidences including genome sequencing have shown that C. graminicola causes anthracnose in maize, while C. sublineolum is the causal agent for anthracnose in sorghum (Vaillancourt and Hanau 1992; Sherriff et al. 1995; O’Connell et al. 2012). Anthracnose is one of the most destructive diseases affecting sorghum production world-wide, especially under warm and humid conditions (Thakur 2007) because the fungus can infect all aerial parts of the plant: the stalk, foliage, panicle, and grain, impacting the quality of both grain, and stalk although the most serious is foliar infection. Both resistant and susceptible genotypes display hypersensitive response upon infection, but only resistant genotypes can reduce pathogen growth through increased production of phytoalexins (Tenkouano et al. 1998). Loss of grain yield ranges from 47 to 67 % in susceptible varieties (Harris et al. 1964; Thomas et al. 1996; Pande et al. 2003; Gwary and Asala 2006), while the loss of stalk is lower at 23 % (Pande et al. 2003). Although fungicide spray may reduce the impact of the disease (Thomas et al. 1996; Gwary and Asala 2006), host resistance is the most economical means for control of anthracnose.
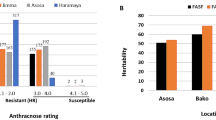
Resistance to C. sublineolum in sorghum is controlled by monogenes (Le Beau and Coleman 1950; Erpelding 2007). However, multiple sources of genetic resistance exist among different resistant genotypes, and resistance is controlled by single dominant or recessive genes depending on crosses (Mehta et al. 2000, 2005; da Costa et al. 2011). Interestingly, whether resistance is dominant or recessive also depends on pathotypes (races) of the pathogen used (da Costa et al. 2011), which means that a genotype resistant in one location may be susceptible in another. Thakur et al. (2007) evaluated 15 sorghum genotypes in 14 locations in India, Thailand, Ethiopia, Kenya, Zambia, Nigeria, and Mali for 4–7 years. They found that anthracnose severity varies significantly among genotypes, years, and locations, and that disease reaction was influenced by location. The differential reaction of sorghum accessions due to variation in virulence within pathogen populations presents challenges in breeding for durable anthracnose resistance (Marley et al. 2001; Valèrio et al. 2005). For example, 9 isolates from 8 locations in India belonged to 9 races (Pande et al. 1991); 34 isolates from 4 regions in Brazil were classified to 22 races (Valèrio et al. 2005), and 18 isolates from the US were found to be of 17 races (Prom et al. 2012). DNA fingerprinting of isolates from the US (Prom et al. 2012) and Ethiopia (Chala et al. 2011) found significant variation among isolates, suggesting that different races predominated at each location. For example, BTx378 was resistant to all 17 races tested by Prom et al. (2012), but for eight races tested by Cardwell et al. (1989), it was resistant only to races 1, 2, 4, and 8. In other words, BTx378 may be resistant to the pathogen in Orange Grove, TX, but not in Griffin, GA (Cardwell et al. 1989). Similar result was reported by Thakur et al. (2007) that IS 8354 can be completely resistant in Indore, India, but susceptible in Pantnagar, India or Mansa, Zambia. The differential reaction of resistant accessions to different races/pathotypes of the pathogen also suggests that resistance to C. sublineolum in sorghum is controlled by different genes in different genotypes.
Genetic mapping of resistance to anthracnose has been conducted. Boora et al. (1998) mapped the recessive resistance gene in SC326-6 with a RAPD marker. Another recessive resistance gene was mapped in G 73 also with RAPD markers, one at 3.26 cM (Singh et al. 2006a) and one at 6.03 cM (Singh et al. 2006b). A dominant resistance gene (Cg1) in SC748-5 was mapped with four AFLP markers by Perumal et al. (2009), one of which, Xtxa6227, was mapped within 1.8 cM and a simple sequence repeat (SSR) marker, Xtxp549, was mapped within 3.6 cM of the gene toward the end of sorghum linkage group LG-05 (Perumal et al. 2009). Anthracnose resistance was also mapped as a quantitative trait. Klein et al. (2001) mapped one anthracnose resistance QTL distal to the SSR marker Xtxp95 and plcolor on chromosome 6. Murali Mohan et al. (2010) mapped four loci using SSR markers: three on chromosome 6, one on chromosome 4, and one of the chromosome 6 QTL was between Xtxp95 and plcolor, close to the position mapped by Klein et al. (2001). cDNA-AFLP technique was used to identify transcripts differentially expressed in a resistant genotype from Uganda (Biruma et al. 2012). Among 126 identified genes, 15 were biotic stress related including two NB-LRR resistance proteins (Cs1A, Cs2A), a lipid transfer protein (SbLTP1), a zinc finger-like transcription factor (SbZnTF1), a rice defensin-like homolog (SbDEFL1), a cell death related protein (SbCDL1), and an unknown gene harboring a casein kinase 2-like domain (SbCK2). Down-regulation of these genes was shown to significantly compromise the resistance response. Cs1A and Cs2A genes are located in two different loci on chromosome 9 (Biruma et al. 2012).
Our goal is to provide more genetic markers that help identify anthracnose resistance genes which can then be used to develop more effective disease control measures. The approach was to use association mapping on a sorghum mini-core collection consisting of 242 accessions with SNP markers. The mini-core is the representative of ICRISAT’s world-wide sorghum landrace collection (Upadhyaya et al. 2009). We have genotyped the 242 accessions of the mini-core with 14,739 SNP markers (Wang et al. 2013). This paper presents markers closely linked to anthracnose phenotypes and potential resistance genes.
Materials and methods
Seed source
Seed of the 242 germplasm accessions of the sorghum mini-core (Upadhyaya et al. 2009) were obtained from the Genetic Resources Division, ICRISAT, Patancheru, India. The mini-core encompasses all five basic races [bicolor (20), caudatum (39), durra (30), guinea (29), and kafir (21)] and 10 intermediate races [caudatum-bicolor (30), durra-bicolor (7), durra-caudatum (19), guinea-bicolor (2), guinea-caudatum (27), guinea-durra (2), guinea-kafir (3), kafir-bicolor (2), kafir-caudatum (7), and kafir-durra (4)]. Races in sorghum are agronomic subtypes based on mostly panicle morphology.
Phenotyping for anthracnose reaction
Field evaluation of the mini-core for anthracnose reaction (resistant/susceptible) was as described previously (Sharma et al. 2012). In brief, the replicated screening was conducted in the anthracnose screening nursery at Patancheru, Andhra Pradesh, India, during the 2009 and 2010 rainy seasons. The C. sublineolum inoculum was isolated on potato dextrose agar (PDA) medium from the naturally infected leaves of susceptible line H 112 collected from sorghum fields at ICRISAT, Patancheru, India. The inoculum was multiplied on autoclaved sorghum grains. Plants were whorl-inoculated with infested sorghum grains at 3–4 grains/plant at 30 days after emergence. Anthracnose severity was rated at the soft-dough stage using a progressive 1–9 scale, where 1 = no disease and 9 = 76–100 % leaf area covered with lesions (Sharma et al. 2012). Plant height and days to 50 % of flowering were also measured. Anthracnose disease score data are in Supplementary Table S1.
Genotyping
In brief, 500 ng of genomic DNA was digested with the CpG methylation-sensitive restriction endonuclease FseI (New England Biolabs, Ipswich, MA) ligated with adapters containing 4 bp identifier tags and sequenced on an Illumina GAIIx (Illumina, San Diego, CA). Raw sequences were aligned to the sorghum genome sequence and analyzed for SNPs using the CLC Bio Genomics Workbench software (CLC Bio, Cambridge, MA) (Wang et al. 2013). SNPs were named based on the chromosome on which they mapped followed by the physical location in bp (e.g., SNP chr1_46978664, resides on SBI-01 at position 46,978,664 bp). Only markers scored in at least 50 % of the 242 accessions were retained. A total of 14,739 SNPs were developed and used in this study.
Data analysis
Marker-trait associations were calculated using two mixed linear models (MLM; Yu et al. 2006): kinship matrix (K model) and K with PCA matrix (K + P model) (Casa et al. 2008; Huang et al. 2010; Setter et al. 2011; Yang et al. 2011; Pasam et al. 2011) implemented in TASSEL 3.0 (Bradbury et al. 2007) available from http://www.maizegenetics.net/. The PCA matrix was performed in R (version 2.15.1, 64 bit; www.r-project.org/). SNP data was transformed manually, and PCA was performed using the prcomp() procedure in R. The K matrix was generated in TASSEL with 168 evenly distributed unlinked SNP markers. Significance of associations between markers and traits was based on threshold p < 6.78 × 10−6, a stringent Bonferroni correction calculated by dividing 0.10 (Setter et al. 2011) by 14,739, the total number of markers used in this study. However, associations with slightly higher p values (10−4 as in Famoso et al. 2011) were also presented to show association patterns, as p value alone may not always be the best indicator for the strength of an association (Heron et al. 2011).
Results
Mini-core evaluation for anthracnose reaction
Tremendous genetic variation for reaction to anthracnose pathogen has been demonstrated in the mini-core collection (Sharma et al. 2012). The resistance was also consistent between the testing environments with Pearson’s correlation coefficient of 0.96 between 2009 and 2010 resistance scores, the evidence that anthracnose resistance is mostly influenced by genes rather than environmental conditions. Resistance to anthracnose is under monogenic control as described previously (Le Beau and Coleman 1950; Coleman and Stokes 1954; Mehta et al. 2000, 2005; Erpelding 2007; da Costa et al. 2011), although disease scores approach normal distribution (Fig. 1; Murali Mohan et al. 2010).
It was also found that plant height was negatively correlated with anthracnose severity with the correlation coefficients of −0.28 and −0.34 for 2009 and 2010, respectively. This means taller plants showed less severe foliar symptoms as compared to shorter plants. Similarly, maturity as measured by the number of days to 50 % flowering was also negatively correlated with the disease severity, with the correlation coefficients of −0.62 and −0.61 for 2009 and 2010, respectively, indicating that late-maturing plants showed reduced foliar symptoms compared to early-maturing plants. All the above correlation coefficients were significant at p < 0.00001.
Association mapping
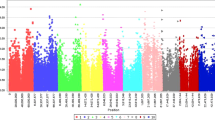
Comparison of K and K + P models showed that both produced identical p values (data not shown), suggesting that using one model was sufficient. Because of this, only results from K model were presented. To further minimize the possibility of false positive associations, other criteria in addition to the Bonferroni correction p value were also used. These were (a) the association had to be present in both environments and (b) multiple markers had to show similar strength of association in one locus whenever there were multiple markers in the same locus (Beló et al. 2008). Minimum p value for (b) was 10−4 as in Famoso et al. (2011). Based on criteria (a) and (b), we found eight marker loci associated with anthracnose resistance and two of these had the p values lower than the Bonferroni threshold (Fig. 2). Overall, associations were stronger in 2010 season than in 2009. But the trend was similar in both years as the data from both years were almost perfectly correlated as mentioned above.
Association p values of marker loci with anthracnose resistance in sorghum in 2009 and 2010 growing seasons. Panels a–h are loci 1–8, respectively. SNP markers are on X-axis. Dashed lines represent the p value of 10−4 and asterisks indicate p values lower than the Bonferroni threshold. On the leftmost and rightmost are markers flanking a locus in each panel. Flanking markers are not considered part of each locus. All markers analyzed in each locus are presented
Among the eight loci, two (loci 2 and 3) only contained one marker each and two (loci 4 and 6) had marker with association p values below that of the Bonferroni threshold. While loci 4 and 7 each consisted of two markers, locus 1 included three markers and loci 5 and 6 each contained seven and eight markers, respectively. For loci with more than one marker, 1, 4, 5, and 7 contained two or more markers all associated with the resistance phenotype with similar strength as measured by p values. Based on consistency of association strength across the testing environments and markers (when possible), loci 1, 2, 3, 4, and 5 showed strong linkage to the resistance phenotype (Fig. 2). Four of the eight loci (2, 3, 5, and 6) are on chromosome 1, two (loci 1 and 8) on chromosome 6, and loci 4 and 7 are located on chromosomes 8 and 10, respectively. Locus 8 only had one marker with p value lower than 10−4 (Fig. 2).
Resistance-related genes around linked marker loci
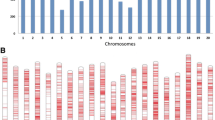
The physical distance used to localize anthracnose resistance-related genes was based on the size of linkage disequilibrium (LD) block in sorghum as in Famoso et al. (2011). LD blocks range from 10–15 (Hamblin et al. 2005) to 50–100 kb (Bouchet et al. 2012). So in most cases, only genes close to the marker loci are presented. In loci 1, 4, and 7, more than one homologs of a particular gene were found. These include four homologs of menthone:neomenthol reductase 1 (MNR1) in locus 1, three homologs of harpin-induced protein 1 (Hin1) in locus 4, and two NB-ARC (nucleotide-binding adaptor shared by APAF-1, R proteins, and CED-4) genes in locus 7. Other potential resistance-related genes are thaumatin in locus 2, a RAV transcription factor in locus 3, an oxysterol-binding protein in locus 5, and an autophagy-related protein in locus 6 (Fig. 3). No genes related to plant disease resistance were found in locus 8 (Table 1).
Annotated genes around marker loci associated with anthracnose resistance in sorghum. Loci 1–7 correspond to panels (a–g) in Fig. 1. See Table 1 for genes in Locus 8. All genes in the genomic region shown around each locus are presented. Gene location and annotation were from www.phytozome.net
The physical distance of the above genes to their respective linked loci also varied. In locus 1, it ranged from 9.1 kb for Sb06g023590 to 41 kb for Sb06g023560. In locus 2, the thaumatin gene was 19 kb from the linked marker and the RAV transcription factor was about 13 kb from chr1_72159383 in locus 3. In locus 4, the closest Hin1 homolog (Sb08g003710) was just 23 bp from the marker locus and farthest (Sb08g003690) was 14 kb. Oxysterol-binding protein in locus 5 was about 46 kb from the marker locus. In locus 6, autophagy-related protein 3 gene was about 2.6 kb from the linked markers. Finally, the two NB-ARC genes were 31.4 and 49.8 kb from the linked markers. These physical distances were within the LD sizes defined by Hamblin et al. (2005) and Bouchet et al. (2012).
Co-localization with previous mapped markers
To determine the effectiveness of association mapping of anthracnose resistance using the sorghum mini-core collection, we compared the mapped marker location to previously mapped anthracnose resistance loci and the most comprehensive of those is by Murali Mohan et al. (2010). Murali Mohan et al. (2010) mapped four anthracnose resistance loci and three of those are on chromosome 6: QAnt1 between Mrco and GlumeT (proximal to Xtxp145), QAnt2 between Fdnhsbm1 (distal to Xtxp274) and Fdnhsbm8 (proximal to Xtxp95), and QAnt3 between Xtxp95 and Xtxp95. QAnt3 was also mapped by Klein et al. (2001). The approximate physical location of the three QTL is shown in Fig. 4. Both SNP loci identified on chromosome 6 were colocalized with two of the three QTL mapped by Murali Mohan et al. (2010). Locus 8 was most likely covered by QAnt3 and locus 1 was close to QAnt2 (Fig. 4).
Approximate physical location in Mbp of sorghum anthracnose resistance QTL (dark rectangles to the left) and markers mapped in Murali Mohan et al. (2010) and the two SNP loci (1 and 8; arrows on the right) mapped in this study. Markers that can not be physically localized were not shown
Discussion
In this study, we used PCA as a covariate to measure population structure in the K + P model, replacing the Q matrix commonly calculated using STRUCTURE (Pritchard et al. 2000). PCA has been used in other studies (Famoso et al. 2011; Setter et al. 2011; Zhao et al. 2011) in association mapping. In our study, we found that K and K + P models produced identical p values which are the measure of association strength. Similar conclusions are also reached by Yang et al. (2011) and Pasam et al. (2011). This was the reason that only results from K model were presented in this study. However, such stringent models may over-correct for population structure and miss potential target genes (false negative association). For example, SNPs near height-controlling genes such as OsBAK1 on chromosome 8, DGL1 on chromosome 1 were only associated with height by naïve model without controlling for K, and both genes were missed by models controlled for K as demonstrated in rice (Zhao et al. 2011). Therefore, the flexible use of association p values (Famoso et al. 2011) and association mapping incorporating biological knowledge of the trait under study will provide more power to marker identification and QTL detection (Wang et al. 2010).
Markers mapped in this study colocalized with previously mapped anthracnose resistance genes. Two SNP loci mapped on chromosome 6 colocalized with anthracnose resistance QTL mapped by Murali Mohan et al. (2010) and by Klein et al. (2001), but we did not find markers linked to the resistance on chromosome 5 on which a dominant resistance gene (Cg1) in SC748-5 was mapped with four AFLP markers (Perumal et al. 2009). Singh et al. (2006a) mapped a recessive resistance gene on chromosome 8. However, a search of GenBank using BLASTN with their sequence did not identify any homologous sequence in sorghum. Therefore, the exact location of their marker could not be determined as was another mapped marker (Singh et al. 2006b) linked to anthracnose resistance.
All identified disease resistance-related genes potentially play a role in countering pathogen attack in sorghum. One mechanism in defense by plants is through the hypersensitive response (HR), the rapid death of plant cells at the site of pathogen infection. HR is triggered upon recognition of a pathogen avirulence (Avr) protein by a plant resistance (R) protein (Martin et al. 2003; Jones and Dangl 2006). Therefore, a restricted HR through rapid cell death is a hallmark of R protein-mediated resistance. Two NB-ARC R genes found in locus 7 on chromosome 10 (Sb10g021850 and Sb10g021860) were partially homologous to Pib (accession number BAA76281) which confers resistance to rice blast (Wang et al. 1999). Interestingly, the two only share about 20 % homology. A gene (Sb01g029070) coding for autophagy-related protein 3 (SbATG3) was closest to locus 6 markers. SbATG3 is 77 % identical and 85 % similar to the tobacco homolog ATG3 (AAW80629). Silencing ATG3 in tobacco resulted in unrestricted TMV-induced hypersensitive cell death due to increased pathogen propagation (Liu et al. 2005). In the R protein-mediated resistance pathway, Hin1 has been shown to mediate the elicitation of HR in the presence of Avr protein in tobacco and tomato (Gopalan et al. 1996) and is a well-known HR marker gene (Pontier et al. 1999). The sorghum homologs (Sb08g003690, Sb08g003705, Sb08g003710, and Sb08g003720; locus 4 in Fig. 3) were the most homologous in tobacco to tobacco Hin1 (CAA68848) and also shared homology with the Arabidopsis Hin1 homologs of AtNHL10, AtNHL3, and AtNHL1. AtNHL10 has been found to be induced during HR in response to infection (Zheng et al. 2004). Overexpression of AtNHL3 enhances resistance to infection by Pseudomonas syringae pv. tomato DC3000 in Arabidopsis (Varet et al. 2003).
Other genes in the R protein-mediated pathway were also found and these include a RAV transcription factor (Sb01g049150 in locus 3). The tomato RAV homolog mediates resistance to bacterial wilt caused by Ralstonia solanacearum and silencing RAV abolishes the resistance (Li et al. 2011). Similarly, the pepper RAV homolog also functions in disease resistance pathway and is rapidly induced by the bacterial pathogen Xanthomonas axonopodis pv. glycines 8ra (Kim et al. 2005; Sohn et al. 2006). Overexpression in Arabidopsis enhances resistance to infection by Pseudomonas syringae pv. tomato DC3000 and to osmotic stresses by high salinity and dehydration (Sohn et al. 2006). Defense genes independent of R protein-mediate pathway were also found. Sb01g010720 protein sequence in locus 5 was 35 % identical and 52 % similar to the potato oxysterol-binding protein StOBP1 (AAR25799). StOBP1 is rapidly induced by Phytophthora infestans infection and seems to be a component of the general, non-specific defense pathway (Avrova et al. 2004). In locus 1, four homologs of menthone:neomenthol reductase (MNR) gene were found. MNR catalyzes a menthone reduction with reduced nicotinamide adenine dinucleotide phosphate as a cofactor to produce the antimicrobial neomenthol. Silencing MNR in pepper (Capsicum annuum) significantly increases its susceptibility to Xanthomonas campestris pv vesicatoria and Colletotrichum coccodes infection (Choi et al. 2008). Another class of proteins with antimicrobial function in plants are thaumatin-like proteins (TLPs) coded by pathogenesis-related (PR)-5 genes such as PR-5 in rice (AAB53368) whose protein sequence shares 32 % identity and 43 % similarity with that of Sb01g021450 in locus 2 (Fig. 3). TLPs are sweet-tasting and can inhibit the growth of pathogenic fungi by permeabilizing the fungal cell membranes (Little et al. 2012). Overexpressing rice PR-5 enhances resistance to Rhizoctonia solani, the causal agent of sheath blight (Datta et al. 1999). In sorghum, protein expression level of one TLP, sormatin, correlates with resistance to grain mold (Rodríguez-Herrera et al. 1999; Bueso et al. 2000).
Finally, we also noticed in our data and in the literature that anthracnose resistance could be related to maturity, i.e., late-maturing plants showed reduced foliar symptoms compared to early-maturing plants. Similar results have also been reported in the literature. Hess et al. (2001) found among 19 sorghum varieties that early flowering was weakly, but significantly correlated with increased severity of foliar (r = 0.151, P < 0.05) and panicle anthracnose (r = 0.152, P < 0.01). Among 92 sorghum varieties reported by Wang et al. (2006), maturity was rated at a scale of 1–6, with 1 being the early and 6 being late-maturing, and anthracnose was rated in a scale of 1–5 with 1 being resistant and 5 being susceptible. The average maturity among the varieties was 4.3 with a variance of 2.44, while the average resistance was 1.5 with a variance of 0.86. This indicates that the group is mostly late-maturing and anthracnose resistant (Wang et al. 2006). Coincidentally, we also found a flowering time (SbFT1) homolog in locus in the opposite side of the two NB-ARC genes (Fig. 3). SbFT1 protein is 62 % identical and 77 % similar to rice Hd3a. Rice plants overexpress Hd3a flowers in 2 weeks compared to 7 weeks in the wild type (Tamaki et al. 2007). Therefore, if SbFT1 was associated with anthracnose resistance, it was because of the correlation between the two traits.
In conclusion, to identify markers linked to anthracnose resistance, we have performed association mapping in the sorghum mini-core collection of 242 accessions with 14,739 SNP markers. Eight marker loci (loci 1–8) were associated with anthracnose resistance in both environments. Annotated disease resistance-related genes were found in all loci based on their physical distance from linked SNP markers except in locus 8. These include two NB-ARC class of R genes, two HR-related genes, a transcription factor that functions in the R gene pathway, a gene that functions in the non-specific host resistance, and a gene for antimicrobial compound production. The results presented here will provide genes and markers for application in molecular breeding of anthracnose resistance in sorghum by developing PCR-based markers through characterizing the relevant genomic regions and further testing in additional varieties.
References
Avrova AO, Taleb N, Rokka VM, Heilbronn J, Campbell E, Hein I, Giloy EM, Cardle L, Bradshaw JE, Stewart HE, Fakim YJ, Loake G, Birch PR (2004) Potato oxysterol binding protein and cathepsin B are rapidly upregulated in independent defense pathways that distinguish R gene-mediated and field resistances to Phytophthora infestans. Mol Plant Pathol 5:45–56
Beló A, Zheng P, Luck S, Shen B, Meyer DJ, Li B, Tingey S, Rafalski A (2008) Whole genome scan detects an allelic variant of fad2 associated with increased oleic acid levels in maize. Mol Genet Genomics 279:1–10
Biruma M, Martin T, Fridborg I, Okori P, Dixelius C (2012) Two loci in sorghum with NB-LRR encoding genes confer resistance to Colletotrichum sublineolum. Theor Appl Genet 124:1005–1015
Boora KS, Frederiksen R, Magill C (1998) DNA-based markers for a recessive gene conferring anthracnose resistance in sorghum. Crop Sci 38:1708–1709
Bouchet S, Pot D, Deu M, Rami JF, Billot C, Perrier X, Rivallan R, Gardes L, Xia L, Wenzl P, Kilian A, Glaszmann JC (2012) Genetic structure, linkage disequilibrium and signature of selection in Sorghum: lessons from physically anchored DArT markers. PLoS One 7(3):e33470
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Bueso FJ, Waniska RD, Rooney WL, Bejosano FP (2000) Activity of antifungal proteins against mold in sorghum caryopses in the field. J Agri Food Chem 48:810–816
Cardwell KF, Hepperly PR, Frederiksen RA (1989) Pathotypes of Colletotrichumgraminicola and seed transmission of sorghum anthracnose. Plant Dis 73:255–257
Casa AM, Pressoir GH, Brown PJ, Mitchell SE, Rooney WL, Tuinstra MR, Franks CD, Kresovich S (2008) Community resources and strategies for association mapping in sorghum. Crop Sci 48:30–40
Chala A, Tronsmo AM, Brurberg MB (2011) Genetic differentiation and gene flow in Colletotrichum sublineolum in Ethiopia, the centre of origin and diversity of sorghum, as revealed by AFLP analysis. Plant Pathol 60:474–482
Choi HW, Lee BG, Kim NH, Park Y, Lim CW, Song HK, Hwang BK (2008) A role for a menthone reductase in resistance against microbial pathogens in plants. Plant Physiol 148:383–401
Coleman OH, Stokes IE (1954) The inheritance of resistance to stalk red rot in sorghum. Agron J 46:61–63
da Costa RV, Zambolim L, Cota LV, da Silva DD, Rodrigues JAS, Tardin FD, Casela CR (2011) Genetic control of sorghum resistance to leaf anthracnose. Plant Pathol 60:1162–1168
Datta K, Velazhahan R, Oliva N, Ona I, Mew T, Khush GS, Muthukrishnan S, Datta SK (1999) Over-expression of the cloned rice thaumatin-like protein (PR-5) gene in transgenic rice plants enhances environmental friendly resistance to Rhizoctonia solani causing sheath blight disease. Theor Appl Genet 98:1138–1145
Erpelding JE (2007) Inheritance of anthracnose resistance for the sorghum cultivar redlan. Plant Pathol J 6:187–190
Famoso AN, Zhao K, Clark RT, Tung CW, Wright MH, Bustamante C, Kochian LV, McCouch SR (2011) Genetic architecture of aluminum tolerance in rice (Oryza sativa) determined through genome-wide association analysis and QTL mapping. PLoS Genet 7:e1002221
Gopalan S, Wei W, He SY (1996) hrp gene-dependent induction of hin1: a plant gene activated rapidly by both harpins and the avrPto gene-mediated signal. Plant J 10:591–600
Gwary DM, Asala SW (2006) Cost-benefit of fungicidal control of anthracnose on sorghum in Northern Nigeria. Intl J Agri Biol 8:306–308
Hamblin MT, Salas Fernandez MG, Casa AM, Mitchell SE, Paterson AH, Kresovich S (2005) Equilibrium processes cannot explain high levels of short- and medium-range linkage disequilibrium in the domesticated grass Sorghum bicolor. Genetics 171:1247–1256
Harris HB, Johnson BJ, Dobson JW Jr, Luttrell ES (1964) Evaluation of anthracnose on grain sorghum. Crop Sci 4:460–462
Heron EA, O’dushlaine C, Segurado R, Gallagher L, Gill M (2011) Exploration of empirical Bayes hierarchical modeling for the analysis of genome-wide association study data. Biostatistics 12:445–461
Hess DE, Bandyopadhyay R, Sissoko I (2001) Reactions of sorghum genotypes to leaf, panicle and grain anthracnose (Colletotrichum graminicola) under field conditions in Mali. Towards Sustainable Sorghum Production, Utilization, and Commercialization in West Africa. West and Central African Sorghum Research Network (WCASRN), Bamako, Mali, 163–178. International Crops Research Institute for the Semi-Arid Tropics, Patancheru, Andhra Pradesh, India
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y, Li C, Zhu C, Lu T, Zhang Z, Li M, Fan D, Guo Y, Wang A, Wang L, Deng L, Li W, Lu Y, Weng Q, Liu K, Huang T, Zhou T, Jing Y, Li W, Lin Z, Buckler ES, Qian Q, Zhang QF, Li J, Han B (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Kim SY, Kim YC, Lee JH, Oh SK, Chung E, Lee S, Lee YH, Choi D, Park JM (2005) Identification of a CaRAV1 possessing an AP2/ERF and B3 DNA-binding domain from pepper leaves infected with Xanthomonas axonopodis pv. glycines 8ra by differential display. Biochim Biophys Acta 1729:141–146
Klein RR, Rodriguez-Herrera R, Schlueter JA, Klein PE, Yu ZH, Rooney WL (2001) Identification of genomic regions that affect grain-mould incidence and other traits of agronomic importance in sorghum. Theor Appl Genet 102:307–319
Le Beau FJ, Coleman OH (1950) The inheritance of resistance in sorghum to leaf anthracnose. Agron J 42:33–34
Li CW, Su RC, Cheng CP, Sanjaya You SJ, Hsieh TH, Chao TC, Chan MT (2011) Tomato RAV transcription factor is a pivotal modulator involved in the AP2/EREBP-mediated defense pathway. Plant Physiol 156:213–227
Little C, Perumal R, Tesso T, Prom LK, Odvody GN, Magill CW (2012) Sorghum pathology and biotechnology-A fungal disease perspective: part I. Grain mold, head smut, and ergot. Eur J Plant Sci Biotech 6:10–30
Liu Y, Schiff M, Czymmek K, Talloczy Z, Levine B, Dinesh-Kumar SP (2005) Autophagy regulates programmed cell death during the plant innate immune response. Cell 121:567–577
Marley PS, Thakur RP, Ajayi O (2001) Variation among foliar isolates of Colletotrichum sublineolum of sorghum in Nigeria. Field Crops Res 69:133–142
Martin GB, Bogdanove AJ, Sessa G (2003) Understanding the functions of plant disease resistance proteins. Ann Rev Plant Biol 54:23–61
Mehta PJ, Collins SD, Rooney WL, Frederiksen RA, Klein RR (2000) Identification of different sources of genetic resistance to anthracnose in sorghum. Int Sorghum Millets Newsl 41:51–54
Mehta PJ, Wiltse CC, Rooney WL, Collins SD, Frederiksen RA, Hess DE, Chisi M, TeBeest DO (2005) Classification and inheritance of genetic resistance to anthracnose in sorghum. Field Crops Res 93:1–9
Murali Mohan S, Madhusudhana R, Mathur K, Chakravarthi DVN, Rathore S, Nagaraja Reddy R, Satish S, Srinivas G, Sarada Mani N, Seetharama N (2010) Identification of quantitative trait loci associated with resistance to foliar diseases in sorghum [Sorghum bicolor (L.) Moench]. Euphytica 176:199–211
O’Connell RJ, Thon MR, Hacquard S, Amyotte SG, Kleemann J, Torres MF, Damm U, Buiate EA, Epstein L, Alkan N, Altmüller J, Alvarado-Balderrama L, Bauser CA, Becker C, Birren BW, Chen Z, Choi J, Crouch JA, Duvick JP, Farman MA, Gan P, Heiman D, Henrissat B, Howard RJ, Kabbage M, Koch C, Kracher B, Kubo Y, Law AD, Lebrun MH, Lee YH, Miyara I, Moore N, Neumann U, Nordström K, Panaccione DG, Panstruga R, Place M, Proctor RH, Prusky D, Rech G, Reinhardt R, Rollins JA, Rounsley S, Schardl CL, Schwartz DC, Shenoy N, Shirasu K, Sikhakolli UR, Stüber K, Sukno SA, Sweigard JA, Takano Y, Takahara H, Trail F, van der Does HC, Voll LM, Will I, Young S, Zeng Q, Zhang J, Zhou S, Dickman MB, Schulze-Lefert P, Loren Ver, van Themaat E, Ma LJ, Vaillancourt LJ (2012) Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat Genet 44:1060–1065
Pande S, Mughogho LK, Bandyopadhyay R, Karunakar RI (1991) Variation in pathogenicity and cultural characteristics of sorghum isolates of Colletotrichum graminicola in India. Plant Dis 75:778–783
Pande S, Bandyopadhyay R, Blümmel M, Narayana Rao J, Thomas D, Navi SS (2003) Disease management factors influencing yield and quality of sorghum and groundnut crop residues. Field Crops Res 84:89–103
Pasam RK, Sharma R, Malosetti M, van Eeuwijk FA, Haseneyer G, Kilian B, Graner A (2011) Genome-wide association studies for agronomical traits in a world wide spring barley collection. BMC Plant Biol 12:16
Perumal R, Menz M, Mehta P, Katilé S, Gutierrez-Rojas L, Klein R, Klein P, Prom L, Schlueter J, Rooney W, Magill C (2009) Molecular mapping of Cg1, a gene for resistance to anthracnose (Colletotrichum sublineolum) in sorghum. Euphytica 165:597–606
Pontier D, Gan S, Amasino RM, Roby D, Lam E (1999) Markers for hypersensitive response and senescence show distinct patterns of expression. Plant Mol Biol 39:1243–1255
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Prom LK, Perumal R, Erattaimuthu SR, Little CR, No EG, Erpelding JE, Rooney WL, Odvody GN, Magill CW (2012) Genetic diversity and pathotype determination of Colletotrichum sublineolum isolates causing anthracnose in sorghum. Eur J Plant Path 133:671–685
Rodríguez-Herrera R, Waniska RD, Rooney WL (1999) Antifungal proteins and grain mold resistance in sorghum with nonpigmented testa. J Agri Food Chem 47:4802–4806
Setter TL, Yan J, Warburton M, Ribaut JM, Xu Y, Sawkins M, Buckler ES, Zhang Z, Gore MA (2011) Genetic association mapping identifies single nucleotide polymorphisms in genes that affect abscisic acid levels in maize floral tissues during drought. J Exp Bot 62:701–716
Sharma R, Upadhyaya HD, Manjunatha SV, Rao VP, Thakur RP (2012) Resistance to foliar diseases in a mini-core collection of sorghum germplasm. Plant Dis 96:1629–1633
Sherriff C, Whelan MJ, Arnold GM, Bailey JA (1995) rDNA sequence analysis confirms the distinction between Colletotrichum graminicola and C. sublineolum. Mycological Res 99:475–478
Singh M, Chaudhary K, Singal HR, Magill CW, Boora KS (2006a) Identification and characterization of RAPD and SCAR markers linked to anthracnose resistance gene in sorghum [Sorghum bicolor (L.) Moench]. Euphytica 149:179–187
Singh M, Chaudhary K, Boora KS (2006b) RAPD-based SCAR marker SCA 12 linked to recessive gene conferring resistance to anthracnose in sorghum [Sorghum bicolor (L.) Moench]. Theor Appl Genet 114:187–192
Sohn KH, Lee SC, Jung HW, Hong JK, Hwang BK (2006) Expression and functional roles of the pepper pathogen-induced transcription factor RAV1 in bacterial disease resistance, and drought and salt stress tolerance. Plant Mol Biol 61:897–915
Tamaki S, Matsuo S, Wong HL, Yokoi S, Shimamoto K (2007) Hd3a protein is a mobile flowering signal in rice. Science 316:1033–1036
Tenkouano A, Miller FR, Fredericksen RA, Nicholson RL (1998) Ontogenetic characteristics and inheritance of resistance to leaf anthracnose in sorghum. Afri Crop Sci J 6:249–258
Thakur RP (2007) Anthracnose. In: Thakur RP, Reddy BVS, Mathur K (eds) Screening techniques for sorghum diseases. Information Bulletin No. 76. ICRISAT, Andhra Pradesh, p 53–57
Thakur RP, Rao VP, Wu SM, Subbarao KV, Mathur K, Tailor HC, Kushwaha US, Dwivedi RR, Krisnaswamy R, Hiremath RV, Indira S (2007) Genetic resistance to foliar anthracnose in sorghum and pathogenic variability in Colletotrichum graminicola. Indian Phytopath 60:13–23
Thomas MD, Sissoko I, Sacko M (1996) Development of leaf anthracnose and its effect on yield and grain weight of sorghum in West Africa. Plant Dis 80:151–153
Upadhyaya HD, Pundir RPS, Dwivedi SL, Gowda CLL, Reddy VG, Singh S (2009) Developing a mini core collection of sorghum for diversified utilization of germplasm. Crop Sci 49:1769–1780
Vaillancourt LJ, Hanau RM (1992) Genetic and morphological comparisons of Glomerella (Colletotrichum) isolates from maize and from sorghum. Exp Mycol 16:219–229
Valèrio H, Rèsende M, Weikert-Oliveira R, Casela C (2005) Virulence and molecular diversity in Colletotrichum graminicola from Brazil. Mycopathologia 159:449–459
Varet A, Hause B, Hause G, Scheel D, Lee J (2003) The Arabidopsis NHL3 gene encodes a plasma membrane protein and its overexpression correlates with increased resistance to Pseudomonas syringae pv. tomato DC3000. Plant Physiol 132(4):2023–2033
Wang ZX, Yano M, Yamanouchi U, Iwamoto M, Monna L, Hayasaka H, Katayose Y, Sasaki T (1999) The Pib gene for rice blast resistance belongs to the nucleotide binding and leucine-rich repeat class of plant disease resistance genes. Plant J 19:55–64
Wang ML, Dean R, Erpelding J, Pederson G (2006) Molecular genetic evaluation of sorghum germplasm differing in response to fungal diseases: rust (Puccinia purpurea) and anthracnose (Collectotrichum graminicola). Euphytica 148:319–330
Wang K, Li M, Hakonarson H (2010) Analysing biological pathways in genome-wide association studies. Nat Rev Genet 11:843–854
Wang Y-H, Upadhyaya HD, Burrell AM, Sahraeian SME, Klein RR, Klein PE (2013) Genetic structure and linkage disequilibrium in a diverse, representative collection of the C4 model plant, Sorghum bicolor. G3: Genes Genomes Genet. doi:10.1534/g3.112.004861
Yang X, Gao S, Xu S, Zhang Z, Prasanna BM, Li L, Li J, Yan J (2011) Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol Breeding 28:511–526
Yu JM, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhao K, Tung CW, Eizenga GC, Wright MH, Ali ML, Price AH, Norton GJ, Islam MR, Reynolds A, Mezey J, McClung AM, Bustamante CD, McCouch SR (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467
Zheng MS, Takahashi H, Miyazaki A, Hamamoto H, Shah J, Yamaguchi I, Kusano T (2004) Up-regulation of Arabidopsis thaliana NHL10 in the hypersensitive response to Cucumber mosaic virus infection and in senescing leaves is controlled by signalling pathways that differ in salicylate involvement. Planta 218:740–750
Acknowledgments
This study was supported in part by the International Crops Research Institute for the Semi-Arid Tropics and the University of Louisiana at Lafayette. We thank Karl H. Hasenstein and two reviewers for suggestions to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. Godwin.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Upadhyaya, H.D., Wang, YH., Sharma, R. et al. Identification of genetic markers linked to anthracnose resistance in sorghum using association analysis. Theor Appl Genet 126, 1649–1657 (2013). https://doi.org/10.1007/s00122-013-2081-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-013-2081-1