Abstract
Forage quality combines traits related to protein content and energy value. High-quality forages contribute to increase farm autonomy by reducing the use of energy or protein-rich supplements. Genetic analyses in forage legume species are complex because of their tetraploidy and allogamy. Indeed, no genetic studies of quality have been published at the molecular level on these species. Nonetheless, mapping populations of the model species M. truncatula can be used to detect QTL for forage quality. Here, we studied a crossing design involving four connected populations of M. truncatula. Each population was composed of ca. 200 recombinant inbred lines (RIL). We sought population-specific QTL and QTL explaining the whole design variation. We grew parents and RIL in a greenhouse for 2 or 3 seasons and analysed plants for chemical composition of vegetative organs (protein content, digestibility, leaf-to-stem ratio) and stem histology (stem cross-section area, tissue proportions). Over the four populations and all the traits, QTL were found on all chromosomes. Among these QTL, only four genomic regions, on chromosomes 1, 3, 7 and 8, contributed to explaining the variations in the whole crossing design. Surprisingly, we found that quality QTL were located in the same genomic regions as morphological QTL. We thus confirmed the quantitative inheritance of quality traits and tight relationships between quality and morphology. Our findings could be explained by a co-location of genes involved in quality and morphology. This study will help to detect candidate genes involved in quantitative variation for quality in forage legume species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Feeding value is of major importance in forage crops because it contributes to meat or milk production by ruminants. When high-quality forages are available for the animal diet, the use of protein and energy-rich supplements is limited. These supplements are costly and their purchase reduces the autonomy of the farm. Besides agronomic practices that may improve forage quality, genetic progress is required. In legume forage such as alfalfa (Medicago sativa), when a breeder includes quality traits in its selection criteria, he focuses on the energy value and the protein content. These traits have a quantitative inheritance and their heritability has moderate values (Neff and Simon 1986; Julier et al. 2000; Guines et al. 2002). The genetic analysis of quantitative traits is complex in alfalfa because of its heterozygosity and autotetraploidy. The use of a related model species (M. truncatula) that has recently been sequenced (Young et al. 2011), is expected to provide information on genetic inheritance of traits of agronomic interest (Young and Udvardi 2009).
Feeding value has been extensively studied in alfalfa. It results from the combination of plant architecture, stem histology and plant composition. It is well known that energy value (or digestibility) and protein content are explained by the leaf-to-stem ratio in the forage (the leaves having higher energy value and protein content than the stems) and by the digestibility of the stem that decreases as it elongates (Mowat et al. 1965; Terry and Tilley 1969; Heinrichs et al. 1969).
A stem consists in a succession of phytomers, each constituted of one internode and the corresponding leaf. The development of the vascular system during internode maturation is characterised by secondary wall thickening, deposition of polysaccharides such as hemicellulose and cellulose and finally lignification of xylem and primary phloem (Wilson 1993; Vallet et al. 1996; Engels and Jung 1998; Jung and Engels 2002), thus resulting in the decrease of stem digestibility (Jung and Engels 2002). Young internodes contain non-lignified tissues whereas mature internodes, whose proportion in the stem increases as plant grows, have lignified tissues. Genetic variation in stem anatomy has been reported in alfalfa (Guines et al. 2003; Julier et al. 2008). It was proposed that reduction of xylem proportion in the stem and increase of the proportion of non-lignified tissues such as pith parenchyma and cortex could be targets to improve forage quality in this forage legume (Guines et al. 2003). Pith parenchyma, in the middle of the stem cross-section, has a low cell-wall density. Contrastingly, the cortex that includes tissues between epidermis and cambium has a high cell-wall density.
At the chemical level, energy value is often measured by the digestibility (or solubility) of dry or organic matter in laboratory conditions (Aufrère 1982). The plant cell wall is a complex biological structure. Its content and its composition are measured because they contribute to the fate of the polymers during digestion. The van Soest analysis (Goering and Van Soest 1970) gives access to the total cell wall, or Neutral Detergent Fiber content (NDF, hemicellulose, cellulose and lignin, except pectins), the Acid Detergent Fiber content (ADF, cellulose and lignin) and the Acid Detergent Lignin content (ADL, lignin). Lignin is a major component of the cell wall that is recognized as limiting digestion of the wall polysaccharides in the rumen (Jung et al. 1994). It reinforces plant cell walls, providing rigidity, impermeability to water, and protection against pathogens but also reduces the accessibility of the microbes of the rumen to the cell walls (Wilson 1993; Vallet et al. 1996; Engels and Jung 1998, 2005; Jung and Engels 2002). It has been observed that a decrease in digestibility with increased stem maturity was correlated to an increase in lignin content and an increase in the ratio of syringyl/guaiacyl, the two major methylated and di-methylated, respectively, monomers that form lignin polymer (Guo et al. 2001a). Over a set of genotypes, digestibility was correlated to cell-wall contents (NDF, ADF and ADL) (Julier et al. 2000). A large genetic variation for digestibility or cell-wall contents was established among varieties (Heinrichs et al. 1969; Buxton et al. 1987; Lenssen et al. 1991; Julier et al. 1996) and more strikingly among genotypes (Julier et al. 2000). The correlation between forage yield and digestibility is negative, especially during plant growth (Lemaire and Allirand 1993; Julier and Huyghe 1997), but the genetic variation around this correlation is large enough to select high-yielding varieties with high-energy value (Julier et al. 2000).
The lignin biosynthesis pathway is composed of many genes with complex regulatory control (Boerjan et al. 2003; Boudet et al. 2003; Zhao and Dixon 2011). Some key genes were identified and targeted to downregulate them. For example, reduction of syringyl monomer content in lignin, obtained by downregulation of COMT (caffeic acid 3-O-methyltransferase), reduced lignin content in alfalfa stems and increased digestibility (Guo et al. 2001b), but a greater improvement in digestibility was attained with a downregulation of CCoAOMT (caffeoyl coenzyme A 3-O-methyl transferase) that induced a decrease in lignin content without change in syringyl content (Guo et al. 2001b). Downregulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase (HCT) induced a reduction in fiber content and an increase in dry matter digestibility (Shadle et al. 2007).
A quantitative inheritance of quality traits was expected: numerous genes of lignin pathway and regulation may explain genetic variation for forage quality and growth, itself quantitatively inherited, influences forage quality. Traits related to feeding value in alfalfa can easily be measured in M. truncatula and they have a similar biological meaning (Schnurr et al. 2007). QTL approaches aim at identifying the regions of the genome that explain phenotypic variation in the traits. In this paper, we present QTL detection in the model species M. truncatula for traits related to energy value, protein content and stem histological traits. A crossing design composed of four connected mapping populations made from four unrelated parental lines was used. This design has already been studied for QTL detection of morphogenetic traits (Lagunes Espinoza et al. 2012). It allows a greater precision for QTL positioning (Jourjon et al. 2005) and fine mapping of candidate genes annotated on the M. truncatula sequence (Young et al. 2011).
Materials and methods
RIL populations
Four RIL populations of M. truncatula that share common parents were used. Populations LR1 (233 RILs), LR4 (199 RILs), LR5 (173 RILs) and LR6 (179 RILs) were, respectively, obtained from the crosses DZA315.26 × DZA45.6, Jemalong6 × DZA315.16, Jemalong6 × F83005.5 and Jemalong6 × A20. DZA315.26 and DZA45.6 originate from Algeria, F83005.5 from France, and Jemalong6 and A20 from Australia. DZA315.26 and DZA315.16 lines were extracted from the same population and seem phenotypically identical, especially for flowering date (Pierre et al. 2008). All experiments were conducted in a greenhouse in Lusignan (France) in 2002, 2003 and 2004 for the LR4 population, spring and autumn 2005 for the LR1 and LR5 populations and 2007 and 2008 for the LR6 population. The four parents were included in each experiment.
Plant cultivation and sample analysis
Seeds of RIL populations were scarified and sown into Petri dishes, imbibed for 24 h at room temperature and vernalized at 4 °C for 48 h. Germinated seeds were transplanted in individual pots and grown in greenhouse in INRA Lusignan (France); the experimental conditions were described in previous publications (Julier et al. 2007; Pierre et al. 2008; Lagunes Espinoza et al. 2012). Each replication was composed of one plant (Table 1), to maximise the number of genotypes under study, as recommended to optimise QTL detection. For LR4 in 2002 and 2003 years, the RILs were arranged in a randomized complete block design with three repetitions. For the other populations or for LR4 in 2004, 15 RILs per population and the parental lines were randomly taken and repeated three times. The repeated lines and the non-repeated ones were randomly displayed in a single experimental design.
When all the lines had flowered and before onset of senescence, all RILs were individually harvested. In 2002 and 2003, on LR4 population, the two longest primary branches were harvested and leaves and stem were separated. In all other experiments, the whole plant was harvested. Samples were dried in an oven at 60 °C and ground to pass through a 1-mm sieve and weighed. In 2002 and 2003, leaf-to-stem ratio was calculated as the ratio between the leaf and the stem dry weights. Leaf-to-stem ratio is an estimate of forage quality because leaves have a higher protein content and digestibility than stems. Enzymatic solubility (Aufrère 1982), hereafter called digestibility, of stems and of leaves and protein content (Dumas method) of stems and of leaves were predicted by near infrared spectroscopy (NIRS) with equations developed at INRA Lusignan (Julier et al. 2000). The standard error of cross validation and the coefficient of determination of the equations were 0.86 and 0.98 for ADF content, 1.58 and 0.96 for digestibility, 0.51 and 0.99 for protein content, and 0.13 and 0.79 for leaf-to-stem ratio, respectively. The whole plant digestibility and protein content were estimated from leaf and stem composition, taking into account the dry weight of leaves and stems. In the other years, digestibility, ADF (Acid Detergent Fibre) content (Goering and Van Soest 1970; Aufrère 1982) and protein content of whole plant and leaf-to-stem ratio were predicted by NIRS. A subset of about 5 % of the samples was analysed in wet chemistry each year to verify the NIRS predictions.
The histological structure of the stem was analysed on LR4 population in 2002 and 2003. A 2-cm long segment was sampled at the mid-point of the basal portion of two mature primary branches per plant considered as the most lignified stem portions (Julier et al. 2008). Each segment was fixed in a glacial acetic acid/95° ethanol fixative solution. A histological analysis was performed according to the methodology used by Guines et al. (2003). Fifty micrometer stem cross-sections performed using a Vibratome® (series 1000) were stained with Fasga (Tolivia and Tolivia 1987) staining lignin in red and cellulose in blue. Sections were mounted on slides in distilled water and examined by stereo microscopy with 30× and 80× magnification, and by light microscopy with 200× magnification. Colour image analysis of stained stem cross-section was carried out with a video camera (3CCD colour camera, CV-M90) installed on a microscope and on a stereomicroscope. A semi-automated image analysis was developed using the 6.1 version of Optimas™ (Media Cybernetics 1996). In the area of the whole histological stem cross-section, the proportions of cortex, of xylem and of pith parenchyma were calculated and expressed as a ratio to the radius of the stem cross-section (Guines et al. 2003).
Analyses of variance were performed for each RIL population in each season or year using the GLM procedure of SAS (SAS Institute Inc. 2000). For LR4 in 2002 and 2003, the effects of genotypes and repetitions were tested for all recorded traits. For the other experiments, only the effect of genotypes was tested. Variances of the random effects (genotype and error) were estimated with the VARCOMP procedure of SAS. Broad sense heritability of each trait was calculated as \( h^{2} = \frac{{\sigma_{\text{L}}^{2} }}{{\sigma_{\text{L}}^{2} + \sigma_{\text{R}}^{2} /b}} \), with \( \sigma_{\text{L}}^{2} \) = variance of lines, considered as random effect, \( \sigma_{\text{R}}^{2} \) = variance of error and b = number of repetitions. Means were calculated for the repeated lines in each experiment. For the non-repeated lines, the raw data were used. Correlations among traits were calculated using the CORR procedure of SAS, on the means of RIL per population. As morphological traits were also recorded in these experiments (Lagunes Espinoza et al. 2012), correlation between quality and morphological traits (length of primary branches, branch elongation rate and flowering time) was calculated.
QTL mapping
Framework maps were already available (Lagunes Espinoza et al. 2012) and comprised between 60 and 62 SSR markers each. QTL mapping was performed using the method of composite interval mapping (CIM) implemented in QTL Cartographer (Basten et al. 1994; Basten et al. 2002). The threshold for adding a QTL, determined at 5 % risk by a permutation test method (1,000 replications), corresponded to LOD ≥ 2.46. QTL positions were estimated where the LOD score reached its maximum in the region under consideration. A LOD support interval was constructed for each QTL based on a LOD drop-off equal to 1 (Lander and Botstein 1989).
The BioMercator software (Arcade et al. 2004) was used to draw the QTL on the map of each population and to build a consensus map by iterative projection of loci using a homothetic function (Lagunes Espinoza et al. 2012). A multi-population QTL analysis performed with MCQTL software package (Jourjon et al. 2005) was carried out to better estimate the position of the QTL that were common to different populations and their effects. Adjusted means of the RIL (Blanc et al. 2006) and the consensus map were used to launch the multi-population QTL analysis with the “connected” option. An additive connected model was chosen, with the iterative QTL mapping method (iQTLm) using genetic cofactors, and a windows size of 5.5 cm. Cofactor selection and test of QTL effects were performed with F test. F thresholds were determined with 1,000 permutations to correspond to a global type I risk of 1 % (across all populations and the total genome).
Mapping of candidate gene related to lignin biosynthesis
The candidate genes described to be involved in lignin biosynthesis were listed from literature (Boerjan et al. 2003; Boudet et al. 2003; Zhao and Dixon 2011). Their names were used on http://medicago.jcvi.org/, in the page Putative function name of the Search tab, to identify M. truncatula contigs that contained genes annotated by these names. Each contig number was then entered in the Keywords Search on http://www.medicagohapmap.org/ to obtain the BAC number. The position of the BAC on the M. truncatula genome assembly version 3.5 was obtained on the contig viewer proposed by the UMN Assembly Browser. This position was compared to the position of the markers of our map on the genome and summarized on the map (Fig. 2) to analyse co-location with QTL.
Results
Genetic variation among RILs of the four mapping population
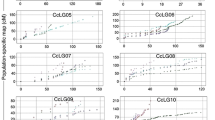
Significant differences among lines in the populations LR5 (except for protein content in autumn) and LR4 (years 2002 and 2003) were observed for forage quality and stem histology (Supplementary Materials 1, 2, 3 and 4). There were no significant differences among lines in LR1 and LR4 in 2004. Broad-sense heritability ranged from 0.62 to 0.88 for traits with a significant genotype effect (Tables 2, 3). Similar values and ranges of variation in all the populations and experiments were observed for ADF, whole plant, stem and leaf digestibility (Table 4). Compared to the other experiments, whole plant protein content and leaf-to-stem ratio were lower in LR4 in 2002 and 2003, probably because only two mature stems were harvested instead of the whole plant in the other experiments. However, large transgressive variations were observed for each of these traits in the RIL populations, indicating that positive and negative alleles are shared between the parents in each RIL population (Fig. 1; Table 5).
Correlations among quality, histological and morphological traits
In all populations, quality traits were highly correlated among themselves (Table 6). Digestibility of whole plant was strongly correlated to plant fiber content (ADF), with correlation values between −0.95 and −0.97. Leaf-to-stem ratio was correlated to both protein content and digestibility of whole plant. Protein content and digestibility were positively correlated. In the LR4 population evaluated in 2002 and 2003, whole plant digestibility was correlated to stem digestibility, but whole plant protein content was correlated to both leaf and stem protein contents (Table 7). Leaf-to-stem ratio was positively correlated to protein content and digestibility of whole plant. Stem width, measured by the area of the whole stem cross section, was negatively correlated to protein content and digestibility of whole plant, but the correlation was not very high. Stem digestibility and stem protein content were positively correlated to the proportion of cortex. The other histological traits were weakly correlated to chemical composition. Stem width was positively correlated to the proportion of parenchyma but negatively correlated to the proportion of cortex, and the proportion of xylem was negatively correlated to the proportion of parenchyma. Consequently, plants with a high stem digestibility had stems with a high proportion of cortex but this increase of cortex proportion was mainly observed in thin stemmed plants.
Most correlations between morphology and quality traits were significant (Table 8). The length of primary branches or their elongation rate was negatively correlated to the whole plant digestibility (and positively correlated to ADF content), protein content and leaf-to-stem ratio. As the length of primary branches was negatively correlated to flowering time, flowering time was conversely positively correlated to digestibility, protein content and leaf-to-stem ratio.
QTL analysis
For QTL detection, all traits were analysed, even when their heritability was low. With a composite interval mapping procedure implemented on each population, a total of 86 QTL were identified across years or seasons in the four RIL populations for quality traits (Supplementary Material 5, Fig. 2), and 20 QTL for stem histology (Supplementary Material 6, Fig. 3). Sixteen, 29, 18 and 13 QTL were revealed in the LR1, LR4, LR5 and LR6 population, respectively.
QTL positions and interval confidences for quality traits on a consensus map of LR1, LR4, LR5 and LR6 RIL populations of M. truncatula. Gene positions are indicated by blue arrows. CAD Cinnamyl alcohol dehydrogenase, CCoAOMT caffeoyl CoA O-methyltransferase, COMT caffeic acid 3-O-methyltransferase, CCR cinnamoyl CoA reductase, HCT hydroxycinnamoyl CoA shikimate/quinate hydroxycinnamoyltransferase, F5H ferulate 5-hydroxylase, PAL phenylalanine ammonia-lyase (color figure online)
Seventeen QTL for ADF were detected on all chromosomes except chromosome 3, corresponding to nine genomic regions in the four RIL populations. They accounted for 7.0–23.7 % of the genetic variation. Additive effects of Jemalong6 alleles were positive in the LR4 and LR5 populations and negative in the LR6 population. DZA315.26 alleles in the LR1 RIL population showed either positive or negative effects.
Eighteen QTL were detected for whole plant digestibility on chromosomes 1, 4, 5, 6, 7 and 8, corresponding to eight genomic regions. Genetic variation explained by each QTL accounted for 5.3–33.1 % of the variation. Each parent had alleles contributing positively or negatively to the variation.
In the LR4 population, six QTL were detected for stem digestibility on chromosomes 1 and 7, corresponding to two genomic regions. Positive and negative effects were shown by Jemalong6. In this same population, seven QTL, on chromosomes 3, 4, 5 and 7, controlled leaf digestibility. QTL accounted for 8.9–20.1 % of the variation. The effect of the Jemalong6 allele was positive for each QTL.
Nine QTL controlled whole plant protein content on chromosomes 1, 2, 3, 7 and 8, corresponding to six genomic regions. QTL on these regions explained from 2.9 to 12.9 % of genetic variation. Jemalong6 alleles induced negative effects in LR4 and LR5 populations but a positive effect in LR6 population. DZA315.26 allele induced negative or positive effects in LR1 population. In LR4 population, leaf protein content was controlled by two QTL, on chromosomes 1 and 8, and stem protein content was controlled by three QTL, on chromosomes 4, 7 and 8. These QTL explained from 13.3 to 23.2 % of variation for leaf protein content and from 13.2 to 18.1 % for stem protein content. Jemalong6 alleles showed either a positive additive effect or a negative effect.
For leaf-to-stem ratio, 24 QTL were detected on all chromosomes of M. truncatula genome in the four RIL populations, corresponding to ten genomic regions. QTL accounted for from 4.9 to 22.1 % of variation.
For stem histology in LR4 population (Supplementary Material 6, Fig. 3), five QTL corresponding to four genomic regions were detected for area of the whole stem cross section. QTL accounted for 7.9–31.0 % of variation. Six QTL controlled pith parenchyma proportion, corresponding to five genomic regions. QTL accounted for 5.7 to 28.6 % of variation. Five QTL were detected for cortex proportion corresponding to five genomic regions. QTL accounted for 9.5–35.4 % of genetic variation. Finally, four genomic regions were detected on chromosomes 1 and 8 for xylem proportion. For all QTLs for histological traits, Jemalong6 alleles induced either positive additive effects or negative ones.
Multi-population QTL analysis
QTL mapping in this multi-cross design, using MCQTL software, showed 8 QTL for forage quality traits (Table 9). On chromosome 7, a cluster of four QTL (ADF, whole plant digestibility and protein content, leaf-to-stem ratio) was located at the position from 56.5 to 57.4 cm, and explained from 4.5 to 12.0 % of total variation. Jemalong6 and A20 alleles at this QTL induced a poor quality (high ADF, low whole plant digestibility and protein content, leaf-to-stem ratio) while the alleles of the three other parents induced a high quality. Two QTL were detected for ADF and whole plant digestibility at 5 cm on chromosome 8, explaining 8.1 and 7.6 % of genetic variation, respectively. Again, the alleles of Jemalong6 and A20 induced a poor quality (high ADF and low whole plant digestibility) while DZA45.5 and F83005.5 alleles induced a high quality. A QTL on chromosome 1 at 70 cm controlled the whole plant protein content and explained 4.1 % of variation. Positive additive effects of Jemalong6, DZA315.26 and F83005.5 were observed. Finally, a QTL was detected for leaf-to-stem ratio on chromosome 3 at 52.6 cm, accounting for 3.7 % of genetic variation. Alleles of DZA315.26 and DZA45.5 showed positive additive effects. The QTL detected in this multipopulation analysis corresponded to QTL with a strong effect (LOD > 7) detected in one or several populations (on chromosome 8 for ADF and DIGT, on chromosome 7 for leaf-to-stem ratio), or to QTL with a moderate effect (LOD < 7) found in several populations (on chromosome 7 for ADF and whole plant digestibility, on chromosome 3 for leaf-to-stem ratio).
Co-location between QTL and candidate genes
The genes described to belong to lignin pathway and annotated on M. truncatula genome were located on the genetic map (Fig. 2). They were irregularly spread over the genome, with many genes located on chromosome 4. Co-locations of these genes with QTL were evidenced for the QTL at the bottom of chromosome 3 and chromosome 4 and at the top of chromosome 5. For the QTL detected in the multi-population analysis, the single co-location with candidate genes was observed at the bottom of chromosome 3 for a QTL of leaf-to-stem ratio.
Discussion
Genetic variation
Genetic variation was observed for forage quality and stem histology within the four RIL populations of M. truncatula. This result is in accordance with the variation in stem chemical composition between four M. truncatula lines (Schnurr et al. 2007). Similarly, in forage legumes such as alfalfa, genetic variation for quality and histological traits has been observed (Julier et al. 2000; Guines et al. 2003). Mean values of whole plant digestibility were similar to those observed in alfalfa (Julier et al. 2000) and higher than in floral stem of Arabidopsis (Barrière et al. 2005). Schnurr et al. (2007) also found similar stem composition in M. truncatula and alfalfa. M. truncatula has much thinner stems than alfalfa, as indicated by the area of the stem cross-section that ranged from 2 to 6 mm² in M. truncatula and from 10 to 20 mm² in alfalfa (Julier et al. 2008). But the histological composition described in this study for M. truncatula was similar to that described for alfalfa (Guines et al. 2003), especially for the xylem proportion that averaged 18 %. The proportion of parenchyma was somewhat higher in alfalfa than in M. truncatula (68 vs. 60 %) and correlatively, the cortex proportion was lower in alfalfa than in M. truncatula (12 vs. 20 %).
The different estimates of quality traits were strongly correlated among themselves in M. truncatula as they were in alfalfa (Julier et al. 2000). The correlations found between quality traits and morphological and histological traits in M. truncatula were in accordance with those found in alfalfa: high plant digestibility was achieved through high leaf-to-stem proportion and high stem digestibility, itself correlated to high cortex proportion; a high protein content was correlated to leaf-to-stem proportion and to leaf and stem protein contents. As in alfalfa (Guines 2002; Lamb et al. 2007), genotypes with long stems had lower stem digestibility and protein content. The correlations observed between flowering date and quality traits were high. Flowering in M. truncatula, as in alfalfa, leads to the establishment of a very low biomass of new organs (axillary inflorescences) that does not directly impact plant composition. As a consequence, the correlation between flowering date and quality probably originated from the correlation between flowering date and stem length.
QTL detection
This study is the first one reporting QTL detection for forage quality in a forage legume species. Numerous QTL for quality and histological traits were detected, as observed in perennial ryegrass (Lolium perenne) for digestibility and fibre content (Cogan et al. 2005). All the chromosomes were involved. For most combinations of traits and mapping populations, each parent carried both positive and negative alleles. This result is in accordance with the fact that the parental lines were not submitted to any breeding program that could have gathered positive alleles in an improved line. The range of variation of R² for quality and histological traits was similar to that of morphological traits (Lagunes Espinoza et al. 2012). When comparing QTL obtained for protein content in this study with QTL related to nitrogen nutrition in LR4 population (Moreau et al. 2012), similar positions were observed in the middle of chromosome 1, the bottom of chromosome 4 and in the middle of chromosome 8. Chromosome 1 was also identified in an association study, with one SSR allele being linked to digestibility in alfalfa (Li et al. 2011).
The multi-population QTL analyses indicated that four genomic regions, on chromosomes 1, 3, 7 and 8, contributed to explain phenotypic variation in the whole crossing design. The R² obtained in this QTL analyses varied among 3.7 and 12.0 and were similar to those obtained for morphological traits, but lower than those obtained for a major QTL for flowering date on chromosome 7 (Lagunes Espinoza et al. 2012). Quality and morphological traits had a quantitative inheritance, as expected, but flowering date was mainly governed by a major gene in this crossing design.
The annotation of M. truncatula genome was used to compare QTL position to candidate gene position. Most QTL for quality traits did not co-localize with genes related to lignin biosynthesis. It was not possible to map all the genes described to belong to lignin pathway since some genes (4CL, C3H, C4H, C3′H) were not annotated on the M. truncatula genome. In addition, expression of lignin genes is frequently controlled by transcription factors that can be located in other genomic regions than those of the regulated genes, as found in Arabidopsis (Chavigneau et al. 2012). Contrastingly in perennial ryegrass, some QTL for quality traits co-localized with genes of the lignin pathway.
All the QTL for quality and histological traits were located in regions where QTL for morphological traits were also found. Either the same genes were involved in both types of traits or genes involved in quality and in morphological traits co-located in these regions. The first hypothesis means that all variations in quality traits are related to variation in morphological traits. Even if quality and morphology were significantly correlated, in this study as in other studies in alfalfa, several results indicated that changes in quality through traditional breeding can be achieved without changes in morphology or yield. Indeed, even if divergent selection for lignin concentration alone had an impact on yield (Kephart et al. 1989), it was possible to select populations with higher digestibility without altered forage yield (Julier et al. 2003a). In addition, within a set of cultivars, a large variation for digestibility was documented among high-yielding cultivars (Julier et al. 2003b). The transgenesis approach showed that the genes involved in lignin biosynthesis (Guo et al. 2001a, b; Reddy et al. 2005; Shadle et al. 2007) had an effect on tissue composition and digestibility. In these studies, morphogenesis was significantly altered in the transgenic plants. However, it seems that downregulation of lignin synthesis had a positive effect on salicylic acid synthesis, a stress hormone. Downregulation of genes involved in salicylic acid synthesis in lignin-modified plants restored growth potential (Gallego-Giraldo et al. 2011). In addition, epistasis or co-regulations could lead to a correlation between quality and morphological traits. In Medicago species as in others, there are interactions between the genes involved in lignin synthesis, the genetic background in which they are placed and the growing environment. This interaction determines yield and persistency of the plants showing reduced lignin content (Pedersen et al. 2005). These elements indicate that despite the co-location of QTL for quality and for morphogenesis and partly common regulation, genes directly related to quality should be involved. These genes could be identified using fine mapping strategies and the variation in the sequence of their homologues in alfalfa could contribute to explain phenotypic variation for forage quality.
There are several consequences of this work on legume and more specifically on alfalfa breeding. The negative correlation between forage yield components and quality traits explains why joint improvement is difficult. The detection of genes related to quality traits was hampered by the fact that no QTL specific to quality trait was identified. No QTL study on quality traits of alfalfa was published. The identification of common QTL positions in M. truncatula and alfalfa would help to focus on these regions to find the genes that are specifically involved in quality traits. The use of these genes into breeding programs would need additional work to find the polymorphisms that would explain higher quality traits. Transfer of knowledge from a diploid annual model species to an autotetraploid perennial crop species is not simplistic but if candidate genes are identified, their analysis in crop species may give promising results. It was the case of Constans-like gene, a candidate gene isolated after QTL studies in M. truncatula (Pierre et al. 2011) that proved to explain variation for stem length and flowering date in alfalfa (Herrmann et al. 2010; Julier 2012).
References
Arcade A, Labourdette A, Falque M, Mangin B, Chardon F, Charcosset A, Joets J (2004) BioMercator: integrating genetic maps and QTL towards discovery of candidate genes. Bioinformatics 20:2324–2326
Aufrère J (1982) Etude de la prévision de la digestibilité des fourrages par une méthode enzymatique. Ann Zootech 31:111–130
Barrière Y, Laperche A, Barrot L, Aurel G, Briand M, Jouanin L (2005) QTL analysis of lignification and cell wall digestibility in the Bay-0 x Shahdara RIL progeny of Arabidopsis thaliana as a model system for forage plant. Plant Sci 168:1235–1245
Basten CJ, Weir BS, Zeng ZB (1994) Zmap-a QTL cartographer, pp 65-66
Basten CJ, Weir BS, Zeng ZB (2002) QTL Cartographer, Version 1.16. Department of Statistics, North Carolina State University, Raleigh
Blanc G, Charcosset A, Mangin B, Gallais A, Moreau L (2006) Connected populations for detecting quantitative trait loci and testing for epistasis: an application in maize. Theor Appl Genet 113:206–224
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Boudet AM, Kajita S, Grima-Pettenati J, Goffner D (2003) Lignins and lignocellulosics: a better control of synthesis for new and improved uses. Trends Plant Sci 8:576–581
Buxton DR, Hornstein JS, Marten GC (1987) Genetic variation for forage quality of alfalfa stems. Can J Plant Sci 67:1057–1067
Chavigneau H, Goué N, Delaunay S, Courtial A, Jouanin L, Reymond M, Méchin V, Barrière Y (2012) QTL for floral stem lignin content and degradability in three recombinant inbred line (RIL) progenies of Arabidopsis thaliana and search for candidate genes involved in cell wall biosynthesis and degradability. OJGen 2:7–20
Cogan N, Smith K, Yamada T, Francki M, Vecchies A, Jones E, Spangenberg G, Forster J (2005) QTL analysis and comparative genomics of herbage quality traits in perennial ryegrass (Lolium perenne L.). Theor Appl Genet 110:364–380
Engels FM, Jung HG (1998) Alfalfa stem tissues: cell-wall development and lignification. Ann Bot 82:561–568
Engels FM, Jung HG (2005) Alfalfa stem tissues: impact of lignification and cell length on ruminal degradation of large particles. An Feed Sci Tech 120:309–321
Gallego-Giraldo L, Escamilla-Trevino L, Jackson LA, Dixon RA (2011) Salicylic acid mediates the reduced growth of lignin down-regulated plants. Proc Natl Acad Sci USA 108:20814–20819
Goering HK, Van Soest PJ (1970) Forage fiber analysis, USDA Agric handbook, vol 379. USDA, Washington, pp 1–20
Guines F (2002) Bases génétiques des variations pour la structure histologique des tiges de luzerne (Medicago sativa L.). ENSA Rennes
Guines F, Julier B, Ecalle C, Huyghe C (2002) Genetic control of quality traits of lucerne (Medicago sativa L.). Aust J Agric Res 53:401–407
Guines F, Julier B, Ecalle C, Huyghe C (2003) Among- and within-cultivar variability for histological traits of lucerne (Medicago sativa L.) stem. Euphytica 130:293–301
Guo D, Chen F, Inoue K, Blount JW, Dixon RA (2001a) Downregulation of Caffeic Acid 3-O-Methyltransferase and Caffeoyl CoA 3-O-Methyltransferase in transgenic alfalfa; impacts on lignin structure and implications for the biosynthesis of G and S lignin. Plant Cell 13:73–88
Guo DG, Chen F, Wheeler J, Winder J, Selman S, Peterson M, Dixon RA (2001b) Improvement of in-rumen digestibility of alfalfa forage by genetic manipulation of lignin O-methyltransferases. Transgenic Res 10:457–464
Heinrichs DH, Troelsen JE, Warder FG (1969) Variation of chemical constituents and morphology characters within and between alfalfa populations. Can J Plant Sci 49:293–305
Herrmann D, Barre P, Santoni S, Julier B (2010) Association of a CONSTANS-LIKE gene to flowering and height in autotetraploid alfalfa. Theor Appl Genet 121:865–876
Jourjon MF, Jasson S, Marcel J, Ngom B, Mangin B (2005) MCQTL: multi-allelic QTL mapping in multi-cross design. Bioinformatics 21:128–130
Julier B (2012) Alfalfa breeding benefits from genetic analyses on M. truncatula. Salt Lake City, UT. Online proceedings to be published fall 2012, http://mbft.usu.edu/
Julier B, Huyghe C (1997) Effect of growth and cultivar on alfalfa digestibility in a multi-site trial. Agronomie 17:481–489
Julier B, Guy P, Castillo-Acuna C, Caubel G, Ecalle C, Esquibet M, Furstoss V, Huyghe C, Lavaud C, Porcheron A, Pacros P, Raynal G (1996) Genetic variation for disease and nematode resistances and forage quality in perennial diploid and tetraploid lucerne populations (Medicago sativa L.). Euphytica 91:241–250
Julier B, Huyghe C, Ecalle C (2000) Within- and among-cultivar genetic variation in alfalfa: forage quality, morphology, and yield. Crop Sci 40:365–369
Julier B, Barre P, Hébert Y, Huguet T, Huyghe C (2003a) Methodology of alfalfa breeding: a review of recent achievements. CJGPB 39:71–81
Julier B, Guines F, Ecalle C, Emile JC, Lila M, Briand M, Huyghe C (2003b) Eléments pour une amélioration génétique de la valeur énergétique de la luzerne. Fourrages 173:49–61
Julier B, Huguet T, Chardon F, Ayadi R, Pierre JB, Prosperi JM, Barre P, Huyghe C (2007) Identification of quantitative trait loci influencing aerial morphogenesis in the model legume Medicago truncatula. Theor Appl Genet 114:1391–1406
Julier B, Guines F, Poussot P, Ecalle C, Huyghe C (2008) Use of image analysis to quantify histological structure along the stems of alfalfa (Medicago sativa L.). Acta Botanica Gallica 155:485–494
Jung HG, Engels FM (2002) Alfalfa stem tissues: cell wall deposition, composition, and degradability. Crop Sci 42:524–534
Jung HG, Smith RR, Endres CS (1994) Cell wall composition and degradability of stem tissue from lucerne divergently selected for lignin and in vitro dry-matter digestibility. Grass Forage Sci 49:295–304
Kephart KD, Buxton DR, Rhill RR (1989) Morphology of Alfalfa divergently selected for herbage lignin concentration. Crop Sci 29:778–782
Lagunes Espinoza L, Huguet T, Julier B (2012) Multi-population QTL detection for aerial morphogenetic traits in the model legume Medicago truncatula. Theor Appl Genet 124:739–754
Lamb JFS, Jung HJG, Sheaffer CC, Samac DA (2007) Alfalfa leaf protein and stem cell wall polysaccharide yields under hay and biomass management systems. Crop Sci 47:1407–1415
Lander ES, Botstein D (1989) Mapping mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121:185–199
Lemaire G, Allirand JM (1993) Relation entre croissance et qualité de la luzerne : interaction génotype-mode d’exploitation. Fourrages 134:183–198
Lenssen AW, Sorensen EL, Posler GL, Harbers LH (1991) Basic alfalfa germplasms differ in nutritive content of forage. Crop Sci 31:293–296
Li X, Wei Y, Moore KJ, Michaud R, Viands DR, Hansen JL, Acharya A, Brummer EC (2011) Association mapping of biomass yield and stem composition in a tetraploid alfalfa breeding population. Plant Genome 4:24–35
Media Cybernetics LP (1996) Optimas6, user guide and technical reference. Optimas Corporation, Washington
Moreau D, Burstin J, Aubert G, Huguet T, Ben C, Prosperi JM, Salon C, Munier-Jolain N (2012) Using a physiological framework for improving the detection of quantitative trait loci related to nitrogen detection in Medicago truncatula. Theor Appl Genet 124:755–768
Mowat DN, Fulkerson RS, Tossell WE, Winch JE (1965) The in vitro digestibility and protein content of leaf and stem portions of forages. Can J Plant Sci 45:321–331
Neff R, Simon U (1986) Heritability of protein content in vitro digestibility in Medicago sativa L. International Meeting Eucarpia Group “Medicago sativa”, pp 199–207
Pedersen JF, Vogel KP, Funnell DL (2005) Impact of reduced lignin on plant fitness. Crop Sci 45:812–819
Pierre JB, Huguet T, Barre P, Huyghe C, Julier B (2008) Detection of QTLs for flowering date in three mapping populations of the model legume species Medicago truncatula. Theor Appl Genet 117:609–620
Pierre JB, Bogard M, Herrmann D, Huyghe C, Julier B (2011) A CONSTANS-like gene candidate that could explain most of the genetic variation for flowering date in Medicago truncatula. Mol Breed 28:25–35
Reddy MSS, Chen F, Shadle G, Jackson L, Aljoe H, Dixon RA (2005) Targeted down-regulation of cytochrome P450 enzymes for forage quality improvement in alfalfa (Medicago sativa L.). Proc Natl Acad Sci USA 102:16573–16578
SAS Institute Inc. (2000) Version 8.1. SAS Institute, Cary, North Carolina
Schnurr JA, Jung HJG, Samac DA (2007) A comparative study of alfalfa and Medicago truncatula stem traits: morphology, chemical composition, and ruminal digestibility. Crop Sci 47:1672–1680
Shadle G, Chen F, Reddy MSS, Jackson L, Nakashima J, Dixon RA (2007) Down-regulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase in transgenic alfalfa affects lignification, development and forage quality. Phytochemistry 68:1521–1529
Terry RA, Tilley JMA (1969) The digestibility of the leaves and stems of perennial ryegrass, cocksfoot, timothy, tall fescue, lucerne and sainfoin, as measured by an in vitro procedure. J br Grassl Soc 19:363–373
Tolivia D, Tolivia J (1987) Fasga: a new polychromatic method for simultaneous and differential staining of plant tissue. J Microsc 148:113–117
Vallet C, Chabbert B, Czaninski Y, Monties B (1996) Histochemistry of lignin deposition during sclerenchyma differentiation in alfalfa stem. Ann Bot 78:625–632
Wilson JR (1993) Organization of forage plant tissues. In: Jung HG, Buxton DR, Hatfield RD, Ralph J (eds) Forage cell wall structure and digestibility. Madison, pp 1–32
Young ND, Udvardi M (2009) Translating Medicago truncatula genomics to crop legumes. Curr Opin Plant Biol 12:193–201
Young ND, Debelle F, Oldroyd GED, Geurts R, Cannon SB, Udvardi MK, Benedito VA, Mayer KFX, Gouzy J, Schoof H, Van de Peer Y, Proost S, Cook DR, Meyers BC, Spannagl M, Cheung F, De Mita S, Krishnakumar V, Gundlach H, Zhou SG, Mudge J, Bharti AK, Murray JD, Naoumkina MA, Rosen B, Silverstein KAT, Tang HB, Rombauts S, Zhao PX, Zhou P, Barbe V, Bardou P, Bechner M, Bellec A, Berger A, Berges H, Bidwell S, Bisseling T, Choisne N, Couloux A, Denny R, Deshpande S, Dai XB, Doyle JJ, Dudez AM, Farmer AD, Fouteau S, Franken C, Gibelin C, Gish J, Goldstein S, Gonzalez AJ, Green PJ, Hallab A, Hartog M, Hua A, Humphray SJ, Jeong DH, Jing Y, Jocker A, Kenton SM, Kim DJ, Klee K, Lai HS, Lang CT, Lin SP, Macmil SL, Magdelenat G, Matthews L, McCorrison J, Monaghan EL, Mun JH, Najar FZ, Nicholson C, Noirot C, O’Bleness M, Paule CR, Poulain J, Prion F, Qin BF, Qu CM, Retzel EF, Riddle C, Sallet E, Samain S, Samson N, Sanders I, Saurat O, Scarpelli C, Schiex T, Segurens B, Severin AJ, Sherrier DJ, Shi RH, Sims S, Singer SR, Sinharoy S, Sterck L, Viollet A, Wang BB, Wang KQ, Wang MY, Wang XH, Warfsmann J, Weissenbach J, White DD, White JD, Wiley GB, Wincker P, Xing YB, Yang LM, Yao ZY, Ying F, Zhai JX, Zhou LP, Zuber A, Denarie J, Dixon RA, May GD, Schwartz DC, Rogers J, Quetier F, Town CD, Roe BA (2011) The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480:520–524
Zhao Q, Dixon RA (2011) Transcriptional networks for lignin biosynthesis: more complex than we thought? Trends Plant Sci 16:227–233
Acknowledgments
We are grateful to all our colleagues involved in chemical and histological measurements (F. Chataigner, P. Cormenier, C. Lévêque, C. Melin, V. Menanteau). L.C. Lagunes Espinoza thanks Campus Tabasco of Colegio de Postgraduados (Mexico) for granting permission to perform the research at INRA. She received a grant from Region Poitou–Charentes for her post-doctoral sabbatical leave at INRA. We thank J.M. Prosperi (INRA, Montpellier) for providing LR1, LR4 and LR5 populations, T. Huguet for providing the LR3 population and mapping data and J.P. Sampoux for help in statistics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Hackett.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2012_1996_MOESM2_ESM.doc
Supplementary material 2 Mean squares for quality and histological traits in LR4 RIL population of Medicago truncatula (DOC 44 kb)
122_2012_1996_MOESM5_ESM.doc
Supplementary material 5 QTLs detected for forage quality traits in LR1, LR4, LR5 and LR6 RIL populations of M. truncatula (DOC 125 kb)
Rights and permissions
About this article
Cite this article
Lagunes Espinoza, L.C., Julier, B. QTL detection for forage quality and stem histology in four connected mapping populations of the model legume Medicago truncatula . Theor Appl Genet 126, 497–509 (2013). https://doi.org/10.1007/s00122-012-1996-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-012-1996-2