Abstract
Soybean [Glycine max (L.) Merr.] oil typically contains 2–4 % stearic acid. Seed oil with 20 % stearic acid would be useful for solid fat applications, both for its cooking properties and health benefits. Breeding lines with high stearic acid have been developed, but many suffer from agronomic problems. This study identifies a new source of high stearic acid, determines its relationship with another high stearic locus and presents molecular markers for it is use in breeding. TCJWB03-806-7-19, a ‘Holladay’ mutant with high stearic acid, was crossed to two FAM94-41-derived lines that contained a point mutation in a seed-specific isoform of a Δ9-stearoyl-acyl carrier protein-desaturase (SACPD-C). Fatty acid analysis was performed over two growing seasons with F 2-derived lines and transgressive segregation for stearic acid content was observed. Sequencing of SACPD isoforms in TCJWB03-806-7-19 revealed the deletion of an ‘A’ nucleotide in exon 3 of SACPD-B, which results in a protein whose final 28 amino acids are predicted to differ from Williams 82 SACPD-B. Sorting intolerant from tolerant (SIFT) analysis revealed that this frameshift mutation may affect SACPD-B protein function. Allele-specific genotyping for the SACPD-C point mutation and SACPD-B nucleotide deletion was performed in both populations. Additive effects and R 2 for stearic acid were +3.3 and 0.55 for SACPD-C and +1.9 and 0.19 for SACPD-B. Average stearic acid in lines homozygous for both mutations was 14.6 %. This SACPD-B mutation represents a novel high stearic allele.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean [Glycine max (L.) Merr.] is the most widely produced oil seed crop in the world, accounting for over half of worldwide oil crop production (USDA-Economic Research Service 2011). An oxidatively stable oil with a relatively high melting temperature is necessary for solid fat applications (Clemente and Cahoon 2009). Soybean seed oil naturally high in saturates would be suitable for this end use (List and Pelloso 2007; Clemente and Cahoon 2009). Palmitic (16:0) and stearic acid (18:0) are the two saturated fatty acids present in soybean oil. Typical palmitic and stearic acid contents of soybean oil are 11 and 4 %, respectively (Hildebrand et al. 2008).
Short chain saturated fatty acids, such as palmitic acid, are undesirable because their consumption results in an unfavorable lipoprotein profile in blood serum (Mensink and Katan 1990). In contrast, stearic acid appears to have no cholesterolemic effects in humans (Kris-Etherton and Yu 1997) and exhibits similar thrombogenic effects as oleic and linoleic acids (Thijssen et al. 2005).
Stearic acid content in soybean typically represents only 2–5 % of total fatty acids (ILSI 2010; USDA-ARS 2012); however, several germplasm lines have been developed with increased stearic acid. All high stearic soybean germplasm lines have been developed using mutagenesis, with the exception of FAM94-41 (Pantalone et al. 2002). FAM94-41 (9 % stearic acid) has a spontaneously occurring mutation in the SACPD-C gene, a seed-specific isoform of a Δ9-stearoyl-acyl carrier protein-desaturase (SACPD), which gives rise to the elevated stearic phenotype (Zhang et al. 2008). SACPD is responsible for the desaturation of stearic acid to oleic acid (Ohlrogge and Browse 1995). The stearic acid QTL on linkage group B2 identified by Spencer et al. (2003) is likely due to the SACPD-C mutation in FAM94-41, as FAM94-41 was a parent in the mapping population used and the most closely linked marker identified in the study, Satt070, is 0.2 cM (centimorgans) from SACPD-C. The elevated stearic acid phenotype in A6 (28 % stearic acid; Hammond and Fehr 1983) is due to the deletion of SACPD-C (Zhang et al. 2008) and is allelic with the mutations in A81-606085 (19 % stearic acid) and FA41545 (16 % stearic acid; Graef et al. 1985). RG-7 (12 % stearic acid) and RG-8 (10.6 % stearic acid) both have a point mutation in SACPD-C and are presumably allelic with the A6 and FAM94-41 mutations (Boersma et al. 2012). ST1 (29 % stearic acid), ST3 (23 % stearic acid), and ST4 (23 % stearic acid) also have high stearic mutations allelic to the SACPD-C deletion in A6; however, the mutation in ST2 (28 % stearic acid) has been hypothesized to be at a different locus (Bubeck et al. 1989). Other elevated stearic germplasm lines have also been developed with unknown causative loci (Rahman et al. 1995; Hudson 2012).
Two other isoforms of SACPD have been identified in soybean, SACPD-A and SACPD-B (Byfield et al. 2006), but to date neither has been associated with an elevated stearic acid phenotype. SACPD-A and SACPD-B transcripts have been detected in developing seed, root, leaf, and flower tissues while SACPD-C transcript has been primarily detected in developing seed (Byfield et al. 2006; Zhang et al. 2008; Kachroo et al. 2008).
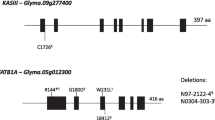
3-Ketoacyl-acyl carrier protein synthase II (KASII) and 18:1-acyl carrier protein thioesterase (18:1-ACP TE) are other genes in which a mutation or differential expression could lead to elevated stearic acid content (Pantalone et al. 2002). KASII is responsible for the elongation of palmitic acid to stearic acid (Ohlrogge and Browse 1995). A mutation in KASII has been found to increase palmitic acid in soybean (Aghoram et al. 2006). Theoretically, overexpression or a mutation of KASII could lead to elevated stearic acid. 18:1-ACP TE catalyzes the hydrolyzation of 18:1-ACP and 18:0-ACP to release fatty acid in order to facilitate their transport to the cytoplasm (Jones et al. 1995). Increased 18:1-ACP TE activity has been shown to increase stearic acid content in sunflower (Cantisán et al. 2000).
Stearic acid QTL not associated with any known enzyme in the fatty acid pathway have also been reported. In addition to the QTL associated with SACPD-C, 12 additional stearic acid QTL have been reported, on 8 different linkage groups (Diers and Shoemaker 1992; Hyten et al. 2004; Panthee et al. 2006; Reinprecht et al. 2006; Li et al. 2011).
Agronomic problems such as lower seed yield, poor germination, and reduced seedling growth rate have been associated with elevated levels of stearic acid (Lundeen et al. 1987; Rahman et al. 1997; Wang et al. 2001a). The effect varies depending on the causative allele, and not every allele results in a yield decrease (Lundeen et al. 1987). The identification of new high stearic QTL and/or novel mutations in known isoforms of SACPD, KASII, or 18:1-ACP TE may provide alternatives to overcome these constraints, and also may lead to a greater ability to fine-tune fatty acid composition (Cardinal 2008). The objectives of this research were to genetically characterize a new source of elevated stearic acid identified in a population developed by mutagenesis, to determine it is relationship to other known high stearic loci, and to develop molecular markers for use in breeding for improved fatty acid composition.
Materials and methods
Plant materials
Two populations were developed by bi-parental crosses performed in 2009 in Clayton, NC. FA-G is a population of 209 F 2-derived lines from the cross LLL-05-1 × TCJWB03-806-7-19. FA-H is a population of 117 F 2-derived lines from the cross LLL-05-14 × TCJWB03-806-7-19. LLL-05-1 (12.0 % stearic acid) and LLL-05-14 (11.7 % stearic acid) are maturity group V F5-derived selections from the cross FAM94-41-3 × N98-4445A. LLL-05-14 matures 3 days later than LLL-05-1 and both lines differ in pubescence color. FAM94-41-3 (8.8 % stearic acid; Pantalone et al. 2002) is a selection from an elevated stearic germplasm line with a natural mutation in SACPD-C (Zhang et al. 2008). N98-4445A (Burton et al. 2006) is a mid-oleic germplasm line. TCJWB03-806-7-19 is a maturity group V elevated stearic selection from a mutagenized population of the cultivar ‘Holladay’ (Burton et al. 1996) that was developed by exposing seed to 250 grays (Gy) of gamma radiation from a Gammacell 220 (MDS Nordion Inc., Ottawa, Ontario, Canada). TCJWB03-806-7-19 has 7 % stearic acid compared to the 4 % stearic acid of ‘Holladay’ (unpublished data).
Field evaluation
In 2010, F 2 plants were grown in Clayton, NC and harvested individually. F 2-derived lines were grown in 3.7-m-long one-row plots in 2011. Soils at Clayton, NC were Norfolk Loamy Sands for both years. In 2011, flower color, pubescence color, and maturity date at the R8 reproductive stage (Fehr and Caviness 1977) were recorded.
Seed oil analysis
Fatty acid methyl ester analysis was performed on a 20-seed sample from each F 2 plant in 2010 and from each F 2:3 line in 2011. Seed samples were crushed and approximately 1 g was extracted for about 24 h in 3 mL of solvent (chloroform:hexane:methanol, 8:5:2 v/v/v) in stoppered glass test tubes. Fatty acid methyl esters of the lipid extracts were prepared and analyzed as outlined by Burkey et al. (2007).
Oil content was measured from a 10-g seed sample from each F 2 plant in 2010 and each F 2:3 line in 2011. Oil content was determined by pulsed proton nuclear magnetic resonance (NMR) using a Maran pulsed NMR (Resonance Instruments, Witney, Oxfordshire, UK) by the field induction decay-spin echo procedure (Rubel 1994).
SACPD gene isoforms sequencing
Sequencing was performed for the three known SACPD gene isoforms (A, B and C) in the three parental lines and Holladay. Primers used for amplification and sequencing are listed in Tables 1 and 2.
Amplification reactions were carried out in 1× New England Biolabs (NEB, Ipswich, Massachusetts, USA) standard reaction buffer with MgCl2 (final MgCl2 1.5 mM), 1.4 Units NEB Taq polymerase, 208 μM dNTPs, 6 pmol forward primer, 6 pmol reverse primer, and 3 µL of 10 ng/µL DNA for a reaction volume of 15 µL. A touchdown PCR reaction was performed with the following parameters—94 °C for 2 min—13 cycles of: 94 °C for 32 s, 63 °C for 30 s,−1 °C/cycle, 72 °C for 2 min—22 cycles of: 94 °C for 32 s, 50 °C for 30 s, and 72 °C for 2 min—72 °C for 7 min. PCR products were cleaned with a thermosensitive alkaline phosphatase (TSAP)/Exol enzyme mix prepared as follows: 10.4 µL TSAP, 10.4 µL Exol, 187.2 µL H2O. 2 µL of the TSAP/Exol mix was used per 8 µL of PCR product and incubated at 37 °C for 45 min followed by inactivation at 80 °C for 15 min. Samples were submitted to GeneWiz (Research Triangle Park, NC) with 5 µM of the sequencing primer.
Molecular marker analysis
A single leaf was collected at random from 20 plants from each F 2-derived line in 2011. A single punch was taken from each leaf using a cork borer and the punches for each F 2-derived line were bulked for DNA extraction. DNA was extracted using a CTAB method (Allen et al. 2006). Primers were designed based on the SACPD-B-1114 SNP and SACPD-C-229 SNP (Zhang et al. 2008) for use in the KBiosciences competitive allele-specific PCR (KASPar) SNP genotyping system (KBiociences, Herts, UK) (Table 3).
The DNA samples were diluted to 2 ng/µL and 3 µL from each sample were transferred to a 384-well plate and dried down at 55 °C for 30 min in an incubator. Total volume per reaction was 4 µL, which consisted of 2 µL 2× KASP reaction mix, 0.11 µL 0.5× assay mix, 0.072 µL 50 mM MgCl2, and 1.818 µL H2O. The 0.5× assay mix was prepared to a total volume of 20 µl, which consisted of 1.2 µL 100 µM of each allele-specific primer, 3 µl 100 µM common reverse primer, and 14.6 µL 10 mM Tris pH 8.3. Thermocycling parameters were—94 °C incubation for 15 min—10 cycles of: 94 °C for 20 s, 65 °C for 60 s, −0.8 °C/cycle—30 cycles of: 94 °C for 20 s and 57 °C for 60 s. Endpoint fluorescence reading was performed using a Roche LightCycler® 480 (Penzberg, Germany). Allele calling was performed using Version 1.5 of the Roche LightCycler® 480 software.
Statistical analysis
Chi-square tests were conducted to determine if the SACPD-C and SACPD-B loci were segregating according to the expected ratios in each population (Snedecor and Cochran 1956). Analysis of variance (ANOVA) for all fatty acids and total oil was conducted using SAS PROC GLM (SAS Institute, Cary, NC, 2009). Means were calculated for each F 2-derived line, using the years as the replicates, and linear regression analysis of maturity versus fatty acid concentration was conducted using PROC REG to determine whether maturity had an effect on fatty acid concentration.
For each population, the F 2-derived line means were used in a PROC GLM ANOVA for the SACPD-B and SACPD-C loci and least squares means were calculated for each genotypic class. Since both populations share the same male parent and the female parents are sister lines, a combined ANOVA for both populations was also conducted. In this combined ANOVA, population and genotypic class by population were also considered as factors. Dunnett pairwise comparison was conducted on the genotypic class least square means to determine if the mutant alleles resulted in a different phenotype than the wild type. Contrast statements were used to estimate the additive and dominant effects of each locus and the additive by additive, additive by dominant, dominant by additive and dominant by dominant epistatic interactions between the SACPD-B and SACPD-C loci (Holland 2001). Additive and additive by additive effects were estimated using the genotypic class least squares means from both years of data (F 2 and F 2:3 lines) while dominant, additive by dominant, dominant by additive, and dominant by dominant effects were estimated using only the data from F 2 plants.
Results
Sequence analysis of SACPD isoforms
The sequencing results were compared to the Williams 82 reference sequence for each isoform (Schmutz et al. 2010). The SACPD-A (glyma07g32850) reference sequence was found to be identical to the Williams 82 reference sequence in all three parents and Holladay.
A silent mutation was identified at nucleotide 930 in SACPD-C (glyma14g27990) in LLL-05-01, LLL-05-14, TCJWB03-806-7-19, and Holladay. In addition, the SNP previously identified in this locus at nucleotide 229 in FAM94-41 (Zhang et al. 2008) was also found in LLL-05-01 and LLL-05-14.
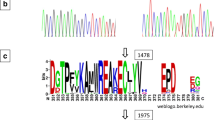
The SACPD-B (glyma02g15600) coding sequence had a silent mutation at nucleotide 76 in TCJWB03-806-7-19 and Holladay. TCJWB03-806-7-19 had a deletion of the A at nucleotide 1,114 (Fig. 1a) that is predicted to alter the location of the stop codon and result in a longer protein whose final 28 amino acids are predicted to differ from those of the SACPD-B protein in Williams 82 (Fig. 1b) and the SACPDs examined by Byfield et al. (2006). The TCJWB03-806-7-19 SACPD-B sequence is available as GenBank no. JQ993842 and an allele designation has been requested to the soybean genetics committee.
Frameshift mutation in the SACPD-B gene of TCJWB03-806-7-19. a Nucleotide sequence comparison of the SACPD-B genes from ‘Williams 82’ (Glyma02g15600) and TCJWB03-806-7-19 (GenBank no. JQ993842) in the region surrounding the ‘A’ deletion at position 1114 (relative to the start codon). b Amino acid comparison showing the frameshift mutation, starting at position 372 (relative to the start amino acid) and predicted altered location of the stop codon in the SACPD-B enzyme from TCJWB03-806-7-19. Amino acids with an asterisk are substitutions predicted to affect enzymatic function by sorting intolerant from tolerant (SIFT) analysis
To further examine the conservation of this position, the sorting intolerant from tolerant (SIFT) algorithm as described by Ng and Henikoff (2001) was used. The Williams 82 SACPD-B reference coding sequence was used as the query sequence and the TCJWB03-806-7-19 amino acid substitutions caused by the nucleotide deletion were the substitutions of interest. The “UniProt-TrEMBL 2009 Mar” protein database was used, with median conservation of sequences specified as 3.00 and sequences >90% identical to the query sequence were removed. 135 related sequences were retrieved by the algorithm and 12 of the 20 amino acid carboxyl-terminus substitutions due to the nucleotide deletion were predicted to affect protein function.
Genetic analysis of the SACPD-C and SACPD-B mutations
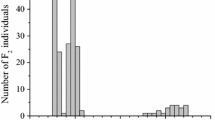
No segregation distortion was detected for the SACPD-C locus or SACPD-B locus in either population (Table 4). There was also no segregation distortion for flower or pubescence color (data not shown). Since the results for both populations were very similar, only the results from the combined analysis are presented. Maturity differed by 16 days among F 2-derived lines and was not a significant factor in the regression analysis; therefore, it was not included as a covariate in the statistical analysis. No significant additive by dominant, dominant by additive, or dominant by dominant epistatic effects were detected. The mutant forms of both genes were associated with an increase in stearic acid and a decrease in oleic acid content in all the analyses (Table 5). The SACPD-C nucleotide 229 mutation had a larger positive additive effect (+3.3) on stearic acid levels than the SACPD-B nucleotide 1,114 deletion (+1.9), while both mutations had comparable negative additive effects on oleic acid (Table 6). Significant epistatic additive by additive gene action of effect +3.3 on stearic acid was also detected. The SACPD-C mutation exhibited a dominance genetic effect (−0.8) on stearic acid levels while the SACPD-B mutation did not (Table 7). Both mutations were associated with a small decrease in palmitic acid and a small decrease in total oil (Table 6). Interestingly, both mutations were associated with opposite significant effects on linolenic acid concentration, a decrease of −0.2 with the mutant form of SACPD-C and an increase of 0.4 with the mutant form of SACPD-B. R 2 values for the SACPD-C and SACPD-B loci are indicated in Table 6.
The double-mutant genotype resulted in average stearic acid of 14.6 % versus the wild-type average of 4.3 %. This occurred largely at the expense of oleic acid where lines with both mutations had 14.8 % oleic acid versus 25.4 % oleic acid for the wild type. The double-mutant genotype also resulted in a small, but significant decrease in total oil, 19.6 % compared to 20.8 % for the wild type (Table 8).
Discussion
The effect of the SACPD-C nucleotide 229 mutation on fatty acid composition was confirmed in FAM94-41-derived lines and a new functional mutation in SACPD-B was identified. The effect of this SACPD-C mutation on stearic acid in these populations is in agreement with the mapping study performed by Spencer et al. (2003); however, this study provides stronger evidence of a dominance genetic effect, likely because of a larger population size and the use of perfect markers. Boersma et al. (2012) recently reported a line with a SACPD-C mutation whose resulting protein product is predicted to be only 63 amino acids long and a separate line with a point mutation resulting in a predicted proline → leucine substitution at amino acid 286. Interestingly, those two lines, FAM-94-41 and the FAM-94-41-derived lines in this study all exhibit comparable levels of stearic acid. This suggests that all characterized elevated stearic mutations involving SACPD-C are due to a complete, rather than partial, loss of function and that one or more additional loci are probably involved in the 28 % stearic acid phenotype observed in A6 (Hammond and Fehr 1983; Zhang et al. 2008). The nucleotide 1,114 deletion in SACPD-B is the first functional mutation in this isoform reported in soybean. This deletion causes a frameshift and is predicted to result in a longer protein whose 28 final amino acids are predicted to differ from Williams 82 SACPD-B (glyma02g15600). The SIFT analysis provides support to the notion that this frameshift results in a functional mutation and that the altered fatty acid composition is due to this mutation and not due to a mutation at a linked locus; however, it is unknown whether the SACPD-B nucleotide 1,114 deletion results in a complete or partial loss of enzymatic function. This mutation has a similar effect on phenotype to that observed with the SACPD-C nucleotide 229 mutation, which is associated with increased stearic acid, primarily at the expense of oleic acid, and small decreases in palmitic acid and total oil. Additionally, the SACPD-B nucleotide 1,114 deletion is associated with a small increase in linolenic acid while the SACPD-C mutation is associated with a small decrease in linolenic acid in these populations. This difference could be due to a low linolenic acid QTL inherited from N98-4445A (Bachlava et al. 2009) being linked to the SACPD-C mutation in the LLL-05-1 and LLL-05-14 parental lines. Although SACPD-B expression is not seed specific (Byfield et al. 2006) and SACPD-B is not as strongly expressed in seed as SACPD-C (Kachroo et al. 2008), this research demonstrates that mutations in this gene do affect seed fatty acid composition and total oil accumulation. There is also significant evidence for an additive by additive epistatic interaction between the SACPD-C and SACPD-B loci. Biologically, this could be due to the wild-type enzyme of one isoform compensating for the mutant enzyme of another isoform, therefore, the observed additive effect of SACPD-C is larger when SACPD-B is mutant than when this locus is wild type.
These mutations bring us closer to the 20 % stearic acid necessary for high stearic oil for use in solid fat applications, but another elevated stearic locus will be needed. Combining these two mutations with a SACPD-A mutation may reduce 18:0 to 18:1 desaturation activity sufficiently to achieve this goal. Alternatively, these mutant genes could be combined with other mutant genes in the fatty acid metabolic pathway that could result in elevated stearic acid concentration, such as KASII *or 18:1-ACP TE. Also, if the SACPD-B nucleotide 1114 deletion only results in a partial loss of function, a mutation with a complete loss of function could also be a candidate for use in developing a high stearic acid soybean oil.
High stearic acid oil, also high in oleic acid, is another oil type desired by industry (Clemente and Cahoon 2009) for its baking qualities and health properties. These data suggest that obtaining both high stearic acid and high oleic acid from a single cultivar may prove difficult due to the increase in stearic acid being obtained at the expense of desaturation to oleic acid. Developing high stearic acid oil and blending it with high oleic acid oil may be a more realistic goal.
Since SACPD-B is expressed in a variety of plant tissues, a mutation in it may have agronomic implications not realized in mutations observed in seed-specific desaturases. Silencing all three known isoforms of SACPD in soybean has been shown to result in adverse morphological differences (Kachroo et al. 2008). Soybean seed with elevated stearic acid has been shown to have increased triacylglycerol and phospholipid melting temperatures (Wang et al. 2001b). Changes in membrane fluidity are known to affect plant metabolic processes (Quinn and Williams 1978; Yamamato et al. 1981). Decreased membrane fluidity could be responsible for the reduction in total seed oil accumulation associated with these SACPD-C and SACPD-B mutations. Because the non-seed-specific SACPD-B had a larger effect on total oil accumulation, the authors propose that metabolic processes occurring in vegetative tissues, rather than seed, are driving this reduction in total seed oil.
It would be of great value to evaluate this new SACPD-B mutation for agronomic effects, such as seed yield, emergence, and cold tolerance to determine if it would be a viable candidate for use in breeding a high-yielding, elevated stearic soybean. The KASPar markers developed in this study would be useful for marker-assisted backcrossing in crosses involving these SACPD-B and SACPD-C mutations to develop a cultivar with elevated stearic acid content.
Abbreviations
- 18:1-ACP TE:
-
18:1-Acyl carrier protein thioesterase
- KASII:
-
3-Ketoacyl-acyl carrier protein synthase II
- KASPar:
-
KBiosciences competitive allele-specific PCR
- NEB:
-
New England biolabs
- SACPD:
-
Δ9-Stearoyl-acyl carrier protein-desaturase
- SIFT:
-
Sorting intolerant from tolerant
- TSAP:
-
Thermosensitive alkaline phosphatase
References
Aghoram K, Wilson RF, Burton JW, Dewey RE (2006) A mutation in a 3-keto-acyl-ACP synthase II gene is associated with elevated palmitic acid levels in soybean seeds. Crop Sci 46:2453–2459
Allen GC, Flores-Vergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325
Bachlava E, Dewey RE, Burton JW, Cardinal AJ (2009) Mapping and comparison of quantitative trait loci segregating for oleic acid seed content in two segregating soybean populations. Crop Sci 49:433–442
Boersma JG, Gillman JD, Bilyeu KD, Ablett GR, Grainger C, Rajcan I (2012) New mutations in a delta-9-stearoyl-acp desaturase gene associated with enhanced stearic acid levels in soybean seed. Crop Sci. doi:10.2135/cropsci2011.08.0411
Bubeck DM, Fehr WR, Hammond EG (1989) Inheritance of palmitic and stearic acid mutants of soybean. Crop Sci 29:652–656
Burkey KO, Booker FL, Pursley WA, Heagle AS (2007) Elevated carbon dioxide and ozone effects on peanut: II seed yield and quality. Crop Sci 47:1488–1497
Burton JW, Carter TE Jr, Huie EB (1996) Registration of ‘Holladay’ soybean. Crop Sci 36:467
Burton JW, Wilson RF, Rebetzke GJ, Pantalone VR (2006) Registration of N98–4445A mid-oleic soybean germplasm line. Crop Sci 46:1010–1012
Byfield GE, Xue H, Upchurch RG (2006) Two genes from soybean encoding soluble Δ9 stearoyl-ACP desaturases. Crop Sci 46:840–846
Cantisán S, Martínez-Force E, Garcés R (2000) Enzymatic studies of high stearic acid sunflower seed mutants. Plant Physiol Biochem 38:377–382
Cardinal AJ (2008) Molecular genetics and breeding for fatty acid manipulation in soybean. Plant Breed Rev 30:259–294
Clemente TE, Cahoon EB (2009) Soybean oil: genetic approaches for modification of functionality and total content. Plant Phys 151:1030–1040
Diers BW, Shoemaker RC (1992) Restriction fragment length polymorphism analysis of soybean fatty acid content. J Am Oil Chem Soc 69:1242–1244
Fehr WR, Caviness CE (1977) Stages of soybean development, special report 80, Ames, IA. Iowa State University, Ext. Service. Agric. Exp. Stn.
Graef GL, Fehr WR, Hammond EG (1985) Inheritance of three stearic acid mutants of soybean. Crop Sci 25:1076–1079
Hammond EG, Fehr WR (1983) Registration of A6 germplasm line of soybean (Reg. No. GP45). Crop Sci 23:192–193
Hildebrand DF, Li R, Hatanaka T (2008) Genomics of soybean oil traits. In: Stacey G (ed) Genetics and genomics of soybean. Springer, New York, pp 185–210
Holland JB (2001) Epistasis and plant breeding. Plant Breed Rev 21:27–92
Hudson K (2012) Soybean Oil-Quality Variants Identified by Large-Scale Mutagenesis. Int J Agron. doi:10.1155/2012/569817
Hyten DL, Pantalone VR, Saxton AM, Schmidt ME, Sams CE (2004) Molecular mapping and identification of soybean fatty acid modifier quantitative trait loci. J Am Oil Chem Soc 81:1115–1118
ILSI (2010) International life sciences institute crop composition database, Version 4.2. International Life Sciences Institute, Washington DC. http://www.cropcomposition.org. Accessed 15 Jan 2012
Jones A, Davies HM, Voelker TA (1995) Palmitoyl-acyl carrier protein (ACP) thioesterase and the evolutionary origin of plant acyl-ACP thioesterases. Plant Cell 7:359–371
Kachroo A, Fu DQ, Havens W, Navarre D, Kachroo P, Ghabrial SA (2008) An oleic acid-mediated pathway induces constitutive defense signaling and enhanced resistance to multiple pathogens in soybean. Mol Plant-Microbe Interact 21:564–575
Kris-Etherton PM, Yu SM (1997) Individual fatty acid effects on plasma lipids and lipoproteins: human studies. Am J Clin Nutr 65:S1628–S1644
Li H, Zhao T, Wang Y, Yu D, Chen S, Zhou R, Gai J (2011) Genetic structure composed of additive QTL, epistatic QTL pairs and collective unmapped minor QTL conferring oil content and fatty acid components of soybeans. Euphytica 182:117–132
List GR, Pelloso T (2007) Zero/low trans margarine, spreads and shortening. In: List GR, Kritchevsky D, Ratnayke N (eds) Trans Fats in Foods. AOCS Press, Champaign, IL, pp 155–176
Lundeen PO, Fehr WR, Hammond EG, Cianzio SR (1987) Association of alleles for high stearic acid with agronomic characters of soybean. Crop Sci 27:1102–1105
Mensink RP, Katan MB (1990) Effect of dietary trans fatty acids on high-density and low-density lipoprotein cholesterol levels in healthy subjects. N Engl J Med 323:439–445
Ng PC, Henikoff S (2001) Predicting deleterious amino acid substitutions. Genome Res 11:863–874
Ohlrogge J, Browse J (1995) Lipid Biosynthesis. Plant Cell. 7:957–970
Pantalone VR, Wilson RF, Novitzky WP, Burton JW (2002) Genetic regulation of elevated stearic acid concentration in soybean oil. J Am Oil Chem Soc 79:549–553
Panthee DR, Pantalone VR, Saxton AM (2006) Modifier QTL for fatty acid composition in soybean oil. Euphytica 152:67–73
Quinn PJ, Williams WP (1978) Plant lipids and their role in membrane function. Prog Biophys Mol Biol 34:109–173
Rahman SM, Takagi Y, Miyamoto K, Kawakita T (1995) High stearic acid soybean mutant induced by X-ray irradiation. Biosci Biotech Biochem 59:922–923
Rahman SM, Takagi Y, Kinoshita T (1997) Genetic control of high stearic acid content in seed oil of two soybean mutants. Theor Appl Genet 95:772–776
Reinprecht Y, Poysa VW, Yu K, Rajcan I, Ablett GR, Pauls KP (2006) Seed and agronomic QTL in low linolenic acid, lipoxygenase-free soybean (Glycine max (L.) Merrill) germplasm. Genome 49:1510–1527
Rubel G (1994) Simultaneous determination of oil and water contents in different oilseeds by pulsed nuclear magnetic resonance. J Am Oil Chem Soc 71:1057–1062
SAS Institute Inc (2009) Base SAS® 9.2 Procedures Guide. Cary, NC: SAS Institute Inc
Schmutz J, Cannon SB, Schlueter J (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Snedecor GW, Cochran WG (1956) Statistical methods applied to experiments in agriculture and biology. The Iowa State College Press, Ames, IA
Spencer MM, Pantalone VR, Meyer EJ, Landau-Ellis D, Hyten DL Jr (2003) Mapping the Fas locus controlling stearic acid contents in soybean. Theor Appl Genet 106:615–619
Thijssen MA, Hornstra G, Mensink RP (2005) Stearic, oleic, and linoleic acids have comparable effects on markers of thrombotic tendency in healthy human subjects. J Nutr 135:2805–2811
USDA-ARS (2012) National genetic resources program. Germplasm resources information network—(GRIN). National germplasm resources laboratory, Beltsville, Maryland. http://www.ars-grin.gov/cgi-bin/npgs/html/obvalue.pl?51084. Accessed 15 Jan 2012
USDA-Economic research service (2011) Oil crops yearbook 2011. World oilseed production. USDA economics, statistics and market information system, Albert R. Mann Library, Cornell University. http://usda.mannlib.cornell.edu/MannUsda/viewDocumentInfo.do?documentID=1290. Accessed 15 Jan 2012
Wang T, Harp T, Hammond EG, Burris JS, Fehr WR (2001a) Seed physiological performance of soybeans with altered saturated fatty acid contents. Seed Sci Res 11:93–97
Wang T, Hammond EG, Fehr WR (2001b) Neutral and polar lipid phase transition of soybeans with various saturated fatty acid contents. J Am Oil Chem Soc 78:1139–1144
Yamamato Y, Ford R, Barber J (1981) Relationship between thylakoid membrane fluidity and the functioning of pea chloroplasts: effect of cholesteryl hemisuccinate. Plant Physiol 67:1069–1072
Zhang P, Burton JW, Upchurch RG, Whittle E, Shanklin J, Dewey RE (2008) Mutations in a Δ9-stearoyl-ACP-desaturase gene are associated with enhanced stearic acid levels in soybean seeds. Crop Sci 48:2305–2313
Acknowledgments
This research was supported in part by the United Soybean Board and the North Carolina Soybean Producers Association. The authors thank Dr. Thomas Carter Jr. for supplying TCJWB03-806-7-19 for use in this study and for reviewing this manuscript. The authors also thank W. Novitzky for training and assistance provided in the seed oil analysis and the undergraduate part-time workers at the Soybean and Nitrogen Fixation Unit (USDA-ARS, NC State University, Raleigh) for assistance in collecting leaf tissue for DNA extraction. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Diers.
Rights and permissions
About this article
Cite this article
Ruddle, P., Whetten, R., Cardinal, A. et al. Effect of a novel mutation in a Δ9-stearoyl-ACP-desaturase on soybean seed oil composition. Theor Appl Genet 126, 241–249 (2013). https://doi.org/10.1007/s00122-012-1977-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-012-1977-5