Abstract
Legumes provide essential micronutrients that are found only in low amounts in the cereals or root crops. An ongoing project at CIAT has shown that the legume common bean is variable in the amount of seed minerals (iron, zinc, and other elements), vitamins, and sulfur amino acids that they contain and that these traits are likely to be inherited quantitatively. In this study we analyzed iron and zinc concentrations in an Andean recombinant inbred line (RIL) population of 100 lines derived from a cross between G21242, a Colombian cream-mottled climbing bean with high seed iron/zinc and G21078, an Argentinean cream seeded climbing bean with low seed iron/zinc. The population was planted across three environments; seed from each genotype was analyzed with two analytical methods, and quantitative trait loci (QTL) were detected using composite interval mapping and single-point analyses. A complete genetic map was created for the cross using a total of 74 microsatellite markers to anchor the map to previously published reference maps and 42 RAPD markers. In total, nine seed mineral QTL were identified on five linkage groups (LGs) with the most important being new loci on b02 and other QTL on b06, b08, and b07 near phaseolin. Seed weight QTL were associated with these on b02 and b08. These Andean-derived QTL are candidates for marker-assisted selection either in combination with QTL from the Mesoamerican genepool or with other QTL found in inter and intra-genepool crosses, and the genetic map can be used to anchor other intra-genepool studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron deficiency anemia (IDA) and other micronutrient deficiencies affect large numbers of people worldwide and in many parts of Africa and Latin America are one of the top health concerns especially among the poor (Graham et al. 2001). Worldwide, over three billion people are affected by IDA while zinc deficiency in the human diet is probably almost as widespread as the lack of sufficient iron (Frossard et al. 2000). IDA causes loss in work productivity and complications in childbirths while zinc deficiency causes stunting and impaired disease immunity (Welch and Graham 1999, 2004). Biofortification addresses these constraints from a diet perspective improving staple crops with higher amounts of essential minerals (Pfeiffer and McClafferty 2007).
Food legumes provide essential micronutrients that are found only in low amounts in the cereals or root crops and in general contain appreciable quantities of iron, zinc, and other minerals (Wang et al. 2003). Legumes usually contain higher concentrations of minerals than cereals due to greater partitioning of these elements into their grain. It is notable that iron homeostasis in both the plant and the grain is mediated by transport and storage mechanisms, with the protein iron reductase being key for uptake by roots and ferritin being key to storage in vegetative tissue, nodules and finally in grain (Briat and Lobreaux 1997). Zinc uptake is often limited by soil availability, and the transport and storage of this micronutrient is less clearly understood although the mineral is important as a co-factor for many proteins including transcription factors. The poorly studied process of seed loading of iron and zinc is key to accumulation of these minerals in legume seeds and involves mobilization from vegetative to reproductive organs (Frossard et al. 2000).
Common bean (Phaseolus vulgaris L.) is the most important food legume for direct human consumption, being especially important in Eastern and Southern Africa and Latin America (Broughton et al. 2003). Two basic types of common beans exist: large-seeded beans like kidney beans which are from the Andean genepool and small-seeded beans like navy beans from the Mesoamerican genepool. An ongoing project at the International Center for Tropical Agriculture (CIAT) as part of the Harvest Plus challenge program on Biofortification has worked on the improvement of both genepools for iron and zinc concentration (Graham et al. 1999; Beebe et al. 2000; Blair et al. 2009a).
Among common bean varieties the large-seeded types of the Andean genepool are the most commonly consumed in Eastern and Southern Africa (Wortmann et al. 1998) and are important in other regions and for international trade. This genepool includes the commercial classes of red-mottled beans and cream-mottled or sugar beans, both popular in a large number of countries from Ethiopia to Malawi. Cranberry beans produced in Canada, China and the United States are also of the cream-mottled commercial class, and are widely traded on international markets into South Africa, Colombia, Europe and the Middle East. Large red beans of both the radical and dark red kidney type are a third commercial type from the Andean genepool found throughout Eastern Africa but concentrated in Kenya and Tanzania. These beans are also important in international trade and have become an important health food in salad bars around the world. In terms of growth habit, Andean beans are very diverse ranging from type I determinate bush beans to very tall type IV indeterminate climbing beans (Singh et al. 1991a).
Given that CIAT’s objective in participating in the Biofortification Project has been to assess the feasibility of improving common beans for micronutrient concentration, especially iron and zinc, we have engaged in extensive analysis of the inheritance and diversity of iron and zinc concentrations in the two genepools of common beans. Furthermore, since CIAT’s breeding program is structured along genepool lines, we have conducted analysis with recombinant inbred line populations in each genepool so as to understand which genes influence the accumulation of iron and zinc and how these vary in different bean types. The development of genetic maps within the Andean genepool is limited by low polymorphism (Blair et al. 2006a; Cichy et al. 2009).
With this in mind, the goals of this study were (1) to construct a new genetic map for a high × low iron Andean recombinant inbred line (RIL) population and (2) to identify quantitative trait loci (QTL) for iron and zinc concentrations in this population. The population was derived from G21242, a Colombian cream-mottled bean with high seed iron/zinc concentration and G21078, an Argentinean cream seeded bean with low seed iron/zinc. These genotypes were the most contrasting Andean beans in the evaluation of a core collection by Islam et al. (2002) and are both climbing beans unlike our previous study of an Andean bush bean cross as described in Cichy et al. (2009). The QTL analysis was based on seed analyzed from three trial sites using two analytical methods and involved the creation of the first intra-genepool genetic map ever-created for Andean climbing beans all of which can assist Andean bean breeding programs to improve seed micronutrient concentration in this genepool. We also mapped the seed proteins and ferritin phaseolin based on contrasting parental alleles to determine association with iron concentration.
Materials and methods
Plant materials
A population of 100 recombinant inbred lines was developed from the cross of two climbing beans, G21242 × G21078, at the International Center for Tropical Agriculture (CIAT). The population size was established at 100 due to the smaller fields and wider spacing used for climbing beans and due the higher expense and manual labor involved in experiments involving this type of bean. In general RIL populations in common bean are limited by the large seed size which makes for a reduced number of seeds produced per plant compared with other legumes or to the cereals.
The parents were identified by Islam et al. (2002) as the most contrasting genotypes for seed iron and zinc concentration within the Andean genepool based on a survey of the CIAT core collection consisting of over 600 accessions evaluated. G21078 is a low iron and zinc genotype from Argentina with medium-sized, cream-colored seed with type IVa climbing bean growth habit. G21242 is a high iron and zinc genotype from Colombia with medium to large, cream-mottled seed with purple stripes and is also a climbing bean with type IVa growth habit. The population was created by advancing F1 plants from the simple cross to the F2 generation and then through single-seed descent from the F2 to the F5 generation. The individual F5 selections were planted for seed increase to the F7 generation and the resulting lines used for further experiments.
Genetic mapping
DNA was extracted for each of the lines in the F5:7 generation using the method of Afanador et al. (1993) and used for all marker analysis described below. Microsatellite (SSR or simple sequence repeat) markers from Yu et al. (2000), Metais et al. (2002), Blair et al. (2003, 2009b, c) and Grisi et al. (2007) were surveyed for polymorphism on the parents of the population and then used for genetic mapping of segregants. A set of RAPD (randomly amplified polymorphic DNA) markers were generated with 10 decamer primers from Islam et al. (2002) analyzed on the entire set of RILs. In addition, a gene-based DNA marker for the protein ferritin was mapped based on amplification with a primer set For- 5′ TCGCCGGGAAAGTTGCCAGT 3′ and Rev- 5′ TTGCATAAAATGCTTGCAGTC 3′ for the ferritin gene from common bean based on Spence et al. (1991) (Genbank entry X58724). Amplification and band detection conditions for these DNA based markers were as described in the previous SSR studies with the ferritin marker mapped with the following PCR reaction: 92°C for 1 min, 55°C for 1.5 min, 72°C for 2 min for 35 cycles and 5 min extension. In addition to the DNA based markers, alleles for one protein marker for the seed storage protein phaseolin was determined using a total extract of seed proteins run on SDS-PAGE gels as described in Singh et al. (1991b). Phaseolin map location was easily determined as the population segregated for the “C” and “T” alleles which are easily distinguished by the number and placement of bands present on the protein gels (Brown et al. 1981). Polymorphisms and segregation patterns were scored for each of the markers, bands or protein gels and a genetic map was constructed using Mapmaker 3.0 (Lander et al. 1987) and a minimum LOD of 3.0. The Kosambi function was used to convert recombination fractions into genetic distances in centiMorgans (cM). Linkage groups (LGs) were oriented and named according to the conventions in Blair et al. (2003) based on SSR loci.
Experimental sites
The population was analyzed over three typical bean-growing locations in Colombia: (1) Darién, Valle which is at 1,450 m elevation and has an average yearly temperature of 20°C average yearly temperature, an annual rainfall of 1,650 mm, and an Udand soil type; (2) Palmira, Valle which is at 1,000 m elevation and has an average yearly temperature of 24°C, an annual rainfall of 905 mm and a Haplustoll soil type; and (3) Popayán, Cauca which is at 1,730 m elevation and has an average yearly temperature of 18°C, an annual rainfall of 2,124 mm, and a Dystrudepts soil type. Native, HCl, and H2SO4 extractable mineral concentrations in the topsoil of Popayán and Darién were low, averaging 2.40 and 4.39 mg/kg for iron and 3.56 and 0.76 mg/kg for zinc; however, total soil iron levels at lower profiles were adequate averaging, 7.88 and 6.84 mg/kg. These conditions are typical of acid soils (pH 5.6–6.1) where Andean beans are grown. Meanwhile iron and zinc soil concentration in Palmira were low at 1.74 and 3.44 mg/kg for iron and zinc, respectively, but pH levels were higher (pH 7.8). The trial in Popayán, where adaptation was best due to the origin of the Andean genotypes, consisted in a lattice design with three replications for the full set of 100 RILs. The trials in Darién and Palmira, which are lower elevation sites with higher temperatures than Popayán, consisted in randomized complete block experiments with two replications each with only the most adapted 83 and 65 genotypes harvested, respectively. All the experiments were planted with trellis supports since the population is predominantly made up of climbing bean genotypes. Agronomic management consisted in best practices as per Checa and Blair (2008) with fertilization of 45 kg of P ha−1 as superphosphate in Popayán and Darién, but not in Palmira. At all sites three foliar applications of zinc and boron as microelements (300 g ha−1 as chelates) were carried out at 14 and 21 days after planting. Protective fungicide and insecticide treatments were applied as needed. In all seasons, plots were hand-harvested to avoid contamination by metal machinery. Seed weight (SW) was measured across experimental locations for each genotype by weighing 100 seeds.
Mineral analysis
Two methods of mineral analysis were implemented for the same seed harvest from Popayán: namely, (1) inductively coupled plasma (abbreviated ICP) analysis at the University of Adelaide using an ARL 3580 ICP with optical emission spectrometry and (2) atomic absorption spectroscopy (AAS) using a Unicam SOLAAR 969 mass spectrophotometer at the CIAT analytical services lab. The samples from the other two sites (Darién and Palmira) were only analyzed with the AAS technique to save on processing costs as this method is only a third as costly as ICP sampling. Sample preparation for the ICP technique involved grinding 10 g of beans (about 30 seeds) in a coffee mill, while for the AAS technique a separate 5 g of beans (about 15 seeds) were ground using a Retsch mill and aluminum grinding balls and elliptical chambers. The seeds were surface cleaned with ethanol to remove soil and dust and oven dried at 45°C before grinding. Details of the ICP and AAS procedures using nitric/perchloric acid digested samples are as described in Blair et al. (2009a) and as in that research we were interested in validating the less expensive AAS assay to complement ICP analysis.
Data analysis
Analyses of variance (ANOVA) and Pearson’s correlations between mineral averages and SW of the RILs were carried out using the program Statistix version 8.0 (Analytical Software, Tallahasse, FL, USA). QTL were detected with composite interval mapping (CIM) analysis that was carried out using the software program QTL Cartographer v. 1.21 (Basten et al. 2001) and the genetic map for the G21242 × G21078 populations as described earlier. The following parameters were used for CIM analysis: 10 cM window size, 1 cM walkspeed, five significant background markers, analysis by forward and backward multiple linear regression for each chromosomal position with a global significance level of 5% and probability thresholds of 0.05 for the partial F test for both marker inclusion or exclusion. Determination coefficients were calculated for each interval separately (R 2) and for each interval given the background markers (TR2) to determine the phenotypic variance explained by a single QTL (either alone or in conjunction with all other significant intervals). Population distributions were evaluated for normality with QTL Cartographer and LOD (log of the odds) thresholds for the individual QTL for each trait were determined by the generation of 1,000 permutations of the data for that trait (Churchill and Doerge 1994). Results were displayed using QTL Cartographer and represented graphically with standard drawing software, to designate genomic regions that proved to be significant in the analysis described above.
Results
Population distributions and parental differences
Iron and zinc concentrations in the RILs presented a continuous distribution for all three locations (Darién, Palmira, and Popayán) and for both AAS and ICP in Popayán, suggesting that mineral concentration behaved as a quantitative trait (Fig. 1). The range and averages for iron concentration for AAS was higher in Darién (28.0–95.1 mg/kg with average of 54.5 mg/kg) than in Popayán (30.1–88.4 mg/kg with average of 48.3 mg/kg), while the zinc concentration range and average for AAS was lower in Darién (16.9–37.0 mg/kg with average of 26.6 mg/kg) than in Popayán (22.2–56.8 mg/kg with average of 31.5 mg/kg). Results with ICP analysis had similar population distributions as with AAS; however, G21078 was lower in iron concentration in the ICP analysis than in the AAS analysis, even though G21242 was similar. For zinc, the average of all the RILs was higher for ICP than for AAS, but the population distribution was similar.
Population distributions for seed iron (Fe) and zinc (Zn) concentrations (in mg/kg) among RILs of the G21242 × G21078 population grown in Darién, Palmira and Popayán as determined by atomic absorption spectroscopy (AAS) or inductively coupled plasma–optical emission spectrometry (abbreviated ICP). Maternal (A) and paternal (B) mineral values indicated by arrows
The parents of the population showed significant differences and were contrasting in terms of mineral concentration in both of the mineral analyses (Fig. 1). G21242, the high mineral parent, was always higher in mineral concentration than G21078, the low mineral parent. In the case of iron concentration, G21078 tended to have values lower than the mean of the population (24.1–52.2 mg/kg) while G21242 was closer to the upper extreme of the population (83.1–98 mg/kg) distribution. This was especially true in the case of ICP analysis where G21242 was higher than any of the RILs while in AAS analysis the parents tended to be near the edges of the population distribution except for G21078 which in Palmira (52.2 mg/kg) was not as low in iron as in the other sites, Darién (35.5 mg/kg) and Popayán (24.1 mg/kg). Meanwhile in the case of zinc concentration the parents were more intermediate but still contrasting with G21242 always near the high end of the distribution (32.5–41.2 mg/kg) but G21078 more intermediate (21.8–27.3 mg/kg). This was not surprising since G21078 was selected for being very low in iron but less so in zinc. ICP values for zinc were higher in range for the RILs, but G21242 was lower when measured with AAS. Overall, transgressive segregation for low iron and for both high and low zinc was evident in the population across the three locations.
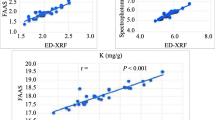
Comparison of mineral detection methods and correlations between minerals
Among the methods used for mineral analysis the ICP and AAS were highly reliable and were significantly correlated (r = 0.849 for iron and r = 0.860 for zinc). The high correlations between methods showed the reliability of each method in determining iron and zinc seed concentrations. The AAS technique provided a savings in processing and reagent costs and so was the preferred method compared with the ICP technique. Reliability of the AAS method was high with low standard deviation for parental genotypes and coefficients of variation averaging 6.8% for iron and 5.6% for zinc per genotype in the analysis of variance conducted for each location. Correlations between minerals were also high in Darién (r = 0.61 for AAS), and Popayán (r = 0.75 for AAS, r = 0.63 for ICP), but not in Palmira (Table 1). Correlations between seed weight and minerals were mostly non-significant.
Genetic map construction
A full genome map was constructed for the G21242 × G21078 cross, but low polymorphism required the screening of a large number of SSR markers. For example, a total of 700 primer pairs were screened on the parents, but only 74 mappable markers were found in that survey resulting in an overall polymorphism rate of 10.6% across all the SSR types. Among the individual types of SSR markers, the polymorphism rate was higher for BM series markers from Gaitán et al. (2002) with 30 out of 68 markers mappable (44.1% of total) perhaps because they target long GA repeats. Polymorphism rate was low for CA repeat based markers from Grisi et al. (2007) with only 13 loci placed (6.7%), for gene-based markers from Yu et al. (2000) with only 4 loci placed (10.8%) and for new BM and BMc series markers from Blair et al. (2009b, c) with only three and ten markers placed (3.8 and 4.0%, respectively). This could be due to the fact that the markers from the Blair et al. (2009b) and Yu et al. (2000) are from gene-based cDNA sequences. BMd and ATA series markers from Blair et al. (2003, 2008) were intermediate in polymorphism with 5 and 9 loci placed for each. The higher polymorphism of these markers could be due to their being based on AT-rich SSR sequences. RAPD polymorphisms per primer were also intermediate to low; however, a total of 42 RAPD markers could be mapped on the genetic map.
Among the noticeable characteristics of the genetic map was that all 11 chromosomes of the common bean genome were represented by the LGs identified in the mapping and that these did not vary greatly in size with most being from 40 to 80 cM in size (Table 2). The largest LGs by genetic distance were b02 (151.2 cM) and b06 (104.2 cM) while the smallest was b01 (31.9 cM). Average linkage group size was 66 cM and the total map length was 726 cM which represents about 75–80% the map coverage of other genetic maps created for common bean. The average number of SSR loci per LG was 6.7, and these tended to be well distributed (χ 2 = 0.009, P = 0.9227) which provided sufficient anchor points to orient and align the new genetic map with those of Blair et al. (2003, 2006b, 2008) and Yu et al. (2000). The average number of RAPD loci per LG was only 3.8 but these tended to cluster on certain LGs (χ 2 = 6.64, P < 0.01).
Overall marker distribution considering both SSR and RAPD markers was fairly uniform with 6–18 markers placed on each LG. As a result, the distance between DNA-based markers was 6.25 cM while this varied between LGs with b08 and b10 having high saturation of one marker every 4.01 and 4.09 cM, respectively, compared with b11, b02 and b06 with one marker every 10.37, 8.89, and 8.86 cM, respectively. In contrast, b07, with one marker every 4.38 cM benefited from further saturation of two gene-based markers, including the Phaseolin protein marker (Phs) which was closely linked with BM185 and at approximately 10 cM from the STS marker for the Ferritin gene (Fer). SSR markers were over-represented on b02 as has been found in many previous studies (Blair et al. 2003, 2008) but were uniformly distributed with 6–8 markers on all other LGs except b05 with 4. Meanwhile, RAPD markers were overrepresented on b02, b06, b07, and b10 and under-represented on b01 and b11. Segregation distortion was observed in parts of four LGs but was not as notable as in inter-genepool crosses.
QTL identification
Using CIM analysis, in total 9 QTL were detected for mineral concentration in the population across 5 of the 11 LGs (Table 3, Fig. 2). Of these, six QTL were for iron concentration and three QTL were for zinc concentration. Among the iron concentration QTL, three were from the AAS analysis and three from the ICP analysis. The QTL on b02 for iron detected with ICP and AAS overlapped near the markers H1902b and E0403A. Separate QTL for iron were detected with AAS at two positions on b06 near the markers BM137 and N0401B, respectively. Another QTL for iron was detected with ICP at the Phaseolin locus and overlapped with the marker BM185, while another iron QTL detected with ICP was found near the marker BMc65 in the middle of b09. Among the zinc QTL none were detected for ICP analysis only for AAS analysis. All the QTL were detected in Palmira which may indicate genotype × environment interaction for zinc accumulation and more variability for this trait in Palmira than in the other locations.
Quantitative trait loci (QTL) for seed iron (Fe) and zinc (Zn) concentrations and seed weight (SW) in the G21242 × G21078 recombinant inbred line mapping population (refer to Table 3 for QTL names). Vertical lines for each composite interval mapping QTL to the left of each linkage group (b01 to b11) represent the range of the QTL that are above the LOD threshold (thin line represents lower LOD QTL and thick line represents higher LOD QTL as confirmed with permutation tests); horizontal marks on the lines indicate the LOD peak for the QTL. Number of boxes with the same shading to the right of the linkage groups indicates the single-point analysis QTL and its significance for each of the trait × location combinations; where atomic absorption spectroscopy is abbreviated as AAS and inductively coupled plasma–optical emission spectrometry is abbreviated ICP and the locations are indicated by letters (a) for Popayán, (b) for Darién and (c) for Palmira. Bolded markers are on the same linkage group as in the previous studies of Blair et al. (2003, 2006b, 2008) and Yu et al. (2000), while italicized markers are on different linkage group
The significant QTL for zinc concentration with CIM analysis were found at the bottom of b02 near the markers PV25 and PV11, at the bottom of b07 near the marker BM239 and in the middle of b08 near the markers BM153 and BM165. Seed size QTL were detected at three locations with CIM analysis. These were at the same locus that was significant for iron concentration under AAS and ICP analyses at the top of b02 and at separate sites in the middle of b03 (between the markers BMc33 and X1101B) and b11 (associated with bMd22). The most significant of these QTL was the one on b03 with a determination coefficient of 19.4% while the others were slightly less significant at 17.5 and 15.0%, respectively.
Meanwhile, SPA analysis was conducted using qGene (Nelson 1997), and a total of 32 markers were significantly associated at P < 0.05 with mineral concentration in the G21242 × G21078 population (Fig. 2). In addition, 20 markers were significantly associated at P < 0.05 with seed size. For the markers associated with minerals, significant associations were about even for zinc (21) and iron (17) concentrations across the analytical techniques analyzed. In comparing the CIM and SPA analyses, more QTL loci were identified with the latter technique compared with the first one, although there was good alignment between the QTL detected with the two methods. SPA analysis showed that even when CIM analysis did not detect significance for both minerals at a certain locus, SPA analysis usually did detect significance, confirming the physiological relationship for iron and zinc accumulation for the majority of QTL.
As measured by the determination coefficients in the CIM analysis, the most important QTL detected in this study were the iron QTL detected with AAS on b02 and b06 in Popayán and Palmira with 21.7 and 25.5% of variance explained, respectively; and the zinc QTL found with AAS on b02 and b07 for Palmira with 18.6 and 27.8% variance explained, respectively. The positive allele for all these QTL came from G21242 the high mineral parent. The lack of QTL from Darién could be due to smaller number of RILs tested there. The iron QTL found near the Phaseolin locus with CIM/SPA analysis of ICP and AAS results were important due to their close association with the seed protein locus and relatively high determination coefficients of 13.0–27.8%, the latter in Palmira. However, curiously the source of higher iron was G21078 at this locus. No QTL was directly associated with the newly mapped ferritin locus.
Discussion
The principal achievement of this research was to identify QTL for iron and zinc concentration on a newly created, full-genome genetic map for an intra-genepool cross involving a high-mineral Andean parent and a low-mineral Andean parent. For the genetic map, all 11 chromosomes/LGs were identified by comparative mapping with microsatellite markers mapped by Blair et al. (2003, 2006b, 2008). Since multiple sources of microsatellite markers were used in this study, the resulting genetic map integrated various types of markers together including various sorts of microsatellites as well as RAPDs, and this appeared to help with better saturation and wider distribution of loci than has been observed in previous studies. The average distance of 6.15 cM between markers was considered to be useful for the QTL analysis of phenotypic results on micronutrient concentration across the three locations.
In terms of microsatellite marker mapping in the present map and in previous maps, marker placement to each LG was largely conserved, especially with ATA markers for Blair et al. (2008), the BM and BMd markers of Blair et al. (2003) and the PV markers of Grisi et al. (2007). Compared with the latter study, which integrated the BM and PV marker types, PV14 that had not been mapped before was added to b06 (chromosome 1 according to designation of Pedrosa et al. 2003) while all other 13 PV markers were found on the correct chromosomes/LGs. Meanwhile, a total of 24 BM markers from Blair et al. (2003, 2006b) were correctly assigned to linkage groups.
Similarly, eight new BMc markers (BMc32, BMc33, BMc34, BMc65, BMc115, BMc121, BMc123 and BMc350) from Blair et al. (2009b) and three new BM markers (BM229, BM238, and BM239) from Blair et al. (2009c) were placed to a linkage group for the first time on any bean genetic map, especially relative to previously mapped markers. Seven out of nine BMd markers (BMd22, BMd26, BMd42, BMd53, BMd78, BMd81, and BMd85) were placed for the first time relative to BM or PV markers in this study as were four BM markers (BM153, BM171, BM188, BM202) not mapped in Blair et al. (2003, 2006b, 2008). Meanwhile, the Yu et al. (2000) markers PV-at001, PV-at008, PV-ctt001 and PV-gaat001 were all in the same positions as in Blair et al. (2003) and four ATA markers (ATA03, ATA06, ATA07, and ATA10) were placed in the same positions as in Blair et al. (2008) except locus BN found on b11 instead of b03.
Another result of this study was that the relative polymorphism of different microsatellite types could be compared and the GA-based genomic markers of Gaitán et al. (2002) were found to be superior in polymorphism within the Andean genepool compared with CA-based genomic markers from Grisi et al. (2007) or gene-based markers from Yu et al. (2000) or Blair et al. (2009b). Some ATA-based markers and Genbank-derived microsatellites from Blair et al. (2003, 2008), respectively, were of moderate polymorphism and would be recommended for other intra-genepool mapping projects certainly within the Andean genepool and perhaps for the Mesoamerican genepool as well.
Apart from the single-copy SSR markers, two gene-based markers were integrated into chromosome b07. These were for the phaseolin and ferritin genes. The mapping of phaseolin was based on segregation of the seed protein patterns with the “C” phaseolin pattern present in G21242 and the “T” phaseolin pattern present in G21078. Meanwhile, the ferritin locus was mapped based on segregation of a dominant band. The placement of Phaseolin in this chromosome has long been known (Gepts 1999) while the mapping of the ferritin locus is a novel aspect of the present study since ferritin loci in common beans have not been characterized, although a multi-gene family is known to exist on various chromosomes for other legumes (Wardrop et al. 1999).
The genetic map was found to be useful for a detailed study of the genetics of iron and zinc concentrations. As in previous studies, the inheritance of seed iron and zinc concentration in this Andean × Andean climbing bean population was found to be quantitative; confirming results from Blair et al. (2009a) for an inter-genepool Andean × Mesoamerican bush bean population and Cichy et al. (2009) for an intra-genepool Andean bush bean population. These results were confirmed by the normal distribution observed for each combination of mineral × location or mineral × analytical method and the fact that the distribution means were close to the bi-parental means in each case. Given the large difference in mineral concentration (especially for seed iron between the parents) it was expected that the RIL individuals would have a wide range of values and this was the case. The segregation for the iron and zinc traits allowed us to find QTL for each of these characteristics.
Among the noticeable results of the QTL analysis was the significance of QTL for iron concentration at the top of b02, in the middle of two other LGs (b06 and b08) and at the top and middle of b09. The QTL on b02 appear to be novel compared to QTL found in previous studies while there was some alignment with iron QTL on b06 and b08 found by Blair et al. (2009a) and iron QTL found on b06 and b09 by Cichy et al. (2009). The lack of QTL for iron and zinc concentration on b01 in this study contrasts with results from Cichy et al. (2009) for another Andean population, but could be due to the fact that the present population did not segregate for the fin gene since the new population was created from two indeterminate, Andean climbing beans.
The phaseolin locus and adjacent markers on b07 were found to have some association with iron and zinc concentrations as was found also by Blair et al. (2009a). The reason for phaseolin’s role in accumulation of minerals is not clear unless accumulation of overall seed storage protein is correlated with increase in other proteins such as ferritin. It is tempting to assign a role for the ferritin gene which mapped within 10 cM of the Phaseolin locus, but phenotypic associations were not significant for this trait at this marker. One other surprise from this study was that the LG b11 had no effect on iron or zinc concentration in this population despite the fact that major QTL for iron and zinc concentrations have now been localized on that chromosome in two previous studies (Blair et al. 2009a; Cichy et al. 2009). As in those previous studies there was evidence for a common mechanism of seed iron and zinc accumulation given overlapping QTL for the two minerals.
Further analysis of linkage group b11 and b07 would be useful, especially to map other ferritin genes as the proteins encoded by this gene family are the principal storage proteins for iron in plant seeds and vegetative organs (Frossard et al. 2000). The differential expression of ferritin in different genotypes or different organs might explain aspects of iron homeostasis which leads to high iron accumulation. Similarly, iron reductase may be important in iron uptake and may lead to higher seed iron (personal communication, M. Grusak, USDA). In contrast there are few candidate genes for zinc accumulation.
Another unique aspect of this research was that we used a high iron source (G21242 with over 85 mg/kg iron) crossed with a low iron source (G21078 with under 35 mg/kg concentration). As was expected, the positive allele for higher iron came from the high mineral parent in the majority of cases except at the phaseolin locus which may indicate that higher iron alleles may sometimes be associated with the low mineral parent. Another iron QTL on b06 came from G21078 which may be similar to QTL found in a Mesoamerican × Mesoamerican mapping population (CIAT, unpublished), although a linked QTL on b06 for iron was from G21242. The relatively low number of QTL found for each mineral may be the result of more oligogenic inheritance in the intra-genepool cross or may be a result of the number of RILs used in the study. Oligogenic inheritance is supported by results of Guzman-Maldonado et al. (2003) who found two QTL explaining 25% of variance for iron and one QTL explaining 15% of variance for zinc; and by Gelin et al. (2007) who found only a single QTL for seed zinc concentration which explained 17.8% of variability for the trait.
Previous studies have also shown variability in iron and zinc concentrations for seed grown in different environments or years or analyzed by different methods (Blair et al. 2009a; Cichy et al. 2009) so it was not surprising to find some QTL specific for one environmental site or methodology versus another. QTL for ICP and AAS tended to overlap especially for iron on b02 and b07. There have also been differing results on the relationship of mineral and seed weight QTL with no association found between these in the inter-genepool cross from Blair et al. (2009a) except possibly at the phaseolin locus which in these types of populations has an effect on seed size. Meanwhile, in the analysis of an Andean × Andean bush bean population by Cichy et al. (2009) seed weight QTL overlapped with mineral QTL on b01 and were closely linked to mineral QTL on b06, but were not overlapping with mineral concentration QTL on b05, b08, b09 or b11. Here, for climbing beans we find only a loose association of seed weight QTL on b02 and b08 with seed mineral concentration QTL. The relationship between seed weight and mineral concentration should be studied further as previous authors have found that the proportion that the seed coat represents to the entire seed may influence iron concentration (Moraghan and Grafton 2002; Moraghan 2004; Ariza-Nieto et al. 2007).
In conclusion, we analyzed micronutrient QTL in the first intra-genepool genetic map for Andean climbing beans based on two parents that were highly contrasting for their seed mineral concentration. The use of multiple sources of microsatellites allowed us to build an unbiased map with uniform distribution of SSR markers. Meanwhile, the use of previously mapped markers allowed the genetic map to be linked to those of Blair et al. (2003) while integrating never-before mapped SSRs together with gene-based markers such as phaseolin and ferritin and established anchor points. In the CIM-QTL analysis we found that most positive alleles (7 out of 9) came from the unique high mineral climbing bean parent and did not show G × E interaction or association with seed size QTL. Meanwhile, only two mineral QTL were contributed by the low mineral parent.
Among the QTL discovered, the important and environmentally stable locus on linkage group b02 will be useful for molecular breeding while the association of mineral concentration with the phaseolin locus will be explored further, especially in relationship to protein concentrations and the linked ferritin locus. Significant and positive correlations between environments (r = 0.42–0.87, P = 0.001) for iron showed that G × E effects are manageable for this mineral while more careful analysis is needed for zinc where correlations between environments are more variable. Significant correlations between iron and zinc, meanwhile, may reveal a physiological relationship between the uptake of the two minerals in low pH soils as has been observed before for iron and zinc concentrations (Blair et al. 2009a, b, c; Cichy et al. 2009). Given that Andean beans are a predominant crop in many areas of Eastern and Southern Africa as well as parts of Latin America (Broughton et al. 2003; Wortmann et al. 1998), the discovery of Andean QTL for micronutrient concentration is expected to make a large contribution to Biofortification breeding of common beans for these regions.
References
Afanador L, Haley SD, Kelly JD (1993) Adoption of a mini-prep DNA extraction method for RAPD marker analysis in common bean. Ann Rep Bean Improv Coop 35:11
Ariza-Nieto M, Blair MW, Welch RM, Glahn RP (2007) Screening of bioavailability patterns in eight bean (Phaseolus vulgaris L.) genotypes using the Caco-2 cell in vitro model. J Agr Food Chem 55:7950–7956
Basten CJ, Weir BS, Zeng ZB (2001) QTL Cartographer: a reference manual and tutorial for QTL mapping. Department of Statistics, North Carolina State University, Raleigh
Beebe S, Gonzalez AV, Rengifo J (2000) Research on trace minerals in the common bean. Food Nutr Bull 21:387–391
Blair MW, Pedraza F, Buendia HF, Gaitán-Solís E, Beebe SE, Gepts P, Tohme J (2003) Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor Appl Genet 107:1362–1374
Blair MW, Giraldo MC, Buendia HF, Tovar E, Duque MC, Beebe SE (2006a) Microsatellite marker diversity in common bean (Phaseolus vulgaris L.). Theor Appl Genet 113:100–109
Blair MW, Iriarte G, Beebe S (2006b) QTL analysis of yield traits in an advanced backcross population derived from a cultivated Andean × wild common bean (Phaseolus vulgaris L.) cross. Theor Appl Genet 112:1149–1163
Blair MW, Buendia HF, Giraldo MC, Métais I, Peltier D (2008) Characterization of AT-rich microsatellites in common bean (Phaseolus vulgaris L.). Theor Appl Genet 118:91–103
Blair MW, Astudillo C, Grusak M, Graham R, Beebe S (2009a) Inheritance of seed iron and zinc content in common bean (Phaseolus vulgaris L.). Mol Breed 23:197–207
Blair MW, Muñoz-Torres M, Giraldo MC, Pedraza F (2009b) Development and diversity assessment of Andean-derived, gene-based microsatellites for common bean (Phaseolus vulgaris L.). BMC Plant Biol 9:100
Blair MW, Muñoz M, Pedraza F, Giraldo MC, Buendía HF, Hurtado N (2009c) Development of microsatellite markers for common bean (Phaseolus vulgaris L.) based on screening of non-enriched small insert genomic libraries. Genome 52:772–782
Briat J-F, Lobreaux S (1997) Iron transport and storage in plants. Trends Plant Sci 2:187–193
Broughton WJ, Hernandez G, Blair M, Beebe S, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp.)––model food legumes. Plant Soil 252:55–128
Brown JWS, Ma Y, Bliss FA, Hall TC (1981) Genetic variation in the subunits of globulin-1 storage protein of French bean. Theor Appl Genet 59:83–88
Checa OE, Blair MW (2008) Mapping QTL for climbing ability and component traits in common bean (Phaseolus vulgaris L.). Mol Breed 22:201–215
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Cichy KA, Caldas GV, Snapp SS, Blair MW (2009) QTL analysis of seed iron, zinc and phosphorus levels in an Andean bean population. Crop Sci 49:1742–1750
Frossard E, Bucher M, Machler F, Mozafar A, Hurrell R (2000) Potential for increasing the content and bioavailability of Fe, Zn and Ca in plants for human nutrition. J Sci Food Agric 80:861–879
Gaitán E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in common bean (Phaseolus vulgaris): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Sci 42:2128–2136
Gelin JR, Forster S, Grafton KF, McClean P, Rojas-Cifuentes GA (2007) Analysis of seed-zinc and other nutrients in a recombinant inbred population of navy bean (Phaseolus vulgaris L.). Crop Sci 47:1361–1366
Gepts P (1999) Development of an integrated linkage map. In: Singh SP (ed) Common bean improvement for the twenty-first century. Kluwer, Dordrecht, pp 389–400
Graham R, Senadhira D, Beebe S, Iglesias C, Monasterio I (1999) Breeding for micronutrient density in edible portions of staple food crops: conventional approaches. Field Crops Res 60:57–80
Graham RD, Welch RM, Bouis HE (2001) Addressing micronutrient malnutrition through enhancing the nutritional quality of staple foods: principals, perspectives and knowledge gaps. Adv Agron 70:77–144
Grisi MCM, Blair MW, Gepts P, Brondani C, Pereira PAA, Brondani RPV (2007) Genetic mapping of microsatellite markers in common bean (Phaseolus vulgaris) population BAT93 × Jalo EEP558. Genet Mol Res 6:691–706
Guzman-Maldonado SH, Martínez O, Acosta-Gallegos J, Guevara-Lara FJ, Paredes-Lopez O (2003) Putative quantitative trait loci for physical and chemical components of common bean. Crop Sci 43:1029–1035
Islam FMA, Basford KE, Jara C, Redden RJ, Beebe SE (2002) Seed compositional and disease resistance differences among gene pools in cultivated common bean. Genet Resour Crop Evol 49:285–293
Lander ES, Green P, Abrahamson J, Barlow A, Daly M, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Metais I, Hamon B, Jalouzot R, Peltier D (2002) Structure and level of genetic diversity in various bean types evidenced with microsatellite markers isolated from a genomic enriched library. Theor Appl Genet 104:1346–1352
Moraghan JT (2004) Accumulation and within-seed distribution of iron in common bean and soybean. Plant Soil 264:287–297
Moraghan JT, Grafton K (2002) Distribution of selected elements between the seed coat and embryo of two black bean cultivars. J Plant Nutr 25:169–176
Nelson JC (1997) QGENE: software for marker-based genomic analysis and breeding. Mol Breed 3:229–235
Pedrosa A, Vallejos CE, Bachmair A, Schwizer D (2003) Integation of common bean (Phaseolus vulgaris L.) linkage and chromosomal maps. Theor Appl Genet 106:205–212
Pfeiffer W, McClafferty B (2007) HarvestPlus: breeding crops for better nutrition. Crop Sci 47:S88–S105
Singh SP, Gepts P, Debouck DG (1991a) Races of common bean (Phaseolus vulgaris, Fabaceae). Econ Bot 45:379–396
Singh SP, Gutierrez JA, Molina A, Urrea C, Gepts P (1991b) Genetic diversity in cultivated common bean: II. Marker-based analysis of morphological and agronomic traits. Crop Sci 31:23–29
Spence MJ, Henzl MT, Lammers PJ (1991) The stucture of a Phaseolus vulgaris cDNA encoding the iron storage protein ferritin. Plant Mol Biol 17:499–504
Wang TL, Domoney C, Hedley CL, Casey R, Grusak MA (2003) Can we improve the nutritional quality of legume seeds? Plant Phys 131:886–891
Wardrop AJ, Wicks RE, Entsch B (1999) Occurrence and expression of members of the ferritin gene family in cowpeas. Biochem J 337:523–530
Welch RM, Graham RD (1999) A new paradigm for world agriculture: productive, sustainable and nutritious food systems to meet human needs. Field Crops Res 60:1–10
Welch RM, Graham RD (2004) Breeding for micronutrients in staple food crops from a human nutrition perspective. J Exp Bot 55:353–364
Wortmann CS, Kirkby RA, Eledu CA, Allen DJ (1998) Atlas of common bean (Phaseolus vulgaris L.) production in Africa. CIAT Pan-African Bean Research Alliance, p 133
Yu K, Park J, Poysa V, Gepts P (2000) Integration of simple sequence repeat (SSR) markers into a molecular linkage map of common bean (Phaseolus vulgaris L.). J Hered 91:429–434
Acknowledgments
This work was supported principally by a Harvest Plus challenge program grant to MWB, and funds from DANIDA to MWB and SEB used for early map development and ICP analysis. We are grateful to A. Hoyos and O. Mosquera for excellent field and laboratory assistance, respectively and to T. Fowles at Waite lab for ICP analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Bervillé.
Rights and permissions
About this article
Cite this article
Blair, M.W., Astudillo, C., Rengifo, J. et al. QTL analyses for seed iron and zinc concentrations in an intra-genepool population of Andean common beans (Phaseolus vulgaris L.). Theor Appl Genet 122, 511–521 (2011). https://doi.org/10.1007/s00122-010-1465-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-010-1465-8