Abstract
The aphid transmitted Turnip yellows virus (TuYV) has become a serious pathogen in many rapeseed (Brassica napus L.) growing areas. Three-years’ field trials were carried out to get detailed information on the genetics of TuYV resistance derived from the resynthesised B. napus line ‘R54’ and to develop closely linked markers. F1 plants and segregating doubled-haploid (DH) populations derived from crosses to susceptible cultivars were analysed using artificial inoculation with virus-bearing aphids, followed by DAS-ELISA. Assuming a threshold of E 405 = 0.1 in ELISA carried out in December, the results led to the conclusion that pre-winter inhibition of TuYV is inherited in a monogenic dominant manner. However, the virus titre in most resistant lines increased during the growing period, indicating that the resistance is incomplete and that the level of the virus titre is influenced by environmental factors. Bulked-segregant marker analysis for this resistance locus identified two closely linked SSR markers along with six closely linked and three co-segregating AFLP markers. Two AFLP markers were converted into co-dominant STS markers, facilitating efficient marker-based selection for TuYV resistance. Effective markers are particularly valuable with respect to breeding for TuYV resistance, because artificial inoculation procedures using virus-bearing aphids are extremely difficult to integrate into practical rapeseed breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oilseed rape, or canola (Brassica napus L.; genome AACC, 2n = 38) is today the world’s third-leading source of both vegetable oil and oil extraction meal, behind oil palm and soybean. In Europe, North America, Asia, and Australia the area of oilseed rape production is continuing to rise greatly, due to an increasing demand for rapeseed oil both as healthy edible oil and as renewable resource for the biodiesel and oleochemical industries. This increased production has resulted in a higher disease pressure for a number of oilseed rape fungal pathogens and for major viral pathogens, e.g. Turnip yellows virus (TuYV). TuYV belongs to the Poleroviruses within the family of the Luteoviridae. This virus is transmitted by aphids—mainly Myzus persicae (Schliephake et al. 2000)—in a persistent manner (Randles and Rathjen 2000) and is widespread worldwide (Graichen and Rabenstein 1996; Thomas et al. 1993; Smith and Hinckes 1985). TuYV is very closely related to Beet western yellows virus (BWYV), Beet Chlorosis virus (BChV) and Beet mild yellowing virus (BMYV). The differentiation between these viruses was only achieved by extensive host range analyses (Graichen and Rabenstein 1996), investigations on the sequence level (Veidt et al. 1988) and phylogenetic studies of their protein sequences (Beuve et al. 2008; Torres et al. 2005). The infection rate of TuYV depends on the vector activity, which increases with mild autumn and winter temperatures. Therefore, it might be assumed that this virus will gain importance as a consequence of global warming. Chemical measures against the vector, i.e. insecticide applications or seed treatment, are neither efficient nor ecologically sustainable (Read and Hewson 1988; Walsh et al. 1989). Yield losses in oilseed rape/canola caused by TuYV have been estimated in the range of 12–34% in Germany (Graichen 1997) and as high as 37–46% in Australia (Jones et al. 2007). The main symptom of TuYV infection, namely a red or violet discoloration of leaves accompanied by stunted growth, closely resembles nutrient deficiency symptoms, often resulting in unnecessary application of fertilizers that cannot be taken up by infected plants (Graichen 1999). In terms of reducing insecticide and fertilizer usage, TuYV resistance is therefore an important breeding goal with regard to the long-term sustainability of rapeseed production on increasing production areas.
Brassica napus is a young species that originated through spontaneous interspecific hybridisations between B. rapa L. (syn. campestris; genome AA, 2n = 20) and B. oleracea L. (CC, 2n = 18). Due to an extremely strong bottleneck selection for zero seed erucic acid and low seed glucosinolate content, current winter oilseed rape and spring-sown canola cultivars have a comparably narrow genetic basis (Hasan et al. 2006). In particular, the availability of novel germplasm for resistance breeding is often extremely limited. On the other hand, resistance for important diseases of oilseed rape can be transferred via resynthesis of B. napus from the diploid progenitor species B. oleracea and B. rapa (e.g. Mithen and Magrath 1992; Diederichsen and Sacristan 1996; Rygulla et al. 2007).
The main prerequisite for efficient breeding of resistance to TuYV is a detailed knowledge of the resistance genetics. Resistance against TuYV was detected for the first time in 1992 in the resynthesized rapeseed line ‘R54’ (Graichen 1994). Results on the genetics of this TuYV resistance are somewhat contradictory. Although resistance had been assumed to be monogenically inherited, other studies indicated the involvement of additional genes and an influence of environmental factors on the expression of the resistance (Graichen 1994, 1998). Dreyer et al. (2001) identified a single quantitative trait locus (QTL) on B. napus chromosome N04 (A04) explaining up to 50% of the phenotypic variance for the TuYV resistance derived from ‘R54’. TuYV resistance derived from ‘R54’ has been incorporated into the cultivar ‘Caletta’, released in Italy in 2001.
As for other aphid-transmitted luteoviruses, e.g. Barley yellow dwarf virus (BYDV, cf. Niks et al. 2004), reliable resistance screening on the phenotypic level requires rearing of virus-bearing aphids followed by artificial inoculation and serological virus detection. This procedure—especially the rearing of aphids and controlled inoculation of individual plants—is impractical for applied breeding programs. Therefore, effective molecular markers would be very useful tools for resistance breeding. The goals of this study were therefore (1) to obtain detailed information on the mode of inheritance of resistance to TuYV, and (2) to develop molecular markers suitable for marker-based selection.
Materials and methods
Plant material
The genetic analysis and marker development was performed using 111 doubled haploid (DH) lines derived from F1 plants of three different crosses of TuYV-resistant (r) × TuYV-susceptible (s) parental lines. 63 DH lines derived from the cross ‘SW01T5128’ (r) × ‘SW01T1000’ (s), 37 lines from the cross ‘Caletta’ (r) × ‘SW01T1000’ (s) and 11 lines from the cross ‘Caletta’ (r) × ‘SW01T4065’ (s). Resistance of all parental lines traces back to ′R54′. All lines were kindly provided by SW Seed GmbH, Hadmersleben, Germany.
In order to get detailed information on the mode of inheritance, 11 half-sister F1-hybrids were produced by pollinating a susceptible male-sterile maternal line (MSL) with 9 different resistant DH Lines which derived resistance from ‘R54’ and displayed a virus titre lower than E 405 = 0.1, as well as with ‘Caletta’ and ‘R54’. A total of 10 F1-plants of each cross were tested for their resistance reaction. The MSL-line was kindly provided by Norddeutsche Pflanzenzucht Hans-Georg Lembke KG, Hohenlieth, Germany.
Testing for TuYV-resistance
To obtain reliable and uniform TuYV infection under field conditions, virus-bearing aphids were reared for artificial inoculation. Virus-free M. persicae were reared for at least 4 days on rapeseed plants carrying the TuYV isolate BN5 (Graichen and Rabenstein 1996). Subsequently, TuYV carrying aphids were transferred to rapeseed plants in the cotyledonous stage for 4 days to infect these plants, and were then removed by insecticide treatment. The TuYV-infected plants were grown in the greenhouse for a few weeks before being used for mass production of virus-bearing aphids. In early October, virus-carrying leaves were taken to the field and at least 10 aphids were transferred using a brush to each DH and F1 plant.
The resistance reaction of the DH lines was determined by a DAS-ELISA procedure adapted from Clark and Adams (1977). ELISA was performed in December, April and late May/early June in each of three successive growing seasons (2004/2005, 2005/2006, 2006/2007). Due to a limited amount of seeds, DH lines were tested in a single replication in 2004/2005, three replications in 2005/2006 and two replications in 2006/2007. Each year, 10 plants per DH line were tested by ELISA for each replication. Due to the homozygosity of DH lines, i.e. each individual plant represents a genetic replication, and with the artificial infection procedure, we used, reliable results were obtained on the analyses of 10 plants per DH line and replication. This procedure has been successfully applied for other insect-transmitted viruses already, e.g. Barley yellow dwarf virus (Scheurer et al. 2001). The F1 hybrids were grown and infected in the growing seasons 2003/2004 and 2006/2007, respectively, in two replications each. These plants were tested by DAS-ELISA in December and April in the 2003/2004 season and in December, April, and late May in the 2006/2007 growing season.
DNA extraction and preparation of bulks
DNA was extracted from the DH-lines and the corresponding parents using a CTAB extraction protocol method modified from Murray and Thompson (1980). Pools for bulked segregant analysis (BSA, Michelmore et al. 1991) were assorted separately for the two larger DH populations, i.e. ‘SW01T5128’ (r) × ‘SW01T1000’ (s) and ‘Caletta’ (r) × ‘SW01T1000’ (s). Bulk A consisted of those DH lines which in the first year had a virus titre lower than E 405 = 0.1 (threshold determined according to Graichen 1998) until the last sampling in June, while bulk B consisted of those lines whose virus titre was initially lower than E 405 = 0.1 but exceeded this value at the last sampling date. The intention of bulk B was to exclude the possibility that lines in bulk A had escaped infection. Bulk C consisted of lines already showing a high virus titre (E 405 > 0.1) at the first sampling date in December.
Simple sequence repeat (SSR) marker analysis
A total of 200 Brassica SSR primer combinations amplifying genome-wide markers in B. napus were screened for polymorphisms among the three bulks. Most of the primers were derived from the BBSRC microsatellite program (Lowe et al. 2004; Piquemal et al. 2005), while further primers developed by the Celera AgGen Brassica Consortium were used. A focus was put on SSRs previously mapped in B. napus (e.g. Piquemal et al. 2005).
The 10 μl volume PCR reaction for the SSR analysis contained 0.5 μM of each primer, using forward primers with an M13-tail (5′-TTTCCCAGTCACGACGTT-3′) at the 5′ end and a third primer (M13uniFi, Cy5-labelled: 5′-AGGGTTTTCCCAGTCACGACGTT-3′) along with 50 ng of template DNA, 0.2 mM of each dNTP, 2.5 mM MgCl2, 0.25 U Taq polymerase (Qiagen) and the 10× PCR reaction buffer supplied with the polymerase enzyme. The PCR reactions were performed in a GeneAmp PCR Systems 9700 (Applied Biosystems) according to a touchdown protocol described by Oetting et al. (1995), using annealing steps at 68, 60, 55 or 50°C depending on the individual SSR. Fragment analyses were performed on a CEQ 8000 capillary sequencing system (Beckman Coulter).
AFLP marker analysis
Digestion and ligation of genomic DNA bulks for AFLP analysis (Vos et al. 1995) was performed using the AFLP Core Reagent Kit (Invitrogen) according to the manufacturer′s instructions, followed by preamplification with EcoRI + 0/MseI + 0 and selective amplification with EcoRI + A/MseI + C and a total of 256 Cy5-labelled EcoRI + ANN/MseI + CNN primer combinations. PCR conditions were set as described by Vos et al. (1995). Separation and detection of AFLPs was carried out on a LI-COR 4300 DNA-Analyzer (LI-COR Biosciences).
Development of sequence-tagged site (STS) markers
AFLP fragments that were polymorphic among the bulks were separated on 4.3% polyacrylamide gels (50 cm, ‘Rothiphorese Gel30’, Roth) and fragments were visualised by silver staining according to Bassam and Caetano-Anollés (1993). Polymorphic fragments were cut out of the gel and dissolved in 40 μl TE-buffer (10 mM Tris, 1 mM EDTA, pH 8).
The AFLP fragments were cloned into XL2 bacteria with tetracycline resistance (Stratagene) using competent cells developed using a protocol adapted from Inoue et al. (1990). Bidirectional sequencing was carried out on a 3130xl Genetic Analyzer (Applied Biosystems) using the BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems) with 200 ng plasmid preparation and a 1:8 dilution of the Terminator Ready Reaction Mix for each plasmid preparation. Sequence-specific primers for each fragment of interest were subsequently developed using the software primer3 (Rozen and Skaletsky 2000). The STS markers were amplified in a standard 10 μl PCR reaction.
Genetic linkage analysis
All SSR, AFLP and STS markers showing polymorphisms among the three bulks were screened in the individual plants of the three DH populations. Linkage analysis taking into account results obtained on all three populations was performed using the software JoinMap 4.0 (Van Ooijen 2006) applying the Kosambi-Transformation (Kosambi 1944).
Statistical analysis
In order to gain information on the reproducibility of the resistance tests between years, Spearman rank correlation coefficients were calculated as described by Lozán and Kausch (2007) using the software Excel (Microsoft Corporation 2003). The significance level for the correlation was calculated at n ≥ 30 and alpha = 0.05. ANOVA was carried out using Proc GLM of SAS 9.2.
Results
Genetic analyses of TuYV resistance
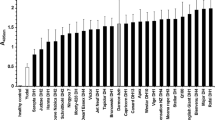
As seen in Fig. 1, the ELISA titre increased in all plants during the growing season. However, the threshold value of E 405 = 0.1 for distinguishing resistant and susceptible lines (Graichen 1994) was found to be appropriate for scoring pre- and post-winter virus levels. A good fit to a 1r:1s segregation was observed prior to winter in the first two growing seasons (December 2004 χ 21r:1s = 2.6; December 2005 χ 21r:1s = 0.73), and this segregation persisted until April in the second growing season (April 2006 χ 21r:1s = 0.01). In repeated samplings between April and June, a continuous increase of the virus titre was observed, but those lines that were scored as resistant in December never reached the virus titre of the susceptible plants. In both years, some DH lines showed a virus titre lower than E 405 = 0.1 until the last sampling in June (10 lines in June 2005 and 19 lines in May 2006). This suggests that resistance to TuYV is determined by a single major gene but is also influenced by additional environmental factors (presumably temperature) and additional minor genes.
Resistance reaction of the 111 DH-lines derived from crosses of TuYV resistant to susceptible genotypes in the years 2004/2005 (a), 2005/2006 (b) and 2006/2007 (c). Samples were collected in December 2004, April 2005, May 2005, and early June 2005, while in the following 2 years, the June samplings were omitted. The threshold for pre-winter resistance was set at a virus titre value of E 405 = 1.0, as suggested by Graichen (1998). The pre-winter (December) virus titre correspond to a 1r:1s-segregation [(χ 21r:1s = 2.6, 2004, χ 21r:1s = 0.73, 2005)] in the first two sampling years, whereas high temperatures in 2006/2007 led to increased pre-winter titre values and gave no clear segregation
The different distribution of virus titre levels in the third growing season was probably caused by the exceptionally mild autumn and winter in 2006/2007. This probably led to a long inoculation period by the aphid vectors and particularly good conditions for virus development. As a result, virus titres in December 2006 were already as high as those normally detected in June, and even DH lines with putative resistance based on their virus levels in 2004/2005 and 2005/2006 revealed extinction-values up to E 405 = 0.5 in April 2007. At the last sampling in late May 2007, all DH lines had a high virus titre (Fig. 1c).
ANOVA carried out separately for each sampling date revealed significant differences between DH lines for each sampling date [date1: MSQ line = 0.655, MSQ error = 0.102, F = 6.37; date 2: MSQ line = 1.341, MSQ error = 0.064, F = 21.04; date 3: MSQ line = 2.995, MSQ error = 0.081, F = 37.06].
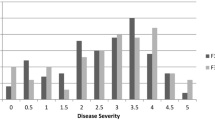
As seen in Fig. 2, the F1 hybrids also showed generally “resistant” titre levels (E 405 < 0.1) in December, indicating that the pre-winter inhibition of TuYV is inherited in a dominant manner. As for the DH lines, the virus titre increased after winter and the pre-winter virus titre levels in the F1 hybrids were higher in December 2006/2007 than in the previous growing season.
Spearman rank correlation analyses showed a high and significant correlation of the virus titres in April for the 3 years (average r s = 0.841; min r s = 0.830 for April 2006 vs. April 2007; max r s = 0.861 for April 2005 vs. April 2006). The correlations in December (Fig. 3) were slightly lower but also significant (average r s = 0.733; min r s = 0.688 for December 2004 vs. December 2005; max r s = 0.771 for December 2004 vs. December 2006). In contrast, the rank correlations among the last sampling dates in May/June were much lower, with an average of r s = 0.524 (min r s = 0.454 for May/June 2006 vs. May/June 2007; max r s = 0.561 for May/June 2005 vs. May/June 2006). From these results, it may be concluded that by the infection method used reliable phenotypic data for the virus titre estimated in December and April were obtained.
Identification of TuYV resistance-linked SSR markers
From 200 SSR primer combinations screened, only two showed polymorphisms between the resistant and the susceptible bulks. The marker Na10-D09 was polymorphic only in the 48 DH lines from the ‘Caletta’ crosses, where a 299 bp fragment was present in resistant lines and a 307 bp fragment in susceptible lines. Two recombinants were observed within these 48 genotypes.
The marker CB10347 was polymorphic in all of the crosses analysed, with a 223 bp fragment being amplified in resistant lines and a 213 bp fragment in susceptible lines. Only one recombinant was observed among 111 DH lines, indicating a tight linkage with the resistance locus. A marker amplified by the primer CB10196 was polymorphic among the bulks but showed only weak linkage to the resistance locus (Fig. 4).
Identification of TuYV resistance-linked AFLP markers
Of the 256 selective +3-AFLP primer combinations tested, a total of 42 revealed polymorphisms between the resistant and susceptible bulks. After testing of these combinations on all DH lines, six markers showed close linkage to the TuYV-resistance locus, while the AFLP fragments E31M48470, E33M50400 and E36M54380 co-segregated with the resistance (Fig. 4). The AFLP markers E31M48470, E32M50450, E33M50400, E35M52240, E36M54380, E43M56110 and E44M50380 were amplified in the resistant genotypes, whereas the markers E31M53450 and E44M52350 amplified in susceptible genotypes.
Conversion of AFLP markers to STS markers
Two of the resistance-linked AFLP markers were successfully converted into co-dominant STS markers (Fig. 4). Based on the 447 bp sequence obtained from the AFLP fragment E32M50450, the following primers were designed: forward: 5′-GATCCGTTTGGGTCTTGGTA-3′; reverse: 5′-TTGATGTGAAACGCACATTG-3′. Sequencing of the resulting amplicon STS3-e32m50-447-320 in resistant and susceptible lines revealed a 41 bp deletion in the resistant genotypes, resulting in a 344 bp fragment compared to the 385 bp amplicon from susceptible genotypes. This marker can therefore be easily scored by agarose gel electrophoresis. STS3-e32m50-447-320 co-segregated with the AFLP fragment it was developed from (Fig. 4).
Similarly, the 470 bp fragment of E31M48470 was converted into STS1-e31m48-437 bp using the following primer sequences: forward 5′-ATCGGACATTGGTCAGGTTC-3′; reverse 5′-CATACCCCACTGGTTCTTGG-3′). These primers amplify a 437 bp fragment in resistant lines, whereas a 61 bp deletion in susceptible lines results in a 376 bp fragment. This STS also co-segregates with the original AFLP fragment and also with the TuYV resistance locus.
Genetic linkage analysis
The polymorphic markers identified by the BSA were used for linkage analysis of the chromosome region surrounding the TuYV resistance locus. The relatively large distance of 4.78 cM between the SSR marker Na10-D09 and the TuYV-resistance locus is because this SSR was polymorphic only in the 48 DH lines derived from the resistance donor ‘Caletta’. In contrast, the SSR marker CB10347 was localised only 0.91 cM from the TuYV-resistance locus. A further SSR marker, CB10196, is located 13.18 cM from the resistance locus. All of the AFLP markers identified by BSA flank the resistance locus in a maximum distance of 3.73 cM, while three AFLP markers and the STS marker STS1-e31m48-437 bp co-segregate with the resistance locus. The second STS marker, STS3-e32m50-447-320, was localised 0.91 cM from the TuYV resistance. Markers Na10-D09, STS-e32m50-447-320 and STS1-e31m48-437 were analysed on additional lines derived from breeding companies and close linkage to TuYV resistance was confirmed (data not shown). Based on known locations of SSR markers linked to the TuYV-resistance locus, the chromosome containing the resistance gene is presumed to be B. napus chromosome N04 (A04).
Discussion
A good reproducibility of the resistance reaction of the DH lines was observed in the first two experiments in 2004/2005 and 2005/2006. In contrast, the 2006/2007 growing season was characterised by an unusually warm autumn and winter (up to 5.5°C higher average monthly temperature than the previous 2 years), and all plants showed greatly increased virus titres after winter. Because high temperature is known to promote the proliferation of the virus and its vector, we concluded that the resistance to TuYV is primarily determined by a dominant resistance gene but also influenced by environmental factors, particularly temperature. A potential influence of temperature on the development of the virus titre in infected plants and/or a possible breakdown of the TuYV-resistance due to temperature stress was already addressed by Dreyer et al. (2001). Similarly, Graichen (1998) reported that environmental conditions, and particularly high temperatures, could lead to a higher level of TuYV susceptibility of rapeseed. An influence of temperature and other environmental factors on resistance has also been described for other poleroviruses, e.g. Beet western yellows virus (BWYV) of lettuce (Walkey and Pink 1990) and Potato leaf roll virus (PLRV) of potato (Barker and Harrison 1984).
The high virus titres of apparently resistant lines in the 2006/2007 growing season demonstrated that the TuYV resistance of ‘R54’ is not a complete type of resistance as e.g. found for bymoviruses (cf. Ordon and Friedt 1993). The TuYV resistance rather represents a quantitative reduction of the virus titre as found in other luteoviruses, e.g. Barley yellow dwarf virus (Niks et al. 2004). Clear indications were found for a monogenic dominant inheritance of pre-winter inhibition of TuYV levels. However, the increased virus titre in plants showing effective pre-winter inhibition indicates that additional factors are involved in this quantitative resistance. These results correspond to the observations of Graichen (1998) and Dreyer et al. (2001), who also assumed that TuYV-resistance is controlled by a single major gene along with additional contributing genes.
Quantitative resistances are also well known in other pathosystems involving poleroviruses. For example, two independent dominant genes in potato together confer a strong resistance against PLRV. Individually these genes are thought to control either virus reproduction or development or short distance transport. In combination, these genes lead to a continuous variation in ELISA values for PLRV (Barker et al. 1994; Barker and Solomon 1990; Derrick and Barker 1997).
Based on previous reports together with observations made in the current work, we propose the following hypothesis with regard to the genetics of TuYV resistance of the resynthesized rapeseed line ‘R54’: (1) low virus titres persisting until the end of the growing period result from the identified dominant major gene combined with the presence of all positive alleles of minor contributing genes. (2) Rising post-winter virus titres may be due to a breakdown of the suppression of virus reproduction or development due to rising temperatures during post-winter plant development. This breakdown may be slowed down by the presence of positive alleles at minor resistance loci. (3) Plants with the recessive allele at the major gene locus encoding resistance to TuYV are able to inhibit neither pre-winter nor post-winter virus proliferation.
Comparable observations concerning a reduced virus titre that increases during the growing period have been observed for BWYV resistance in lettuce (Walkey and Pink 1990) and soybean (Duffus and Milbrath 1977). However, the main influence on yield reductions caused by TuYV seems to be the pre-winter viral inhibition. The resistant cv. ‘Caletta’ may exhibit a rising post-winter virus titre without significant yield losses, which can be considered as virus tolerance (results not shown). Therefore, for practical breeding an incorporation of the major resistance gene seems to be sufficient. As for other aphid transmitted viruses like BYDV (Scheurer et al. 2001), an incorporation of phenotypic testing using virus-bearing aphids is not practical in applied breeding programs, meaning that the availability of molecular markers is of prime importance.
The SSR, AFLP and STS markers developed in this study are closely linked to the major TuYV resistance locus and therefore represent a valuable resource for practical rapeseed breeding. The co-dominant SSR and STS markers, in particular marker STS1-e31m48-437 bp, which co-segregates with the resistance in different genetic backgrounds, are especially well suited for marker assisted selection procedures. In this respect, grouping of respective DH-lines according to the marker genotypes revealed significant differences between resistant and susceptible genotypes at the first and second sampling date (third sampling not tested, see above). At the first sampling date mean ELISA values for the resistant DH-lines were E 405 = 0.07, E 405 = 0.03 and E 405 = 0.24 in autumn 2004, 2005, and 2006, respectively, while ELISA values of those DH lines showing the susceptible marker genotype were E 405 = 0.84, E 405 = 0.28, and E 405 = 1.05. Similar results were obtained for the sampling date in spring, i.e. E 405 = 0.19, E 405 = 0.04, and E 405 = 0.47 for the resistant DH lines and E 405 = 1.60, E 405 = 1.27, and E 405 = 1.08 for the susceptible DH lines in spring 2005, 2006, 2007, respectively. Marker-assisted breeding for resistance to aphid-transmitted viruses is also broadly applied, e.g. for BYDV resistance (Ford et al. 1998; Scheurer et al. 2001; Niks et al. 2004). On the one hand, these markers facilitate selection in a very early developmental stage, e.g. DH plants may be directly screened in vitro. On the other hand, they avoid the need to rear virus-bearing aphids, which is otherwise essential for a reliable resistance screening.
BLASTn searches using the AFLP sequences with close linkage to the TuYV resistance locus gave ambiguous results. A BLASTn match to Arabidopsis thaliana genomic sequences gave significant matches (e < −50) of four markers to different positions on A. thaliana chromosomes 1, 4 and 5. According to Parkin et al. (2005), two of the relevant chromosome regions on A. thaliana chromosomes 1 and 4 could have rough synteny to B. napus chromosome N04 (A04), however, Parkin’s comparative map does not allow a definitive link between A. thaliana and other B. napus genetic maps. Using the BrGSP BLAST Server at the Brassica Genome Gateway (http://brassica.bbsrc.ac.uk/), seven of nine AFLP sequences matched significantly (e < −50) to completed B. rapa BAC clone sequences from the B. rapa genome sequencing project (status May 27, 2009). Four of the BACs that were hit originate from B. rapa chromosome A03, whereas the others derive variously from A02, A06, A08 and A10. Hence, it is difficult to draw conclusions from the sequence alignments. Although the B. rapa parent of the resistance donor ‘R54’ was also a Chinese cabbage (B. rapa ssp. pekinensis; Girke 2002), like the accession used for the B. rapa sequencing, chromosome rearrangements or deletions influencing sequences surrounding the resistance locus cannot be discounted.
On the other hand, the putative chromosomal location of the TuYV resistance in the B. napus genetic map could be confirmed using linked SSR markers from previously published maps. Two SSR markers linked to the TuYV resistance locus, CB10347 and Na10-D09, were mapped to B. napus chromosome N04 (A04) by Oregin (2005), Kim et al. (2006), Basunanda et al. (2007), Radoev et al. (2008), Sun et al. (2007) and Choi et al. (2007). Both markers were located on B. napus chromosome N09 (A09) by Piquemal et al. (2005), which could indicate homologous loci. However, the linkage block containing both markers in Piquemal’s map was separated by a gap of over 30 cM from the rest of N09, therefore, this chromosome assignment should be treated with caution. To confirm the identity of the linkage group containing the resistance locus we tested 30 additional SSR primer combinations with known loci on chromosomes N04 or N09, respectively (data not shown). None of the markers from N09 showed linkage to the TuYV resistance locus, whereas the marker CB10196 was found to be linked to the resistance locus (Fig. 4). This marker is located on chromosome N04 (A04) in all cases, we could find (Piquemal et al. 2005; Choi et al. 2007; Basunanda et al. 2007; Radoev et al. 2008). Therefore, we conclude that the major TuYV resistance gene is located on chromosome N04 (A04). This corresponds with the localisation of the major QTL for TuYV resistance by Dreyer et al. (2001) on the same chromosome. Localisation of the TuYV resistance-linked markers developed in this study into other existing B. napus genetic maps will further confirm the chromosomal location and represent a further step towards map-based cloning of the responsible resistance gene.
References
Barker H, Harrison BD (1984) Polygenic resistance to Potato leafroll virus (PLRV). In: Annual Report Scottish Crop Research Institute, pp 194–195
Barker H, Solomon RM (1990) Indication of simple genetic control of one aspect of multigenic resistance to Potato leafroll virus. In: Abstract 11th Trien Conference on EAPR, Edinburgh, p 473
Barker H, Solomon-Blackburn RM, McNicol JW, Bradshaw JE (1994) Resistance to Potato leaf roll virus multiplication in potato is under major gene control. Theor Appl Genet 88:754–758
Bassam BJ, Caetano-Anollés G (1993) Silver staining of DNA in polyacrylamide gels. Appl Biochem Biotechnol 42:181–188
Basunanda P, Spiller TH, Hasan M, Gehringer A, Schondelmaier J, Lühs W, Friedt W, Snowdon R (2007) Marker-assisted increase of genetic diversity in a double-low seed quality winter oilseed rape genetic background. Plant Breeding 126:581–587
Beuve M, Stevens M, Liu H-Y, Wintermantel WM, Hauser S, Lemaire O (2008) Biological and molecular characterization of an American sugar beet-infecting Beet western yellows virus isolate. Plant Dis 92:51–60
Choi SR, Teakle GR, Plaha P, Kim JH, Allender CJ, Beynon E, Piao ZY, Soengas P, Han TH, King GJ, Barker GC, Hand P, Lydiate DJ, Batley J, Edwards D, Koo DH, Bang JW, Park B-S, Lim YP (2007) The reference genetic linkage map for the multinational Brassica rapa genome sequencing project. Theor Appl Genet 115:777–792
Clark MF, Adams AN (1977) Characteristics of the microplate method of enzyme-linked immunosorbent assay for the detection of plant viruses. J Gen Virol 4:475–483
Derrick PM, Barker H (1997) Short and long distance spread of Potato leafroll luteovirus: effects of host genes and transgenes conferring resistance to virus accumulation in potato. J Gen Virol 78:243–251
Diederichsen E, Sacristan MD (1996) Disease response of resynthesized Brassica napus L. lines carrying different contributions of resistance to Plasmodiophora brassicae Wor. Plant Breeding 115:5–10
Dreyer F, Graichen K, Jung C (2001) A major quantitative trait locus for resistance to Turnip yellows virus (TuYV, syn. Beet western yellows virus, BWYV) in rapeseed. Plant Breeding 120:457–462
Duffus JE, Milbrath GM (1977) Susceptibility and immunity in soybean to Beet western yellows virus. Phytopathology 67:269–272
Ford CM, Paltridge NG, Rathjen JP, Moritz RL, Simpson RJ, Symons RH (1998) Rapid and informative assay for Yd2, the barley yellow dwarf virus resistance gene, based on the nucleotide sequence of a closely linked gene. Mol Breeding 4:23–31
Girke A (2002) Neue Genpools aus resynthetisiertem Raps (Brassica napus L.) für die Hybridzüchtung. Dissertation, Georg August University, Göttingen
Graichen K (1994) Nachweis von Resistenz gegenüber dem Turnip yellows luteovirus (TuYV) in Winterraps und verwandten Arten. Vortr Pflanzenzüchtung 30:132–143
Graichen K (1997) Befallssituation mit Viruskrankheiten in Winterraps im Anbaujahr 1995/96 und erste Befunde zum Jahr 1996/97. Phytomedizin 27:57–59
Graichen K (1998) Einfluß von Umweltbedingungen auf die Symptom- und Resistenzausprägung von Winterraps nach Inokulation mit dem Turnip yellows virus. Mitt Biol Bundesanst 357:291–292
Graichen K (1999) Häufig verkannt: Die Wasserrübenvergilbung an Winterraps. Der Pflanzenarzt 9–10:8–11
Graichen K, Rabenstein F (1996) European isolates of Beet western yellows virus (BWYV) from oilseed rape (Brassica napus L. ssp. napus) are non-pathogenic on sugar beet (Beta vulgaris L. var. altissima) but represent isolates of Turnip yellows virus (TuYV). J Plant Dis Prot 103:233–245
Hasan M, Seyis F, Badani AG, Pons-Kuhnemann J, Lühs W, Friedt W, Snowdon RJ (2006) Surveying genetic diversity in the Brassica napus gene pool using SSR markers. Genet Res Crop Evol 53:793–802
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28
Jones RAC, Coutts BA, Hawkes J (2007) Yield-limiting potential of Beet western yellows virus in Brassica napus. Aust J Agric Res 58:788–801
Kim JS, Chung TY, King GJ, Jin M, Yang T-J, Jin Y-M, Kim H-I, Park B-S (2006) A sequence-tagged linkage map of Brassica rapa. Genetics 174:29–39
Kosambi DD (1944) The estimation of map distance from recombination values. Ann Eugen 12:172–175
Lowe AJ, Moule C, Trick M, Edwards KJ (2004) Efficient lagre-scale development of microsatellites for marker and mapping applications in Brassica crop species. Theor Appl Genet 108:1103–1112
Lozán JL, Kausch H (2007) Angewandte Statistik für Naturwissenschaftler, 4th edn. Wissenschaftliche Auswertungen, Hamburg, pp 214–215
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: A rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Mithen RF, Magrath R (1992) Glucosinolates and resistance to Leptosphaeria maculans in wild and cultivated Brassica species. Plant Breeding 108:60–68
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acid Res 19:4321–4325
Niks RE, Habekuß A, Bekele B, Ordon F (2004) A novel major gene on chromosome 6H for resistance of barley against the Barley yellow dwarf virus. Theor Appl Genet 109:1536–1543
Oetting WS, Lee HK, Flanders DJ, Wiesner GL, Sellers TA, King RA (1995) Linkage analysis with multiplexed short tandem repeat polymorphisms using infrared fluorescence and M13 tailed primers. Genomics 30:450–458
Ordon F, Friedt W (1993) Mode of inheritance and genetic diversity of BaMMV resistance of exotic barley germplasms carrying genes different from ‘ym4’. Theor Appl Genet 86:229–233
Oregin (2005) http://www.oregin.info/Stakeholders/meetings/220305/pres/ teakle220305.pdf. Cited 22 Feb 2008
Parkin IA, Gulden SM, Sharpe AG, Lukens L, Trick M, Osborn TC, Lydiate DJ (2005) Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics 171:765–781
Piquemal J, Cinquin E, Couton F, Rondeau C, Seignoret E, Doucet I, Perret D, Villenger M-J, Vincourt P, Blanchard P (2005) Construction of an oilseed rape (Brassica napus L.) genetic map with SSR markers. Theor Appl Genet 111:1514–1523
Radoev M, Becker HC, Ecke W (2008) Genetic analysis of heterosis for yield and yield components in rapeseed (Brassica napus L.) by quantitative trait locus mapping. Genetics 179:1547–1558
Randles JW, Rathjen JP (2000) Genus Luteovirus. In: Murphy FA, Fauquet CM, Bishop DHL, Ghabrial SA, Jarvis AW, Martelli GP, Mayo MA, Summers MD (eds) Virus taxonomy. Sixth report of the international committee on taxonomy of viruses. Springer, New York, pp 379–383
Read MA, Hewson RT (1988) Prevention of Beet western yellows virus (BWYV) in winter oilseed rape by control of aphid vectors with deltamethrin. Brighton crop protection conference—pest and diseases, pp 989–997
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Rygulla W, Snowdon RJ, Eynck C, von Tiedemann A, Lühs W, Friedt W (2007) Broadening the genetic basis of Verticillium longisporum resistance in Brassica napus by interspecific hybridisation. Phytopathology 97:1391–1396
Scheurer KS, Friedt W, Huth W, Waugh R, Ordon F (2001) QTL analysis of tolerance to a German strain of BYDV-PAV in barley (Hordeum vulgare L.). Theor Appl Genet 103:1074–1083
Schliephake E, Graichen K, Rabenstein F (2000) Investigations on the vector transmission of the Beet mild yellowing virus (BMYV) and the Turnip yellows virus (TuYV). J Plant Dis Prot 107:81–87
Smith HG, Hinckes JA (1985) Luteovirus interactions between oilseed rape and sugar beet. In: Proceedings of the 1984 British Crop Protection Conference—Pest and Diseases, pp 831–835
Sun Z, Wang Z, Tu J, Zhang J, Yu F, McVetty PBE, Li G (2007) An ultradense genetic recombination map for Brassica napus, consisting of 13551 SRAP markers. Theor Appl Genet 114:1305–1317
Thomas PE, Hang AN, Reed G, Gilliland GC, Reisenauer G (1993) Potential role of winter rapeseed culture on the epidemiology of potato leaf roll disease. Plant Dis 77:420–423
Torres MW, Corrêa RL, Schrago CG (2005) Analysis of differential selective forces acting on the coat protein (P3) of the plant virus family Luteoviridae. Genet Mol Res 4:790–802
Van Ooijen JW (2006) JoinMap® 4, Software for the calculation of genetic linkage maps in experimental populations. Kyazma B. V., Wageningen
Veidt I, Lot H, Leiser M, Scheidecker D, Guilley H, Richards K, Jonard G (1988) Nucleotide sequence of Beet western yellows virus RNA. Nucleic Acids Res 16:9917–9932
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Walkey DGA, Pink DAC (1990) Studies on resistance to Beet western yellows virus in lettuce and the occurence of field sources of the virus. Plant Pathol 39:141–155
Walsh JA, Perrin RM, Miller A, Laycock DS (1989) Studies on Beet western yellows virus in winter oilseed rape (Brassica napus ssp. oleifera) and the effect of insecticidal treatment on its spread. Crop Prot 8:137–143
Acknowledgments
We gratefully acknowledge SW Seed GmbH Hadmersleben for providing the DH-lines and Norddeutsche Pflanzenzucht (NPZ) for placing the F1-hybrids at our disposal. This work was supported by grants of the Gemeinschaft zur Förderung der privaten deutschen Pflanzenzüchtung e.V. (GFP) and Arbeitsgemeinschaft industrieller Forschungsvereinigungen “Otto von Guericke” e.V. (AiF).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Visser.
Rights and permissions
About this article
Cite this article
Juergens, M., Paetsch, C., Krämer, I. et al. Genetic analyses of the host-pathogen system Turnip yellows virus (TuYV)—rapeseed (Brassica napus L.) and development of molecular markers for TuYV-resistance. Theor Appl Genet 120, 735–744 (2010). https://doi.org/10.1007/s00122-009-1194-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1194-z