Abstract
DNA polymorphisms such as insertion/deletions and duplications affecting genome segments larger than 1 kb are known as copy-number variations (CNVs) or structural variations (SVs). They have been recently studied in animals and humans by using array-comparative genome hybridization (aCGH), and have been associated with several human diseases. Their presence and phenotypic effects in plants have not been investigated on a genomic scale, although individual structural variations affecting traits have been described. We used aCGH to investigate the presence of CNVs in maize by comparing the genome of 13 maize inbred lines to B73. Analysis of hybridization signal ratios of 60,472 60-mer oligonucleotide probes between inbreds in relation to their location in the reference genome (B73) allowed us to identify clusters of probes that deviated from the ratio expected for equal copy-numbers. We found CNVs distributed along the maize genome in all chromosome arms. They occur with appreciable frequency in different germplasm subgroups, suggesting ancient origin. Validation of several CNV regions showed both insertion/deletions and copy-number differences. The nature of CNVs detected suggests CNVs might have a considerable impact on plant phenotypes, including disease response and heterosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromosomal alterations such as insertions, deletions, and differences in copy-number of genomic regions larger than 1 kb are known as copy-number variants (CNVs) or structural variations (SVs) (Feuk et al. 2006). Human CNV analysis began in the early 1990s with comparative genomic hybridization (CGH; Kallioniemi et al. 1992). It scaled up to whole-genome analysis of CNVs with the use of microarrays to perform CGH (aCGH) (Geschwind et al. 1998; Pinkel et al. 1998; Pollack et al. 1999; Solinas-Toldo et al. 1997). In this technique, genomic DNA is labeled with fluorescent dyes and hybridized to microarrays. Microarray hybridization signals that deviate significantly from the expected 1:1 ratio between sample and reference genomes indicate copy-number differences (Feuk et al. 2006). However, when multiple copies of a sequence are present in the genomes, absolute quantification of the number of copies and precise location of the polymorphic DNA segments are not possible by aCGH. Different array platforms have been used for aCGH experiments (Gresham et al. 2008), including arrays primarily designed for applications such as SNP genotyping (Zhao et al. 2004) and gene expression (Auer et al. 2007; Skvortsov et al. 2007).
Copy-number variants have been detected by aCGH in humans (Iafrate et al. 2004; McCarroll et al. 2008; Redon et al. 2006; Sebat et al. 2004), chimpanzee (Perry et al. 2008), rat (Guryev et al. 2008), mouse (Lakshmi et al. 2006; She et al. 2008), Drosophila (Emerson et al. 2008), yeast (Infante et al. 2003), and E. coli (Skvortsov et al. 2007). It has also been shown that rat and humans share orthologous genes in CNV regions (Guryev et al. 2008). Many CNVs detected by aCGH were associated with cancer (Beroukhim et al. 2007; Pinkel et al. 1998; Pollack et al. 1999, 2002; Shlien et al. 2008; Solinas-Toldo et al. 1997; Zhao et al. 2004; reviewed in Kallioniemi 2008), autoimmune susceptibility (Aitman et al. 2006; Fanciulli et al. 2007), including HIV susceptibility (Gonzalez et al. 2005), schizophrenia (Xu et al. 2008), and others human diseases.
In addition to causing diseases, effects of CNVs in fitness and gene expression have been reported. Amylase genes exhibit variation in their number of copies in humans (Iafrate et al. 2004) and their copy-number in the genome has been under selection (Perry et al. 2007). Individuals from populations with a higher starch-based diet had more copies of amylase genes in the genome and higher amylase protein level than individuals from populations with low-starch diet (Perry et al. 2007). CNVs detected among 15 female isolines of Drosophila have been under purifying selection (Emerson et al. 2008). Additionally, an example of the impact of gene copy-number has been illustrated in yeast where a co-inducer and a galactokinase arose from a common bi-functional ancestor gene and acquired different regulation, thereby increasing the fitness of the organism (Hittinger and Carroll 2007). Genes with higher copy-number had higher gene expression in rat (Guryev et al. 2008). A dramatic phenotypic change due to a CNV affecting gene regulation was described in tomatoes where an insertion of 6–8 kb down-regulated the expression of a transcription factor resulting in the increase of tomato fruit (Cong et al. 2008). In a comparison between the effect of SNPs and CNVs in human gene expression, CNVs were found to capture almost 18% of the gene expression variation with little overlap with the variation captured by SNPs (Stranger et al. 2007).
The majority of the CNVs detected in Drosophila were duplications rather than deletions, and they varied in frequency when contained in intergenic regions, introns, exons or entire genes (Emerson et al. 2008). The number of base-pairs delimited by CNVs might be higher than the number of bases in SNPs for any particular genome (Feuk et al. 2006). The proportion of nucleotides contained within CNVs is approximately 5% of the mouse genome (20 Mb among 15 mouse inbreds tested) (She et al. 2008), 2% of the Drosophila genome (Emerson et al. 2008), 0.8% (22 Mb) between two strains of rat (Guryev et al. 2008), and 5% of the human genome considering regions larger than 50 kb in 270 individuals (McCarroll et al. 2008). The number of CNVs detected varied among different studies depending on the species, samples, and aCGH platforms used. In humans for example, a first report using aCGH in 270 individuals of the HapMap population identified 1,447 CNVs (Redon et al. 2006) while a second report using the same population detected 3,048 CNVs using a new aCGH platform built to detect CNVs and SNPs (McCarroll et al. 2008). CNVs can arise in somatic tissues, creating somatic mosaicism. They were identified by two different aCGH platforms in a study comparing 19 pairs of both normal and diseased human monozygotic twins (Bruder et al. 2008).
In plants, large genome structural variations affecting the intraspecific collinearity have been reported (Fu and Dooner 2002; Brunner et al. 2005). aCGH has been used in plants to detect single-feature polymorphisms in Arabidopsis (Borevitz et al. 2003) and rice (Kumar et al. 2007), to detect structural variations created by mutagenesis in rice (Bruce et al. 2009) and Citrus (Ríos et al. 2008), and as a molecular marker for genotyping in Arabidopsis (Salathia et al. 2007).
To investigate the extent of CNVs in the maize genome, we performed aCGH using a microarray designed for gene expression studies (102,000 60-mer oligonucleotide probes) focusing the analysis on genic regions. Thirteen maize inbred lines were compared to the B73 reference inbred. Conservative analysis allowed us to detect CNVs with high validation rate. Several CNVs identified were detected in more than one inbred, suggesting their appreciable allelic frequency in the maize populations. Some CNVs represented large inserted/deleted regions. The same variants were present in different heterotic groups, suggesting that the events responsible for the creation of these CNVs predate this population stratification.
Materials and methods
Genomic DNA preparation
Genomic DNA was isolated from fresh-frozen leaf samples (Table 1) with DNeasy Plant Mini Kits (Qiagen, Valencia, CA, USA), including an incubation with RNAse A, following the instructions of the manufacturer. Total DNA was quantified with a spectrophotometer and electrophoresed on 0.7% agarose gel for integrity checking.
aCGH experiments
Custom Agilent (Agilent Technologies, Santa Clara, CA, USA) 2 × 105 K microarrays containing 102,353 unique 60-mer oligonucleotides designed from 45,028 maize ESTs and unigenes were used (M. Beatty, Pioneer Hi-Bred, unpublished results). This array was originally designed for gene expression, and contains multiple probes per maize ESTs or unigenes. For each aCGH hybridization, 2 μg of genomic DNA was digested with the restriction enzymes AluI and RsaI (Promega, Madison, WI, USA). After 2 h of incubation, the samples were heated to 65°C for 20 min to inactivate the enzymes. The fragmented DNA was labeled via a random primed labeling reaction (Agilent Oligonucleotide Array-Based CGH for Genomic DNA Analysis, v4.0) that incorporated Cy3-UTP into the product. The labeled DNA was filtered with a Microcon YM-30 column (Millipore, Billerica, MA, USA) to remove unincorporated nucleotides and quantified with a spectrophotometer to measure yield and dye incorporation rates. Hybridization and blocking buffers (Agilent Technologies, Santa Clara, CA, USA) were added to the samples prior to denaturation at 95°C for 3 min and incubation at 37°C for 30 min. Each sample was hybridized to an array for 40 h at 65°C while rotating at 10 rpm. The arrays were disassembled and washed in Oligo aCGH Wash Buffer 1 (Agilent Technologies, Santa Clara, CA, USA) at room temperature for 5 min. A second wash was performed in Oligo aCGH Wash Buffer 2 (Agilent Technologies, Santa Clara, CA, USA) for 1 min at 37°C. Slides were then dipped in acetonitrile and air dried. An Agilent G2505B DNA microarray scanner was utilized to capture TIF images at two laser power settings (100 and 10%).
Data analysis
The microarray images were visually inspected for image artifacts. Feature intensities were extracted, filtered, and normalized with Agilent’s Feature Extraction Software (version 9.5.1). Further quality control was performed using data analysis tools in Resolver Database (Rosetta Biosoftware, Seattle, WA, USA), and ratios between the normalized intensities of all maize lines and B73 were computed. The genomic position of each microarray probe was determined by searching for their sequence in the B73 BAC sequences using BLAST (Altschul et al. 1990). Specific coordinates for each BAC were obtained from a B73 BAC contig map (Fengler et al. 2007). Probes with fewer than ten matches to the B73 genome, at least 58 out of 60 matched nucleotides, no gaps, and signal above the background (~ 100 rfu) in at least one maize line being compared, were used in subsequent data analysis. Probes with more than one match to the same contig had their coordinates averaged. For each probe, the log2 of signal ratio between the two genomes was plotted against its genomic position. Conservatively, a CNV was declared only when at least three probes in the same genomic position exhibited twofold or higher hybridization ratio. The identification of CNVs along the chromosomes was facilitated by using R (R Development Core Team 2009) scripts developed for this purpose. Correlation between number of aCGH probes and number of CNVs detected along the genome were calculated by dividing each maize chromosome in ten bins and counting the number of aCGH probes and CNVs present in each bin. The correlation is presented as an average over all maize chromosomes.
Validation of CNVs
Validation of CNVs were performed by PCR with primers externally flanking the microarray probe sequences. This provided a test that is independent of the DNA sequences present in the microarray probes. In addition, to reduce the possibility of interference from SNPs in the primer annealing sites, we designed two non-overlapping pairs of primers to span each of the 110 probe sequences and neighboring regions, distributed in 15 CNVs. The pair of primers with most reproducible amplification was selected for analysis (Supplementary Table 1).
The genome positions of the IDP markers (Fu et al. 2006) in the regions of candidate CNVs were determined by matching their primer sequences to the B73 sequenced BACs and sorting them based on physical map positions. IDP markers without sequence matches to B73 BACs were placed within the set using their genetic positions.
Results
The sequence search of the 102,353 60-mer microarray probes in the B73 reference genome (version 3a.50, http://www.maizesequence.org) resulted in the match of 60,472 (59% of the probes) probes to 75,564 loci (1.25 matches per probe), of which 17,501 were unique. We identified CNVs in the maize genome as defined by three or more probes in the same genomic position showing differences equal or greater than two fold between genotypes.
A total of 2,109 putative CNVs were identified, including 248 between Mo17 and B73 (Table 1). Of those, 878 (42%) were detected only once, including one present only in B73. The remaining 1,231 (58%) of the CNVs were detected at least twice (Fig. 1). CNVs detected in at least three of the inbreds represent 37% (790) of the total. These CNVs represent an average of 438 ± 186 SD (standard-deviation) CNVs between each maize inbred line in comparison to B73. Considering only CNVs detected in at least two maize inbred lines, the average is 371 ± 146 SD. A more stringent criterion of identifying CNVs with a minimum of four probes detected averages of 144 ± 66 SD for all CNVs and 116 ± 48 SD CNVs present in at least two maize genomes.
CNVs were dispersed along the entire maize genome with a higher density toward the telomers and lower density toward peri-centromeric regions in a similar pattern in relation to probe density (correlation = 0.9565; Fig. 2). The majority of the CNVs detected between B73 and other inbreds (57%) represent more copies in B73 or presence in B73 and absence in the other inbreds. This reflects the fact that most of the probes were designed from B73 sequences; therefore, the sequences deleted in B73 were not represented on the array. The extremes were 36 and 82% when PHW52 and Mo17 were compared to B73, respectively. The number of consecutive probes indicative of CNVs varied, suggesting that the regions encompassing CNVs are of variable size (Supplementary Fig. 1). In particular, the short arm of chromosome six has been identified as a region of extreme size differences among the inbreds tested (Supplementary Fig. 1).
CNVs detected along the maize genome by aCGH. For each of the maize chromosomes the first track shows CNVs detected between Mo17 and B73. CNV regions are labeled according to the presence of more copies in Mo17 (green arrows pointing up) or B73 (red arrows pointing down). The second track shows CNVs detected between all inbred lines and B73 (black vertical bars). The third track shows the density of the microarray probes along the chromosome. Centromeres are shown as gray boxes and CNVs selected for PCR validation are identified by blue triangles. Coordinates along the chromosomes are given as bands in a maize physical map (Fengler et al. 2007)
Variation in aCGH experiments and CNV detection
In order to investigate the reproducibility of CNV detection by aCGH, we first compared the hybridization results of two independent sample preparations of the maize inbred line PHP38. Using stringent criteria, no CNVs were detected between these two samples of PHP38 (Table 2; Supplementary Fig. 2). With less stringent criteria of at least two or a single probe located at the same genomic position and showing at least twofold differences in hybridization intensities, one or 129 false positives, were detected, respectively.
In a second experiment, we performed aCGH of three independent samples of B73 and Mo17. From those, one B73 hybridization was excluded from further analysis due to excessive signal variation. The remaining hybridizations were compared both between and within genotypes. Even though the correlations between intensities of different hybridizations were high, considerable variability was identified among the aCGH comparisons (Table 2). A total of 695 CNVs were detected in the two comparisons of Mo17 to B73, of which 341 were detected in both comparisons. These numbers of CNVs are higher than the number of CNVs we detected in our first comparison between Mo17 and B73 (Table 1). The comparisons between aCGH slides of the same inbred, PHP38 or Mo17, showed no or low number of CNVs detected. However, the comparison of two B73 samples revealed 58 presumed false CNVs. While this is considerably fewer than detected in between-genotype comparisons (Table 2), stringent validation of CNVs was required.
Validation of CNVs by PCR
We used PCR to further analyze 15 putative CNVs detected by aCGH between Mo17 and B73, dispersed along the maize genome and showing high hybridization ratios (Fig. 2; Table 3). We confirmed 12 of those CNVs as insertion/deletion polymorphisms (Table 3) and identified their allelic versions in the remaining inbreds. Some of the validated CNV regions were defined by few probes and represent relatively short DNA fragments while others encompass at least several hundreds of kilobases.
The largest CNV was found in the short arm of chromosome six (Fig. 3; Supplementary Fig. 1). Of the 18 probes matching the region, eight have two or more fold-differences in hybridization signal between Mo17 and B73 (Fig. 3a). PCR primers spanning these eight probes amplified expected products in B73 but not in Mo17 (Fig. 3b). This suggests the whole DNA segment is absent in Mo17. Additionally, one of the remaining ten probes with fold-changes lower than two did not amplify in Mo17. Other six of those ten probes match more than one region in the B73 genome (Fig. 3b); therefore, their observed hybridization ratios represent an average over all matching regions. This CNV region spans several BACs comprising a DNA segment up to a few megabases in length (Fig. 3b). EST data on the BACs encompassing the eight PCR amplicons suggest that there are at least 23 genes or pseudo-genes in this DNA segment (data not shown). To further validate and delimit this CNV, we also tested 15 previously described indel (IDP) markers (Fu et al. 2006) present in and around 16 loci of the putative deletion and obtained an equivalent result, i.e., the deleted allele was always in Mo17 (Fig. 3b). Interestingly, the IDP markers span a region significantly larger than the region identified by aCGH. We physically and genetically mapped a total of 61 consecutive IDP markers on the short arm of chromosome six (data not shown) that have been reported to amplify fragments in B73 but not in Mo17 (Fu et al. 2006). It is unlikely that 15 or more consecutive markers present in B73 but absent in Mo17 occur by chance alone (P ≤ 5 × 10−4), considering that about 60% of the IDP markers are of the type present in B73/absent in Mo17. In addition to Mo17, this large deletion occurs in four other inbreds from two distinct heterotic groups: non-stiff stalk (NSS) and stiff-stalk (SSS) (Fig. 4).
Large CNV detected in the short arm of chromosome six at 69.2 cM. a Mo17/B73 hybridization intensity ratios of microarray probes as a function of the probe position on maize physical map. Probes selected for PCR validation are shown as red circles and probes matching the B73 genome more than once are shown as blue circles. Solid and dashed horizontal red lines correspond to one and two standard-deviations of log2 of intensity ratios, respectively, calculated for all probes in that experiment. The blue curve is a loess function (R Development Core Team 2009) based on the aCGH ratio. b Detailed view of the CNV region showing B73 BACs, microarray probes with corresponding ID-numbers (log2 ratio and number of matches to B73 genome in parentheses), IDP markers, and PCR results of B73 (left lane) and Mo17 (right lane) for both PCR spanning microarray probes and IDP markers. Probes with more than twofold log2 ratio are shown in red. *Sequence of PCR amplicons and IDP markers match the CNV region more than once
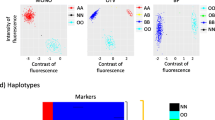
PCR validation of the biggest CNV identified on chromosome six (Fig. 3) in different maize inbred lines. For each probe the log2(aCGH ratio) and PCR amplification is shown. Inbred lines were classified as belonging to non-stiff-stalk synthetic (NSS) or stiff-stalk synthetic (SSS) heterotic groups
A second large CNV detected in the short arm of chromosome six also spans several BACs (Fig. 5). PCR amplification of fragments spanning five microarray probes produced expected results in B73, but no PCR products were detected in Mo17 (Fig. 5b). This deletion variant was also present in the same four inbreds that lack the largest CNV described above. The PCR results were in agreement with the observed probe ratios (Fig. 6).
A second large CNV detected in the short arm of chromosome six at 69.2 cM, adjacent to the CNV in Fig. 3. a aCGH view. Probes selected for PCR validation are shown as red circles and probes matching the B73 genome more than once are shown as blue circles. Solid and dashed horizontal red lines correspond to one and two standard-deviations of log2 of intensity ratios, respectively, calculated for all probes in that experiment. The blue curve is a loess function (R Development Core Team 2009) based on the aCGH ratio. Coordinates along the chromosomes are given as bands in a maize physical map (Fengler et al. 2007). b Detailed view showing B73 BACs, microarray probes with corresponding ID numbers (number of matches to B73 genome in parenthesis), IDP markers, and PCR validation between B73 (left lane) and Mo17 (right lane). Probes with more than twofold log2 ratio are shown in red
PCR validation of a CNV on chromosome six (Fig. 5) in other inbreds. For each probe the log2(aCGH ratio) and PCR amplification is shown. Inbred lines were classified as belonging to non-stiff-stalk synthetic (NSS) or stiff-stalk synthetic (SSS) heterotic groups
A CNV located on chromosome five (530 cM) generated PCR products with two distinct results (Supplementary Fig. 3). Of the 11 PCR products located in the center of the CNV region, five amplified in both B73 and Mo17 while the other six only amplified in B73 (Supplementary Fig. 3B). The five amplicons present in both B73 and Mo17 match from five to eight places the B73 genome and were also amplified in other inbreds (Supplementary Fig. 3C). The six amplicons, which did not amplify in Mo17, produced amplification products of varying sizes in other inbreds (Supplementary Fig. 3C). Three of these amplicons also match other loci in the B73 genome. These PCR results suggest the presence of two CNVs in the region: one with a DNA segment present in B73 and not in Mo17, and other with different number of copies between B73 and Mo17.
The three CNV regions that could not be validated by PCR showed expected products from both B73 and Mo17. They may represent DNA sequences differing in copy-number between the two genomes, either in tandem or in several dispersed loci throughout the genome. Two of these CNV regions, one on chromosomes eight (206 cM) and other on chromosome nine (300.2 cM), are related by sequence and share four microarray probes with 3–5 matches to the B73 genome (Supplementary Fig. 4). Array CGH data indicates that Mo17 is missing at least one but not all of these copies.
Discussion
We detected 2,109 dispersed as well as clustered CNVs in the maize genome among 14 maize inbred lines with aCGH (Fig. 2). Several factors affected our estimation of the number of CNVs detected between different maize inbred lines against B73 (Table 1). First, the microarray platform used for aCGH was primarily developed for gene expression. It contains probes spanning introns and in some cases representing sequences of transcripts from inbreds other than B73. Consequently, we were able to match 60,472 of the 102,353 microarray features to 75,564 locations on the presently available incomplete B73 genome sequence (version 3a.50, http://www.maizesequence.org). In addition, the distribution of genes along the maize genome is not uniform (Fengler et al. 2007); therefore, microarray probes and the CNVs we detected showed a similar distribution with fewer probes in the paracentromeric regions (Fig. 2). Second, the majority of the probes were designed to be complementary to the B73 allele, and therefore we could not detect sequences present in the other maize inbreds but absent from B73. Third, hybridization signals of some of the microarray probes may deviate from the expected 1:1 ratio due to DNA polymorphisms between the maize genomes used in our experiments. This could have affected the hybridization of DNA fragments containing SNPs, even though the 60-mer probes are not highly sensitive to up to two mismatches within the target sequence (Hughes et al. 2001). To reduce errors in CNV detection due to SNPs, we called a CNV only when three or more probes within the same physical position deviated significantly from the expected 1:1 ratio. Thus, SNPs affecting one or two probes did not lead to false calling of a CNV. As a consequence of this approach and the relatively low probe density along the genome, our method to detect CNVs is biased toward detection of large CNVs. Fourth, the majority of the B73 BACs are in phase one of sequencing, i.e., they contain gaps and unordered contigs. This limited our ability to accurately assign physical genome coordinates for the microarray probes. Consequently, some of the algorithms developed for human aCGH analysis assuming high-density ordered probes (Olshen et al. 2004; Price et al. 2005) could not be used. Finally, the detection of CNVs in repeated preparations of the same genotypes (Table 2) showed an appreciable variability despite the precautions taken. Together, these considerations suggest that the number of CNVs identified (Table 1) is an underestimate, especially with respect to small CNVs, and that the methodology favors detection of large insertion–deletion variants.
The application of a moderate-density oligonucleotide-based array to a highly polymorphic large plant genome containing nearly 80% of repetitive sequences poses special challenges. In the future, the availability of a finished genome sequence and of a high-density microarray platform specifically designed for aCGH would greatly improve the detection and estimation of CNVs. Next-generation sequencing techniques are rapidly improving, and offer advantages over aCGH by allowing direct detection of DNA variations and detection of their recombination breakpoints (Campbell et al. 2008; Chen et al. 2008). They also offer the advantage of detecting inversions and translocations that would not be detected by aCGH. Its use for CNV detection might be a reasonable option, when a reference genome is available to facilitate assembly and costs are not a limitation. However, aCGH might still perform better in genomic regions with several copies, where the assembly of next-generation sequence reads with low-to-moderate genome coverage might be difficult. Poor sequence assembly would also compromise the detection of CNVs whose sequences are deleted in the reference genome.
Of the 12 maize CNV regions validated by PCR (Fig. 2; Table 3) two are large insertions/deletions spanning at least hundreds of kilobases (Figs. 3, 5) and the others are CNVs smaller than 10 kb. Similarly, in the human genome long CNVs are less frequent (McCarroll et al. 2008). Although we cannot formally exclude the possibility that the indel CNVs validated by PCR result from extremely high-polymorphic regions of the genome affecting PCR primer annealing, this is unlikely. The design of two independent pairs of primers to amplify several DNA fragments in each CNV region greatly reduced this risk. Another potential problem would be the presence of densely scattered micro-indels present in one of the alleles along the CNV region. This could lead to the validation of deleted fragments by PCR that do not represent a single CNV; however, we believe this is unlikely for all CNVs validated. The three CNVs that could not be validated by PCR (Table 3) are likely to be DNA fragments present in both B73 and Mo17 with different number of copies in their genomes (Supplementary Fig. 4). Quantitative methods such as quantitative PCR would have to be used to validate those CNVs and to estimate their copy-number in the maize genome.
About half of the CNVs identified occur in only one of the 13 inbreds compared to B73 (Fig. 1). This apparent high rare allele frequency may suggest non-neutral fitness or recent origin, but more data is necessary for quantitative analysis. On the other hand, detection of many CNVs in several different maize inbreds (Figs. 1 2) suggests that they are present at an appreciable allelic frequency in the current cultivated maize. Although at present little is known about the mutation rate of CNVs, the detection of many CNVs in both major heterotic group suggests that at least some of them represent ancestral events predating the development of these germplasm groups (Figs. 5, 6). This has significant implications for plant genome sequencing, genetics, and breeding. Several diverse maize inbreds will have to be sequenced to identify all genes and to estimate the time of appearance of such sequence variants. As an alternative, DNA sequencing of CNV regions could be used to identify such new DNA sequences in relation to the reference genome. Further characterization of CNVs by DNA sequencing would generate important information about their DNA content and flanking sequences. This data may reveal the mechanisms of CNV creation.
CNVs might affect diversity and recombination within and around directly affected regions. In primates, rodents, fruitfly, rice, and yeast, single-nucleotide mutation rate increases close to insertions/deletions (Tian et al. 2008). In maize, repetitive regions have lower recombination rates when compared to other genomic regions (Fu et al. 2002), and insertion/deletion polymorphisms decrease recombination in their vicinity (Dooner and Martínez-Férez 1997), affecting genetic to physical distance ratios. These examples support the need to detect CNVs and understand the mechanisms involved in their creation and their relationship with recombination. It was recently demonstrated in humans that most CNVs are in linkage disequilibrium (LD) with SNPs and that LD decay of the two happen at similar rates (McCarroll et al. 2008). Based on this study, the utility of CNVs as genetic markers would be redundant to other molecular markers. However, many plants, including maize, have very active transposon systems, which continuously create new polymorphisms. It has been demonstrated that transposable elements can cause macrotransposition generating large structural variations (Huang and Dooner 2008). Such recent genomic rearrangements on the background of ancestral haplotypes could be easily detected by aCGH.
The relative importance of CNVs will be higher if they contain regulatory regions and genes. Therefore, extensive characterization and experiments dedicated to evaluate their direct effect on plant phenotypes would have to be performed. There is evidence that many of the non-colinear sequences in maize are repeats or transposons, some of them containing pseudogenes (Fu and Dooner 2002; Brunner et al. 2005) and also that CNVs, including genic insertions/deletions, contribute to phenotype (Cong et al. 2008; Grant et al. 1995, 1998). It has been shown in Arabidopsis and Brassica that the RPM1 gene conferring resistance to Pseudomonas syringae was absent in susceptible lines (Grant et al. 1995, 1998). In maize, a major QTL for resistance to sugar-cane mosaic virus was identified between the inbreds Siyi (resistant) and Mo17 (susceptible) (Mingliang Xu, personal communication) in the biggest CNV region reported here (Fig. 3). Rcg1, a gene conferring resistance to Colletotrichum graminicola in maize has been recently cloned (Petra Wolters, personal communication). Rcg1 is located on a 230-kb insertion absent in most of the North American maize germplasm. These examples show the importance of CNVs to plant disease response by direct affecting causative genes.
Global plant phenotypes such as hybrid vigor (heterosis) could also be affected by CNVs. The complementation of deleted or inactive genes in hybrids is one of the possible mechanisms of heterosis according to the dominance hypothesis (Springer and Stupar 2007). In addition, the dominance was recently demonstrated to be the predominant genetic component of yield QTLs in maize (Garcia et al. 2008). As the genes present in the inserted/deleted CNVs are probably not essential for viability, it is likely that functional members of partially redundant gene families and/or genes related to environmental response are located in these regions. Based on these observations, complementation of such genes or regulatory regions in hybrids by the presence of a large number of CNVs could contribute to hybrid vigor.
References
Aitman TJ, Dong R, Vyse TJ, Norsworthy PJ, Johnson MD, Smith J, Mangion J, Roberton-Lowe C, Marshall AJ, Petretto E, Hodges MD, Bhangal G, Patel SG, Sheehan-Rooney K, Duda M, Cook PR, Evans DJ, Domin J, Flint J, Boyle JJ, Pusey CD, Cook HT (2006) Copy number polymorphism in Fcgr3 predisposes to glomerulonephritis in rats and humans. Nature 439:851–855
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Auer H, Newsom DL, Nowak NJ, McHugh KM, Singh S, Yu C, Yang Y, Wenger GD, Gastier-Foster JM, Kornacker K (2007) Gene-resolution analysis of DNA copy number variation using oligonucleotide expression microarrays. BMC Genomics 8:111
Beroukhim R, Getz G, Nghiemphu L, Barretina J, Hsueh T, Linhart D, Vivanco I, Lee JC, Huang JH, Alexander S, Du J, Kau T, Thomas RK, Shah K, Soto H, Perner S, Prensner J, Debiasi RM, Demichelis F, Hatton C, Rubin MA, Garraway LA, Nelson SF, Liau L, Mischel PS, Cloughesy TF, Meyerson M, Golub TA, Lander ES, Mellinghoff IK, Sellers WR (2007) Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc Natl Acad Sci USA 104:20007–20012
Borevitz JO, Liang D, Plouffe D, Chang H, Zhu T, Weigel D, Berry CC, Winzeler E, Chory J (2003) Large-scale identification of single-feature polymorphisms in complex genomes. Genome Res 13:513–523
Bruce M, Hess A, Bai J, Mauleon R, Diaz MG, Sugiyama N, Bordeos A, Wang GL, Leung H, Leach J (2009) Detection of genomic deletions in rice using oligonucleotide microarrays. BMC Genomics 10:129
Bruder CEG, Piotrowski A, Gijsbers AACJ, Andersson R, Erickson S, de Stahl TD, Menzel U, Sandgren J, von Tell D, Poplawski A, Crowley M, Crasto C, Partridge EC, Tiwari H, Allison DB, Komorowski J, van Ommen GB, Boomsma DI, Pedersen NL, den Dunnen JT, Wirdefeldt K, Dumanski JP (2008) Phenotypically concordant and discordant monozygotic twins display different DNA copy-number-variation profiles. Am J Hum Genet 82:763–771
Brunner S, Fengler K, Morgante M, Tingey S, Rafalski A (2005) Evolution of DNA sequence nonhomologies among maize inbreds. Plant Cell 17:343–360
Campbell PJ, Stephens PJ, Pleasance ED, O’Meara S, Li H, Santarius T, Stebbings LA, Leroy C, Edkins S, Hardy C, Teague JW, Menzies A, Goodhead I, Turner DJ, Clee CM, Quail MA, Cox A, Brown C, Durbin R, Hurles ME, Edwards PAW, Bignell GR, Stratton MR, Futreal PA (2008) Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat Genet 40:722–729
Chen W, Kalscheuer V, Tzschach A, Menzel C, Ullmann R, Schulz MH, Erdogan F, Li N, Kijas Z, Arkesteijn G, Pajares IL, Goetz-Sothmann M, Heinrich U, Rost I, Dufke A, Grasshoff U, Glaeser B, Vingron M, Ropers HH (2008) Mapping translocation breakpoints by next-generation sequencing. Genome Res 18:1143–1149
Cong B, Barrero LS, Tanksley SD (2008) Regulatory change in YABBY-like transcription factor led to evolution of extreme fruit size during tomato domestication. Nat Genet 40:800–804
Dooner HK, Martínez-Férez IM (1997) Recombination occurs uniformly within the bronze gene, a meiotic recombination hotspot in the maize genome. Plant Cell 9:1633–1646
Emerson JJ, Cardoso-Moreira M, Borevitz JO, Long M (2008) Natural selection shapes genome-wide patterns of copy-number polymorphism in Drosophila melanogaster. Science 320:1629–1631
Fanciulli M, Norsworthy PJ, Petretto E, Dong R, Harper L, Kamesh L, Heward JM, Gough SCL, de Smith A, Blakemore AIF, Froguel P, Owen CJ, Pearce SHS, Teixeira L, Guillevin L, Graham DSC, Pusey CD, Cook HT, Vyse TJ, Aitman TJ (2007) FCGR3B copy number variation is associated with susceptibility to systemic, but not organ-specific, autoimmunity. Nat Genet 39:721–723
Fengler K, Allen SM, Li B, Rafalski A (2007) Distribution of genes, recombination, and repetitive elements in the maize genome. Crop Sci 47:S83–S95
Feuk L, Carson AR, Scherer SW (2006) Structural variation in the human genome. Nat Rev Genet 7:85–97
Fu H, Dooner HK (2002) Intraspecific violation of genetic colinearity and its implications in maize. Proc Natl Acad Sci USA 99:9573–9578
Fu H, Zheng Z, Dooner HK (2002) Recombination rates between adjacent genic and retrotransposon regions in maize vary by 2 orders of magnitude. Proc Natl Acad Sci USA 99:1082–1087
Fu Y, Wen T, Ronin YI, Chen HD, Guo L, Mester DI, Yang Y, Lee M, Korol AB, Ashlock DA, Schnable PS (2006) Genetic dissection of intermated recombinant inbred lines using a new genetic map of maize. Genetics 174:1671–1683
Garcia AAF, Wang S, Melchinger AE, Zeng Z (2008) QTL mapping and the genetic basis of heterosis in maize and rice. Genetics 180:1707–1724
Geschwind DH, Gregg J, Boone K, Karrim J, Pawlikowska-Haddal A, Rao E, Ellison J, Ciccodicola A, D’Urso M, Woods R, Rappold GA, Swerdloff R, Nelson SF (1998) Klinefelter’s syndrome as a model of anomalous cerebral laterality: testing gene dosage in the X chromosome pseudoautosomal region using a DNA microarray. Dev Genet 23:215–229
Gonzalez E, Kulkarni H, Bolivar H, Mangano A, Sanchez R, Catano G, Nibbs RJ, Freedman BI, Quinones MP, Bamshad MJ, Murthy KK, Rovin BH, Bradley W, Clark RA, Anderson SA, O’connell RJ, Agan BK, Ahuja SS, Bologna R, Sen L, Dolan MJ, Ahuja SK (2005) The influence of CCL3L1 gene-containing segmental duplications on HIV-1/AIDS susceptibility. Science 307:1434–1440
Grant M, Godiard L, Straube E, Ashfield T, Lewald J, Sattler A, Innes R, Dangl J (1995) Structure of the Arabidopsis RPM1 gene enabling dual specificity disease resistance. Science 269:843–846
Grant M, McDowell J, Sharpe A, de Torres Zabala M, Lydiate D, Dangl J (1998) Independent deletions of a pathogen-resistance gene in Brassica and Arabidopsis. Proc Natl Acad Sci USA 95:15843–15848
Gresham D, Dunham MJ, Botstein D (2008) Comparing whole genomes using DNA microarrays. Nat Rev Genet 9:291–302
Guryev V, Saar K, Adamovic T, Verheul M, van Heesch SAAC, Cook S, Pravenec M, Aitman T, Jacob H, Shull JD, Hubner N, Cuppen E (2008) Distribution and functional impact of DNA copy number variation in the rat. Nat Genet 40:538–545
Hittinger CT, Carroll SB (2007) Gene duplication and the adaptive evolution of a classic genetic switch. Nature 449:677–681
Huang JT, Dooner HK (2008) Macrotransposition and other complex chromosomal restructuring in maize by closely linked transposons in direct orientation. Plant Cell 20:2019–2032
Hughes TR, Mao M, Jones AR, Burchard J, Marton MJ, Shannon KW, Lefkowitz SM, Ziman M, Schelter JM, Meyer MR, Kobayashi S, Davis C, Dai H, He YD, Stephaniants SB, Cavet G, Walker WL, West A, Coffey E, Shoemaker DD, Stoughton R, Blanchard AP, Friend SH, Linsley PS (2001) Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nat Biotechnol 19:342–347
Iafrate AJ, Feuk L, Rivera MN, Listewnik ML, Donahoe PK, Qi Y, Scherer SW, Lee C (2004) Detection of large-scale variation in the human genome. Nat Genet 36:949–951
Infante JJ, Dombek KM, Rebordinos L, Cantoral JM, Young ET (2003) Genome-wide amplifications caused by chromosomal rearrangements play a major role in the adaptive evolution of natural yeast. Genetics 165:1745–1759
Kallioniemi A (2008) CGH microarrays and cancer. Curr Opin Biotechnol 19:36–40
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, Pinkel D (1992) Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 258:818–821
Kumar R, Qiu J, Joshi T, Valliyodan B, Xu D, Nguyen HT (2007) Single feature polymorphism discovery in rice. PLoS One 2:e284
Lakshmi B, Hall IM, Egan C, Alexander J, Leotta A, Healy J, Zender L, Spector MS, Xue W, Lowe SW, Wigler M, Lucito R (2006) Mouse genomic representational oligonucleotide microarray analysis: detection of copy number variations in normal and tumor specimens. Proc Natl Acad Sci USA 103:11234–11239
McCarroll SA, Kuruvilla FG, Korn JM, Cawley S, Nemesh J, Wysoker A, Shapero MH, de Bakker PIW, Maller JB, Kirby A, Elliott AL, Parkin M, Hubbell E, Webster T, Mei R, Veitch J, Collins PJ, Handsaker R, Lincoln S, Nizzari M, Blume J, Jones KW, Rava R, Daly MJ, Gabriel SB, Altshuler D (2008) Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat Genet 40:1166–1174
Olshen AB, Venkatraman ES, Lucito R, Wigler M (2004) Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5:557–572
Perry GH, Dominy NJ, Claw KG, Lee AS, Fiegler H, Redon R, Werner J, Villanea FA, Mountain JL, Misra R, Carter NP, Lee C, Stone AC (2007) Diet and the evolution of human amylase gene copy number variation. Nat Genet 39:1256–1260
Perry GH, Yang F, Marques-Bonet T, Murphy C, Fitzgerald T, Lee AS, Hyland C, Stone AC, Hurles ME, Tyler-Smith C, Eichler EE, Carter NP, Lee C, Redon R (2008) Copy number variation and evolution in humans and chimpanzees. Genome Res 18:1689–1710
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D, Collins C, Kuo WL, Chen C, Zhai Y, Dairkee SH, Ljung BM, Gray JW, Albertson DG (1998) High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 20:207–211
Pollack JR, Perou CM, Alizadeh AA, Eisen MB, Pergamenschikov A, Williams CF, Jeffrey SS, Botstein D, Brown PO (1999) Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat Genet 23:41–46
Pollack JR, Sorlie T, Perou CM, Rees CA, Jeffrey SS, Lonning PE, Tibshirani R, Botstein D, Borresen-Dale A, Brown PO (2002) Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc Natl Acad Sci USA 99:12963–12968
Price TS, Regan R, Mott R, Hedman A, Honey B, Daniels RJ, Smith L, Greenfield A, Tiganescu A, Buckle V, Ventress N, Ayyub H, Salhan A, Pedraza-Diaz S, Broxholme J, Ragoussis J, Higgs DR, Flint J, Knight SJL (2005) SW-ARRAY: a dynamic programming solution for the identification of copy-number changes in genomic DNA using array comparative genome hybridization data. Nucleic Acids Res 33:3455–3464
Redon R, Ishikawa S, Fitch KR, Feuk L, Perry GH, Andrews TD, Fiegler H, Shapero MH, Carson AR, Chen W, Cho EK, Dallaire S, Freeman JL, Gonzalez JR, Gratacos M, Huang J, Kalaitzopoulos D, Komura D, MacDonald JR, Marshall CR, Mei R, Montgomery L, Nishimura K, Okamura K, Shen F, Somerville MJ, Tchinda J, Valsesia A, Woodwark C, Yang F, Zhang J, Zerjal T, Zhang J, Armengol L, Conrad DF, Estivill X, Tyler-Smith C, Carter NP, Aburatani H, Lee C, Jones KW, Scherer SW, Hurles ME (2006) Global variation in copy number in the human genome. Nature 444:444–454
Ríos G, Naranjo M, Iglesias D, Ruiz-Rivero O, Geraud M, Usach A, Talon M (2008) Characterization of hemizygous deletions in citrus using array-comparative genomic hybridization and microsynteny comparisons with the poplar genome. BMC Genomics 9:381
Salathia N, Hana NL, Sangster TA, Morneau K, Landry CR, Schellenberg K, Behere AS et al (2007) Indel arrays: an affordable alternative for genotyping. Plant J 51:727–737
Sebat J, Lakshmi B, Troge J, Alexander J, Young J, Lundin P, Maner S, Massa H, Walker M, Chi M, Navin N, Lucito R, Healy J, Hicks J, Ye K, Reiner A, Gilliam TC, Trask B, Patterson N, Zetterberg A, Wigler M (2004) Large-scale copy number polymorphism in the human genome. Science 305:525–528
She X, Cheng Z, Zöllner S, Church DM, Eichler EE (2008) Mouse segmental duplication and copy number variation. Nat Genet 40:909–914
Shlien A, Tabori U, Marshall CR, Pienkowska M, Feuk L, Novokmet A, Nanda S, Druker H, Scherer SW, Malkin D (2008) Excessive genomic DNA copy number variation in the Li-Fraumeni cancer predisposition syndrome. Proc Natl Acad Sci USA 105:11264–11269
Skvortsov D, Abdueva D, Stitzer ME, Finkel SE, Tavaré S (2007) Using expression arrays for copy number detection: an example from E. coli. BMC Bioinformatics 8:203
Solinas-Toldo S, Lampel S, Stilgenbauer S, Nickolenko J, Benner A, Döhner H, Cremer T, Lichter P (1997) Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 20:399–407
Springer NM, Stupar RM (2007) Allelic variation and heterosis in maize: how do two halves make more than a whole? Genome Res 17:264–275
Stranger BE, Forrest MS, Dunning M, Ingle CE, Beazley C, Thorne N, Redon R, Bird CP, de Grassi A, Lee C, Tyler-Smith C, Carter N, Scherer SW, Tavare S, Deloukas P, Hurles ME, Dermitzakis ET (2007) Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315:848–853
R Development Core Team (2009) R: a language and environment for statistical computing. http://www.R-project.org. Accessed 24 Mar 2009
Tian D, Wang Q, Zhang P, Araki H, Yang S, Kreitman M, Nagylaki T, Hudson R, Bergelson J, Chen J (2008) Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes. Nature 455:105–108
Xu B, Roos JL, Levy S, van Rensburg EJ, Gogos JA, Karayiorgou M (2008) Strong association of de novo copy number mutations with sporadic schizophrenia. Nat Genet 40:880–885
Zhao X, Li C, Paez JG, Chin K, Jänne PA, Chen T, Girard L, Minna J, Christiani D, Leo C, Gray JW, Sellers WR, Meyerson M (2004) An integrated view of copy number and allelic alterations in the cancer genome using single nucleotide polymorphism arrays. Cancer Res 64:3060–3071
Acknowledgments
We are thankful to Scott Tingey for frequent challenging discussions and to DuPont Crop Genetics for providing financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Heslop-Harrison.
Contribution to the special issue “Heterosis in Plants”.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2009_1128_MOESM2_ESM.pdf
Result of aCGH between Mo17 and B73. Probes highlighted in green and red are those detecting a CNV region with more copies or presence in Mo17 and B73, respectively. Solid and dashed horizontal red lines correspond to one and two standard-deviations, respectively, calculated for all probes in the experiment. The blue curve is a loess function (R Development Core Team 2009) based on the aCGH ratio. Coordinates along the chromosomes are given as bands in a maize physical map (Fengler et al. 2007). (PDF 2,581 kb)
122_2009_1128_MOESM3_ESM.pdf
Result of aCGH experiment between two independent preparations of the inbred line PHP38. Solid and dashed horizontal red lines correspond to one and two standard-deviations, respectively, calculated for all probes in the experiment. The blue curve is a loess function (R Development Core Team 2009) based on the aCGH ratio. Coordinates along the chromosomes are given as bands in a maize physical map (Fengler et al. 2007). (PDF 2,497 kb)
122_2009_1128_MOESM4_ESM.png
PCR of a CNV region on chromosome five at 530 cM. A. aCGH results. B. Detailed view of CNV regions showing B73 BACs, microarray probes (number of matches to B73 genome in parentheses), and PCR results. Left lane: PCR from B73; right lane: PCR from Mo17. C. PCR of probe regions in other maize inbreds. (PNG 308 kb)
122_2009_1128_MOESM5_ESM.png
PCR of two CNV regions on chromosomes eight (206 cM) and nine (300.2 cM). A. aCGH results. B. Detailed view of CNV regions showing B73 BACs, microarray probes (number of matches to B73 genome in parentheses) and PCR results. Left lane: PCR from B73; right lane: PCR from Mo17. C. PCR of probe regions in other maize inbreds. (PNG 193 kb)
Rights and permissions
About this article
Cite this article
Beló, A., Beatty, M.K., Hondred, D. et al. Allelic genome structural variations in maize detected by array comparative genome hybridization. Theor Appl Genet 120, 355–367 (2010). https://doi.org/10.1007/s00122-009-1128-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1128-9