Abstract
High erucic acid rapeseed (HEAR) oil is of interest for industrial purposes because erucic acid (22:1) and its derivatives are important renewable raw materials for the oleochemical industry. Currently available cultivars contain only about 50% erucic acid in the seed oil. A substantial increase in erucic acid content would significantly reduce processing costs and could increase market prospects of HEAR oil. It has been proposed that erucic acid content in rapeseed is limited because of insufficient fatty acid elongation, lack of insertion of erucic acid into the central sn-2 position of the triaclyglycerol backbone and due to competitive desaturation of the precursor oleic acid (18:1) to linoleic acid (18:2). The objective of the present study was to increase erucic content of HEAR winter rapeseed through over expression of the rapeseed fatty acid elongase gene (fae1) in combination with expression of the lysophosphatidic acid acyltransferase gene from Limnanthes douglasii (Ld-LPAAT), which enables insertion of erucic acid into the sn-2 glycerol position. Furthermore, mutant alleles for low contents of polyunsaturated fatty acids (18:2 + 18:3) were combined with the transgenic material. Selected transgenic lines showed up to 63% erucic acid in the seed oil in comparison to a mean of 54% erucic acid of segregating non-transgenic HEAR plants. Amongst 220 F2 plants derived from the cross between a transgenic HEAR line and a non-transgenic HEAR line with a low content of polyunsaturated fatty acids, recombinant F2 plants were identified with an erucic acid content of up to 72% and a polyunsaturated fatty acid content as low as 6%. Regression analysis revealed that a reduction of 10% in polyunsaturated fatty acids content led to a 6.5% increase in erucic acid content. Results from selected F2 plants were confirmed in the next generation by analysing F4 seeds harvested from five F3 plants per selected F2 plant. F3 lines contained up to 72% erucic acid and as little as 4% polyunsaturated fatty acids content in the seed oil. The 72% erucic acid content of rapeseed oil achieved in the present study represents a major breakthrough in breeding high erucic acid rapeseed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
World vegetable oil markets are highly competitive requiring a steady improvement in oil quality to maintain or increase market shares. Genetic improvement of oilseeds has the objective of increasing oil yields with a uniform fatty acid composition for nutritional, pharmaceutical and industrial purposes (Roscoe 2005). Oil from traditional rapeseed (Brassica napus L.) differs significantly from most other vegetable oils by its high contents of erucic acid (22:1) and eicosenoic acid (20:1; Jönsson 1977; Becker et al. 1999). They are the major very long chain fatty acids (VLCFAs) in the seed oil, together accounting for 45–60% of the total fatty acid mixture. High levels of erucic acid in cooking and salad oil extracted from rapeseed have been associated with health problems (Beare et al. 1963). Following identification of a spontaneous rapeseed mutant with low erucic acid content, genetic studies have shown that this trait is under control of two loci that act in an additive manner (Harvey and Downey 1964; Kondra and Stefansson 1965). Major efforts in the 1960s led to the development of low erucic acid rapeseed (LEAR) varieties. Nowadays, double zero (00) or Canola quality cultivars with a low erucic acid content in the seed oil and a low glucosinolate content in the seeds are predominantly being grown in most parts of the world.
However, High Erucic Acid Rapeseed (HEAR) types retained some importance. HEAR cultivars are presently cultivated to a small extent in Europe (up to 40,000 hectares in 2006/2007) and in USA/Canada as an identity preserved crop (Möllers 2004; Scarth and Tang 2006). Erucic acid and its derivatives are important renewable raw materials used in plastic film, nylon, lubricant, cosmetic and emollient industries (Leonard 1994; Sonntag 1995; Piazza and Foglia 2001). Currently available HEAR cultivars contain only about 50% erucic acid in the seed oil. A substantial increase of the erucic acid content of the rapeseed oil would significantly reduce processing costs and could increase market prospects. However, possibilities to increase erucic acid by classical breeding are limited, because in rapeseed and related Brassica species erucic acid is inserted only in the first (sn-1) and third (sn-3) position of the glycerol backbone. This limits erucic acid content to a maximum of 67%. The reason for this limitation lies in the specificity of the B. napus sn-2 acyltransferase (LPAAT—lysophosphatidic acid acyltransferase), which does not accept erucoyl-CoA as a substrate (Cao et al. 1990; Frentzen 1993). To overcome this limitation, the gene for an erucoyl-CoA preferring sn-2 acyltransferase was isolated from Limnanthes species. Expression of this gene in transgenic rapeseed altered seed oil sn-2 proportions of erucic acid, but did not lead to an increase in erucic acid content of the seed oil (Lassner et al. 1995; Brough et al. 1996; Weier et al. 1997).
In a next step, interest focussed on the fatty acid elongation mechanism from oleic acid to erucic acid. This elongation is the result of two cycles of a four-step mechanism, in which first oleoyl-CoA and then eicosenoyl-CoA serve as substrates (Puyaubert et al. 2005). In the first step, the β-ketoacyl-CoA synthase (KCS; fae1 gene) catalyses the condensation reaction of oleoyl-CoA or eicosenoyl-CoA with malonyl-CoA. It is believed that this initial reaction is the rate-limiting step of the four-step mechanism (Cassagne et al. 1994). The fae1 genes from Arabidopsis and from rapeseed were cloned and over expressed under control of a seed specific promoter in transgenic HEAR (Katavic et al. 2001; Han et al. 2001). However, this led only to a minor increase in erucic acid content. Even the combination with the expression of the Ld-LPAAT gene from Limnanthes douglasii did not result in a substantial increase of the erucic acid content in transgenic HEAR (Han et al. 2001).
There is some evidence that the cytosolic pool of available oleoyl-CoA or malonyl-CoA may limit fatty acid elongation (Bao et al. 1998; Domergue et al. 1999). Crossing conventional HEAR with oilseed rape with reduced contents of linoleic acid (18:2) and linolenic acid (18:3) did result in recombinant high erucic low polyunsaturated fatty acid (HELP) F3 plants which, however, did not show an increased erucic acid content compared to the parental HEAR genotype (50% erucic acid; Sasongko and Möllers 2005). An obvious explanation for this result was that in the HELP material the activity of the β-ketoacyl-CoA synthase activity (fae1.1 and fae1.2 genes) was too low to allow for enhanced erucic acid synthesis.
The first objective of the present study was to repeat the approach of Han et al. (2001) to increase erucic acid content by expressing the Ld-LPAAT gene and over expressing the Bn-fae1 gene in transgenic HEAR. The second objective was to study in this material, the effect of genetically reduced polyunsaturated fatty acids content on erucic acid content.
Materials and methods
Plant material
6575-1 HELP is a winter rapeseed F4 line with 27% oleic acid, 7% linoleic and linolenic acid and 50% erucic acid content obtained from a cross between the winter rapeseed cultivar Maplus and the high oleic acid doubled haploid winter rapeseed line DH XXII D9 (for details see Sasongko and Möllers 2005). BGRV2 is a UK high erucic acid winter oilseed rape breeding line from Nickerson UK Ltd with about 52% erucic acid in the seed oil (Wilmer et al. 2003). This line was used in the transformation experiments to produce transgenic line 361.2B (see below). F1 plants were obtained by crossing transgenic line 361.2B with line 6575-1 HELP. Segregating F2 and F3 plant generations were produced by growing the plants in the greenhouse and self pollination of F1 and F2 plants, respectively.
Performance of greenhouse experiments
In August 2005, 220 randomly selected F2 seeds along with eight seeds from each parent were sown for producing the F2 plant population. F2 seeds were sown in a multipot tray containing T-soil (Fruhstorfer Erde; pH 5.9). F2 plants were allowed to grow for three weeks in the greenhouse. For vernalisation, plantlets were transferred to 4°C temperature with 8 h light for 8 weeks. Afterwards, plantlets were transferred to 9 cm diameter pots containing normal compost soil. Plantlets were cultivated in the greenhouse providing 16 h light by using 400 W sodium-steam lamps. Temperature varied during the day from 20 to 25°C and during the night from 10 to 15°C. Hakaphos fertilizer containing N:P:K (15:11:15) + 0.2% trace minerals was added at two week intervals to the top soil (100 mg) of each pot until the plants matured. Insecticide was applied when aphid and thrips attack was recognized. Sulphur vapour supply was constant during the whole experiment to avoid fungal diseases. Self-fertilisation was ensured by covering the flowers of the main raceme with micropore plastic bags. F3 seeds were harvested at maturity from the main raceme in February 2006 and stored at 4°C.
During the period October 2006 to April 2007, the greenhouse experiment was repeated with F3 seeds (from sowing to harvest). For this experiment 41 F2 plants with an erucic acid content higher than the mean of parent 361.2B (>62.5%) were selected. Five F3 seeds from each of the F2 plants and of the parents were sown. The greenhouse experiment was conducted as described above, following a randomized complete block design. The five F3 plants from each F2 plant and the parents were considered as five replicates. Five separate tables inside the green house represented the complete blocks consisting of all genotypes (F3 plants and parents). Following self-fertilisation F4 seeds were harvested from main raceme.
Methods
Binary plasmid construction
The fae1-1 (C-genome) and fae1-2 (A-genome) genes were amplified from BGRV2 using primers BnFAE-F (5′ CCTCATGACGTCCATTAACGTAAAGCTCC 3′) and BnFAE-R (5′ GTGAGCTCTTATTAGGACCGACCGTTTGGG 3′), cloned into pBluescript II (KS+) (Stratagene), and sequenced. Next, the fae genes were transferred as RcaI–EcoICRI fragment into pAR4 (Biogemma UK) NcoI–SmaI sites to place the gene between the Pnapin and CHS polyA sequences. The expression cassettes were then lifted out as a SalI–SacI fragment and ligated into pT7Blue2 SalI–SacI sites, before transfer of an EcoRI–EcoICRI fragment into the binary vector pSCVnos144 (Biogemma UK) EcoRI–SmaI adjacent to a similar Pnapin::lat2 LPAAT::CHS poly A cassette (Brough et al. 1996) to create pEW13 and pEW14 (Wilmer et al. 2001).
Agrobacterium tumefaciens transformation and generation of T1-plants
Agrobacterium tumefaciens strain C58 pMP90 carrying above described binary plasmid pEW13 or pEW14 were used to transform rapeseed line BGRV2 essentially following the protocol of Moloney et al. (1989). Ten transgenic plants (T1) carrying pEW13 and 18 transgenic plants with pEW14 were regenerated and oil composition was analysed following self pollination of T1 and T2 plants. Erucic acid content in these lines varied between less than 30 and about 65%. Line 361.2B, transformed with pEW14—fae1-2, was selected as it had the highest erucic acid content in T3 seeds (Wilmer et al. 2001). T4 seeds of 361.2B were used in the crossing experiment described above.
DNA isolation and PCR analysis to follow transgene segregation in the F2 population
DNA was isolated from leaf material frozen in liquid nitrogen (−196°C) following the protocol of Ishizawa et al. (1991) with some modifications as described in Nath et al. (2007). Multiplex-PCR was performed using specific primers for the Ld-LPAAT gene and locus specific primers for the endogenous single copy Bn-fad2 gene. Amplification of the single copy Bn-fad2 gene served as positive control for the presence of DNA in sufficient quantity and quality. PCR primers, PCR conditions and gel electrophoresis were as described in Nath et al. (2007).
Seed quality analysis
Trierucin analysis
Seed samples (150 mg) obtained from F2 and F3 plants were analysed for trierucin (C69; EEE) content by high temperature gas liquid chromatography (HT-GLC) analysis according to the method described by Möllers et al. (1997). The analysis was done using silicon capillary column RTX-65TG (Restek no. 17,005) 15 m × 0.25 mm i.d. (0.1 μm film thickness). Trierucin is expressed as percent of the sum of all triglycerides.
Analysis of total fatty acid composition
Following trierucin analysis the remaining part of each sample was transferred to a new tube and left on a hot plate at 37.5°C over night to evaporate. The fatty acid profile was determined by analysis of methyl esters by gas liquid chromatography according to Rücker and Röbbelen (1996). Fatty acids are expressed as percent of the sum of all fatty acids. The following fatty acids were determined: palmitic acid (16:0), stearic acid (18:0), oleic acid (18:1), linoleic acid (18:2), linolenic acid (18:3), eicosenoic acid (20:1), erucic acid (22:1) and nervonic acid (24:1). Saturated fatty acids (SFA; 16:0 + 18:0), polyunsaturated fatty acids (PUFA; 18:2 + 18:3) and monounsaturated fatty acids (MUFA; 18:1 + 20:1 + 22:1) were calculated from the contents of individual fatty acids.
Analysis of the fatty acid composition at the sn-2 position of triacylglycerols
A total of 15 mg mixed seed samples from 5 replicates of the F3 population (F4 seeds) were collected in a 5 ml plastic tube. A measure of 0.5 ml iso-octane:iso-propanol (9:1) was added to each sample and seeds were homogenised with a steel rod. The supernatant was transferred to a new tube and the solvent evaporated by using a stream of warm air. The dried oil residue was mixed with 500 µl buffer (50 mM KH2PO4-, pH 7.2 with 0.5% Triton X-100) and kept in a supersonic bath for 2 min. The mixture was incubated at 30°C for 1 h by adding 25 µl (250 units) lipase from Rhizopus arrhizus (Sigma-Aldrich). Then 200 µl petroleum ether-70:iso-propanol (3:2) was added and the supernatant was collected in a 1 ml glass tube by centrifugation at 150 × G (1,000 rpm) for 5 min. The extraction was repeated twice and the supernatants were merged and evaporated by using a stream of warm air. 25 µl petroleum ether-70:iso-propanol (3:2) was added and mixed well by vortex. A measure of 20 µl from this mixture was taken and transferred to a thin layer chromatography (TLC) plate (F 1,500/LS 254, 20 × 20 cm) using a 20 µl syringe. TLC plates were placed in the eluent diethylether/petroleumether (3:1) and allowed to run for 30 min. Afterwards dry TLC plates were stained by subjecting them to iodine vapour. Monoacylglycerol spots (representing the fatty acids at the sn-2 position) were marked with a pencil. Following evaporation of the iodine, the marked areas were scraped out using a scalpel and transferred for fatty acid extraction in a separate 1 ml glass tube. The scrape was mixed with 250 µl iso-octane, incubated for 20 min and the supernatant was collected by centrifugation in a new 1 ml glass tube. This procedure was repeated twice. Iso-octane from the collected supernatant was evaporated and then the fatty acids were extracted and analysed as described above.
Analysis of seed oil and protein content
Oil and protein content of seed samples were determined by Near-Infrared-Reflectance-Spectroscopy (NIRS) using the equation raps2001.eqa (Tillmann 2007). Spectra were recorded using a standard ring cup equipped with a 14 mm PVC adapter. Values were adjusted using previously developed regression equations. Oil and protein contents are expressed on seed dry matter basis.
Statistical analysis
Spearman’s rank correlation coefficients were calculated using the Plant Breeding Statistical Program PLABSTAT (Version 2N; Utz 2007). Multiple mean comparisons were made with Fisher’s least significant difference (LSD) procedure using Stat Graphics Plus for Windows 3.0 (Statistical Graphics Corp., Rockville, USA). Hypotheses for transgene copy number segregation in the F2 population were tested using Chi-square (χ 2) test as described by Gomez and Gomez (1976).
Results
Variation of seed quality traits and segregation of the transgenes in the F2 population
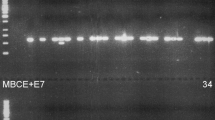
The F2 population derived from the cross between the transgenic line 361.2B and the high erucic acid, low polyunsaturated fatty acid line 6575-1 HELP showed a large quantitative variation for erucic acid content and other traits (Table 1). Erucic acid (22:1) was the most prominent fatty acid, accounting for 58.8% of the total fatty acid content. Its content ranged from 44 to 72% and showed large transgressive segregation compared to the higher parent 361.2B. Frequency distribution of the erucic acid content showed a continuous variation indicating polygenic inheritance (Fig. 1). Polyunsaturated fatty acid content (PUFA; 18:2 + 18:3) varied from six to 26%, in some F2 plants thereby being as low as in the 6575-1 HELP parent. Trierucoylglycerol (Trierucin) content showed a discontinuous variation from 0.0 to 25% (Table 1 and Fig. 2), indicating segregation and functioning of the Ld-LPAAT transgene. From the 220 F2 plants analysed, 211 contained trierucin (Fig. 2), suggesting the presence of two transgene copies in transgenic parent 361.2B and a 15:1 segregation in the F2 population (Chi-square χ2 = 2.52). Using PCR primers directed against the Ld-LPAAT transgene, four out of 57 randomly selected F2 plants were negative in PCR (see an example in Fig. 3), again confirming 15:1 segregation of two transgene copies in the F2 population (Chi-square χ2 = 0.06). All F2 plants positive in PCR for the Ld-LPAAT gene contained trierucin, whereas those negative in PCR did not contain trierucin. The clearly separated group of segregating non-transgenic F2 plants (Fig. 2) lacking the Ld-LPAAT + Bn-fae1 transgenes contained between 45 and 57% erucic acid in the seed oil (see Fig. 4), with a mean of 53.6%. Parent 361.2B contained 62.5% erucic acid in the seed oil, indicating that the two transgenes in the homozygous state in this population increased erucic acid content by 8.9%.
Correlations between traits in the F2 population
Spearman’s rank correlation coefficient analysis of the F2 population (F3 seeds) showed negative correlations between erucic acid and protein, SFA, oleic acid, PUFA and eicosenoic acid content (Table 2). Correlations between erucic acid content and oil, trierucin and MUFA content were positive (see also Fig. 4). The calculation of the regression between erucic acid content and PUFA content (y = −0.65x + 69.63) showed that a reduction of 10% in PUFA content led to an increase of 6.5% in erucic acid content.
Performance of selected F3 lines
From the F2 population, 41 plants showed a higher erucic acid content (>62.5%) than the mean of parent 361.2B. Five F3 plants from each of those 41 F2 plants were cultivated in the greenhouse and F4 seeds were harvested. In the F3 population, erucic acid content varied from 50 to 72% (Fig. 5) with the mean of 64.8%. The scatter plot for the mean erucic acid content of F3 lines (F4 seeds) versus F2 plants (F3 seeds) shows a significant positive correlation (r s = 0.57**; Fig. 5), proving effective selection of high erucic acid plants amongst single plants of the F2 population. The fatty acid composition of the six F3 lines (F4 seeds) with the highest erucic acid contents is shown in Table 3. F3 line III-G-7 contained with 72.3% the highest amount of erucic acid. Four of the F3 lines had an equal to or lower PUFA content than parent 6575-1 HELP, indicating homozygosity of the genes causing low PUFA content. All six F3 lines showed higher trierucin contents compared to transgenic parent 361.2B.
From the six F3 lines with the highest erucic acid content and the parental lines, the fatty acid composition at the sn-2 position of the triacylglycerols was analysed. In the F3 lines the erucic acid content at the sn-2 position varied from 36.8 to 65.3%, which compares favourably to the 31.6% of transgenic parent 361.2B (Table 3). Erucic acid at sn-2 position was only detected in case of the presence of Ld-LPAAT transgene. No, or only in one case very little, eicosenoic acid (20:1) was detected at the sn-2 position (data not shown).
Discussion
Transformation of the winter rapeseed breeding line BGRV2 with the Bn-fae1+Ld-LPAAT construct resulted in the regeneration of 18 primary transgenic lines, from which 361.2B was selected in preliminary experiments as the transgenic line with the highest erucic acid content. Crossing of 361.2B to 6575-1 HELP revealed in the F2 population the segregation of two transgene copies. Comparing erucic acid contents of 361.2B with the mean erucic acid content of non-transgenic F2 plants segregating in the population showed that, the two transgene copies led to an increase in erucic acid content from 53.6 to 62.5%, i.e. 8.9% (Fig. 4). This result is in contrast to the results of Han et al. (2001), who did not observe a significant increase in erucic acid content following transformation of a resynthesized high erucic acid line with principally the same construct. It could be that the Bn-fae1 gene in the transgene construct of Han et al. (2001) was not functional (Nath 2008).
The frequency distribution for the erucic acid content of the F2 population showed a large and continuous variation as expected for a polygenic inherited trait (Fig. 1). Erucic acid content varied from 44 to 72%. In addition to the effect caused by the two transgene copies, this variation may be due to segregation of loci of parent 6575-1 HELP responsible for low contents of polyunsaturated fatty acids (PUFA). The low PUFA content in parent 6575-1 HELP is caused by a mutation in the oleic acid desaturase fad2 gene and by two to three other unknown genes having minor effects (Schierholt et al. 2001; Sasongko and Möllers 2005). According to the regression, a 10% reduction in PUFA content leads to a 6.5% increase in erucic acid content. Nath (2008) reported similar results for a high erucic acid doubled haploid winter rapeseed population segregating for PUFA content. Furthermore, environmental effects, different effective alleles at the two endogenous erucic acid loci fae1.1 (A-genome, B. rapa) and fae1.2 (C-genome, B. oleracea) as well as other unknown genetic factors may have contributed to the variation found in the F2 population (Stefansson and Hougen 1964; Jönsson 1977; Ecke et al. 1995; Zhao et al. 2008).
Selection for erucic acid content amongst segregating F2 plants proved to be efficient as shown by comparing the erucic acid contents of F2 plants with those of the derived F3 plants (Fig. 5). F2 plants with about 70% erucic acid content in the seed oil were confirmed in the F3 generation (F4 seeds, Table 3). It is likely that the F3 plants are already homozygous for the two transgene copies and for the endogenous two erucic acid genes. However, F4 plant generation need to be tested to confirm this. Some of the high erucic acid lines had PUFA contents as low as in parent 6575-1 HELP (Table 3), indicating homozygosity for the genes causing low PUFA content.
The F3 line III-G-7 had with 72.3% the highest erucic acid content in the seed oil as a mean of five plants (Table 3). This is about 9 and 23% more than that of transgenic parent 361.2B and non-transgenic parent 6575-1 HELP. The 72.3% erucic acid content can be regarded as relatively stable, because the two parental lines had similar erucic acid contents at two different seasons in the greenhouse (compare results in Tables 1 and 3). Furthermore, parent 6575-1 HELP was tested previously in field experiments and likewise had about 50% erucic acid in the seed oil (Sasongko and Möllers 2005).
The formation of trierucoylglycerol (Trierucin, C69) proved that the Limnanthes douglasii lysophosphatidic acid acyltransferase (Ld-LPAAT) functioned in transgenic oilseed rape. Amongst the six F3 lines with the highest erucic acid content, line VI-D-9 showed the highest trierucin content (23.5%; Table 3). Variation amongst the lines might be due to differences in homozygosity of the transgenes and in the availability of erucoyl-CoA. Assuming random distribution of erucoyl-CoA to all three triacylglycerol positions, one would expect a trierucin content of 36% for F3 line VI-D-9 with 71.4% erucic acid content. However, this line showed only 23.5% trierucin content, indicating that Ld-LPAAT activity may not be strong enough in comparison to the resident rapeseed Bn-LPAAT activity, which has a strong preference for oleic acid (Frentzen 1998). The sn-2 fatty acid mixture of the seed oils of the F3 lines contained up to 65% erucic acid (Table 3). However, lines with similar erucic acid content contained very different amounts of erucic acid in the sn-2 position, indicating that beside the Ld-LPAAT gene there may be other genes involved in the expression of this trait. No, or very limited amounts of erucic acid at sn-2 position were found in non-transgenic parental line 6575-1 HELP (Table 3). Similar results were also reported by Weier et al. (1997) and Han et al. (2001) in Ld-LPAAT expressing transgenic rapeseed lines. The lack of significant amounts of eicosenoic acid at the sn-2 position confirms the pronounced preference of the Ld-LPAAT enzyme for erucoyl-CoA (Han et al. 2001).
The around 72% erucic acid content achieved in the present experiments represents a milestone in the breeding of high erucic acid oilseed rape. Further increases in erucic acid content can be expected from progress in reducing the contents of the remaining fatty acids, mainly oleic acid, polyunsaturated fatty acids and eicosenoic acid. Reduction in PUFA content probably could be achieved more easily following an RNAi antisense approach (Stoutjesdijk et al. 2002). Integration of the antisense fad2 gene into the same T-DNA as the Ld-LPAAT and the Bn-fae1 gene would greatly simplify the genetics and breeding of high erucic acid cultivars. The material developed in the present study should be of interest for the oleochemical industry but also for further studies aimed at identifying other physiological limitations in erucic acid biosynthesis.
References
Bao X, Pollard M, Ohlrogge J (1998) The biosynthesis of erucic acid in developing embryos of Brassica rapa. Plant Physiol 118:183–190
Beare JL, Campbell JA, Youngs CG, Craig BM (1963) Effects of saturated fat in rats fed rapeseed oil. Can J Biochem Physiol 41:605–612
Becker HC, Löptien H, Röbbelen G (1999) Breeding: an overview. In: Gomez-Campo C (ed) Biology of Brassica coenospecies. Elsevier, Amsterdam, pp 413–460
Brough CL, Coventry JM, Christie WW, Kroon JTM, Brown AR, Barsby TL, Slabas AR (1996) Towards the genetic engineering of triacylglycerols of defined fatty acid composition: major changes in erucic acid content at the sn-2 position affected by the introduction of a 1-acyl-sn-glycerol-3-phosphate acyltransferase from Limnanthes douglasii into oil seed rape. Mol Breed 2:133–142
Cao YZ, Oo KC, Huang AHC (1990) Lysophosphatidate acyl transferase in the microsomes from maturing seeds of meadowfoam (Limnanthes alba). Plant Physiol 94:1199–1206
Cassagne C, Lessire R, Bessoule JJ, Moreau P, Creach A, Schneider F, Sturbois B (1994) Biosynthesis of very long chain fatty acids in higher plants. Prog Lipid Res 33:55–69
Domergue F, Chevalier S, Santarelli X, Cassagne C, Lessire R (1999) Evidence that oleoyl-CoA and ATP-dependent elongations coexist in rapeseed (Brassica napus L.). Eur J Biochem 263:464–470
Ecke W, Uzunova M, Weissleder K (1995) Mapping the genome of rapeseed (Brassica napus L.) II. Localization of genes controlling erucic acid synthesis and seed oil content. Theor Appl Genet 91:972–977
Frentzen M (1993) Acyltransferases and triacylglycerols. In: Moore TS Jr (ed) Lipid metabolism in plants. CRC Press, Boca Raton, pp 195–220
Frentzen M (1998) Acyltransferases from basic science to modified seed oils. Fett/Lipid 100:161–166
Gomez KA, Gomez AA (1976) Statistical procedures for agricultural research with emphasis on rice. IRRI, Los Banos
Han J, Lühs W, Sonntag K, Zähringer U, Borchardt DS, Wolter FP, Heinz E, Frentzen M (2001) Functional characterization of β-ketoacyl-CoA synthase genes from Brassica napus L. Plant Mol Biol 46:229–239
Harvey BL, Downey RK (1964) The inheritance of erucic acid content in rapeseed (Brassica napus). Can J Plant Sci 44:104–111
Ishizawa M, Kobayashi Y, Miyamura T, Matsuura S (1991) Simple procedure of DNA isolation from human serum. Nucleic Acids Res 19:5792
Jönsson R (1977) Erucic acid heredity in rapeseed (Brassica napus L. and Brassica campestris L.). Hereditas 86:159–170
Katavic V, Friesen W, Barton DL, Gossen KK, Giblin EM, Luciw T, An J, Zou J, MacKenzie SL, Keller WA, Males D, Taylor DC (2001) Improving erucic acid content in rapeseed through biotechnology: what can the Arabidopsis FAE1 and the yeast SLC1–1 genes contribute? Crop Sci 41:739–747
Kondra ZP, Stefansson BR (1965) Inheritance of erucic and eicosenoic acid content of rapeseed oil (Brassica napus). Can J Genet Cytol 7:500–510
Lassner MW, Levering CK, Davies HM, Knutzon DS (1995) Lysophosphatidic acid acyltransferase from meadowfoam mediates insertion of erucic acid at the sn-2 position of triacylglycerol in transgenic rapeseed oil. Plant Physiol 109:1389–1394
Leonard C (1994) Sources and commercial applications of high erucic vegetable oils. Lipid Technol 4:79–83
Möllers C (2004) Potential and future prospects for rapeseed oil. In: Gunstone FD (ed) Rapeseed and canola oil-production, processing, properties and uses. Blackwell, Oxford, pp 186–217
Möllers C, Lühs W, Schaffert E, Thies W (1997) High-temperature gas chromatography for the detection of trierucoylglycerol in the seed oil of transgenic rapeseed (Brassica napus L.). Fett/Lipid 99:356–362
Moloney MM, Walker JM, Sharma KK (1989) High efficiency transformation of Brassica napus using Agrobacterium vectors. Plant Cell Rep 8:238–242
Nath UK, Iqbal MCM, Möllers C (2007) Early, non-destructive selection of microspore-derived embryo genotypes in oilseed rape (Brassica napus L.) by molecular markers and oil quality analysis. Mol Breed 19:285–289
Nath UK (2008) Increasing erucic acid content in the seed oil of rapeseed (Brassica napus L.) by combining selection for natural variation and transgenic approaches. PhD thesis Georg-August-Universität Göttingen, Faculty of Agriculture. http://webdoc.sub.gwdg.de/diss/2008/nath/nath.pdf
Piazza GJ, Foglia TA (2001) Rapeseed oil for oleochemical uses. Eur J Lipid Sci Technol 103:405–454
Puyaubert J, Garcia C, Chevalier S, Lessire R (2005) Acyl-CoA elongase, a key enzyme in the development of high-erucic acid rapeseed? Eur J Lipid Sci Technol 107:263–267
Roscoe TJ (2005) Identification of acyltransferases controlling triacylglycerol biosynthesis in oilseeds using a genomic based approach. Eur J Lipid Sci Technol 107:162–256
Rücker B, Röbbelen G (1996) Impact of low linolenic acid content on seed yield of winter oilseed rape (Brassica napus L.). Plant Breed 115:226–230
Sasongko ND, Möllers C (2005) Toward increasing erucic acid content in oilseed rape (Brassica napus L.) through the combination with genes for high oleic acid. J Am Oil Chem Soc 82:445–449
Scarth R, Tang J (2006) Modification of Brassica oil using conventional and transgenic approaches. Crop Sci 46:1225–1236
Schierholt A, Rücker B, Becker HC (2001) Inheritance of high oleic acid mutations in winter oilseed rape (Brassica napus L.). Crop Sci 41:1444–1449
Sonntag NOV (1995) Industrial utilization of long-chain fatty acids and their derivatives. In: Kimber DS, McGregor DI (eds) Brassica oilseeds. CAB International, Oxon, pp 339–352
Stoutjesdijk PA, Singh SP, Liu Q, Hurlstone CJ, Waterhouse PA, Green AG (2002) hpRNA-mediated targeting of the Arabidopsis FAD2 gene gives highly efficient and stable silencing. Plant Physiol 129:1723–1731
Stefansson BR, Hougen FW (1964) Selection of rape plants (Brassica napus) with seed oil practically free from erucic acid. Can J Plant Sci 44:359–364
Tillmann P (2007) http://www.vdlufa.de/nirs. (Site visited 24 June 2008)
Utz HF (2007) PLABSTAT (Version 2N), a computer program for the computation of variances and covariances. Institute of Plant Breeding, Seed Science, and Population Genetics, University of Hohenheim, Stuttgart, Germany, (http://www.uni-hohenheim.de/ipspwww/soft.html) (Site visited 24 June 2008)
Weier D, Hanke C, Eickelkamp A, Lühs W, Dettendorfer J, Schaffert E, Möllers C, Friedt W, Wolter FP, Frentzen M (1997) Trierucoylglycerol biosynthesis in transgenic plants of rapeseed (Brassica napus L.). Fett/Lipid 99:160–165
Wilmer JA, Wallington EJ, Slabas AR (2001) Oil biosynthesis—Very high erucic acid rapeseed, WO 03/033713
Wilmer JA, Wallington EJ, Slabas AR (2003) Very high erucic acid rape: a dream or reality. In: Proceedings of 11th Intl Rapeseed Congress, 6–10 July 2003, Copenhagen, Denmark 2:583–585
Zhao J, Dimov Z, Becker HC, Ecke W, Möllers C (2008) Mapping QTL controlling fatty acid composition in a doubled haploid rapeseed population segregating for oil content. Mol Breed 21:115–125
Acknowledgments
UKN gratefully acknowledges German Academic Exchange Service (DAAD), Bonn, Germany, for providing a scholarship, Uwe Ammermann and Rosemarie Clemens for excellent technical assistance. Technical assistance for B. napus plant transformation was provided by Andrea Grey and Ruth Bates (Biogemma UK).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. Rajcan.
Dedicated to professor Gerhard Röbbelen on the occasion of his 80th birthday.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Nath, U.K., Wilmer, J.A., Wallington, E.J. et al. Increasing erucic acid content through combination of endogenous low polyunsaturated fatty acids alleles with Ld-LPAAT + Bn-fae1 transgenes in rapeseed (Brassica napus L.). Theor Appl Genet 118, 765–773 (2009). https://doi.org/10.1007/s00122-008-0936-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-008-0936-7