Abstract
The small genome size (740 Mb), short life cycle (3 months) and high economic importance as a food crop legume make chickpea (Cicer arietinum L.) an important system for genomics research. Although several genetic linkage maps using various markers and genomic tools have become available, sequencing efforts and their use are limited in chickpea genomic research. In this study, we explored the genome organization of chickpea by sequencing approximately 500 kb from 11 BAC clones (three representing ascochyta blight resistance QTL1 (ABR-QTL1) and eight randomly selected BAC clones). Our analysis revealed that these sequenced chickpea genomic regions have a gene density of one per 9.2 kb, an average gene length of 2,500 bp, an average of 4.7 exons per gene, with an average exon and intron size of 401 and 316 bp, respectively, and approximately 8.6% repetitive elements. Other features analyzed included exon and intron length, number of exons per gene, protein length and %GC content. Although there are reports on high synteny among legume genomes, the microsynteny between the 500 kb chickpea and available Medicago truncatula genomic sequences varied depending on the region analyzed. The GBrowse-based annotation of these BACs is available at http://www.genome.ou.edu/plants_totals.html. We believe that our work provides significant information that supports a chickpea genome sequencing effort in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Leguminosae family constitutes about 650 genera and 18,000 species and ranks third among families of flowering plants. Legumes are unique because of their capability of fixing atmospheric nitrogen in soil through symbiosis. Among legumes there is extensive literature on genetic studies for several species, but the existence of a wide range of genomic tools and genome information is limited only to Medicago truncatula (Mt), Lotus japonicus (Lj) and Glycine max.
Chickpea (Cicer arietinum L.) is an economically important crop with a life cycle of 3–4 months, and a genome size of 740 Mbp that is only 1.5 times larger than M. truncatula. Although genomic research in chickpea is feasible, it was not until the beginning of twenty-first century that this aspect of research began in this cool season legume crop. Since then, several genomic tools that include bacterial artificial chromosome (BAC) libraries (Rajesh et al. 2004; Lichtenzveig et al. 2005), expressed sequences tags (EST) (Buhariwalla et al. 2005; Corum and Pang 2005), Targeted Induced Local Lesions IN the Genome (TILLING) mutants (Rajesh P. N. et al. unpublished data) and gene technology (Sarmah et al. 2004; Sanyal et al. 2005) have been developed in chickpea, and made available to the research community. Utilization of these tools for crop improvement should be the next focus as it will provide precedence for effective application of genomics for crop improvement of crop legumes. With the availability of suitable genomic tools, genetic materials and also the shorter life cycle with relatively smaller genome than other crop legumes, chickpea can be considered as a model crop legume.
Current knowledge of the chickpea genome is based mostly on seven linkage maps that are comprised the less informative RAPD, ISSR, AFLP and SSR markers. No gene specific markers have been mapped to the genome and thus these maps have limited use in comparative genomic studies.
Genome organization and composition analysis between other model legumes, such as Mt and Lj, has revealed several interesting features of their genomes. For example, 18,844 and 20,800 genes (excluding transposons) were predicted to be present in Mt and Lj, respectively. Also, the average gene density was estimated to be 12.6 genes per 100 kb in Mt and 17.4 genes per 100 kb for Lj. In addition, 38 and 19% of the Mt and Lj genomes, respectively, were predicted to contain repetitive sequences (Cannon et al. 2006). Availability of genome sequences in these legumes (149 and 121 Mbp in Mt and Lj, respectively) facilitated the analysis of synteny and colinearity among them, which will have long-term application in transferring genetic information between different species. Phylogenetically, Mt and Lj belong to galegoid phylum and were separated by 37–38 million years ago (MYA) (Choi et al. 2004). Microsynteny analysis between Mt and Lj genome sequences discovered colinearity at different synteny blocks (Cannon et al. 2006).
Since chickpea is closer phylogenetically to Mt than Lj is to Mt, and both belong to the galegoid phylum, microsynteny analysis between these two legumes will be useful for transferability of genetic information between them. The lack of genomic sequences has limited the information on genome organization and microsynteny in chickpea. Also, the absence of common markers, or cross-species specific markers, in genetic linkage maps has restricted the macrosynteny establishment between chickpea and model legumes.
Ascochyta blight [caused by Ascochyta rabiei (Pass). Lab.] is a devastating disease worldwide and has the potential to cause up to 100% yield loss to chickpea. Earlier genetic studies identified two QTL (ABR-QTL1 and ABR-QTL2) that confer resistance to this disease (Santra et al. 2000). To dissect ascochyta blight resistance QTL and narrow the genetic distance between the flanking markers, it is necessary to increase the marker density at this genomic region.
Therefore, in this study, we sequenced 471,638 bp from 11 BAC clones that include three clones representing ABR-QTL1 and analyzed genome organization and composition in this economically important crop legume. Microsynteny analysis between chickpea with M. truncatula was also performed. In addition to genome organization and microsynteny analysis, we characterized ABR-QTL1 genomic region by increasing marker density developed from the ends of BAC clones. Our results provide new insight into the chickpea genome, and the sequences representing different genomic regions will be a potential resource for the development of additional markers to increase marker density in important regions of the chickpea genetic map.
Materials and methods
BAC selection and sequencing
The BAC clones used in this study (Table 1) can be classified into the following categories: (i) targeted selection and (ii) random selection. BAC clones in the first category represent QTL1 accounting for ascochyta blight resistance. BAC clones representing the latter category were selected randomly.
The detailed procedures for cloned, large insert genomic DNA isolation, random shot-gun cloning, fluorescent-based DNA sequencing and subsequent analysis were used as described earlier (Bodenteich et al. 1993; Pan et al. 1994; Chissoe et al. 1995; Roe et al. 1996; Roe 2004). Fifty microgram portions of purified BAC DNA was randomly sheared and made blunt-ended, and after kinase treatment and gel purification, fragments in the 2–4 kb range were ligated into SmaI-cut, bacterial alkaline phosphatase (BAP)-treated pUC18 (Pharmacia) and transformed by electroporation into Escherichia coli, strain XL1BlueMRF’ (Stratagene) competent cells. A random library of approximately 1,200 colonies was picked from each transformation, grown in terrific broth (TB) medium (Roe 2004) supplemented with 100 μg of ampicillin for 14 h at 37°C with shaking at 250 rpm in a HiGro incubator (Genomic Solutions), and the sequencing templates were isolated by a semi-automated cleared lysate-based procedure on a Capilar Life Sciences Sciclone ALH 3000 workstation equipped with a Twister II robotic arm and four microtiter plate shakers.
Sequencing reactions were performed as previously described (Chissoe et al. 1995; Roe et al. 1996; Roe 2004) using the Amersham ET Terminator kit (US-81070) or Applied Biosystem BigDyes version 3.1 (4336921) sequencing reaction mixes diluted 1:32 with TM buffer (20 ml Tris-HCl pH9.0, 0.5 ml 1 M MgCl2 and 29.5 ml ddH2O). The reactions were incubated for 60 cycles in a Perkin–Elmer Cetus DNA Thermocycler 9600 under the cycle conditions recommended by the manufacturer. Any unincorporated dye terminators were removed by ethanol precipitation at room temperature, and after dissolving the fluorescent-labeled nested fragment sets in 0.1 mM EDTA pH 7.4, the nested fragment sets were resolved by electrophoresis on an ABI 3730 Capillary DNA Sequencer. After base calling with the ABI Analysis software, the analyzed data were transferred to a Sun Workstation Cluster, and assembled using Phred and Phrap (Ewing and Green 1998; Ewing et al. 1998). Overlapping sequences and contigs were analyzed using Consed (Gordon et al. 1998). Gap closure and proofreading were performed either using custom primer walking or using PCR amplification of the region corresponding to the gap in the sequence followed by sequencing directly using the amplification or nested primers, or by sub-cloning into pUC18 and cycle sequencing with the universal pUC-primers (Roe 2004). In some instances, additional synthetic custom primers and PCRs with 7deaza-dGTP replacing dGTP (Roe 2004) or by rolling circle amplification (Detter et al. 2002) were necessary to obtain at least threefold coverage for each base. The sequenced BACs along with their GenBank accession numbers are listed in Table 1 and the genome browser-based annotation (Gbrowse) is available at http://genome.ou.edu/plants_totals.html.
Sequence annotation
The chickpea BACs sequenced were annotated using Repeatmasker (Jurka 2000), Genscan (Burge and Karlin 1997) and FgenesH (Salamov and Solovyev 2000), and gene predictions were compared to the Arabidopsis genome and the Plant unigene database (Wheeler et al. 2005). Results from this are displayed for each BAC on the Advanced Center for Genome Research’s web site (http://www.genome.ou.edu) using Gbrowse (Stein et al. 2002).
Further analysis was performed to determine genomic organization of the BACs sequenced. Genic information was determined using the FgenesH dicot model, as FgenesH has been shown to be the most accurate gene modeling program for plants (Yu et al. 2002). Identification of these predicted genes were performed via a basic local alignment search tool (BLAST) (Zhang and Madden 1997) against the NCBI databases. Genes that have no significant homology (an e value of less than e−10) were designated as “unknown,” if they have EST support, “hypothetical” if the gene is conserved in multiple organisms and “putative” if there is no support other than prediction by a gene model. Other genomic organization information, such as GC content and repetitive sequences present (simple repeats, retroviral insertions, and transposons) were determined using Repeatmasker (Jurka 2000), against the Arabidopsis matrix. The values of calculated parameters are in Table 1 and the predicted genes in Table 2.
Plant materials
Chickpea accession FLIP84-92C and wild relative PI599072 that are resistant and susceptible to Ascochyta rabiei (Pass). Lab. respectively were used in this study. The F2 population from this cross was advanced by single seed descent to the F8 to produce F7-derived RILs (Santra et al. 2000; Tekeoglu et al. 2002). A population of 142 F7:8 derived recombinant inbred lines (RILs) from the FLIP 84-92C (resistant) x ×C. reticulatum PI 599072 (susceptible) cross was used for genetic mapping. DNA was extracted from leaf tissue of each RIL and the parental lines according to Doyle and Doyle (1987).
Linkage analysis
Segregation of marker loci was tested for goodness of fit to the expected Mendelian ratio of 1:1 using Chi-square analysis (P < 0.05). Markers with distorted distribution were also used for linkage analysis. Linkage analysis was performed using Mapmaker/Exp 3.0 (Lander et al. 1987). Linkage groups were established at a constant LOD score of 4.0 and a recombination value of 0.25 by two point analyses using the ‘group’ command. The most possible order of loci within a group was determined using multipoint ‘compare’ command, and these orders were verified using the ‘ripple’ command. The Kosambi mapping function was used to determine cM distances between markers (Kosambi 1944). Double crossovers were checked by ‘double crossover’ command in Map Manager QTb (version 2.8) (Manly 1998). QTL analysis for blight resistance was carried out with the simple interval mapping function using Qgene (Nelson 1997) at an LOD score of 3.0. Single-point regression analysis was used to identify markers significantly associated with blight resistance.
Results
BAC selection and genetic analysis
The BAC clones AC161102, AC145766 and AC145456 were selected by targeted screening of the BAC library. The BAC clone AC161102 was identified by screening our BAC library with a single locus SCAR733b marker derived from a RAPD marker UBC733b. AC145766 and AC145456 clones were identified with a single locus marker OPS06-01 (Rakshit et al. 2003). UBC733b and OPS06-01 were genetically proved to be flanking ABR-QTL1 in earlier studies (Santra et al. 2000; Tekeoglu et al. 2002; Rakshit et al. 2003). In this study, we generated six markers from the ends of identified BAC clones and a gene (Flavonoid 3-0-galactosyl transferase) present in AC145456. Since these PCR-based markers are monomorphic in amplicon length between the parental lines (FLIP84-92C and PI599072), we developed cleaved amplified polymorphic site (CAPS) and derived CAPS (dCAPS) markers by utilizing the SNPs to increase marker density at QTL1 (Rajesh et al. 2005).
Genome organization
FgenesH program predicted 51 genes in 11 BAC clones representing 471,638 bp of chickpea genome (Table 1). Of these 51 predicted genes, 20 were with known function, 18 were putative genes and the remaining 13 were retro elements. Of 51 predicted genes, 12 genes identified ESTs with other plants at http://www.ncbi.nlm.nih.gov database. The average gene length of 2.5 kb, protein length of 452aa, 8.68% of repetitive elements and the one long terminal repeat (LTR) for every 13.5 kb were estimated (Table 2). The gene density in chickpea is estimated to be ranging from 1 gene for 5.3 kb to 1 gene per 63.3 kb and averaged 1 gene for every 9 kb (11 genes per 100 kb) which is similar to Mt in which 12.6 genes per 100 kb were found and relatively higher than Lj in which 17.4 genes per 100 kb were found (Cannon et al. 2006). No gene duplication was detected among these chickpea BAC clones. Interestingly, the overall percentage of repetitive elements such as simple and low complexity repeats is small (8.68%) in chickpea genome ranging from 2.12 to 20.31%. In our study, we calculated that 25% of the genes were retro elements in chickpea which are commonly considered to be abundant in the genome and also considered to play a role in genome expansion (Neumann et al. 2006). Among the 11 BAC clones, unequal distribution of gene density was detected from a relatively high density of 1 gene per 5.3 kb (AC145766) to a relatively low density of 1 gene in 63.5 kb (AC145459).
We characterize these BAC clones into two main categories based on selection.
Targeted selection: BAC clones representing ABR-QTL1 region
Three BAC clones (AC161102, AC145766 and AC145456) represent a genomic region where ABR-QTL1 was mapped previously (Tekeoglu et al. 2002). Interestingly, these BAC clones of 80 kb consist of only 6.55% of repetitive sequences, a gene density of 1 gene per 5.7 kb and 14 genes. Of all the genes present in this agronomically important genomic region, only one retrotransposon was detected.
Random selection
The remaining randomly selected BAC clones were further sub-divided into three types based on the presence of retroelements.
BAC clones containing high retrotransposons
In AC161103 (74.4 kb) and AC161104 (56.6 kb), gene islands where genes surrounded by retro elements were observed. This 130,965 bp revealed gene density averaging 1 gene per 6.8 kb which is slightly lower than ABR-QTL1 genomic region and higher percentage level of short tandem repetitive elements (16%).
BAC clones containing low retrotransposons
Three BAC clones (AC145458, AC161105 and AC145459) consist of relatively low amount of retrotransposons with four retroelements of seven genes predicted. The average gene density of one gene for every 11.1 kb which is lower than previous two categories was observed in this 123,087 bp. Also, the moderate percentage level of short tandem repetitive elements (11.2%) was detected.
BAC clones containing no retrotransposons
AC161101 (68.3 kb), AC145457 (23.3 kb), and AC145781 (45.8 kb) are the BAC clones that contained no retrotransposons and had a very low level of short tandem repetitive elements as well (5.4%). These four BAC clones contained an average low gene density of 1 gene per 31.4 kb ranging from 1 gene per 5.8 kb to 1 gene per 65.8 kb.
Microsynteny
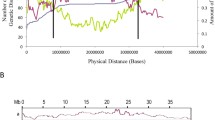
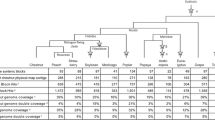
Blast analyses of gene sequences from each chickpea BAC clone with Mt genome sequences and expressed sequence tags (ESTs) were performed for microsynteny studies at http://www.medicago.org and http://www.tigr.org (currently http://www.jcvi.org), respectively. Significant similarities of the genes, representing ABR-QTL1 genomic region, such as protein phosphatase, 40S ribosomal protein, flavanoid galactosyl transferase and eukaryotic transcription factor with e-values ranging from 2e-28 to 1e-134, were observed (Fig. 1a). The physically linked genes in AC145456 and AC145766 represent one chromosome in chickpea while the genes in these BAC clones represented different chromosomes in Mt on blastn analysis of genes from ABR-QTL1 (Fig. 1a). Nevertheless, three of four genes from AC161102 displayed a high degree of microsynteny between chickpea and M. truncatula. Although the genes structures were different, more than 85% homology was detected among these three genes (Fig. 1b).
a Blastn analysis of the genes present in BAC contigs (AC161102, AC145456 and AC145766) representing ABR-QTL1 region with Mt genome sequences at http://www.medicago.org/genome. b Microsynteny analysis of chickpea BAC clone AC161102 and Mt BAC clone AC146861
No significant similarities were detected for genes from the remaining eight BAC clones (AC161103, AC161104, AC145457, AC161101, AC145781, AC145458, AC161105 and AC145459). Twenty-one genes out of 51 predicted genes in chickpea did not identify homologs in genome sequences and ESTs of Mt.
Discussion
Genome organization and composition
Sequencing of 471,638 bp from 11 BAC clones provides significant new insights into chickpea genomic organization. Considering an average gene density of one gene per 9.2 kb and the genome size of 740 Mb, if the genes were evenly distributed throughout the genome, 80,000 genes could be present in chickpea genome. However, since the size of the heterochromatic region in the chickpea genome is unknown, it is clear that the actual number of genes will be much less than this number. Considering the average length of a gene and estimated number of genes (80,000) if they are evenly distributed, gene-rich region is predicted to be occupying 27% of chickpea genome. On the basis of these BAC sequences, the average exon per gene in chickpea is 4.7, which is close to Lj (4.8) and Arabidopsis (5.2) but higher than Mt (3.1). The average predicted exon size (401 bp) in these chickpea BACS is significantly larger than Mt (318 bp), Lj (296 bp) and Arabidopsis (268 bp) and the average intron size of 316 bp, is close to Mt (364 bp), Lj (378 bp) and significantly larger than Arabidopsis (165 bp). Taken together with other parameters such as gene length, gene density, GC content and protein length, chickpea, Mt, Lj and Arabidopsis, all closely resemble one another with the exceptions of gene density and intron length values of Arabidopsis (Table 3). The smaller intron length in Arabidopsis is due to its small genome size, as species with smaller genomes tend to have smaller introns (Dubcovsky et al. 2001). Although the amount of sequence analyzed to predict the genome organization of chickpea (~500 kb) is less than Mt (149 Mbp) and Lj (121 Mbp), the estimated values of the three different, legume genomes appear not to be significantly different from one another (Table 3). This would indicate that the extrapolation of the results obtained from the representative 500 kb sequences, especially the estimates of gene-rich region to the whole chickpea genome, is probably correct. As a first step, gene-rich regions need to be identified in chickpea genome by genetically mapping expressed gene tags (ESTs) and/or by integrating genetic and physical maps.
Genetic analysis, genome organization and composition of ABR-QTL1 region
Our efforts to clone ABR-QTL1 using map-based cloning approach have unraveled the architecture of this agronomically important genomic region. Our genetic analysis using six markers derived from AC161102, AC145766 and AC145456 increased the marker density at QTL1. Marker-trait association using QGene software program discovered 20(T)l12-Right, a CAPS marker derived from the end of AC161102 that accounted 56% of the variation in ascochyta blight resistance at LOD value 19.98, an improvement of the previously reported (35% and LOD 13.40) (Rajesh and Muehlbauer 2008). Low repetitive sequences, high gene density and 14 candidate genes are the characteristics of ABR- QTL1.
Another interesting observation was that none of the genes at this resistant locus resemble NBS-LRR type genes which may hypothesize that the ascochyta blight resistance in chickpea involves a different mechanism other than classical R genes where NBS-LRR motifs are present. This could be due to the fact that Ascochyta rabiei is a necrotrophic fungus and resistance to necrotrophic fungus is poorly understood unlike resistance to biotrophic fungi. However, several candidate genes were predicted at ABR-QTL1 such as Flavonoid 3-0-galactosyl transferase, protein phosphatase and putative eukaryotic transcription initiation factor (TIF6) and are interesting candidate genes for ascochyta blight resistance. The former two genes (Flavonoid 3-0-galactosyl transferase and protein phosphatase) identified ESTs from the database and there were no corresponding ESTs for putative eukaryotic transcription initiation factor (TIF6). There are reports available on possible involvement of flavonoids in disease resistance in other plant systems (Jeandet et al. 2002; Yu et al. 2003); however, further investigation is required to correlate the function of these genes with blight resistance using suitable reverse genetic approaches or genetic transformation.
Microsynteny
The genes in Mt were annotated by Eugene gene prediction program which uses a combined ab initio and homology-based approach while FgenesH program was used in chickpea. Comparison of the Gbrowse displays of Mt at http://www.genome.ou.edu and http://www.tigr.org (currently http://www.jcvi.org) that used Eugene and the final annotation to the raw FgenesH predictions have found that the differences were at either end in the 5′ and/or 3′ non-translated regions that did not effect the gene call.
Choi et al. (2004) estimated the macrosynteny among legumes by genetically mapping orthologous genes and discovered broad genome conservation among them. However, genes that are genetically close to one another might physically be far away from one another (Goff et al. 2002). Although macrosynteny identifies the orthologous genomic location for the genes and determines the gene order along the chromosomes, microsynteny based on genomic sequences unravels the actual evolutionary relationship among the plant species.
A high level of microsynteny was reported among Mt and Lj by comparing the genome sequences of 149 and 121 Mbp, respectively (Cannon et al. 2006). In their study, they estimated a minimum of ten large-scale synteny blocks with substantial colinearity between these two genomes. Phylogenetically, the tribe Cicereae, where chickpea belongs, falls within Trifolieae, which includes both Medicago and Loteae. Hence, we expected microsynteny between chickpea and Mt as found between Mt and Lj.
Surprisingly, our comparison of 11 chickpea BAC clones reveals that Mt and chickpea display limited synteny in this 471,638 kb of genomic sequences. This lack of synteny might be due to one or more of the following reasons: (1) Sampling bias—the representative BAC clones selected are not orthologous between chickpea and M. truncatula. (2) Chickpea BAC selection—8 of 11 chickpea BAC clones are short genomic sequences representing different genomic regions and are not actually contiguous. (3) Incompleteness of the Mt genome—the gene-rich regions of the Mt genome are not sequenced completely and therefore extensive microsynteny was not found between chickpea and M. truncatula. (4) Genomic differences—although, time wise, Lj diverged earlier than chickpea, it is possible that chickpea has undergone evolutionary processes that have caused divergence since its split with Mt, which has resulted in less microsynteny, than Mt has with Lj. Perhaps, by specifically choosing orthologous regions, as well as longer contigs, microsynteny between chickpea and M. truncatula can be investigated in more detail.
In contrast to clear absence of microsynteny with most of the BAC sequences between chickpea and Mt, we observed one syntenous block between chickpea BAC clone AC161102 and Mt BAC clone AC146861. If the orientation of the sequences AC161102 of chickpea is reversed, the gene direction and order will be identical in both chickpea and Mt (Fig. 1b). All four predicted genes in AC161102 of chickpea identified ESTs of Mt from the database at http://www.ncbi.nlm.nih.gov which indicates that these genes are present in the genome. We identified homologs to three of four genes (AC161102.2.fg.1, AC161102.2.fg.2 and AC161102.2.fg.4) from the 23 kb BAC clone, AC161102, in the 125.3 kb Mt BAC clone, AC146861 (Fig. 2). Although these three chickpea genes showed between 88 and 92% homology with their Mt counterparts, they differed in structure with respect to the number of exons in the protein phosphatase gene and the position of exons and introns in the unknown and hypothetical protein genes (Fig. 2). However, it should be noted that these discrepancies could be due to the gene prediction software, and not due to actual differences in intron/exon borders. As the matrix was not specifically designed for chickpea, it could be splitting the genes incorrectly.
One of the four predicted chickpea genes (AC161102.2.fg.3) failed to identify its homolog in Mt and similarly the predicted gene AC146861.24.fg.8 in Mt did not have its homolog in chickpea. Since the sequences that are flanking AC161102 are not available in chickpea, and only “one gene loss and one gene gain” was observed between chickpea and Mt, it is impossible to infer if they are the result of genomic rearrangements. However, by genetically mapping the orthologous genes that are predicted in the flanking Mt BAC sequences (AC146861) in chickpea, microsynteny can be analyzed in depth between chickpea and Mt. Although sequence analysis using 10 of 11 BAC clones did not exhibit microsynteny between chickpea and Mt, the chickpea BAC clone AC161102 showed that comparative genomics between chickpea and Mt should not be completely ruled out.
In conclusion, our analysis of approximately 500 kb of genomic sequence from 11 BAC clones revealed several interesting features of the chickpea genome architecture. Sequencing of BAC clones representing ABR-QTL1 identified candidate genes for future experiments to elucidate the genes that control this agronomically important trait. Also, microsynteny was established between chickpea and Mt at ABR-QTL1 genomic region, suggesting that the genomic data from Mt may be a useful source for identification of additional candidate genes in chickpea. The new markers flanking ABR-QTL1 with increased contribution to resistance can be exploited in marker-assisted selection. Having discovered information on genome composition and organization, our representative work will have significant impact on chickpea genome sequencing efforts in the future.
References
Bodenteich A, Chissoe S, Wang YF, Roe BA (1993) Shotgun cloning as the strategy of choice to generate templates for high-throughput dideoxynucleotide sequencing. In: Venter JC (ed) Automated DNA sequencing and analysis techniques. Academic Press, London, pp 42–50
Buhariwalla HK, Jayashree B, Eshwar K, Crouch JH (2005) Development of ESTs from chickpea roots and their use in diversity analysis of the Cicer genus. BMC Plant Biol 17(5):16
Burge C, Karlin S (1997) Prediction of complete gene structures in human genomic DNA. J Mol Biol 268:78–94
Cannon SB, Sterck L, Rombauts S, Sato S, Cheung F, Gouzy J, Wang X, Mudge J, Vasdewani J, Schiex T, Spannagl M, Monaghan E, Nicholson C, Humphray SJ, Schoof H, Mayer KF, Rogers J, Quétier F, Oldroyd GE, Debellé F, Cook DR, Retzel EF, Roe BA, Town CD, Tabata S, Van de Peer Y, Young ND (2006) Legume genome evolution viewed through the Medicago truncatula and Lotus japonicus genomes. Proc Natl Acad Sci USA 103:14959–14964
Chissoe SL, Bodenteich A, Wang YF, Wang YP, Burian Dennis CSW, Crabtree J, Freeman A, Iyer K, Jian L, Ma Y, McLaury HJ, Pan HQ, Sharan O, Toth S, Wong Z, Zhang G, Heisterkamp N, Groffen J, Roe BA (1995) Sequence and analysis of the human ABL gene, the BCR gene, and regions involved in the Philadelphia chromosomal translocation. Genomics 27:67–82
Choi HK, Mun JH, Kim DJ, Zhu H, Baek JM, Mudge J, Roe B, Ellis N, Doyle J, Kiss GB, Young ND, Cook DR (2004) Estimating genome conservation between crop and model legume species. Proc Natl Acad Sci 101:15289–15294
Corum T, Pang ECK (2005) Isolation and analysis of candidate ascochyta blight defence genes in chickpea. Part I. Generation and analysis of an expressed sequence tag (EST) library. Physiol Mol Plant Pathol 66:192–200
Detter JC, Jett JM, Lucas SM, Dalin E, Arellano AR, Wang M, Nelson JR, Chapman J, Lou Y, Rokhsar D, Hawkins TL, Richardson PM (2002) Isothermal strand-displacement amplification applications for high-throughput genomics. Genomics 80:691–698
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small amount of fresh leaf tissue. Phytochem Bull 19:11–15
Dubcovsky J, Ramakrishna W, SanMiguel PJ, Busso CS, Yan L, Shiloff BA, Bennetzen JL (2001) Comparative sequence analysis of collinear barley and rice bacterial artificial chromosomes. Plant Physiol 125:1342–1353
Ewing B, Green P (1998) Basecalling of automated sequencer traces using phred. II. Error probabilities. Genome Res 8:186–194
Ewing B, Hillier L, Wendl M, Green P (1998) Basecalling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res 8:175–185
Feng Q, Zhang Y, Hao P, Wang S, Fu G, Huang Y, Li Y, Zhu J, Liu Y, Hu X et al (2002) Sequence and analysis of rice chromosome 4. Nature 420:316–320
Gill KS, Gill BS, Endo TR, Taylor T (1996) Identification and high-density mapping of gene-rich regions in chromosome group 1 of wheat. Genetics 144:1883–1891
Goff SA et al (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296:92–100
Gordon D, Abajian C, Green P (1998) Consed: a graphical tool for sequence finishing. Genome Res 8:195–202
Jeandet P, Douillet-Breuil AC, Bessis R, Debord S, Sbaghi M, Adrian M (2002) Phytoalexins from the Vitaceae: biosynthesis, phytoalexins gene expression in transgenic plants, antifungal activity and metabolism. J Agric Food Chem 50:2731–2741
Jurka J (2000) Repbase update: a database and an electronic journal of repetitive elements. Trends Genet 16:418–420
Kosambi DD (1944) The estimation of map distance from recombination values. Ann Eugen 12:172–175
Kato T, Sato S, Nakamura Y, Kaneko T, Asamizu E, Tabata S (2003) Structural analysis of a Lotus japonicus genome: V. Sequence features and mapping of sixty-four TAC clones which cover the 6.4 Mb regions of the genome. DNA Res 10:277–285
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Lichtenzveig J, Scheuring C, Dodge J, Abbo S, Zhang HB (2005) Construction of BAC and BIBAC libraries and their applications for generation of SSR markers for genome analysis of chickpea, Cicer arietinum L. Theor Appl Genet 110:492–510
Manly KF (1998) User’s manual for map manager classic and map manager QT
Nelson JC (1997) QGENE: software for maker-based genomic analysis and breeding. Mol Breed 3:239–245
Neumann P, Koblížková A, Navrátilová A, Macas JI (2006) Significant expansion of Vicia pannonica genome size mediated by amplification of a single type of giant retroelement. Genetics 173:1047–1056
Pan HQ, Wang YP, Chissoe SL, Bodenteich A, Wang Z, Iyer K, Clifton SW, Crabtree JS, Roe BA (1994) The complete nucleotide sequence of the SacBII domain of the P1 pAD10-SacBII cloning vector and three cosmid cloning vectors: pTCF, svPHEP, and LAWRIST16. Genetic Anal Techniques Appl 11:181–186
Rajesh PN, Muehlbauer FJ (2008) Discovery and detection of single nucleotide polymorphism (SNP) in coding and genomic sequences in chickpea (Cicer arietinum L.). Euphytica (in press)
Rajesh PN, Coyne C, Meksem K, Sharma KD, Gupta V, Muehlbauer FJ (2004) Construction of a HindIII bacterial artificial chromosome library and its use in identification of clones associated with disease resistance in chickpea. Theor Appl Genet 108:663–669
Rajesh PN, McPhee K, Muehlbauer FJ (2005) Detection of polymorphism using CAPS and dCAPS markers in two chickpea genotypes. Int Chickpea Pigeonpea Newsl 12:4–6
Rakshit S, Winter P, Tekeoglu M, Juarez Muñoz J, Pfaff T, Benko-Iseppon AM, Muehlbauer FJ, Kahl G (2003) DAF marker tightly linked to a major locus for Ascochyta blight resistance in chickpea (Cicer arietinum L.). Euphytica 132:23–30
Roe BA (2004) Shotgun library construction for DNA sequencing in methods in molecular biology, vol 255: Bacterial artificial chromosomes, vol 1: Library construction, physical mapping and sequencing. Human Press, Totowa, pp 171–187
Roe B, Crabtree J, Khan A (1996) In: Rickwood D (eds) DNA isolation and sequencing: essential techniques series. Wiley, New York
Salamov AA, Solovyev VV (2000) Ab initio gene finding in drosophila genomic DNA. Genome Res 10:516–522
Sandhu D, Gill KS (2002) Gene-containing regions of wheat and the other grass genomes. Plant Physiol 128:803–811
Santra DK, Tekeoglu M, Ratnaparkhe MB, Gupta VS, Ranjekar PK, Muehlbauer FJ (2000) Identification and mapping of QTLs conferring resistance to ascochyta blight in chickpea. Crop Sci 40:1606–1612
Sanyal I, Singh AK, Kaushik M, Amla DV (2005) Agrobacterium-mediated transformation of chickpea (Cicer arietinum L.) with Bacillus thuringiensis cry-IAc gene for resistance against pod borer insect Helicoverpa armigera. Plant Sci 168:1135–1146
Sarmah BK, Moore A, Tate W, Molvig L, Morton RL, Rees DP, Chiaiese P, Chrispeels MJ, Tabe LM, Higgins TJV (2004) Transgenic chickpea seeds expressing high levels of a bean α-amylase inhibitor. Mol Breeding 14:73–82
Stein L et al (2002) The generic genome browser: a building block for a model organism system database. Genome Res 12:1599–1610
Tekeoglu M, Rajesh PN, Muehlbauer FJ (2002) Integration of sequence tagged microsatellite sites to the chickpea genetic map. Theor Appl Genet 105:847–854
Ware D, Stein L (2003) Comparison of genes among cereals. Curr Opin Plant Biol 6:121–127
Wheeler DL et al (2005) Plant genome resources at the National Center for Biotechnology Information. Plant Physiol 138:1280–1288
Yu O et al (2002) A draft sequence of the rice genome. Science 296:79–91
Yu O, Shi J, Hession AO, Maxwell CA, McGonigle B, Odell JT (2003) Metabolic engineering to increase isoflavone biosynthesis in soybean seed. Phytochemistry 63:753–763
Zhang J, Madden T (1997) PowerBLAST: a new network BLAST application for interactive and automated sequence analysis and annotation. Genome Res 7:649–656
Acknowledgments
PNR wishes to thank McKnight Foundation, USA, for the financial support. MB wishes to thank National Science Foundation, USA, for supporting genome sequencing research. We thank Hongshing Lai and Steve Kenton of bioinformatics group at OU for implementing the Genome Browser for the Chickpea BACs and for their help and advice during their analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by D. A. Hoisington.
P. N. Rajesh and M. O’Bleness have contributed equally.
Rights and permissions
About this article
Cite this article
Rajesh, P.N., O’Bleness, M., Roe, B.A. et al. Analysis of genome organization, composition and microsynteny using 500 kb BAC sequences in chickpea. Theor Appl Genet 117, 449–458 (2008). https://doi.org/10.1007/s00122-008-0789-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-008-0789-0