Abstract
‘Express’, a hard red spring wheat cultivar that has been widely grown in the western United States, is used to differentiate races of Puccinia striiformis f. sp. tritici, the causal fungal pathogen of wheat stripe rust. To identify genes conferring race-specific, overall resistance to stripe rust, Express was crossed with ‘Avocet S’. The parents and F1, F2, F3 and F5 populations were tested with races PST-1, PST-21, PST-43, and PST-45 of P. striiformis f. sp. tritici in the seedling stage under controlled greenhouse conditions. Two dominant genes for resistance to stripe rust were identified, one conferring resistance to PST-1 and PST-21, and the other conferring resistance to all four races. Linkage groups were constructed for the resistance genes using 146 F5 lines to establish resistance gene analog and chromosome-specific simple sequence repeat marker polymorphisms. The gene for resistance to races PST-1 and PST-21 was mapped on the long arm of chromosome 1B, and that conferring resistance to all four races was mapped on the long arm of chromosome 5B. We temporarily designate the gene on 1BL as YrExp1 and the gene on 5BL as YrExp2. Polymorphism of at least one of the two markers flanking YrExp2 was detected in 91% of the 44 tested wheat genotypes, suggesting that they would be useful in marker-assisted selection for combining the gene with other resistance genes into many other wheat cultivars. Knowledge of these genes will be useful to understand recent virulence changes in the pathogen populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe rust (yellow rust), caused by Puccinia striiformis Westend. f. sp. tritici Eriks., is one of the most important diseases of wheat (Triticum aestivum L.) worldwide (Stubbs 1985; Chen et al. 1996; Chen 2005). The disease is preferably controlled through the release of resistant cultivars. However, rapid virulence changes in pathogen populations can render resistant cultivars susceptible. Isolates of the pathogen with the same virulence/avirulence pattern on a set of wheat genotypes are considered to be the same race or pathotype. Several sets of wheat genotypes are used to differentiate races of P. striiformis f. sp. tritici in different regions of the world, each set having been chosen to distinguish races meaningful for local wheat production and breeding programs. In the United States, a set of 20 wheat genotypes are used to differentiate races of P. striiformis f. sp. tritici (Chen et al. 2002; Chen 2005, 2007). Among them only ‘Express’ has unknown genes for resistance.
Express (PI 573003), a hard red spring wheat developed by Western Plant Breeders Inc., has been widely grown in the western United States since its release in 1991 (http://www.ars-grin.gov/cgi-bin/npgs/acc/search.pl?accid=PI+573003). The cultivar was highly resistant to stripe rust until 1997, when stripe rust samples were collected from it in California. The stripe rust isolates could not be differentiated from previous races without considering the susceptible reaction on seedlings of Express. Therefore, Express was added to the differential set in order to distinguish races PST-58 and PST-59 from others (Line 2002).
The North American P. striiformis f. sp. tritici population has undergone dramatic changes in virulence structure since 2000 (Chen et al. 2002; Chen 2005, 2007). Pathogen races that had been identified before 2000 are considered “the old group” whereas races identified since 2000 and sharing virulences on Lemhi (Yr21), Lee (Yr7, Yr22, Yr23), Fielder (Yr6, Yr20), Express (Yr?), AVS/6*Yr8 (Yr8), AVS/6*Yr9 (Yr9), Clement (Yr9, YrCle) and Compair (Yr8, Yr19) are considered “the new group” of races. The new group has become predominant in North America and has caused widespread epidemics in the recent years (Chen 2005; Chen and Penman 2005, 2006). Virulence on seedlings of Express has been detected in most isolates of “new group” races. The identification of gene(s) for resistance in Express should lead to a better understanding of the virulence changes and development of better strategies for more effective control of stripe rust.
Although Express is susceptible to the recent predominant races in the seedling growth stage, it has remained moderately resistant in the field. Stripe rust damage on Express has been much less than on susceptible spring wheat cultivars such as Fielder, Zak and Jubilee (X. M. Chen, unpublished data). Stripe rust response data from commercial fields, experimental plots, and controlled greenhouse tests show that Express has non-race specific, high-temperature adult-plant (HTAP) resistance, as well as race-specific, overall (also called seedling) resistance to stripe rust (Chen 2005; Chen and Lin 2007). The mapping of quantitative trait loci (QTL) for HTAP resistance in Express will be published separately. Here, we report the genetics of genes conferring race-specific, overall resistance to stripe rust in Express and mapping them to chromosomal locations using resistance gene analog polymorphism (RGAP) (Chen et al. 1998; Shi et al. 2001) and simple sequence repeat (SSR) (Röder et al. 1998) markers.
Materials and methods
Plant materials
To study the genetics of overall resistance in Express, a cross was made between Express and ‘Avocet S’ (AVS), a line that is susceptible to most races of P. striiformis f. sp. tritici. To determine if Express and Alpowa have the same gene or genes for stripe rust resistance, Express was crossed with ‘Alpowa’, which carries YrAlp on the short arm of chromosome 1B (Lin and Chen 2007). Seeds from the parental plants were used for later phenotype and molecular marker studies. F1, F2 and F3 populations of the two crosses were used to determine the mode of inheritance of overall resistance in Express. F5 lines advanced from individual F2 plants of the AVS/Express cross through single seed-descent were used for genetic analysis and molecular mapping of the resistance genes.
Wheat genotypes ‘Compair’ (Yr8 and Yr19), ‘Druchamp’ (Yr3a, YrDru, YrDru2), ‘Lemhi’ (Yr21), ‘Alpowa’ (YrAlp) and AVS/6*Yr9 (Yr9) were used in molecular marker tests to determine if Express and these cultivars share a resistance gene(s) on chromosomes 1B or 5B; viz., Yr21 in Lemhi, YrAlp in Alpowa, and Yr9 in AVS/6*Yr9 on chromosome 1B and Yr19 in Compair and YrDru in Druchamp on chromosome 5B (Chen et al. 1995; Shi et al. 2001; Chen 2005; Lin and Chen 2007). To determine polymorphism of the markers flanking one of the Express resistance genes, DNA of 44 spring and winter wheat cultivars and genotypes were tested with the markers flanking the resistance gene.
Evaluation of resistance to stripe rust
Seedling tests were conducted under controlled greenhouse conditions as described by Chen and Line (1992a, b). A total of 13 P. striiformis f. sp. tritici races (PST-1, PST-3, PST-7, PST-21, PST-23, PST-43, PST-45, PST-58, PST-59, PST-78, PST-98, PST-100, and PST-111) were initially chosen to confirm the response arrays of Express and AVS. Races that were avirulent on seedlings of Express but virulent on AVS were used to test F1, F2 and F3 populations derived from AVS/Express and Alpowa/Express, and F5 of AVS/Express. Seedlings of parents, and F1, F2, F3 and F5 populations were grown in controlled greenhouse conditions as previously described (Chen and Line 1992a, b). About 15 seeds of each parent, 3 seeds of F1, 300 seeds for F2, and 15 seeds for each of F3 and F5 line were planted in each pot, except for F1s. A total of 146 F3 and 146 F5 lines were used for AVS/Express and 150 F3 lines were used for Alpowa/Express.
Seedlings at the two-leaf stage were inoculated with selected races of P. striiformis f. sp. tritici, grown and evaluated in the seedling stage (Chen and Line 1992a, b). A set of stripe rust differential genotypes was included to confirm the identity of selected races. Infection type (IT) data were collected 18–21 days after inoculation based on the 0–9 scale of Line and Qayoum (1992).
To verify the genotypes of F2 plants showing IT 4–6, the plants were trimmed, transplanted into bigger pots, and grown to obtain F3 seeds. The F3 seedlings were tested with the same races. The IT data of the progenies were used to determine genotypes of F2 plants.
DNA extraction, PCR amplification, electrophoresis and gel visualization
Genomic DNA was isolated from more than 20 plants from two-leaf stage seedlings for each of the parents and F5 lines using the CTAB method (Saghai-Maroof et al. 1984). The RGAP method (Chen et al. 1998; Shi et al. 2001) and the SSR procedure (Röder et al. 1998) were used to target genomic regions.
PCR reactions were performed in a GeneAmp® PCR System 9700 thermo-cycler. A 15 μl reaction mixture consisting of 30 ng of template DNA, 1.5 μl Mg-free 10X PCR buffer (Promega, Madison, WI, USA), 0.6 unit of Taq DNA polymerase (Promega), 5 mM of MgCl2, 0.2 mM each of dCTP, dGTP, dTTP, and dATP (Sigma Chemical Co., St. Louis, MO, USA) and 30 ng of each primer synthesized by Operon Biotechnologies, Inc. (Huntsville, AL, USA). After 5 min of denaturation at 94°C, amplifications were programmed for 40 consecutive cycles, each consisting of 1 min at 94°C, 1 min at either 45, 50, 55, or 60°C (45°C for RGA primers, 50, 55 or 60°C for SSR primers depending on the individual primer pair), 2 min at 72°C and followed by a 7 min extension step at 72°C. After amplification, 6 μl of formamide loading buffer [98% formamide, 10 mM EDTA (pH 8.0), 0.5% (W/V) xylene cyanol and 0.5% (W/V) bromophenol blue] was added to the PCR product. After 4 min denaturation at 94°C, about 7 μl of the PCR product and loading buffer mixture for each sample was loaded for electrophoresis in a 5% polyacrylamide gel as previously described (Chen et al. 1998). After electrophoresis, the gel was silver-stained according to the recommendation of the manufacturer (Promega).
Bulk segregant analysis
The initial resistant and susceptible bulks each consisted of equal amounts of DNA from ten lines based on the phenotypic data from the test with PST-21 that detected two resistance genes in Express. Because these bulks resulted in markers linked only to one of the two genes, additional 20 lines were added to each of the bulks to identify markers for the other gene. The new resistant bulk consisted of ten lines resistant to both races PST-21 and PST-43 that detected only one of the two genes and 20 lines resistant to PST-21 but susceptible to PST-43. The susceptible bulk consisted of all 20 lines susceptible to PST-21 and 10 lines resistant to PST-21, but susceptible to PST-43.
Determination of chromosomal locations of the resistance genes
The nulli-tetrasomic lines of ‘Chinese Spring’ (CS) (Sears 1966) were used to locate RGAP markers on wheat chromosomes using the procedure as previously described (Shi et al. 2001; Lin and Chen 2007). SSR markers from the chromosomes identified with RGAP analysis of the CS nulli-tetrasomic lines were screened to map the resistance genes to chromosomal locations. A total of 15 SSR markers on chromosome 1B and 25 SSR markers on chromosome 5B were screened. The primer sequences of the SSR markers linked with the resistance genes are listed in Table 1.
Data analysis and linkage map construction
Chi-squared analysis was used to test the goodness of fit of observed segregations to specific ratios. The “chitest” procedure in the data analysis of Microsoft Office 2000 was used to calculate P values. Linkage maps were constructed using Mapmaker Macintosh version 2.0 (Lander et al. 1987). Two-point analysis with a logarithm of the odds (LOD) threshold of 3.0 or greater was used to determine linkage relationships among markers whereas multipoint analysis was used for determining the best locus order in linkage groups. Recombination values were converted to map distances using the Kosambi mapping function (Kosambi 1944).
Results
Genetics of resistance in Express to races of P. striiformis f. sp. tritici
The susceptible parent, AVS, was susceptible (IT 9) to all races, whereas Express was susceptible (IT 9) to races PST-58, 59, 78, 98, 100 and 111, but resistant (IT 2) to PST-1, 3, 7, 21, 23, 43 and PST-45 (Table 2). Alpowa was resistant only to PST-1 and PST-21. Based on these results, we chose PST-1, PST-21, PST-43 and PST-45 to evaluate the progenies of AVS/Express and Alpowa/Express to identify genes in Express and determine if Express and Alpowa have a common gene for resistance.
The reactions of F1, F2 and F3 generations of cross AVS/Express to PST-1, PST-21, PST-43 and PST-45 are shown in Table 3. All three F1 plants were resistant (IT 2) to PST-1. Among the 296 F2 plants, 10 were recorded with IT 4 (three plants), 5 (four plants), and 6 (three plants). F3 lines from the 10 plants were tested with PST-1. The progenies of the three plants with IT 4 were homozygous resistant, progenies of the four plant with IT 5 segregated, and progenies of the three plants with IT 6 were homozygous susceptible (IT 7–8). In the analysis in Table 3, seven plants were therefore included in the resistant class and three plants were pooled with the susceptible group. F2 segregation fitted a 15:1 ratio (P = 0.55) for resistant and susceptible plants. The test with PST-21 produced similar results (Table 3). These results suggested that Express had two dominant genes for resistance to PST-1 and PST-21. These results were further confirmed by the segregation of F3 lines into 7:8:1 ratio (P = 0.52) and of the F5 lines in a 175:32:49 ratio (P = 0.17) for homozygous resistant, segregating and homozygous susceptible lines.
When inoculated with PST-43, F1 plants had IT 2, and the F2 population segregated three resistant: 1 susceptible (P = 0.29) without intermediate phenotypes. The segregation among 146 F3 lines fitted a 1:2:1 ratio (P = 0.18) and the 146 F5 lines segregated 7:2:7 (P = 0.23) for homozygous resistant, segregating and homozygous susceptible phenotypic classes. Re-grown plants were tested with PST-45. The results were the same as obtained in the earlier test, thus indicating that resistance to PST-43 and PST-45 was controlled by the same dominant gene.
All 45 F3 lines resistant to PST-43 and PST-45 were also resistant to PST-1 and PST-21, and all 10 lines susceptible to PST-1 and PST-21 also were susceptible to PST-43 and PST-45. Further, all 74 F5 lines resistant to PST-43 and PST-45 were also resistant to PST-1 or PST-21, and all 20 lines susceptible to PST-1 and PST-21 were susceptible to PST-43 and PST-45. These results indicated that the gene for resistance to PST-43 and PST-45 also conferred resistance to PST-1 and PST-21. We tentatively designated the gene conferring resistance only to PST-1 and PST-21 as YrExp1 and the gene conferring resistance to all four races as YrExp2.
Identification and mapping of molecular markers associated with resistance
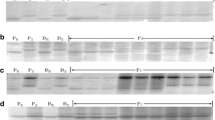
A total of 498 RGA primer pairs were screened for polymorphism between AVS and Express; 128 (25.7%) produced polymorphic bands. Polymorphic RGA primer pairs were tested in the bulk segregant analysis with F5 lines to identify markers associated with resistance genes. A total of 16 RGAP markers were associated with resistance. Using Mapmaker software, the 16 markers were grouped into two linkage groups. Nine were linked to YrExp1 and seven were linked to YrExp2. All were dominant, except, Xwgp82 that was co-dominant. RGAP markers Xwgp78 and Xwgp82 are shown in Fig. 1 as examples.
RGAP markers Xwgp74, Xwgp75, Xwgp 86 and Xwgp87 were assayed on CS and its 21 nulli-tetrasomic lines to determine the chromosomes of the two linkage groups. Marker Xwgp74 was present as an 800-bp band in AVS and CS, but not in Express. Markers Xwgp75, Xwgp86 and Xwgp87 were represented as bands of 810, 480 and 500 bp, respectively, in both Express and CS, but not in AVS. All nulli-tetrasomic lines, except N1BT1A, showed the 800- and 810-bp bands of Xwgp74 and Xwgp75, indicating that YrExp1 is located on chromosome 1B. The 480-bp band of Xwgp86 and 500-bp band of Xwgp87 were detected in all nulli-tetrasomic lines except N5BT5A, indicating that YrExp2 was located on chromosome 5B.
Fifteen SSR markers on 1B and 25 on 5B were screened for polymorphism to confirm the chromosome locations. Two polymorphic SSR markers on the long arm of chromosome 1B and four SSR markers on the long arm of chromosome 5B were found to be associated with the RGAP linkage groups.
All dominant markers, including 15 RGAP and one SSR markers, segregated in 9:7 ratios for presence and absence, and six co-dominant markers (one RGAP and five SSR markers) segregated in 7:2:7 ratios for presence of the Express band, both bands, and the AVS band in each case (Table 4). The results indicated that these markers were single copy and should be reliable for constructing the linkage maps.
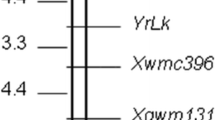
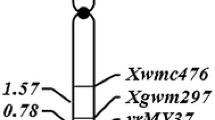
The RGAP and SSR markers were used to construct linkage groups for the resistance genes (Fig. 2). YrExp1 mapped proximal to Xwmc631 outside of the Xwmc631 - Xgwm268 interval. YrExp2 fell in the Xgwm604 - Xgwm639 interval. According to Röder et al. (1998) and Somers et al. (2004), markers Xgwm268 and Xwmc631 are on the long arm of 1B and Xgwm604 and Xgwm639 are on the long arm of 5B; thus YrExp1 and YrExp2 should also be on chromosome 1BL and 5BL, respectively.
Linkage maps for YrExp1 on the long arm of chromosome 1B (a) and YrExp2 on the long arm of chromosome 5B (b) based on the AVS/Express mapping population of 146 F5 lines. All resistant gene analog polymorphism (RGAP) markers were present in Express except for Xwgp74-1B and Xwgp86-5B present in AVS. The linkage groups on 1BL and 5BL were determined by analyzing the nulli-tetrasomic Chinese Spring lines with markers Xwgp74 and Xwgp75 on 1B and Xwgp86 and Xwgp87 on 5B and confirmed with SSR markers Xgwm268 and Xwmc631 on 1BL and Xgwm499, Xgwm604, Xgwm639, and Xwmc118 on 5BL
Relationships of the resistance genes in Express to other genes on chromosomes 1B and 5B
Wheat genotypes Lemhi, AVS/6*Yr9 and Alpowa were reported to have genes Yr21 on 1B (Pahalawatta and Chen 2005a), Yr9 on 1RS (1RS/1BL) (Shi et al. 2001) and YrAlp on 1BS (Lin and Chen 2007), respectively. In the present study, resistance gene YrExp1 was mapped on 1BL. In order to determine the relationships of YrExp1 with Yr21, Yr9 and YrAlp, we tested Express, Lemhi, AVS/6*Yr9 and Alpowa with markers flanking YrExp1 (Xwmc631 and Xwgp78), Yr9 (Xwgp1 and Xwgp15), YrAlp (Xwgp47 and Xwgp48) and markers closely linked to Yr21 (M1 and M2). The flanking markers for Yr9 and YrAlp, and markers closely linked with Yr21 were not detected in Express, indicating that YrExp1 was different from these genes. Eight susceptible plants were observed in the F2 population of the Alpowa/Express cross, which segregated in a 63R:1S ratio, further confirming that YrExp1 was different from YrAlp.
Genes Yr19 in Compair (Chen et al. 1995) and YrDru in Druchamp (Chen et al. 1996) confering race-specific, overall resistance were reported on chromosome 5B. When the wheat genotypes were amplified with the primers for the YrExp2 flanking markers (Xwgp81 and Xwgp82), they did not have the marker bands, indicating that YrExp2 was different from Yr19 and YrDru.
Polymorphism of the YrExp2 flanking markers in wheat cultivars
A total of 44 wheat cultivars were tested for polymorphism of the RGAP markers (Xwgp81 and Xwgp82) flanking YrExp2 (Table 5). Besides Express, both markers were present in four cultivars or breeding lines (Expresso, Blanca Grande, Jeff Pronto and Buck Proto), suggesting that these cultivars may have YrExp2. Six cultivars (Zak, Eden, Lolo, IDO377s, Cashup and Druchamp) had only Xwgp81 whereas Macon and Waikea had only Xwgp82. Thirty-two (73%) had neither marker, suggesting these markers would be useful in marker-assisted selection for combining YrExp2 with other stripe rust resistance genes.
Discussion
Among the 20 wheat genotypes that are used to differentiate races of P. striiformis f. sp. tritici in the United States, Express was the only one without identified resistance genes (Chen 2005). In this study, we determined two genes conferring race-specific, overall stripe rust resistance in Express. Using 13 selected races, we confirmed that Express was resistant to previously identified races when added to the differential set (Line 2002; Chen 2005) and showed that Express is a valuable differential because it has genes different from those in the other 19 differential genotypes (Chen 2005).
Express was developed from the cross Veery/BH1146 (http://www.ars-grin.gov/cgi-bin/npgs/acc/search.pl?accid=PI+573003). Because ‘Veery’ has resistance gene Yr9 in the 1B/1R wheat/rye chromosome translocation (Merker 1982), Express was thought to have Yr9. However, races PST-58 and PST-59 that are virulent on Express are not virulent on genotypes with only Yr9, even though most US Yr9-virulent races detected since 2000 are virulent on Express (Line 2002; Chen 2005). Results from the present study further confirmed that Express does not carry Yr9.
Other stripe rust resistance genes on chromosome 1B include Yr10, Yr15, Yr21, Yr24, YrH52 and YrAlp in Alpowa (Chen 2005; Lin and Chen 2007). Yr10, Yr15, Yr24, Yr26 and YrH52 are located on 1BS whereas YrExp1 is on 1BL. Yr29 confers non-race specific, adult-plant resistance (William et al. 2003) whereas YrExp1 is a gene for race-specific, overall resistance. In wheat genotype Lemhi, gene Yr21 confers resistance only to PST-21 (Chen et al. 1995; Chen 2005), and a closely linked gene, RpsLem, confers resistance to some races of P. striiformis f. sp. hordei (the barley stripe rust pathogen) (Pahalawatta and Chen 2005a). The RGAP markers linked to these genes were absent in Express and markers flanking YrExp1 were absent in Lemhi (data not shown). YrExp1 confers resistance to PST-1, to which Yr21 is not effective, therefore, YrExp1 and Yr21 should be different. YrAlp in Alpowa confers resistance only to races PST-1 and PST-21 (Lin and Chen 2007); however, this gene is located on 1BS whereas YrExp1 is on 1BL. The genetic analysis of Alpowa/Express and the reciprocal marker tests also indicated that they are located at different loci.
Two genes, Yr19 in Compair and YrDru in Druchamp, for race-specific resistance to stripe rust were reported on chromosome 5B (Chen et al. 1995, 1996). Several races, including PST-23 used in this study, are virulent on Druchamp but not on Express (Chen 2005). These observations indicate that YrExp2 on chromosome 5BL is different from YrDru. The relationship between YrExp2 and Yr19 is less clear because Compair also has Yr8 and 27 races virulent on Express also are virulent on Compair (Chen 2005). However, 17 races, including PST-58 and PST-59 used in this study, are virulent on Express, but avirulent on Compair. Five races (PST-68, PST-87, PST-104 and PST-106) are virulent on Express and the Yr8 single gene line, but avirulent on Compair. Further, race PST-121, detected in 2005, is virulent on Compair and avirulent on Express (Chen 2007). Moreover, the YrExp2 flanking markers were not present in Compair and indicated that YrExp2 and Yr19 are different. Based on chromosomal locations, race reactions, and presence/absence of molecular markers, the two genes in Express are different from previously reported genes. Further studies are needed to determine the genetic distances between YrExp1 and those genes on 1BL and between YrExp2 and those on 5BL.
Since YrExp2 confers resistance to more races than YrExp1 (the latter conferring resistance only to the widely avirulent races PST-1 and PST-21), YrExp2 is more useful than YrExp1 for developing resistant cultivars. As YrExp2 confers race-specific overall resistance and is not effective against the current US. P. striiformis f. sp. tritici population, it should be used in combination with other effective genes, especially non-race specific HTAP resistance genes.
One of the major obstacles of marker-assisted selection is lack of polymorphism at marker loci. To determine how useful markers flanking YrExp2 in a wide range of wheat genetic backgrounds, we tested 23 spring and 21 winter wheat genotypes. Among the spring wheat genotypes, Expresso, Blanca Grande, Buck Pronto and Jeff/Pronto had both flanking marker bands, indicating that these genotypes may have YrExp2. The presence of the markers in Expresso was expected because it was developed from Express. However, the presence of YrExp2 in the other three spring wheat cultivars could not be determined because of lack of their pedigree information. In a seedling test of spring wheat germplasms with six races (PST-17, PST-37, PST-43, PST-45, PST-100, and PST-116), all four cultivars and Express were resistant to PST-43 and PST-45 (data not shown), to which YrExp2 is effective as shown in the present study. While the results suggest that these cultivars may have YrExp2, data are not conclusive because some of them were also resistant to races that were virulent on Express, indicating the presence of additional or different resistance genes. The absence of both tightly linked markers in 32 (73%) and one of the markers in eight (18%) of surveyed wheat genotypes indicated potential usefulness of these markers in combining YrExp2 with other genes in wheat cultivars.
References
Chen XM (2005) Epidemiology and control of stripe rust on wheat. Can J Plant Pathol 27:314-337
Chen XM (2007) Challenges and solutions for stripe rust control in the United States. Aust J Agric Res 58:648–655
Chen XM, Jones SS, Line RF (1995) Chromosomal location of genes for stripe rust resistance in spring wheat cultivars Compair, Fielder, Lee, and Lemhi and interactions of aneuploid wheats with races of Puccinia striiformis. Phytopathology 85:375–381
Chen XM, Jones SS, Line RF (1996) Chromosomal location of genes for resistance to Puccinia striiformis in seven wheat cultivars with resistance genes at the Yr3 and Yr4 loci. Phytopathology 86:1228–1233
Chen XM., Lin F (2007) Identification and molecular mapping of genes for all-stage and high-temperature adult-plant resistance to stripe rust in ‘Express’ wheat. Phytopathology 97:S21
Chen XM, Line RF (1992a) Inheritance of stripe rust resistance in wheat cultivars used to differentiate races of Puccinia striiformis in North America. Phytopathology 82:633–637
Chen XM, Line RF (1992b) Identification of stripe rust resistance genes in wheat genotypes used to differentiate North American races of Puccinia striiformis. Phytopathology 82:1428–1434
Chen XM, Line RF, Leung H (1998) Genome scanning for resistance-gene analogs in rice, barley, and wheat by high-resolution electrophoresis. Theor Appl Genet 97:345–355
Chen XM, Moore M, Milus EA, Long DL, Line RF, Marshall D, Jackson L (2002) Wheat stripe rust epidemics and races of Puccinia striiformis f. sp. tritici in the United States in 2000. Plant Dis 86:39–46
Chen XM, Penman L (2005) Stripe rust epidemic and races of Puccinia striiformis in the United States in 2004. Phytopathology 95:S19
Chen XM, Penman L (2006) Stripe rust epidemic and races of Puccinia striiformis in the United States in 2005. Phytopathology 96:S23
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Lander ES, Green P, Abrahamson J, Barlow A, Daly M J, Lincoln SE, Newburg I (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Leister D, Ballvora A, Salamini F, Gebhardt C (1996) A PCR-based approach for isolating pathogen resistance genes from potato with potential for wide application in plants. Nat Genet 14:421–429
Lin F, Chen XM (2007) Genetics and molecular mapping of genes for race-specific all-stage resistance and non-race specific high-temperature adult-plant resistance to stripe rust in spring wheat cultivar Alpowa. Theor Appl Genet 114:1277–1287
Line RF (2002) Stripe rust of wheat and barley in North America: a retrospective historical review. Annu Rev Phytopathol 40:75–118
Line RF, Qayoum A (1992) Virulence, aggressiveness, evolution, and distribution of races of Puccinia striiformis (the cause of stripe rust of wheat) in North America, 1968–87 USDA-ARS, Technical bulletin no. 1788, 44 pp
Merker A (1982) “Veery”—a CIMMYT spring wheat with the 1B/1R chromosome translocation. Cereal Res Comm 10:105–106
Pahalawatta V, Chen XM (2005a) Genetic analysis and molecular mapping of wheat genes conferring resistance to the wheat stripe rust and barley stripe rust pathogens. Phytopathology 95:427–432
Pahalawatta V, Chen XM (2005b) Inheritance and molecular mapping of barley genes conferring resistance to wheat stripe rust. Phytopathology 95:884–889
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–333
Saghai-Maroof MA, Soliman K, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sears ER (1966) Nullisomic–tetrasomic combinations in hexaploid wheat. In: Riley R, Lewis KR (eds) Chromosome Manipulations and Plant Genetics. Oliver and Boyd, Edinburgh, 29–45
Shi ZX, Chen XM, Line RF, Leung H, Wellings CR (2001) Development of resistance gene analog polymorphism markers for the Yr9 gene resistance to wheat stripe rust. Genome 44:509–516
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Stubbs RW (1985) Stripe rust. In: Roelfs AP, Bushnell WR (eds) The Cereal Rusts. Vol. 2, Disease, Distribution, Epidemiology and Control. Academic Press, Orlando, 61–101
William M, Singh RP, Huerta-Espino J, Ortiz Islas S, Hoisington D (2003) Molecular marker mapping of leaf rust resistance gene Lr46 and its association with stripe rust resistance gene Yr29 in wheat. Phytopathology 93:153–159
Yan GP, Chen XM (2007) Molecular mapping of the rps1.a recessive gene for resistance to stripe rust in BBA 2890 barley. Phytopathology 97:668–673
Yan GP, Chen XM, Line RF, Wellings CR (2003) Resistance gene analog polymorphism markers co-segregating with the Yr5 gene for resistance to wheat stripe rust. Theor Appl Genet 106:636–643
Acknowledgments
This research was supported by the US Department of Agriculture, Agricultural Research Service, Vogel Foundation, and Washington Wheat Commission. PPNS no. 0466, Department of Plant Pathology, College of Agricultural, Human, and Natural Resource Sciences Research Center, project no. 13Z-3061-3824 and 13C-3061-3923, Washington State University, Pullman, WA 99164-6430, USA. We are grateful to Dr. Robert A. McIntosh and Dr. Jayaveeramuthu Nirmala for their critical review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. W. Snape.
Rights and permissions
About this article
Cite this article
Lin, F., Chen, X.M. Molecular mapping of genes for race-specific overall resistance to stripe rust in wheat cultivar Express. Theor Appl Genet 116, 797–806 (2008). https://doi.org/10.1007/s00122-008-0713-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-008-0713-7