Abstract
Root growth of Arabidopsis thaliana is inhibited by proton rhizotoxicity in low ionic strength media when the pH of the medium is lower than 5.0. QTL analysis at pH 4.7 revealed that two major QTLs on chromosome 2 and 5 and an additional six epistatic interacting loci pairs control proton resistance in the Ler/Col recombinant inbred population. These genetic factors are independently associated with proton resistance in comparison to the known Al resistant QTL and epistases detected in the same RI population at 4 μM Al at pH 5.0. This indicates that different genetic factors regulate mechanisms of resistance to each stress in this plant species. No correlation was observed between proton resistance and Al resistance among 260 accessions indicating that there is no simple relationship between the genetic factors controlling each trait. Several accessions with different combinations of proton (pH 4.7) and Al (4 μM Al at pH 5.0) resistances were identified by phenotypic cluster analysis. Although this grouping was performed using root growth data, the degree of resistance was correlated with their sensitivity to short-term damage in the root tip, indicating that the same resistance mechanism controls proton resistance at different time scales. Resistant accessions grew better than sensitive ones in acid soil culture. This suggests that proton resistance in hydroponic conditions could be an important index in breeding programs to improve productivity in acid soil, at least in acid sensitive plant species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Growth of crop plants is limited in acid soils due to complex stress factors which consist of a series of toxicities and nutrient deficiencies (Kochian 1995; Salazar et al. 1997). Heavy applications of fertilizer, including calcium carbonate (lime stone) and phosphate fertilizer, are a common means of overcoming stress factors in acid soil (Alva et al. 1986; Speher et al. 1993). However, this approach may not be the ideal solution in either developing or developed countries because of the high cost of fertilizers (Ishitani et al. 2004) and greater energy costs. Hence, improvement of growth capability through enhancement of resistance to stress factors would be a more useful solution to reduce inputs of agricultural fertilizers in acid soil areas. To facilitate effective breeding programs, which include marker assisted selection and genetic modification, it is important to understand the mechanisms of resistance to stress factors at the molecular level.
Rhizotoxicity of aluminum (Al) is among the most important of the complex stress factors that cause severe inhibition of root growth and thus enhancement of drought sensitivity of crop plants (Foy 1988). However, Al in acid soil also induces phosphorous deficiency due to the formation of Al phosphate which shows very limited solubility (López-Bucio et al. 2000). Other stress factors, such as an excess of manganese and a deficiency of calcium, cause growth reduction in most crop plant species (Rengel 1992; Gonzalez et al. 1998; Horst et al. 1999; Rengel and Zhang 2003). In addition, proton toxicity (low pH of soil solution) may also cause damage to plants in some soil types (Knoepp and Swank 1994; Linkes et al. 1997). For example, plant growth is limited by proton toxicity and free Fe2+ toxicity in sulfate soil, but not by Al toxicity. In fact, a screening study of soybean in Brazil indicated that resistance to low Ca, which is connected with low pH resistance (e.g., Kinraide 1998; Koyama et al. 2001), was correlated with growth performance on Cerrado soil containing a large amount of exchangeable Al (Spehar and Sauza 1999). These situations indicate that protons must be a stress factor in acid soils and may exert selection pressure in agricultural fields in some areas. These multiple stress factors drastically reduce the yield of crop plants.
This complex series of stress factors can be simulated under laboratory conditions, and this can be useful for physiological and molecular biological studies aimed at identifying genes coding for resistance (Kobayashi et al. 2005). For example, Al in growth media induces phosphate deficiency due to the formation of a precipitate of Al-phosphate. This precipitation becomes significant when growth medium pH is higher than pH 4.5. High concentrations of phosphate drastically decrease Al toxicity, but exacerbate phosphate deficiency stress (Koyama et al. 1988). On the other hand, low pH growth media (pH < 4.5) enhance Al toxicity because Al in the medium can exist in its phytotoxic form, Al3+ (Kinraide and Parker 1989, 1990); this induces proton rhizotoxicity in some plant species (Kinraide and Parker 1990; Yokota and Ojima 1995; Koyama et al. 1995). This complex interplay between toxic mechanisms makes identification of resistant mechanisms at the molecular level difficult.
Genetic approaches, for instance mutant studies, could be useful for distinguishing among the complex stress factors acting under experimental conditions and could help to clarify the molecular mechanisms of resistance to each stress factor. For example, two types of mutant carrot cell lines have been selected in Al-containing media, namely Al tolerance (Arihara et al. 1991) and efficient P uptake from insoluble Al-phosphate (Koyama et al. 1988). The latter cell line shares characteristics with a capable of efficient P utilization from Al phosphate, namely white lupin (Neumann et al. 1999; Kihara et al. 2003). These studies contributed to separate Al resistance and resistance to Al-phosphate induced P deficiency, which are involved in complex Al stress. Similar genetic approaches could be useful for separating other stress factors in acid soils under experimental conditions, such as Al resistance and proton resistance.
One screening study of proton resistance indicated that these factors are genetically different in soybean (Lazof and Holland 1999), but such genetic analysis is very limited in other plant species. In addition, interference of proton toxicity during the selection and evaluation of Al resistant cultivars was suggested by common bean (Rangel et al. 2005) and spinach (Yang et al. 2005). Hence, it would be difficult to conclude that resistance to proton rhizotoxicity is genetically independent from Al resistance. Genetic studies in Arabidopsis could be a useful means of answering this question, because each stress factor can be clearly distinguished in this plant species. Al and proton rhizotoxicities cause different patterns of cell damage in growing root tips of Arabidopsis thaliana (Koyama et al. 1995). Exposure to low pH growth media damages the plasma membrane of root tip cells irreversibly within a short time (Koyama et al. 2001), whereas exposure to Al solution causes swelling of cells with no visible damage to the plasma membrane (Kobayashi et al. 2005).
Extensive seed collections of natural accessions and mapping populations for QTL analysis are available from public biological resource centers (e.g., NASC, TAIR and RIKEN BRC). These biological resources could be useful for comparing the genetic architecture of these stress factors. In the present study, we used two genetic approaches to compare Al resistance and proton resistance in Arabidopsis. Firstly, using the same recombinant inbred (RI) population that was used in a QTL study of Al resistance (Kobayashi and Koyama 2002), we performed QTL analysis of proton resistance in Arabidopsis. Secondly we conducted a cluster analysis to identify specific groups with differential Al and proton resistance profiles. Together with physiological characterization, we found that Al resistance and proton resistance in Arabidopsis appear to be genetically independent and driven by different functional genes.
Materials and methods
Arabidopsis accessions
Original seed of 260 A. thaliana accessions were obtained from Sendai Arabidopsis seed stock center, and the seed progenies were derived by single seed descent methods (see supplementary Table 1). These accessions are currently available at RIKEN BRC (SASSC; http://www.brc.riken.go.jp/lab/epd/SASSC). The recombinant inbred (RI) population was made by a cross between Col-4 and Ler-0 (Lister and Dean 1993) at the Nottingham Arabidopsis stock center (NASC; http://www.arabidopsis.info/). A subset of the RI population (99 lines) was used for QTL analysis of proton resistance.
Growth conditions
Arabidopsis hydroponic culture was conducted as described previously (Koyama et al. 2001; Kobayashi et al. 2005). Briefly, control culture solutions were prepared by adding 1/50 strength Ca-free and P-free MGRL nutrients (Fujiwara et al. 1992) to 200 μM CaCl2 solution. Al-toxic solutions were prepared by adding an appropriate amount of Al stock (1 mM AlCl3) to the control culture solution at pH 5.0, whereas low pH treatment solutions were adjusted to appropriate pH by adding 0.1 N HCl to the control culture solution. Screening of accessions was carried out in Al solution containing 4 μM total Al at pH 5.0, whereas that for proton resistance was done at pH 4.7 with no Al. In this solution, Al is monomer and {Al3+} at the plasma membrane surface is 14.3 μM (personal communication with Dr. Thomas Kinraide at ARS, USDA). Seedlings were synchronously germinated by pre-incubation in distilled water at 4°C for 3 days and then grown on plastic mesh supported by photo slide mounts as described previously (Toda et al. 1999). Seedlings (10 per accession) were grown in plastic containers containing 6 l of culture solution (maximum 130 lines per container) with 12 h of light per day (PPFD 30 μmol E m−1 s−1) at 25°C. All culture solutions were renewed every 2 days, in order to maintain rhizotoxicities in the test solutions during growth experiments.
Estimation of root growth
Root length was measured on day 7 using a video microscope (Pico Scopeman, Kenis, Tokyo) equipped with a device for measuring length on a TV monitor (MC-300, Kenis). Ten seedlings of each accession for each treatment were grown and scored for root length. The three highest values were used to omit growth-delayed seedlings. This process was repeated three times, and data were pooled and used for calculating mean relative root length (RRL) and other values. As described previously, this could minimize the effect of growth-delayed individuals and is suitable for estimating the potential capacity of accessions under given conditions (Kobayashi et al. 2005). The RRL was defined as RRL (the mean of the root length in toxic solution/that in the control).
QTL analyses
QTL analyses were carried out following the same methods used in previous QTL studies (Kobayashi and Koyama 2002; Kobayashi et al. 2005). Genetic linkage maps of the Ler/Col populations were processed using public data for the genotypes of RI lines obtained from NASC (http://www.arabidopsis.info/) using Mapmaker/EXP version 3.0b (Lander et al. 1987). About 195 genetic makers were used for Ler/Col RI populations. Using RRL as a resistance index, composite interval mapping was carried out using QTL Cartographer, Model 6 (ver. 1.13; Basten et al. 1994), while epistatic interacting loci pairs were analyzed by complete pair-wise search (i.e., automated genome wide two-way ANOVA) using EPISTAT (Chase et al. 1997). The complete pair-wise search allows detection of epistatic interactions between two loci, even if the marker has not been detected as a significant QTL by the CIM method. To reject false positives, a threshold of the logarithm of odd (LOD) score for CIM analysis was calculated by a permutation test as described by Churchill and Doerge (1994) (1,000 times permutation at the significant level of α = 0.05), while the threshold for detection of LLR of EPISTAT was fixed to P < 0.005. The R 2 values were calculated by each program.
Phenotypic cluster analysis
The data sets of phenotypes (root length in control solution and RRLs of low pH treatment and Al treatment) were imported into the CLUSTER program (Eisen et al. 1998; http://www.rana.lbl.gov/EisenSoftware.htm). The data were first mean-centered for each condition, and then hierarchical cluster analysis was performed using an uncentered matrix and average linkage method. Resulting tree figures were displayed using the software package, Treeview (http://www.rana.lbl.gov/EisenSoftware.htm).
Statistical analysis
The coefficient of variation (CV; %) for each condition was calculated by dividing the standard deviation by the mean. Broad sense heritability (H b2 ) was calculated from RRL values (n = 9) of accessions using the following formula
where σ 2g is genetic variance, σ 2e environment variance, and r number of data points employed. Significant differences among the means of all accessions were estimated using one-way ANOVA followed by a least significant difference (LSD) test (P < 0.01). The coefficient of determination (r 2) between Al and low pH treatments was calculated using the RRLs of accessions in each treatment.
Root staining and microscopic observations
Histochemical analyses of the root tips derived from the proton and Al treatments were performed as described previously (proton stress, Koyama et al. 2001; Al stress, Kobayashi et al. 2005). Briefly, growing roots (day 5–7 in the control MGRL medium that contained 1/50 MGRL nutrients and additional Ca to give a final concentration of 200 μM) were transferred to stress conditions (proton stress: 200 μM CaCl2 at pH 4.7 for 1 or 24 h; Al stress: 4 μM Al at pH 5.0 for 24 h). Damage caused by proton stress was visualized using propidium iodide (PI; 4.5 μM of PI for 1 m), while callose and H2O2 accumulation caused by Al stress were visualized using aniline blue [AB; 0.1% (w/v) aniline blue in 0.15 M K2HPO4 at pH 9.5 for 15 min] and 2′,7′-dichlorodihydrofluorescein diacetate (H2DCFDA; 10 μM for 10 min), respectively. Root apexes were stained directly using PI and H2DCFDA, while the roots were pre-fixed in 10% (w/v) formaldehyde, 5% (w/v) glacial acetic acid and 45% (w/v) ethanol in vacuo for 3 h and then used for the AB staining. Fluorescence in the root tip was observed using a fluorescent microscope (IMT-2-21-RFL, Olympus, Tokyo) equipped with appropriate dichroic mirror units (IMT-2-DMG for PI; IMT-2-DMIB for H3DCFDA; and IMT-2-DMV for AB-callose, respectively). Images were photographed using a digital camera (PMDC α/OL-1, Olympus).
Soil culture of specific responses in accessions extracted from cluster analysis
Soil culture was performed with an acid soil, namely Kawatabi soil, which had been employed in several previous studies for estimating the resistance of varieties and transformants in acid soil environments (e.g., Kobayashi et al. 2005; Ma et al. 2005). Basal soil was prepared by adding 250 mg NaH2PO4, 48 mg KCl, 36 mg MgSO4, and 132 mg (NH4)2SO4 to 100 g of air-dried soil. Either 0.06 or 0.25 g of CaCO3 was added to 100 g aliquots of basal soil which was then designated as partially neutralized soil or neutralized soil, respectively. Seedlings were grown for 3 weeks under a 12 h light and dark photoperiod (PPFD; 100 μmol E m−1 s−1) at 20–25°C.
Results
QTL controlling proton resistance in two RI populations
The QTLs controlling Al resistance have been identified in the Ler/Col RI population (Kobayashi and Koyama 2002; Hoekenga et al. 2003). The QTLs controlling proton resistance in this RI population have not been identified. We therefore performed QTL analysis of proton resistance using relative root length (RRL) at pH 4.7 as the resistance index. There was a small difference in RRL between the parental accessions, Ler and Col. The RRLs of Ler and Col were 49.0 and 44.1%, respectively (Table 1). The RI population showed a normal distribution of RRL values (0.025 < P < 0.050) when assessed using the χ 2 test (χ 2 = 12.51) with a coefficient of variation of 20.7% (Ler/Col). The broad sense heritability (H b2 ) of RRL in the RI population was 0.99. Using these data sets, we identified two major QTLs (P < 0.05) by the CIM method and six significant epistatic interacting loci pairs (P < 0.005) using a complete pairwise search (Tables 2, 3). Major QTLs were detected in the middle of chromosome 2 (73.9 cM from the top and linked to genetic marker nga1126) and in the middle of chromosome 5 (109.49 cM from the top and linked to genetic marker mi219), which together explained about 35% of the proton resistance phenotypic variation within the Ler/Col RI population. Although the Ler allele showed a positive effect on the RRL at both QTL positions, three of the heterologous epistatic loci pairs (e.g., CL and LC of nga139 × mi2) had more positive effects on the RRL than the Ler homologous loci pairs. This variation in the genetic influences on RRL could explain transgressive segregation among the Ler/Col RI population. No major QTL detected by the proton resistance test overlapped that associated with Al resistance previously identified in the same mapping population (Kobayashi and Koyama 2002) (Table 2).
Distribution of Al resistance and low pH resistance of Arabidopsis accessions
We used 260 of 300 natural accessions (ecotypes) based on their suitability for hydroponic culture, and grew them in control (pH 5.0), Al (4 μM Al at pH 5.0), and low pH (pH 4.7) solutions. Root growth of the accessions ranged from 8.1 to 20.2 mm in the control solution, while those in the proton (pH 4.7) and the Al (4 μM) toxic solutions ranged from 4.0 to 13.7 mm and from 1.0 to 16.1 mm, respectively (Table 4). In all cases, broad sense heritability was higher than 0.90. Using these data sets, we calculated RRL as the resistance index of accessions for each stress factor and found each to conform to a typical normal distribution judged by χ 2 test (pH, χ 2 = 8.45; Al, χ 2 = 8.77; 0.250 < P < 0.500) (Fig. 1a, b). Coefficients of variation for RRL in low pH and Al solutions were 13.0 and 22.4, respectively. Larger CV values suggested that the genetic factor controlling root growth in the Al solution would be more valuable for breeding purposes than that controlling root growth in the low pH solution. No significant (i.e., r 2 = 0.003, P = 0.393) correlation of RRL in the Al and proton toxic solutions indicated that there was no simple genetic interaction between these two traits (Fig. 1c).
Distribution of relative root lengths of 260 Arabidopsis thaliana accessions in low pH (a) and Al (b) treatments. Root length in control solution (pH 5.0, no Al) was expressed as 100%. Seedlings of accessions were grown hydroponically for 7 days in low pH (pH 4.7) or in Al (4 μM, pH 5.0) solutions. Correlation of the RRL in low pH and Al is shown in c
Cluster analysis of Al resistance and proton resistance among accessions
Cluster analysis of the phenotypes of natural accessions is one approach to analyzing genetic interactions between various traits (Maloof et al. 2001). Using root length in control solution as a reference, we performed hierarchical cluster analysis for the RRL of Al and low pH solutions. As shown in Fig. 2, accessions can be grouped by different combinations of Al-resistance and proton-resistance. Accessions in the right side cluster are mostly sensitive to Al (green at the second line), while the left side cluster contains both Al-resistant and intermediate accessions (red or black). Five intermediate accessions (i.e., both black for Al resistance and proton resistance) were grouped in the right side cluster. The right side cluster also contained one group of Al-sensitive/proton-resistant accessions (SR, 32 accessions; pink bar) and two groups of Al/proton-sensitive accessions (SS, 31 accessions; blue bar). On the other hand, the Al/proton-resistant group (RR, 28 accessions; orange bar) and the Al resistant/proton-sensitive group (RS, 24 accessions; brown bar) were identified in the left side cluster. To examine the reliability of the groping by cluster analysis, we compared root growth of selected accessions (RR: Lö-2 and Co-4; RS: Col-4 and Le-0; SR: Bch-4, Fr-2, and SS: Ler-0 and Gy-0) at the usual screening conditions (i.e., 4 μM Al at pH 5.0 and zero Al at pH 4.7) and at more severe screening conditions (i.e., 6 μM Al at pH 5.0 and zero Al at pH 4.6). As shown in Fig. 3, resistant accessions grew better than sensitive accessions under both the more severe and less severe conditions, indicating that the reliability of the groping under variable degrees of stress.
Hierarchical cluster analysis to group accessions that showed similar responses for proton and Al toxicities. Columns indicate the values for each accession. Root length in control solution is shown in the upper row (control) and followed by the relative root length values of 4 μM Al (Al) and pH 4.7 (low pH). Differences from the mean of each condition are indicated by colors in rows as follows: intermediate (black), less than average (green), and greater than average (red). Typical accession groups are highlighted by bars at the lower side of rows. RR Al and low pH resistant (orange), RS Al resistant but low pH sensitive (brown), SR Al sensitive but low pH resistant (pink), and SS Al and low pH sensitive (blue)
Relative root length of selected accession in low pH (pH 4.7 and 4.6, left panel) and Al (4 and 6 μM at pH 5.0, right panel) solutions. Each two accessions with different combinations of Al resistance and low pH resistance were selected from 260 accessions by the phenotypic cluster analysis in Fig. 2 (RR Al and low pH resistant, RS Al resistant but low pH sensitive, SR Al sensitive but low pH resistant, and SS Al and low pH sensitive). Mean ± SD (n = 9) of the relative root length [(growth in Al or at low pH)/control (no Al at pH 5.0)] from three replications are shown. Different letters indicate significant difference by LSD test (P < 0.01)
Aluminum and proton resistance of accessions judged by root tip damage
To further test the reliability of groping, we performed histochemical analyses using known methods for evaluating Al resistance and low pH resistance. To test Al resistance, we observed accumulation of ROS (i.e., H2O2) and callose in the root tip following 24 h Al treatment; both accumulations are sensitive indicators of Al injury. Both SR and SS groups showed brighter fluorescence than RR and RS, indicating that the Al-sensitive groups accumulate greater amounts of ROS (Fig. 4Aa) and callose (Fig. 4Ab) than Al-resistant groups. On the other hand, the damage in the root tip by proton toxicity can be visualized by propidium iodide staining (Koyama et al. 2001). By this method, proton-sensitive accession groups (RS and SS) showed more damage than resistant groups (RR and SR) after growing in the low pH solution for 1 day (Fig. 4B). This provided confirmation of the reliability of assessments of Al and proton resistance judged by RRL.
Histochemical analyses of the root tip damage caused by Al (A) and low pH (B) treatments. Accessions showing different combinations of Al and low pH resistances (RR Al and low pH resistant, RS Al resistant but low pH sensitive, SR Al sensitive but low pH resistant, and SS Al and low pH sensitive) were exposed to Al (4 μM Al, pH 5.0) for 24 and 12 h, then stained by H2DCFDA (A a; for H2O2) and aniline blue (A b; for callose). On the other hand, the same accessions were incubated in the low pH solution (pH 4.7) for 24 h and then stained by propidium iodide (B; damage in the plasma membrane) to visualize damages caused by protons on the plasma membrane. The bar indicates 50 μm
It would also indicate that long-term resistance judged by RRL at 1 week is correlated with that of short-term resistance at 24 h. Although Al injury cannot be detected in less than 1 day by the current procedures, proton damage can be detected within a shorter term (<1 h) by identifying seedlings that have damaged cells in portions of the root tip. When the same number (15 seedlings) of the growing roots of each accession were exposed to a low pH simple solution for a short-term (pH 4.7, CaCl2 200 μM, 30 min), the number of seedlings with damaged cells in the root tip of sensitive accessions (RS and SS) was significantly greater than in the resistant accessions (RR and SR) (Table 5).
Growth of proton resistance and sensitive accessions on acid soil
To determine whether proton resistance judged by hydroponic culture was correlated with resistance in acid soil, we grew two proton-resistant and two proton-sensitive accessions in acid soil. To minimize the effect of Al toxicity in the soil, accessions were selected to have similar levels of Al-resistance (SR and SS) (Fig. 5a). All four accessions grew similarly in neutralized soil (pH 5.5). However, proton-sensitive accessions, Gy-0 and Ler-0, showed growth inhibition in partially neutralized soil (pH 4.6), while no significant growth reduction was observed in Bch-4 and Fr-2 (Fig. 5b).
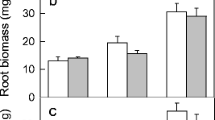
Growth of low pH resistant and sensitive accessions with same level of resistance to Al on a typical acidic andosol of Japan, namely Kawatabi andosol. Plants of SR (Al sensitive but low pH resistance) and SS (Al sensitive and low pH sensitive) were grown for 3 weeks on the neutralized (pH 5.5) and partially neutralized (pH 4.6) andosol which contained different levels of CaCO3 (a). Relative rosette diameter in soil culture (%; pH 4.6/pH 5.5) are also shown (b). Different letters indicate significant difference by LSD test (P < 0.01). The bar indicates 1 cm
Discussion
Physiological and genetic studies to investigate the mechanisms of proton toxicity and resistance are limited when compared to those of Al toxicity. One reason for this may be the uncertain nature of rhizotoxicity of protons in hydroponic culture due to the alleviative effects of other cations in culture solutions (Kinraide et al. 1992). For example, high concentrations of coexisting Ca2+ ion in low pH solutions drastically alleviate proton rhizotoxicity by three separate mechanisms (Kinraide 1998). However, the toxicity is clear in low ionic strength solutions (Koyama et al. 2001). In the present study, we employed a low ionic strength solution to enhance proton rhizotoxicity (Koyama et al. 2001). In this culture solution, growth of accessions were inhibited by lowering pH (Figs. 3, 4B, Table 5). A significant difference in RRL was observed between resistant and sensitive accessions (Fig. 3), indicating that genetic variation must exist to control proton resistance of Arabidopsis.
The solutions used in our study combined higher-than-usual pH for Al studies, low ionic strength, and low Al concentration. This combination of conditions allowed the imposition of Al stress in the absence of proton stress and prevented the formation of polynuclear or solid-phase Al. For the solution that was used for screening of Al sensitivity (4 μM Al, Ca 200 μM at pH5.0), no Al precipitation was expected by GEOCHEM analysis (data not shown), but the Al toxicity to judge by {Al3+} at the plasma membrane surface as estimated by a Gouy–Chapman–Stern model, which is more reliable index for the assessment of Al toxicity than bulk-phase {Al3+} (e.g., Kinraide et al. 1992). The {Al3+}PM in the screening solution is 14.3 μM, which is very similar to that in a common Al test solution (i.e., 100 μM Al in 400 μM Ca at pH 4.4; {Al}PM = 14.7 μM) (personal communication with Dr. Thomas Kinraide at ARS, USDA). In fact, sensitive biochemical indicators of Al toxicity such as callose formation (e.g., Wissemeier et al. 1992) were identified in the same solution (Fig. 4). It was thus we could infer that we could estimate Al sensitivity of accessions by minimizing proton toxicity. In this experimental system, no significant correlations of the RRL in Al and in low pH (Fig. 1c) and differential genetic architecture of QTL in a Ler/Col RI population were observed. These results indicated that proton resistance and Al resistance are mostly regulated by different genetic mechanisms which regulate different biological system.
Cluster analysis with phenotypic values allowed us to group typical accessions that showed different combinations of proton and Al resistance. Because the degree of resistance of accessions was similar even when tested under two degrees of severity, we infer that resistance indices used for phenotypic cluster analysis is reliable (Fig. 2). Resistance of accessions to both toxicities judged by root growth was correlated to that of short-term responses. For example, typical Al damage symptoms, namely callose accumulation (e.g., Schireiner et al. 1994; Kaneko et al. 1999; Sivaguru et al. 2000) and H2O2 generation (Ezaki et al. 2001; Basu et al. 2001) in Al-resistant accessions were less than those in sensitive accessions (Fig. 4Aa, Ab). In addition, an early symptom of proton toxicity in growing roots, namely the short-term damage in the root tip (Koyama et al. 2001), was marked in the proton-sensitive accessions while it was ambiguous in the resistant accessions (Fig. 4B). These results may suggest that the same mechanisms control both long-term and short-term resistance to each stress factor. For example, Arabidopsis Al resistance is mainly regulated by malate release from the roots (Hoekenga et al. 2006), and it can be observed at similar time scales as those employed here to observe callose and H2O2 generation (Yamamoto et al. 2002). Similar relationships are common in other plant species that employ organic acid release as an Al resistance mechanism, such as wheat (Delhaize et al. 1993; Ryan et al. 1995; Sasaki et al. 2004), barley (Ma et al. 2004) and snap bean (Miyasaka et al. 1991).
We were able to identify the genetic architecture of proton resistance in a Ler/Col RI population including two major QTLs detected by CIM and six epistatic interacting loci pairs (Tables 2, 3). The major QTLs for proton resistance did not overlap with previously identified Al-resistant QTLs (Kobayashi and Koyama 2002; Hoekenga et al. 2003). This indicates that the major genetic factors controlling Al resistance and proton resistance are clearly distinct at least in this RI population. In fact, Al responsive malate excretion controlled by the Col allele at the top of chromosome 1 (QTL1 of Al resistance; Hoekenga et al. 2003) was identified as the Al-resistant mechanism in the RI population. However, a T-DNA insertion mutant (KO in AtALMT1) lacks Al-responsive malate excretion and showed no change in low pH sensitivity (pH 4.2; Hoekenga et al. 2006), indicating that malate release is not involved in proton resistance. However, cluster analysis indicates that different mechanisms drive proton resistance, which is distinct from that for Al resistance. At present, there are several possible mechanisms of proton resistance, such as enhanced proton efflux, as a part of integrated reactions (see review Netting 2002) that could contribute phenotypic variations in proton resistance. Comparison of such capacities among accession gropes would be a useful approach for answering this question. In this context, recent advances of public resource centers and databases in A. thaliana could accelerate functional biological approaches for understanding the mechanisms of proton resistance at the molecular level.
In soil culture using acid soil, growth of proton-sensitive accessions was more inhibited than growth of resistant accessions (Fig. 5). Since, in this experiment, we used four accessions with similar levels of Al-resistance, the observed growth differences may be caused by differences in their sensitivity to protons. Although Al toxicity is believed as most the severe stress factor in acid soil, our results suggest that protons can be stress factor in some plant species sensitive to protons such as spinach (Yang et al. 2005) and turnip (Kinraide and Parker 1990). This factor would be one of the important breeding targets for improving crop productivity in acid soil. Further research is needed to examine this possibility.
Reference
Alva AK, Asher CJ, Edwards DG (1986) The role of calcium in alleviating aluminum toxicity. Aust J Agric Res 37:375–382
Arihara A, Kumagai R, Koyama H, Ojima K (1991) Aluminum tolerance of carrot (Daucus carota L.) plants regenerated from selected cell cultures. Soil Sci Plant Nutr 37:699–705
Basten CJ, Weir BS, Zeng Z-B (1994) Zmap-a QTL cartographer. In: Smith C, Gavora JS, Benkel B, Chesnais J, Fairful W, Gibson JP, Kennedy BW, Burnside EB (eds) Proceedings of the 5th World Congress on Genetics Applied to Livestock Production: Computing Strategies and Software 22:65–66
Basu U, Good AG, Taylor GJ (2001) Transgenic Brassicanapus plants overexpressing aluminium-induced mitochondrial manganese superoxide dismutase cDNA are resistant to aluminium. Plant Cell Environ 24:1278–1269
Chase K, Adler FR, Lark KG (1997) Epistat: a computer program for identifying and testing interaction between pairs of quantitative trait loci. Theor Appl Genet 4:724–730
Churchill AG, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Delhaize E, Ryan PR, Randall PJ (1993) Aluminum tolerance in wheat (Triticum aestivum L.). II. Aluminum-stimulated excretion of malic acid from root apices. Plant Physiol 103:685–693
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Ezaki B, Katsuhara M, Kawamura M, Matsumoto H (2001) Different mechanisms of four aluminum (Al)-resistant transgenes for Al toxicity in Arabidopsis. Plant Physiol 127:918–927
Foy CD (1988) Plant adaptation to acid aluminum-toxic soils. Commun Soil Sci Plant Anal 19:959–987
Fujiwara T, Hirai YM, Chino M, Komeda Y, Naito S (1992) Effect of sulfur nutrition on expression of soybean seed storage protein genes in transgenic petunia. Plant Physiol 99:263–268
Gonzalez A, Steffen KL, Lynch JP (1998) Light and excess manganese. Implications for oxidative stress in common bean. Plant Physiol 108:493–504
Hoekenga OA, Vision TJ, Shaff JE, Monforte AJ, Lee GP, Howell SH, Kochian LV (2003) Identification and characterization of aluminum tolerance loci in Arabidopsis (Landsberg erecta (× Columbia) by quantitative trait locus mapping. A physiologically simple but genetically complex trait. Plant Physiol 132:936–948
Hoekenga OA, Maron LG, Pineros MA, Cancado GM, Shaff J, Kobayashi Y, Ryan PR, Dong B, Delhaize E, Sasaki T, Matsumoto H, Yamamoto Y, Koyama H, Kochian LV (2006) AtALMT1, which encodes a malate transporter, is identified as one of several genes critical for aluminum tolerance in Arabidopsis. Proc Natl Acad Sci USA 103:9738–9743
Horst WJ, Fecht M, Naumann A, Wissemeier A, Maier P (1999) Physiology of manganese toxicity and tolerance in Vigna unguiculata (L.) Walp. J Plant Nutr 162:263–274
Ishitani M, Rao I, Wenzlb P, Beebea S, Tohme J (2004) Integration of genomics approach with traditional breeding towards improving abiotic stress adaptation: drought and aluminum toxicity as case studies. Field Crops Res 90:35–45
Kaneko M, Yoshimura E, Nishizawa NK (1999) Time course study of aluminum-induced callose formation in barley roots as observed by digital microscopy and low-vacuum scanning electron microscopy. Soil Sci Plant Nutr 45:701–712
Kihara T, Wada T, Suzuki Y, Hara T, Koyama H (2003) Alteration of citrate metabolism in cluster roots of white lupin. Plant Cell Physiol 44:901–908
Kinraide TB (1998) Three mechanisms for the calcium alleviation of mineral toxicities. Plant Physiol 118:513–520
Kinraide TB, Parker DR (1989) Assessing the phytotoxicity of mononuclear hydroxy-aluminum. Plant Cell Environ 12:479–487
Kinraide TB, Parker DR (1990) Apparent phytotoxicity of mononuclear hydroxy-aluminium to four dicotyledous species. Physiol Plant 79:283–288
Kinraide TB, Ryan PR, Kochian LV (1992) Interaction effects of Al3+, H+ and other cations on root elongation considered in items of cell-surface electric potential. Plant Physiol 99:1461–1468
Knoepp JD, Swank WT (1994) Long-term soil chemistry change in aggrading forest ecosystem. Soil Sci Soc Am J 58:325–331
Kobayashi Y, Koyama H (2002) QTL analysis of Al tolerance in recombinant inbred lines of Arabidopsis thaliana. Plant Cell Physiol 43:1526–1533
Kobayashi Y, Furuta Y, Ohno T, Hara T, Koyama H (2005) Quantitative trait loci controlling aluminium tolerance in two accessions of Arabidopsis thaliana (Landsberg erecta and Cape Verde Islands). Plant Cell Environ 28:1516–1524
Kochian LV (1995) Cellular mechanisms of aluminum toxicity and resistance in plants. Ann Rev Plant Physiol Plant Mol Biol 46:237–260
Koyama H, Okawara R, Ojima K, Yamaya T (1988) Re-evaluation of characteristics of a carrot cell line previously selected as aluminium-tolerant cell line. Physiol Plant 74:683–687
Koyama H, Toda T, Yokota S, Zuraida D, Hara T (1995) Effect of aluminum and pH on root growth and cell viability in Arabidopsis thaliana strain Landsberg in hydroponic culture. Plant Cell Physiol 36:201–205
Koyama H, Toda T, Hara T (2001) Brief exposure to low-pH stress causes irreversible damage to the growing root in Arabidopsis thaliana: Pectin–Ca interaction may play an important role in proton rhizotoxicity. J Exp Bot 52:361–369
Lander ES, Green P, Abrahamson J, Barlow A, Daly M, Lincoln S, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Lazof DB, Holland MJ (1999) Evaluation of the aluminium-induced root growth inhibition in isolation from low pH effects in Glycine max, Pisum sativum and Phaseolus vulgaris. Aust J Plant Physiol 26:147–157
Linkes GE, Driscoll CT, Buso DC (1997) Long-term effects of acid rain: response and recovery of a forest ecosystem. Science 272:244–246
Lister C, Dean C (1993) Recombinant inbred lines for mapping RFLP and phenotypic markers in Arabidopsis thaliana. Plant J 4:745–750
López-Bucio J, Martínez de la Vega O, Guevara-García A, Herrera-Estrella L (2000) Enhanced phosphorus uptake in transgenic tobacco plants that overproduce citrate. Nat Biotech 18:450–453
Ma JF, Nagao S, Sato K, Ito H, Furukawa J, Takeda K (2004) Molecular mapping of a gene responsible for Al-activated secretion of citrate in barley. J Exp Bot 55:1335–1341
Ma JF, Nagao S, Huang CF, Nishimura M (2005) Isolation and characterization of a rice mutant hypersensitive to Al. Plant Cell Physiol 46:1054–1061
Maloof JN, Borevitz JO, Dabi T, Lutes J, Nehring RB, Redfern JL, Trainer GT, Wilson JM, Asami T, Berry CC, Weigel D, Chory J (2001) Natural variation in light sensitivity of Arabidopsis. Nat Genet 29:441–446
Miyasaka SC, Buta JG, Howell RK, Foy CD (1991) Mechanism of aluminum tolerance in snapbeans: root exudation of citric acid. Plant Physiol 96:737–743
Netting AG (2002) pH, abcisic acid and the integration of metabolism in plants under stressed and non-stresses conditions. II. Modifications in modes of metabolism induced by variation in the tension on the water column and by stress. J Exp Bot 53:151–173
Neumann G, Massonneau A, Martinoia E, Romheld V (1999) Physiological adaptations to phosphorous deficiency during proteoid root development in white lupins. Planta 208:373–382
Rangel AF, Mobin M, Rao IM, Horst WJ (2005) Proton toxicity interferes with the screening of common bean (Phaseolus vulgaris L.) genotypes for aluminium resistance in nutrient solution. J Plant Nutr Soil Sci 168:607–616
Rengel Z (1992) Role of calcium in aluminum toxicity. New Phytol 121:499–513
Rengel Z, Zhang W (2003) Role of dynamics of intracellular calcium in aluminium-toxicity syndrome. New Phytol 159:295–314
Ryan PR, Delhaize E, Randall PJ (1995) Characterization of Al-stimulated malate efflux from the root apices of Al-tolerant genotypes of wheat. Planta 196:103–110
Salazar FS, Pandey S, Narro L, Perez JC, Ceballos H, Parentoni SN, Bahia Filho AFC (1997) Diallel analysis of acid-soil tolerant and intolerant tropical maize populations. Crop Sci 37:1457–1462
Sasaki T, Yamamoto Y, Ezaki B, Katsuhara M, Ahn SJ, Ryan PR, Delhaize E, Matsumoto H (2004) A wheat gene encoding an aluminum-activated malate transporter. Plant J 37:645–653
Schireiner KA, Hoddinott J, Taylor GJ (1994) Aluminum-induced deposition of (1,3)-beta-glucans (callose) in Triticum aestivum L. Plant Soil 162:273–280
Sivaguru M, Fujiwara T, Samaj J, Baluska F, Yang ZM, Osawa H, Maeda T, Mori T, Volkmann D, Matsumoto H (2000) Aluminum-induced 1,3-ß-d-glucan inhibits cell-to-cell trafficking of molecules through plasmodesmata. A new mechanism of aluminum toxicity in plants. Plant Physiol 124:991–1005
Speher CR, Monteiro PMFO, Zuffo NL (1993) Soybean breeding in central Brazil. In: Arantes NE, Souza PIM (eds) Proceedings of the Symposium Soybean Cult Braz Cerrados (Savannas), Brazil, pp 229–251
Spehar CR, Sauza LAC (1999) Selecting Soybean (Glycine max L. Merrill) tolerant to low-calcium stress in short term hydroponics experiment. Euphytica 106:35–38
Toda T, Koyama H, Hara T (1999) A simple hydroponic culture method for the development of a highly viable root system in Arabidopsis thaliana. Biosci Biotechnol Biochem 63:210–212
Wissemeier AH, Diening A, Hergenröder A, Horst WJ, Mix-Wagner G (1992) Callose formation as parameter for assessing genotypical plant tolerance of aluminium and manganese. Plant Soil 146:67–75
Yamamoto Y, Kobayashi Y, Devi S, Rikiishi S, Matsumoto H (2002) Aluminum toxicity is associated with mitochondrial dysfunction and the production of reactive oxygen species in plant cells. Plant Physiol 128:63–72
Yang JL, Zheng SJ, He YF, Matsumoto H (2005) Aluminium resistance requires resistance to acid stress: a case study with spinach that exudes oxalate rapidly when exposed to Al stress. J Exp Bot 56:1197–1203
Yokota S, Ojima K (1995) Physiological response of root tip of alfalfa to low pH and aluminium stress in water culture. Plant Soil 171:163–165
Acknowledgments
This research was supported by RITE and JSPS for HK. We wish to thank to Dr. Thomas Kinraide at ARS, USDA for critical reading of the MS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Langridge.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2007_602_MOESM1_ESM.xls
Supplementary Table 1 Al tolerance and low pH tolerance of Arabidopsis 260 accessions. Seedlings were grown hydroponically in Al (pH 5.0, 4 mM), low pH (pH 4.7) and control (pH 5.0, zero Al) solutions for 7 d. Relative root length of each condition is shown (%; Al or low pH / control) (XLS 47 kb)
Rights and permissions
About this article
Cite this article
Ikka, T., Kobayashi, Y., Iuchi, S. et al. Natural variation of Arabidopsis thaliana reveals that aluminum resistance and proton resistance are controlled by different genetic factors. Theor Appl Genet 115, 709–719 (2007). https://doi.org/10.1007/s00122-007-0602-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0602-5