Abstract
The objective of this study was to map new resistance genes against powdery mildew (Blumeria graminis f. sp. hordei L.), leaf rust (Puccinia hordei L.) and scald [Rhynchosporium secalis (Oud.) J. Davis] in the advanced backcross doubled haploid (BC2DH) population S42 derived from a cross between the spring barley cultivar ‘Scarlett’ and the wild barley accession ISR42-8 (Hordeum vulgare ssp. spontaneum). Using field data of disease severity recorded in eight environments under natural infestation and genotype data of 98 SSR loci, we detected nine QTL for powdery mildew, six QTL for leaf rust resistance and three QTL for scald resistance. The presence of the exotic QTL alleles reduced disease symptoms by a maximum of 51.5, 37.6 and 16.5% for powdery mildew, leaf rust and scald, respectively. Some of the detected QTL may correspond to previously identified qualitative (i.e. Mla) and to quantitative resistance genes. Others may be newly identified resistance genes. For the majority of resistance QTL (61.0%) the wild barley contributed the favourable allele demonstrating the usefulness of wild barley in the quest for resistant cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The use of resistant barley varieties has proved an efficient tool to reduce disease severity and to prevent yield losses. Mapping studies in barley, however, have shown that rather few resistance alleles exist in the actual elite germplasm, which constraints the effectiveness of breeding programmes. Williams (2003) argues that a considerable relatedness in cultivated germplasm and re-selection of the same resistance genes may explain the limited number of resistance genes in barley in comparison to non-selected pathosystems such as Arabidopsis. The wide use of a restricted number of qualitative resistance genes has consequently led to a rapid adaptation of pathogens and thus, to a devaluation of these resistance genes. This trend has reinforced the relevance of gene diversity for successful resistance breeding and has directed attention towards quantitative resistance. Quantitative or partial resistance is assumed more durable, because the selection pressure is lower and resistance is more difficult for the fungus to overcome.
The restricted availability of new resistance genes in the gene pool of cultivated barley has directed the focus of research on wild barley as a potential donor of new resistance genes. Qualitative and quantitative resistance genes derived from exotic barley have been identified in different elite genetic backgrounds (Backes et al. 2003; Genger et al. 2003; Ivandic et al. 1998; Qi et al. 1998). Early generations, like F2 and doubled haploid (DH), or recombinant inbred lines (RIL), were primarily used for mapping of exotic resistance genes. The use of advanced backcross (AB) populations, however, has been proposed for more straightforward introgression of QTL from exotic donors (Tanksley and Nelson 1996). Repeated backcrossing with the elite parent reduces the number and size of exotic introgressions, which often carry deleterious traits. The resolution and accuracy of QTL detection is thus increased, and favourable exotic alleles may be rapidly isolated and transferred into elite germplasm. To our knowledge, the study by Wu et al. (2004), who analysed resistance against rice blast in an advanced backcross population, is the only attempt to integrate the mapping of resistance QTL and introgression into high-performing cultivars. In barley, the AB-QTL analysis has been used for QTL detection and introgression of yield related traits from a single exotic donor into two spring cultivars (Pillen et al. 2003, 2004).
Disease-resistance tests are conventionally based either on data from primary detached leaves infected with a single defined isolate or from artificial inoculation under controlled conditions (Ivandic et al. 1998; Sayed et al. 2004). In barley, few people have studied disease severity and infection type of naturally occurring symptoms in the field (Falak et al. 1999; Spaner et al. 1998; Read et al. 2003), although quantitative field resistance of adult plants is assumed more durable than seedling or race-specific resistance. Backes et al. (2003) found different QTL for powdery mildew resistance scored on detached leaves and in the field. The authors concluded that, first, results obtained on primary detached leaves might not be representative for adult plant resistance, and second, factors effective in the field might not be detectable on detached leaves.
In the present study, we attempt to identify favourable exotic alleles, which improve disease resistance in the backcross (BC2)DH population S42 derived from a cross between the spring barley cultivar ‘Scarlett’ and the wild barley accession ISR42-8 from Israel. Here we present data for naturally occurring leaf symptoms of powdery mildew (Blumeria graminis f. sp. hordei L.), leaf rust (Puccinia hordei L.) and scald (Rhynchosporium secalis (Oud.) J. Davis) from field tests in eight environments.
Materials and methods
Plant material
The development of the BC2DH population S42 was conducted according to the advanced backcross strategy of Tanksley and Nelson (1996). The S42 originating from the cross of the German spring barley variety ‘Scarlett’ (S) with the Israeli wild barley accession ISR42-8 (42) was generated via two cycles of backcrossing, followed by doubled haploid production through anther culture (von Korff et al. 2004b). The S42 population counts 301 BC2DH lines derived from 76 BC2F1 and 12 BC1F1 plants.
Molecular characterisation
The BC2DH population was genotyped with 98 SSR markers, which were evenly distributed over the chromosomes. The following prefixes of SSR names indicate the published sources from which the primer sequence information was taken: HVM, Liu et al. (1996); Bmag, Bmac, EBmag, EBmac, Ramsay et al. (2000); Hv, Becker and Heun (1995) and Pillen et al. (2000); GBM, Thiel et al. (2003); GBMS, Li et al. (2003); and MGB, von Korff et al. (2004a). The construction of the SSR consensus map is reported elsewhere (von Korff et al. 2004b). The genotyped markers were assigned to bins according to information by Kleinhofs and Graner (2001) and by the OWB population (Costa et al. 2001, http://barleyworld.org).
Phenotypic evaluation of disease symptoms
Phenotypic evaluation of the BC2DH plants was carried out under field conditions at five different locations in the seasons 2003 (03) and 2004 (04). The test locations were the experimental station Dikopshof of the University of Bonn (D03, D04, West Germany), and the breeders’ experimental stations in Gudow (G03, G04, Nordsaat Saatzucht, North Germany), Morgenrot and Herzogenaurach (M03, M04 and H04, Saatzucht Josef Breun, East and South Germany) and Estrées-Saint-Denis (E04, Saaten Union Recherche, Northern France). The BC2DH population S42 was cultivated without replications in randomised plots (D03, D04, E04 and M04) or in two rows per BC2DH line (M03 and H04) and six rows per BC2DH line (G03 and G04). Plot sizes were 2.25 m2 (D03 and D04), 5 m2 (M04) and 1 m2 (E04). As a control, the recurrent parent ‘Scarlett’ and the barley cultivars ‘Alexis’, ‘Pasadena’ and ‘Chantal’ (as susceptible checks) were tested with eight replications per block. The plots were treated following standard agricultural practices, but without applying fungicides, and no artificial infection with pathogens was carried out. Disease severity was recorded for naturally occurring leaf symptoms of powdery mildew, leaf rust and scald. Disease severity was surveyed at the maximum stage of disease development on a scale from one (resistant) to nine (susceptible).
Statistical analyses
Statistical analyses were carried out with SAS, version 8.0 (SAS Institute 1999). LSMEANS of disease severity for the BC2DH population and the recurrent parent ‘Scarlett’ were calculated for each environment separately. Significant differences between LSMEANS of the S42 population and the recurrent parent were calculated with the procedure GLM using a Tukey–Kramer adjustment for multiple comparisons. The detection of QTL was carried out using the following mixed hierarchical model in the GLM procedure:
Where μ is the general mean, M i the fixed effect of the ith marker genotype, L j (M i ) the random effect of the jth BC2DH line nested in the ith marker genotype, E k the random effect of the kth environment, M i × E k the random interaction effect of the ith marker and kth environment, εm(ijk) is the error of y ijkm . Marker main effects and marker × environment (M × E) interactions are interpreted as a putative QTL, if the P-value calculated by the type III sums of squares is less than 0.01 (Pillen et al. 2003). Groups of linked significant markers (≤20 cM) showing the same effect were interpreted as a single putative QTL, and only the most significant locus from each group of linked loci is recorded. In order to meet the assumption underlying the ANOVA model, the data for leaf rust were transformed by calculating the inverse of the square root. The data for scald were transformed by calculating the inverse. The data for powdery mildew were not transformed. The relative performance of the homozygous exotic genotype (RP [Hsp]) was calculated as follows:
For each trait, Hsp and Hv are the least square means of the homozygous exotic and the homozygous elite genotypes, respectively, calculated across environments. For the traits leaf rust and scald, the LSMEANS of non-transformed data were calculated.
Results and discussion
Powdery mildew
Powdery mildew symptoms were recorded in eight environments, and the mean disease severity in the S42 population ranged from 2.7 to 7.3 (Table 1). Averaged over all environments, the disease severity was significantly lower for the S42 population than for ‘Scarlett’. Although the mean disease severity recorded for the S42 population was below that of the recurrent parent ‘Scarlett’ in each single environment, the difference was only significant in M04.
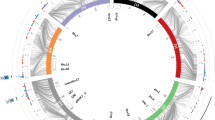
Nine putative QTL were detected for powdery mildew. The QTL were distributed over all chromosomes except for 6H (Fig. 1). The QTL analysis detected seven marker main effects, one of which also displayed a significant M × E interaction effect. At two loci only the M × E interaction effect was significant (Table 2). For six QTL the exotic parent contributed the favourable allele. The strongest effect was measured at locus GMS21 on 1H, where the exotic allele reduced disease severity by 51.5%. This locus corresponds to the Mla locus, which carries a cluster of race-specific powdery mildew resistance genes. Indeed, a large number of the known Mla alleles have already been identified in accessions of H. vulgare ssp. spontaneum (Jorgensen 1994), and several of these have been introduced into elite germplasm (Fischbeck and Jahoor 1991; Zeller 1998).
At the QTL QPm.S42-2H.c, the exotic allele reduced powdery mildew severity by 41.5%. This QTL maps to the same bin as a gene, which encodes chalcone synthase [(CHS) Karakousis et al. 2003)]. CHS belongs to the class of defence response genes and has been shown to accumulate in barley leaves upon inoculation with powdery mildew (Christensen et al. 1998). QPm.S42-2H.c also corresponds to the location of the resistance QTL M1LA introduced from H. laevigatum into barley varieties (Giese et al. 1993). A resistance QTL at this locus, however, has not been reported from studies of crosses with H. vulgare ssp. spontaneum. The QTL QPm.S42-3H.b, for which the M × E effect was significant, maps to the same location as the denso gene for reduced plant height (Laurie et al. 1995). The co-localisation of a QTL for disease resistance with a candidate gene for plant height might indicate disease escape. It is likely that the prostrate growth conditioned by the denso gene, which is present in ‘Scarlett’, favours infection with fungal diseases and, thus, accounts for the QTL detected in this genomic region.
The location of QPm.S42-4H.a corresponds to a quantitative resistance locus against powdery mildew detected under field conditions in an RI population derived from a cross between the cultivar ‘Vada’ and the wild barley accession 1B-87 (Backes et al. 2003). This location on the long arm of chromosome 4H coincides with the position of the mlo gene (Thomas et al. 1998). The recurrent parent ‘Scarlett’, however, is known to carry the Mlg, not the mlo powdery mildew resistance locus on chromosome 4H. The QTL on 4H with the favourable effect of the elite allele might therefore coincide with the Mlg locus mapped to bin 6 by Kürth et al. (2001), although the bin locations do not overlap. At the QTL QPm.S42-5H.a, the exotic allele increased disease susceptibility by 21.0%. The recurrent parent ‘Scarlett’ carries, in addition to the Mlg gene, resistance genes of unknown origin (Anonymous 2004), one of which might be linked to AF043094A on 5H. A previous study by Schönfeld et al. (1996) has already described a resistance gene Mlj derived from H. vulgare ssp. spontaneum close to the significant marker AF043094A on 5H. The same authors reported on a second gene Mlf also derived from H. vulgare ssp. spontaneum. The location of the Mlf gene is close to the QTL QPm.S42-7H.a at HVCHI26A on 7H. At this locus the exotic allele decreased the infection level by 31.9%. The SSR marker HVCHI26A on 7H is derived from a gene sequence, which encodes a chitinase (CHI). Leah et al. (1991) could show that CHI, extracted from barley seeds, has antifungal properties. The CHI might therefore be a candidate for the disease QTL QPm.S42-7H.a.
Leaf rust
Leaf rust symptoms were recorded in four environments, and disease severity ranged from 1.6 to 2.8 in S42 and from 2.0 to 3.9 in ‘Scarlett’ (Table 1). The disease severity averaged over all environments was significantly lower for the S42 population than for ‘Scarlett’. Although the mean disease severity recorded for the S42 population was below that of the recurrent parent ‘Scarlett’ in each single environment, the difference was only significant in D03.
Six putative QTL were identified for leaf rust resistance (Table 2). The QTL for leaf rust resistance were located on 2H, 3H, 4H, 5H and 7H (Fig. 1). At three loci the marker main effect and at three loci the M × E interaction were significant. The exotic allele reduced disease severity at four QTL. The strongest effect was measured for the QTL on the long arm of 4H. Here the exotic allele reduced disease severity by 37.6%.
In barley, about 16 race-specific resistance genes for leaf rust (designated as Rph loci) have been reported (Franckowiak et al. 1997). Four of these genes have been identified in H. vulgare ssp. spontaneum. Feuerstein et al. (1990) detected two resistance genes, Rph10 and Rph11, on chromosomes 3H and 6H, and Ivandic et al. (1998) mapped Rph16 at the centromeric region of 2H. Jin et al. (1996) identified Rph15 on chromosome 2H distal to Rph16. None of these major disease resistance genes, previously identified in wild barley, mapped close to the QTL detected in this study. Park and Karakousis (2002) mapped a major resistance gene (Rph19) in the cross ‘Chebec’ × ‘Harrington’ to locus HVM49 on 7H, which is located 2 cM proximal to the significant QTL QLr.S42-7H.a detected in this study. The QTL QLr.S42-5H.a exhibiting a M × E interaction co-locates with the loci Rph9 and Rph12 detected by Borovkova et al. (1998). The Rph12 resistance originates from the malting barley Triumph and might be active in ‘Scarlett’.
At QTL QLr.S42-2H.b, the exotic allele reduced disease symptoms by 25.9%. On chromosome arm 2HL, a locus for quantitatively inherited resistance against leaf rust (Rphq2) was localised close to the QTL QLr.S42-2H.b found in this study (Qi et al. 1998). Backes et al. (2003) discovered a QTL at the same location in the cross Vada × 1B-87. The QTL QLr.S42-4H.a coincides with the quantitative resistance locus Rphq4, which conferred partial resistance to leaf rust by prolonging the latent infection period (Qi et al. 1998). The correspondence of resistance QTL with the CHS and CHI candidate genes is shown in Table 2 and has already been discussed for powdery mildew.
Scald
Scald symptoms were recorded in four environments, and disease severity ranged from 1.5 to 3.1 in S42 and from 1.0 to 3.1 in ‘Scarlett’ (Table 1). The mean disease severity averaged over all environments was not significantly different between the recurrent parent and the S42 population.
Three putative QTL were detected for scald resistance. At two QTL the M × E interaction effect and at one QTL the marker main effect were significant (Table 2). The pathogen is known to be highly variable in pathogenicity and to overcome rapidly single resistance genes (McDonald and Linde 2002). Different scald pathotypes might have been present in the tested environments. Consequently, race-specific resistance genes might have been active in certain environments resulting in only one QTL detected as a marker main effect. At QRh.S42-2H.a and QRh.S42-3H.a, the exotic allele increased disease severity by 14.3% and 5.3%, respectively. In contrast, the exotic allele decreased disease symptoms by 16.5% at QRh.S42-4H.a with a marker main effect. Spaner et al. (1998) and Jensen et al. (2002) detected a QTL for scald resistance on 4HL. The QTL detected by both authors are located downstream from the QTL discovered in this study. However, considering the large confidence intervals for the localisation of QTL and the difficulty of indirect comparison of map locations, this QTL on 4H might correspond to the one localised by Spaner et al. (1998) and Jensen et al. (2002).
A number of scald resistance genes (Rrs), including some of wild barley, have been mapped predominantly to the centromeric region of chromosome 3H (Abbott et al. 1991, 1995; Schweizer et al. 1995; Graner and Tekauz 1996; Garvin et al. 2000; Genger et al. 2003). The QTL QRh.S42-3H.a, however, was located at the distal end of the long arm of chromosome 3H. Sayed et al. (2004) have also detected two QTL QRS6 and QRS7b on the long arm of chromosome 3H close to the marker HVM62. In addition, Backes et al. (1995) mapped a QTL for scald resistance towards the distal end of chromosome 3H. The QTL QRh.S42-4H.a with a favourable effect of the exotic allele mapped to a genomic region, which to our knowledge has not been identified in previous QTL mapping studies.
General discussion
We were able to demonstrate that QTL for powdery mildew, leaf rust and scald resistance could be detected based on field-scored data. Although the majority of effects were marker main effects, some were M × E interactions, possibly because different pathotypes in different environments may avoid recognition.
Some of the QTL detected in this study correspond to previously identified qualitative genes (i.e. Mla and Mlf); some may be newly detected quantitative resistance factors. In the past, authors have demonstrated that on the basis of field-scored data, previously identified major resistance genes can be verified (Qi et al. 1998; Spaner et al. 1998). On the other hand, some authors have given evidence that additional genetic factors play a crucial role in field resistance (Backes et al. 2003; Qi et al. 2000). Using field-scored data Spaner et al. (1998) were able to map two QTL for powdery mildew within the same bin where the two major resistance genes Mlg and MI(TR) are located. They could hence demonstrate the power and accuracy of QTL analysis, even when based on field-scored data. Williams et al. (2003) found that one QTL of adult plant net blotch resistance mapped to Rph4, a major gene for seedling resistance. The authors reported, however, that QTL on other chromosomes were also important contributors to increased adult plant resistance.
Knowledge of the biological significance underlying quantitative trait loci for disease resistance is generally limited. In recent years, advances in plant–microbe interactions and genome mapping have led to an increased understanding of the genes involved in plant defence and quantitative disease resistance. A large number of defence-related genes have been isolated, and derived functional markers have been applied in QTL analyses for disease resistance (Madsen et al. 2003; Li et al. 1999). Faris et al. (1999) and Wu et al. (2004) reported that candidate defence genes including CHI, thaumatin and CHS had major effects on disease resistance in wheat and rice, respectively. In this study, we found that two loci for CHS on 2H and for CHI on 7H were significant for disease resistance to powdery mildew and leaf rust. At both loci the exotic allele increased disease resistance against powdery mildew and leaf rust. These two loci might be interesting candidates for further investigation into the nature of quantitative resistance.
The distribution of resistance QTL mapped in this study was not random. No QTL was found on chromosome 6H, whereas six resistance QTL were found on chromosome 2H. Five loci on 2HS, 3HS, 4HL, 5HL and 7HL were significant for resistance to at least two diseases (Fig. 1). However, an analysis for correlation between the leaf symptoms of the three investigated diseases did not reveal any significant correlation (data not shown). On the basis of the present results it is difficult to conclude whether the same genes regulate resistance against two diseases or whether tightly linked genes could not be resolved by the current QTL mapping. According to Williams (2003) more than half of the mapped resistance genes occur in clusters, and clusters containing resistance genes for more than one disease are commonly found.
When field-scored data are used, plant growth characteristics may have confounding effects on the study of QTL for resistance. Kicherer et al. (2000) found that a QTL for leaf rust on 2H co-located with a QTL for heading time and may therefore be a pleiotropic effect of earliness. In the present study, chromosome arm 2HS, for example, showed significant effects on powdery mildew, leaf rust and scald resistance. The significant markers were mapped close to a marker associated with the Ppd-H1 gene, which promotes early flowering under long day conditions (Laurie et al. 1995). It is possible that the resistance effect at this locus is a pleiotropic effect of earliness. The earlier plants also showed reduced height (data not shown) and may have therefore been more susceptible to infection.
For the majority of disease QTL (61.0%) the wild barley accession ISR42-8 contributed the favourable allele, demonstrating the usefulness of exotic barley in the quest for resistant cultivars. The identification of markers linked to resistance genes as well as the population structure employed in this study will allow us to rapidly isolate the resistance QTL. Markers closely linked to resistance QTL can be used to select against deleterious wild characters, like head shattering, and to select lines carrying the favourable alleles using marker-assisted selection. Pure introgression lines are currently generated (von Korff et al. 2004b). These will be used to study the molecular basis of disease resistance and to introgress the resistance QTL into adapted germplasm.
References
Abbott DC, Burdon JJ, Jarosz AM, Brown AHD, Mueller WJ, Read BJ (1991) The relationship between seedling infection types and field reactions to leaf scald in Clipper barley backcross lines. Aust J Agric Res 42:801–809
Abbott DC, Lagudah ES, Brown AHD (1995) Identification of RFLPs flanking a scald resistance gene of barley chromosome 6. J Heredity 86:152–154
Anonymous (2004) Bundessortenamt, Beschreibende Sortenliste Getreide, Mais, Ölfrüchte, Leguminosen, Hackfrüchte. Deutscher Landwirtschaftsverlag
Backes G, Graner A, Foroughi-Wehr B, Fischbeck G, Wenzel G, Jahoor A (1995) Localization of quantitative trait loci (QTL) for agronomic important characters by the use of a RFLP map in barley (Hordeum vulgare L). Theor Appl Genet 90:294–302
Backes G, Madsen LH, Jaiser H, Stougaard J, Herz M, Mohler V, Jahoor A (2003) Localization of genes for resistance against Blumeria graminis f. sp. hordei and Puccinia graminis in a cross between a barley cultivar and a wild barley (Hordeum vulgare ssp. spontaneum) line. Theor Appl Genet 106:353–362
Becker J, Heun M (1995) Barley microsatellites: allele variation and mapping. Plant Mol Biol 27:835–845
Borovkova IG, Jin Y, Steffenson BJ (1998) Chromosomal location and genetic relationship of leaf rust resistance genes Rph9 and Rph12 in barley. Phytopathology 88:76–80
Christensen AB, Gregersen PL, Schröder J, Collinge DB (1998) A chalcone synthase with an unusual substrate preference is expressed in barley leaves in response to UV light and pathogen attack. Plant Mol Biol 37:849–857
Costa JM, Corey A, Hayes PM, Jobet C, Kleinhofs A, Kopisch-Obusch A, Kramer SF, Kudrna D, Li M, Riera-Lizarazu O, Sato K, Szucs P, Toojinda T, Vales MJ, Wolfe RI (2001) Molecular mapping of the Oregon Wolfe Barleys: a phenotypically polymorphic doubled-haploid population. Theor Appl Genet 103:415–424
Falak I, Falk DE, Tinker NA, Mather DE (1999) Resistance to powdery mildew in a doubled haploid barley population and association with marker loci. Euphytica 107:185–192
Faris JD, Li WL, Liu DJ, Chen PD, Gill BS (1999) Candidate gene analysis of quantitative disease resistance in wheat. Theor Appl Genet 98:219–225
Feuerstein U, Brown AHD, Burdon JJ (1990) Linkage of rust resistance genes from wild barley (Hordeum spontaneum) with isozyme markers. Plant Breed 104:318–324
Fischbeck G, Jahoor A (1991) The transfer of genes for mildew resistance from Hordeum spontaneum. In: Jorgensen JH (ed) Integrated control of cereal mildews: virulence patterns and their change. Riso National Laboratory, Denmark, pp 247–255
Franckowiak JD, Jin Y, Steffenson BJ (1997) Recommended allele symbols for leaf rust resistance genes in barley. Barley Genet Newsl 27:36–44
Garvin DF, Brown AHD, Raman H, Read BJ (2000) Genetic mapping of the barley Rrs14 scald resistance gene with RFLP, isozyme and seed storage protein markers. Plant Breed 119:193–196
Genger RK, Williams KJ, Raman H, Wallwork H, Burdon JJ, Brown AHD (2003) Leaf scald resistance genes in Hordeum vulgare and Hordeum vulgare ssp. spontaneum: parallels between cultivated and wild barley. Aust J Agric Res 54:1335–1342
Giese H, Holm-Jensen AG, Jensen HP, Jensen J (1993) Localization of the Laevigatum powdery mildew resistance gene to barley chromosome 2 by the use of RFLP markers. Theor Appl Genet 85:897–900
Graner A, Tekauz A (1996) RFLP mapping in barley of a dominant gene conferring resistance to scald (Rhynchosporium secalis). Theor Appl Genet 93:704–771
Ivandic V, Walther U, Graner A (1998) Molecular mapping of a new gene in wild barley conferring complete resistance to leaf rust (Puccinia hordei Otth.). Theor Appl Genet 97:1235–1239
Jensen J, Backes G, Skinnes H, Giese H (2002) Quantitative trait loci for scald resistance in barley localized by a non-interval mapping procedure. Plant Breed 121:124–128
Jin Y, Cui GH, Steffenson BJ, Franckowiak JD (1996) New leaf rust resistance genes in barley and their allelic relationship with other Rph genes. Phytopathology 86:887–890
Jorgensen HJ (1994) Genetics of powdery mildew resistance in barley. Crit Rev Plant Sci 13:97–119
Karakousis A, Gustafson JP, Chalmers KJ, Barr AR, Langridge P (2003) A consensus map of barley integrating SSR, RFLP, and AFLP markers. Aust J Agric Res 54:1173–1185
Kicherer S, Backes G, Walther U, Jahoor A (2000) Localising QTLs for leaf rust resistance and agronomic traits in barley (Hordeum vulgare L.). Theor Appl Genet 100:881–888
Kleinhofs A, Graner A (2001) An integrated map of the barley genome. In: Phillips RL, Vasil IK (eds) DNA-based markers in plants. Kluwer, Dordrecht, pp 187–199
von Korff M, Plümpe J, Michalek W, Léon J, Pillen K (2004a) Insertion of 18 new SSR markers in the Oregon Wolfe Barley map. Barley Genet Newsl 34:1–4
von Korff M, Wang H, Léon J, Pillen K (2004b) Development of candidate introgression lines using an exotic barley accession (H. vulgare ssp. spontaneum) as donor. Theor Appl Genet 109:1736–1745
Kürth J, Kolsch R, Simons V, Schulze-Lefert P (2001) A high-resolution genetic map and a diagnostic RFLP marker for the Mlg resistance locus to powdery mildew in barley. Theor Appl Genet 102:53–60
Laurie DA, Pratchett N, Bezant JH, Snape JW (1995) RFLP mapping of five major genes and eight quantitative trait loci controlling flowering time in a winter × spring barley (Hordeum vulgare L.) cross. Genome 38:575–585
Leah R, Tommerup H, Svensen I, Mundy J (1991) Biochemical and molecular characterisation of three barley seed proteins with antifungal properties. J Biol Chem 266:1564–1573
Li ZK, Luo LJ, Mei HW, Paterson AH, Zhao XZ, Zhong DB, Wang YP, Yu XQ, Zhu L, Tabien R, Stansel JW, Ying CS (1999) A ‘defeated’ rice resistance gene acts as a QTL against a virulent strain of Xanthomonas oryzae pv. oryzae. Mol Gen Genet 261:58–63
Li JZ, Sjakste TG, Röder MS, Ganal MW (2003) Development and genetic mapping of 127 new microsatellite markers in barley. Theor Appl Genet 107:1021–1027
Liu Z-W, Biyashev RM, Saghai Maroof MA (1996) Development of simple sequence repeat DNA markers and their integration into a barley linkage map. Theor Appl Genet 93:869–876
Madsen LH, Collins NC, Rakwalska M, Backes G, Sandal N, Krusell L, Jensen J, Waterman EH, Jahoor A, Ayliffe M, Pryor AJ, Langridge P, Schulze-Lefert P, Stougaard J (2003) Barley disease resistance gene analogs of the NBS–LRR class: identification and mapping. Mol Genet Genomics 269:150–161
McDonald BC, Linde C (2002) The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163–180
Park RF, Karakousis A (2002) Characterization and mapping of gene Rph19 conferring resistance to Puccinia hordei in the cultivar ‘Reka 1’ and several Australian barleys. Plant Breed 121:232–236
Pillen K, Binder A, Kreuzkam B, Ramsay L, Waugh R, Förster J, Léon J (2000) Mapping new EMBL-derived barley microsatellites and their use in differentiating German barley cultivars. Theor Appl Genet 101:652–660
Pillen K, Zacharias A, Léon J (2003) Advanced backcross QTL analysis in barley (Hordeum vulgare L). Theor Appl Genet 107:340–352
Pillen K, Zacharias A, Léon J (2004) Comparative AB-QTL analysis in barley using a single exotic donor of Hordeum vulgare ssp. spontaneum. Theor Appl Genet 108:1591–1601
Qi X, Niks RE, Stam P, Lindhout P (1998) Identification of QTLs for partial resistance to leaf rust (Puccinia hordei) in barley. Theor Appl Genet 96:1205–1215
Qi X, Fufa F, Sijtsma D, Niks RE, Lindhout P, Stam P (2000) The evidence for abundance of QTLs for partial resistance to Puccinia hordei on the barley genome. Mol Breed 6:1–9
Ramsay L, Macaulay M, degli Ivanissevich S, Maclean K, Cardle L, Fuller J, Edwards KJ, Tuvesson S, Morgante M, Massari A, Maestri E, Marmiroli N, Sjakste T, Ganal M, Powell W, Waugh R (2000) A simple sequence repeat-based linkage map of barley. Genetics 156:1997–2005
Read BJ, Raman H, McMichael G, Chalmers KJ, Ablett GA, Platz GJ, Raman R, Genger RK, Boyd WJR, Li CD, Grime CR, Park RF, Wallwork H, Pragnell R, Lance RCM (2003) Mapping and QTL analysis of the barley population Sloop × Halycon. Aust J Agric Res 54:1145–1153
SAS Institute (1999) The SAS system for Windows, release 8.00. Cary
Sayed H, Backes G, Kayyal H, Yahyaoui A, Ceccarelli S, Grando S, Jahoor A, Baum M (2004) New molecular markers linked to qualitative and quantitative powdery mildew and scald resistance genes in barley for dry areas. Euphytica 135:225–228
Schönfeld M, Ragni A, Fischbeck G, Jahoor A (1996) RFLP mapping of three new loci for resistance genes to powdery mildew (Erysiphe graminis f. sp. hordei) in barley. Theor Appl Genet 93:48–56
Schweizer GF, Baumer M, Daniel G, Rugel H, Röder MS (1995) RFLP markers linked to scald (Rhynchosporium secalis) resistance gene Rh2 in barley. Theor Appl Genet 90:920–924
Spaner D, Shugar LP, Choo TM, Falak E, Briggs KG, Legge WG, Falk DE, Ullrich SE, Tinker NA, Steffenson BJ, Mather DE (1998) Mapping of disease resistance loci in barley on the basis of visual assessment of naturally occurring symptoms. Crop Sci 38:843–850
Tanksley SD, Nelson JC (1996) Advanced backcross QTL analysis: a method of the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor Appl Genet 92:191–203
Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L). Theor Appl Genet 106:411–422
Thomas WTB, Baird E, Fuller JD, Lawrence P, Young GR, Russell J, Ramsay L, Waugh R, Powell W (1998) Identification of a QTL decreasing yield in barley linked to Mlo powdery mildew resistance. Mol Breed 4:381–393
Williams KJ (2003) The molecular genetics of disease resistance in barley. Aust J Agric Res 54:1065–1079
Williams KJ, Platz GJ, Barr AR, Cheong J, Willsmore K, Cakir M, Wallwork H (2003) A comparison of the genetics of seedling and adult plant resistance to the spot form of net blotch (Pyrenophora teres f. maculata). Aust J Agric Res 54:1387–1394
Wu J-L, Sinha PK, Variar M, Zheng K-L, Leach JE, Courtois B, Leung H (2004) Association between molecular markers and blast resistance in an advanced backcross population of rice. Theor App Genet 108:1024–1032
Zeller FJ (1998) Nutzung des genetischen Potentials der Hordeum-Wildarten zur Verbesserung der Kulturgerste (Hordeum vulgare L.). J Appl Bot 72:162–167
Zhou FS, Kurth JC, Wei FS EC, Vale G, Yahaiaoui N, Keller B, Somerville S, Wise R, Schulze-Lefert P (2001) Cell-autonomous expression of barley Mla1 confers race-specific resistance to the powdery mildew fungus via Rar1 independent signalling pathway. Plant Cell 13:337–350
Acknowledgements
We thank E. Laubach (Nordsaat Saatzucht), J. Breun (Breun Saatzucht Josef Breun) and V. Lein (Saaten Union Recherche, Estrées) and their teams for conducting the field experiments. The excellent technical assistance of M. Noschinski, C. Golletz, and the team of our experimental station Dikopshof in Wesseling is appreciated. This work was funded by the German Plant Genome Research Initiative (GABI) of the Federal Ministry of Education and Research (BMBF, project 0312278A).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Wenzel
Rights and permissions
About this article
Cite this article
von Korff, M., Wang, H., Léon, J. et al. AB-QTL analysis in spring barley. I. Detection of resistance genes against powdery mildew, leaf rust and scald introgressed from wild barley. Theor Appl Genet 111, 583–590 (2005). https://doi.org/10.1007/s00122-005-2049-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-005-2049-x