Abstract
Retrotransposons have been found to comprise the most common class of transposable elements in eukaryotes and to occur in high copy number in plant genomes. Several of these elements have been sequenced and were found to display a high degree of heterogeneity and insertional polymorphism, both within and between species. The dispersion, ubiquity and prevalence of retrotransposons in plant genomes provide an excellent basis for the development of marker systems and, hence, may be good molecular candidates in distinguishing among apple clones, when they represent bud mutations of the original variety, considering that the random amplified polymorphic DNA (RAPD), amplified fragment length polymorphism (AFLP) and simple sequence repeat (SSR) used thus far in fingerprinting analyses have failed to meet discrimination expectations. The technique called sequence-specific amplified polymorphism (S-SAP), which makes it possible to identify dominant markers for the detection of variation in the DNA flanking the retrotransposon insertion site, was used in the present study to distinguish several clones of the cultivars ‘Gala’ and ‘Braeburn’ in apple fingerprinting. Moreover, our results suggest that the bud mutations, which have generated new patented varieties of ‘Gala’ and ‘Braeburn’, appear to derive from retrotransposon insertion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A number of polymerase chain reaction (PCR) techniques developed over the last few years make it possible to characterise many plant genomes and, hence, represent important tools for fingerprinting. When applied to apple, fingerprinting is usually a routine process by virtue of the gametophytic self-incompatibility of the species and in some cases hybridogenous origin, resulting in a high level of heterozygosity. Indeed, Liebhard et al. (2002) characterised 140 simple sequence repeat (SSR) markers in different apple cultivars, and the literature reports many successes in apple variety fingerprinting using different PCR techniques to identify polymorphisms (Mulcahy et al. 1993; Hokanson et al. 1998). In their review, Wünsch and Hormaza (2002) report that while all the genotypes studied so far can be distinguished with the most widely employed PCR techniques like restriction fragment length polymorphism (RFLP), random amplified polymorphic DNA (RAPD), SSR, inter-simple sequence repeat (ISSR) and amplified fragment length polymorphism (AFLP), they also note that mutations have proved the exceptions, i.e. it is impossible to distinguish apple clones with these tools.
Retrotransposons, the most common class of eukaryotic transposable elements, stand apart from other transposable elements by their ability to transpose via an RNA intermediate, which they convert to DNA by reverse transcription prior to reinsertion. They are generally present in high copy number and occupy a considerable proportion of some plant genomes (Flavell et al. 1992). Attempting to detect polymorphisms at the DNA level, Waugh et al. (1997) combined the general principle of AFLP with sequence-specific PCR in developing a technique capable of revealing the genetic distribution of retrotransposable elements in the barley genome. They called this approach sequence-specific amplified polymorphism (S-SAP) and, given appropriate primer design, described it as potentially applicable to any known sequence.
The last several years have seen a growing number of reports in literature about S-SAP application in plants with short life cycles (from Waugh et al. 1997 to Tahara et al. 2004). In woody plants, Bretó et al. (2001) used S-SAP to distinguish Citrus clementina accessions that had originated from bud mutation during vegetative propagation, or, as the authors report, “selection of fortuitous somatic mutations”, noting further that most discriminative changes among the clementine cultivars correspond to mutations at retrotransposon sequences. Indeed, other researchers studying bud mutation in Citrus frequently report reversal to its “parent” type (Asíns et al. 1999). While most reported examples of this phenomenon can be explained by the chimerical nature of mutations, others cannot (Mendel 1981).
Literature about retrotransposon activity in apple is very scanty. Yao et al. (2001) demonstrated that in several apple mutants (‘Rae Ime’, ‘Spencer Seedless’ and ‘Wellington Bloomless’) a retrotransposon insertion in a MADS-box transcription factor produced apetalous flowers that readily go on to develop into parthenocarpic fruit.
The sports of the apple cultivars (cvs.) ‘Gala’ and ‘Braeburn’ are good cases in point. ‘Gala’ is one of the most important polyclonal varieties and its clones are usually selected on the phenotypic basis of a more intense or diffuse red fruit colour. Indeed, while a handful of its clones with improved skin colour have been released since the 1980s (Dickinson and White 1986; White 1991), these accessions have also generated problems in nursery propagation and varietal discrimination because of phenotype similarity as well as clonal instability due to their chimerical status (Sansavini et al. 1999). Like for Gala, sports of the cv. ‘Braeburn’ are known by their different skin colour intensity, ‘Hillwell’ being an example of selection for its peculiar red colour. Then, too, since clones with entirely yellow fruits or those having the lightest colours do not meet market demands, it is necessary to certify the plant material distributed to growers (Sansavini et al. 1999) via a reliable technique to distinguish among genotypes. The present study thus employed the S-SAP approach to characterise and distinguish ‘Gala’ and ‘Braeburn’ mutated clones via fingerprinting.
Materials and methods
Plant materials
The polyclonal apple cvs. ‘Gala’ and ‘Braeburn’ were used for analysis. The standard ‘Gala’ was compared to its mutant sports ‘Ruby Gala’, ‘IG 31’, ‘Gala Must’, ‘Galaxy’, ‘Royal Gala’ and ‘Mondial Gala’ and cv. ‘Braeburn’ to ‘Hillwell’. All the clones differed from the originals in fruit skin colour.
The DNA was extracted from young leaves harvested in the apple collection at the University of Bologna during early spring, immediately frozen in liquid nitrogen and then stored at −80°C. DNA was extracted from freeze-dried leaves using DNeasy® Plant Mini Kit (Qiagen) and the concentration was assessed by spectrophotometry (Nanodrop, ND-1000, Wilmington, DE, USA). Three different extractions were performed and, consequently, all the protocols were repeated three times for each tested accession so as to check the reproducibility of results.
SSR, AFLP and S-SAP analyses
Twenty-four primer pairs for apple microsatellites (Table 1) that were distributed among the 17 apple chromosomes were analysed according to Liebhard et al. (2002) and Vinatzer et al. (2004). AFLPs were analysed after the standard procedures in Vos et al. (1995). The DNA was digested with EcoRI and MseI and 35 primer combinations (with three selective nucleotides) were tested. For S-SAP analysis, 15 amplifications were carried out using one AFLP primer and one retrotransposon [long terminal repeats (LTRs)] primer according to Waugh et al. (1997), with modifications as per Leigh et al. (2003). The LTR primer sequences employed were LTR1 GCAATTTCGTTCATTCACAG and LTR2 GTGCAAATGGTTACGTCACT (in direction 5′–3′), designed on the only available apple retrotransposon sequence (AJ291492, Yao et al. 2001) but in a different position with respect to those described in that paper.
Amplified fragments were separated and analysed on an LI-COR DNA analyser Gene RedIR 4200 (MWG-Biotech) or on 5% polyacrylamide gels; in the latter case bands were silver-stained. For the Gala clones, S-SAP bands were scored as either present (1) or absent (0) in each sample, and pairwise genetic similarities were calculated with the Dice coefficient (Dice 1945) using only polymorphic markers. The similarity data were used in a UPGMA cluster analysis performed with the NTsys 2.0 software.
Results
SSRs and AFLPs
When the two standard cvs. ‘Gala’ and ‘Braeburn’ were compared, 88% of the SSRs tested (Table 1) produced polymorphisms, with only three markers (CH01C06, CH02B12 and CH03H06) showing the same allele composition within the two cultivars. Similarly, 100% of the AFLP primers produced polymorphic bands between ‘Gala’ and ‘Braeburn’. By contrast, none of the SSR (64 alleles with 24 primer pairs) and AFLP (more than 1,000 bands with 35 primer combinations) markers could distinguish between an original cv. and its clones.
S-SAPs on ‘Gala’ clones
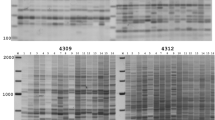
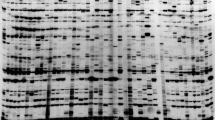
These results, summarized in Table 2 and Fig. 1, indicate that all the tested ‘Gala’ clones are clearly distinguishable by S-SAP markers. The cluster analysis indicates that all the tested clones are different from the standard ‘Gala’ and among themselves (Fig. 1). For cluster analysis only the polymorphic markers were considered so as to emphasize the differences among the sports. The cophenetic matrix correlation (r=0.87) confirmed the stability of the tree nodes. Of the 15 primer combinations tested, 8 produced polymorphic bands among the ‘Gala’ clones and 6 (LTR1M58, LTR1M59, LTR1M60, LTR2M52, LTR2M53, LTR2M55) showed no difference. LTR2M59 clearly distinguished standard ‘Gala’, which has a band of 180 bp that differs from all the sports (Table 2, Fig. 2). LTR1M52 produced a polymorphic band (200 bp) only in the genotype ‘Gala Must’, LTR2M58 one band (180 bp) only in IG 31 and LTR1M61 one band (610 bp) only in ‘Galaxy’ (Table 2). The primer combinations LTR1M53, LTR1M62, LTR2M54 and LTR2M60 produced bands that were shared among more than one genotype: LTR1M53 showed the same marker (about 260 bp) in ‘IG 31’, ‘Galaxy’ and ‘Royal Gala’; LTR1M62 another band (280 bp) in ‘IG 31’, ‘Gala Must’, ‘Galaxy’ and ‘Mondial Gala’; LTR2M54 one band (350 bp) in ‘IG 31’, ‘Gala Must’ and ‘Galaxy’; and LTR2M60 one band (190 bp) in both ‘Gala Must’ and ‘IG 31’. LTR1M54 amplified a band of 400 bp only in ‘Ruby Gala’ and ‘IG 31’, another one of 710 bp in ‘Galaxy’ (Table 2).
Examples of apple fingerprinting by S-SAP analysis: the primer combination LTR2M59 clearly distinguishes 6 ‘Gala’ STD from all its clones, i.e. 1 ‘Ruby Gala’, 2 ‘IG 31’, 3 ‘Gala Must’, 4 ‘Galaxy’, 7 ‘Royal Gala’ and 9 ‘Mondial Gala’. Also included and marked with stars are 8 ‘Braeburn’ and its derivative 5 ‘Hillwell’
S-SAPs on ‘Braeburn’ clones
A single primer combination (LTR2M55) was sufficient to identify a polymorphism between ‘Hillwell’ and the standard ‘Braeburn’: a band of 280 bp, present in only the latter, clearly distinguished it from ‘Hillwell’ (Fig. 3).
Discussion
Our findings demonstrate that S-SAPs were able to produce polymorphisms both among the ‘Gala’ clones and between ‘Braeburn’ and ‘Hillwell’. All the analysed clones, in which the skin colour variation originated via bud mutation (Dickinson and White 1986), can be discriminated from the standard, original variety and among themselves by S-SAPs. The reproducibility of the results was checked. The lack of AFLP polymorphisms, after testing 35 primer combinations, indicates that the polymorphic bands found in our analysis have not been generated by incomplete digestion of DNA or by methylation of restriction sites. Moreover, in the S-SAP analysis, the methylation-insensitive enzyme MseI (Xiong et al. 1999; Yamamoto and Yamamoto 2004) was used to further minimise potential problems due to methylation differences. A sequencing analysis is, however, needed to ascertain whether the polymorphisms really correspond to retrotransposons.
In the literature, several reports suggested retrotransposons as a good model system to characterise apple clones. Sansavini et al. (1999), analysing the instability of some ‘Gala’ clones, reported that 12% of more than 500 trees regressed to yellow or faded red striped skin coloration in the first cropping year (second of orchard life); this instability continued in the next years, indicating that the requisite of stability in patenting new varieties should be reviewed. In Citrus, Asíns et al. (1999) analysed the contribution of retrotransposon activity to generate genetic instability, introducing the idea to use these elements for fingerprinting purpose. Analogously to our results, Bretó et al. (2001) overcame the lack of SSR variability in clementines by using inter-retrotransposon amplified polymorphism (IRAP) markers, discriminating different accessions of this species. The same authors explained these results with the different origin of the polymorphisms of SSR loci and retrotransposons: the former by unequal or asymmetric meiotic recombination within sexual reproduction and by replication slippage, the latter ones by “fortuitous somatic mutations”. Of course mutations that do not involve the bud will not appear in the clonally propagated plants.
The results reported in the present paper are also in agreement with Kobayashi et al. (2004), who report a retrotransposon-induced mutation associated with the loss of pigmentation in white cultivars of Vitis vinifera. Mutations caused by retrotransposon insertions in or near genes can alter gene expression or the structure of the encoded proteins (Kumar and Bennetzen 1999). The possibility that retrotransposon activity may also be related to colour intensity, variation and mutation regression in apple clones has to be considered. In any case, the S-SAP markers are, at present, the only PCR approach that can be used in apple fingerprinting to discriminate the different sports from their standard varieties.
References
Asíns MJ, Monforte AJ, Mestre PF, Carbonell EA (1999) Citrus and Prunus copia-like retrotransposons. Theor Appl Genet 99:503–510
Bretó MP, Ruiz C, Pina JA, Asíns MJ (2001) The diversification of Citrus clementina Hort. ex Tan., a vegetatively propagated crop species. Mol Phylogenet Evol 21:285–293
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Dickinson JP, White AG (1986) Red colour distribution in the skin of Gala apple and some of its sports. NZ J Agric Res 29:695–698
Flavell AJ, Dunbar E, Anderson R, Pearce SR, Hartley R, Kumar A (1992) Ty1-copia group retrotransposons are ubiquitous and heterogeneous in higher plants. Nucleic Acids Res 20:3639–3644
Hokanson SC, Szewc-McFadden AK, Lamboy WF, McFerson JR (1998) Microsatellite (SSR) markers reveal genetic identities, genetic diversity and relationships in a Malus × domestica Borkh. core subset collection. Theor Appl Genet 97:671–683
Kobayashi S, Goto-Yamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304:982
Kumar A, Bennetzen JL (1999) Plant retrotransposons. Annu Rev Genet 33:479–532
Leigh F, Kalendar R, Lea V, Lee D, Donini P, Schulman AH (2003) Comparison of the utility of barley retrotransposon families for genetic analysis by molecular marker techniques. Mol Gen Genomics 269:464–474
Liebhard R, Gianfranceschi L, Koller B, Ryder CD, Tarchini R, Van de Weg E, Gessler C (2002) Development and characterisation of 140 new microsatellites in apple (Malus × domestica Borkh.). Mol Breed 10:217–241
Maliepaard C, Alston FH, van Arkel G, Brown LM, Chevreau E, Dunemann F, Evans KM, Gardiner S, Guilford P, van Heusden AW, Janse J, Laurens F, Lynn JR, Manganaris AG, den Nijs APM, Periam N, Rillerink E, Roche P, Ryder C, Sansavini S, Schmidt H, Tartarini S, Verhaegh JJ, Vrielink van Ginkel M, King GJ (1998) Aligning male and female linkage maps of apple using multi-allelic markers. Theor Appl Genet 97:60–63
Mendel K (1981) Bud mutations in citrus and their potential commercial value. Proc Int Soc Citricult 1:86–89
Mulcahy DL, Cresti M, Sansavini S, Douglas GC, Linskens HF, Bergamini A, Mulcahy G, Vignani R, Pancaldi M (1993) The use of random amplified polymorphic DNAs to fingerprint apple genotypes. Sci Horticult 54:89–96
Sansavini S, Buscaroli C, Stainer R (1999) Instabilità dei mutanti del melo cv Gala. Frutticoltura 10:63–72
Tahara M, Aoki T, Suzuka S, Yamashita H, Tanaka M, Matsunaga S, Kokumai S (2004) Isolation of an active element from a high-copy-number family of retrotransposons in the sweet potato genome. Mol Gen Genomics 272:116–127
Vinatzer BA, Patocchi A, Tartarini S, Gianfranceschi L, Sansavini S, Gessler C (2004) Isolation of two microsatellite markers from BAC clones of the Vf scab resistance region and molecular characterization of scab resistant accessions in Malus germplasm. Plant Breed 123:321–326
Vos P, Hogers R, Bleeker M, Reijans M, Van der Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Xiong LZ, Xu CG, Saghai Maroof MA, Qifa Zhang (1999) Patterns of cytosine methylation in an elite rice hybrid and its parental lines, detected by a methylation-sensitive amplification polymorphism technique. Mol Gen Genet 261:439–446
Waugh R, McLean K, Flavell AJ, Pearce SR, Kumar A, Thomas BB, Powell W (1997) Genetic distribution of Bare-1-like retrotransposable elements in the barley genome revealed by sequence-specific amplification polymorphisms (S-SAP). Mol Gen Genet 253:687–694
White AG (1991) The Gala apple. Fruit Var J 45:2–3
Wünsch A, Hormaza JI (2002) Cultivar identification and genetic fingerprinting of temperate fruit tree species using DNA markers. Euphytica 125:59–67
Yamamoto F, Yamamoto M (2004) A DNA microarray-based methylation-sensitive (MS)-AFLP hybridization method for genetic and epigenetic analyses. Mol Gen Genomics 271:678–686
Yao J, Dong Y, Morris BA (2001) Parthenocarpic apple fruit production conferred by transposon insertion mutations in a MADS-box transcription factor. Proc Natl Acad Sci USA 98:1306–1311
Acknowledgements
We are grateful to Dr. D. Lee, Dr. E. Chiapparino and Dr A. Acquadro for their suggestions during the work done at NIAB, Cambridge (UK). This research was supported with funds from the Emilia-Romagna Regional Government and CAV (Emilia Romagna, Italy).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Nybom
Rights and permissions
About this article
Cite this article
Venturi, S., Dondini, L., Donini, P. et al. Retrotransposon characterisation and fingerprinting of apple clones by S-SAP markers. Theor Appl Genet 112, 440–444 (2006). https://doi.org/10.1007/s00122-005-0143-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-005-0143-8