Abstract
Lycopersicon peruvianum LA2172 is completely resistant to Oidium neolycopersici, the causal agent of tomato powdery mildew. Despite the large genetic distance between the cultivated tomato and L. peruvianum, fertile F1 hybrids of L. esculentum cv. Moneymaker × L. peruvianum LA2172 were produced, and a pseudo-F2 population was generated by mating F1 half-sibs. The disease tests on the pseudo-F2 population and two BC1 families showed that the resistance in LA2172 is governed by one dominant gene, designated as Ol-4. In the pseudo-F2 population, distorted segregation was observed, and multi-allelic, single-locus markers were used to display different marker-allele configurations per locus. Parameters for both distortion and linkage between genetic loci were determined by maximum likelihood estimation, and the necessity of using multi-allelic, single-locus markers was illustrated. Finally, a genetic linkage map of chromosome 6 around the Ol-4 locus was constructed by using the pseudo-F2 population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lycopersicon peruvianum is one of the wild relatives of tomato and provides a vast reservoir of valuable traits for crop improvement, such as disease, pest and virus resistances (e.g. Brüggemann et al. 1996; Ammiraju et al. 2003). L. peruvianum LA2172 is almost immune to Oidium neolycopersici, which was previously named O. lycopersici (Kiss et al. 2001), the causal agent of powdery mildew in tomato (Lindhout et al. 1994). The resistance in L. peruvianum LA2172 is considered as complete, compared to the incomplete resistance conferred by the Ol-1 and Ol-3 genes that map on chromosome 6 and originate from L. hirsutum G1.1560 and G1.1290, respectively (Huang et al. 2000; Van der Beek et al. 1994).
L. peruvianum is reproductively isolated from the esculentum complex by severe crossing barriers (Taylor 1986). These barriers can be partially overcome by using either in vitro techniques or bridge accessions like L. peruvianum LA1708 and LA2172 (Poysa 1990; Rick 1982; Van Heusden et al. 1999; Veremis and Roberts 1996).
L. peruvianum is an out-crossing species, with accessions and individuals within accessions differing in marker alleles at the same locus (Baudry et al. 2001; Ganal and Tanksley 1996; van Ooijen et al. 1994). In several studies, where intra-specific crosses were used for mapping, multi-alleles at a single locus were encountered, and a mapping strategy was used to deal with multi-allelic loci. One example is the study of van Ooijen et al. (1994), in which a restriction fragment length polymorphism (RFLP) linkage map of L. peruvianum was constructed using three reciprocal backcross populations from an intra-specific cross between L. peruvianum LA2157 and LA2172. Multiple alleles were observed within the LA2172 accession, but only RFLP markers that yielded polymorphism between, but not within, the two parents were used. In this way, the multi-allelic loci were excluded. In the study of Ganal and Tanksley (1996), an F1 mapping population was generated by crossing two L. peruvianum plants. For some RFLP probes, more than two alleles were identified between the two parents. Only RFLP markers, which simultaneously segregated in both male and female gametes (Ritter et al. 1990), were included and scored dominantly. Via this strategy, the number of useful RFLP markers was limited, and the co-dominant information usually provided by RFLP markers was neglected.
Distorted marker segregation has been reported frequently in mapping studies of L. peruvianum, as is also reported in other inter- and intra-specific crosses within Lycopersicon (Haanstra et al. 1999; Kaloshian et al. 1998; Sandbrink et al. 1995; Van Heusden et al. 1999; Zamir and Tadmor 1986). In L. peruvianum, this may be caused by both self-incompatibility and unilateral incompatibility, resulting in preferential transmission of certain alleles. One gametophytic self-incompatibility locus in L. peruvianum has been mapped on chromosome 1 (Tanksley and Loaiza-Figueroa 1985). Three loci for the unilateral incompatibility in L. pennellii were identified on chromosomes 1, 6 and 10 (Chetelat and De Verna 1991).
All of these phenomena, crossing barriers, multi-alleles per locus and distorted segregation hamper genetic studies of L. peruvianum. In this study, we characterised the resistance to O. neolycopersici in L. peruvianum LA2172 by using different populations from an interspecific cross between L. esculentum cv. Moneymaker and L. peruvianum LA2172. Multi-allelic, single-locus markers were exploited to unfold the complexities such as multi-allelism and distorted segregation in a pseudo-F2 population. We illustrated the risk of exclusive use of bi-allelic markers.
Materials and methods
Plant and fungal materials
Hundreds of pollinations were made on plants of L. esculentum cv. Moneymaker (MM) with pollen from two plants of L. peruvianum LA2172 (LA2172), and only 11 F1 plants could be raised (Table 1). These F1 plants were self-incompatible, so pseudo-F2 populations were obtained by cross-pollinations between individual F1 plants. Finally, a pseudo-F2 population (hereafter referred to as the F2 population) of 194 plants, derived from a cross between two F1 plants (F1a and F1b) that originated from different individual LA2172 pollen parents, was produced for the present study (Fig. 1). Different backcross populations were generated by using MM as a recurrent parent. Two BC1 families (in total, 80 plants) of the two F1 plants (F1a and F1b) were used in the present study (Fig. 1).
Twenty informative F2 plants of the Ol-1 reference mapping population, which was derived from an interspecific cross of MM × L. hirsutum G1.1560 (Fig. 3b; Huang et al. 2000), were used to position molecular markers linked to the O. neolycopersici resistance in LA2172.
The pathogenic fungus O. neolycopersici, which originated from infected tomato plants (Lindhout et al. 1994), was maintained on MM plants in a growth chamber at a temperature of 21°C with 70% relative humidity (RH).
Disease test
The inoculum preparation and the inoculation were performed as described by Bai et al. (2003). The experimental setup was according to a randomized block design. For the disease test on the F2 population, six blocks were used, and each contained 32–33 F2 plants, three LA2172 plants as resistant control and five MM plants as susceptible control. The inoculated plants were grown in a greenhouse at 20±3°C with 30–70% RH. The plant was scored as resistant (no visible fungal sporulation) or susceptible (with fungal sporulation) at 14, 17 and 21 days post inoculation (dpi). For the disease tests on the BC1, one block contained 80 BC1 plants of the two BC1 families, 20 plants of each MM and LA2172 as susceptible and resistant control, respectively. The inoculated plants grew in a greenhouse at 22°C with 70% RH, and were evaluated as described above at 9 dpi and 12 dpi.
Molecular markers and linkage analysis
As described by Bai et al. (2003), total DNA was extracted from leaves of the F2 and BC1 plants, and the AFLP fingerprints were generated. Bulked segregant analysis (BSA) was performed for AFLP analysis on the resistant and susceptible pools, which were composed of equal volumes of AFLP pre-amplification products of eight resistant and eight susceptible F2 plants, respectively. AFLP markers were converted into cleaved amplified polymorphic sequence (CAPS) markers according to Brugmans et al. (2003). The CAPS marker Aps1 was generated as described by Bai et al. (2003).
The genetic linkage maps were constructed using the software package JoinMap, version 3.0 (van Ooijen and Voorrips 2001). A full account of linkage analysis, using marker data of the F2 population is given in the Appendix.
Results
The resistance to O. neolycopersici in LA2172
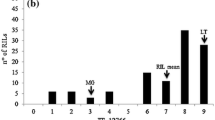
Fertile F1 hybrids of MM × LA2172 were produced despite the large genetic distance between these two species. The F1 progeny was not tested for resistance to O. neolycopersici, since it was very difficult to generate the F1 plants and the development of these F1 plants was very irregular. Instead, two BC1 families from two F1 plants (F1a and F1b) that were used for generating the F2 population (Fig. 1) were analyzed for resistance to O. neolycopersici. As shown in Table 2, all plants of the susceptible control MM showed fungal sporulation, and all LA2172 plants were immune (no symptoms). The BC1 plants could unambiguously be classified as either resistant or susceptible. One BC1 family (MM × F1a) was segregating for resistance (12 resistant:33 susceptible, Table 2), suggesting that the ancestor F1a plant was heterozygously resistant. Thus, resistance in LA2172 should be dominant, although the segregation in this BC1 family did not follow the 1:1 ratio of a monogenic model (χ2=8.889, P=0.003). All BC1 plants (n=35) of MM × F1b were susceptible, indicating that the F1b plant was homozygously susceptible. In the last 10 years, not a single LA2172 plant has been found to be susceptible to O. neolycopersici, despite extensive testing of this accession (data not shown). No susceptible LA2172 plants have been found so far; however, a susceptibility allele was transferred to the F1b plant, indicating that the resistance gene is heterozygously present in the LA2172 parent. To verify whether the resistance in LA2172 is monogenic, a large-scale disease test was performed on the F2 population derived from the cross of F1a × F1b. As shown in Table 2, 100 F2 plants were resistant and 94 were susceptible, which is in agreement with a segregation ratio of 1:1 for a monogenic model (χ2=0.1289, P=0.72). We therefore concluded that the resistance in LA2172 is governed by one dominant gene, designated as Ol-4.
Identification of markers linked to the Ol-4 locus
Since the F2 population was derived from a cross between two F1 plants (F1a × F1b) that originated from different LA2172 plants (Fig. 1), it might harbour different marker alleles at one locus. Assuming this can also be true for markers linked to the Ol-4 locus, the heterozygous resistant plant F1a has one marker allele from LA2172 (coded as ‘p’, linked to the resistance allele of the Ol-4 locus) and one marker allele from MM (coded as ‘e’). The homozygous susceptible plant F1b has one marker allele from LA2172 (coded as ‘p*’, linked to the susceptibility allele of the Ol-4 locus) and one marker allele ‘e’ from MM. Therefore, four marker genotypes (p/p*, e/p, e/p* and e/e) segregating at a 1:1:1:1 ratio were expected in the F2 population. To test for this hypothesis, multi-allelic, single-locus markers were required to differentiate the four marker genotypes in the F2 population.
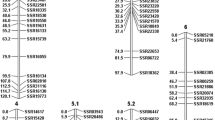
In order to efficiently identify the marker allele p linked in coupling phase to the Ol-4 locus, BSA was performed by using AFLP on the resistant and susceptible pools of the F2 plants (see Materials and methods). By using a total of 256 Pst/Mse primer combinations, 16 AFLP markers were identified that were present only in the resistant pool. Ten of these AFLP markers showed close co-segregation with Ol-4 (Fig. 3a) by testing them on 56 F2 individuals (20 resistant and 36 susceptible). To assign the AFLP markers to a particular chromosome, one AFLP marker (P18 M51–450) was successfully converted into a CAPS marker, designated By-4 (Table 3). By using the Ol-1 reference mapping population (see Materials and methods), By-4 was positioned on chromosome 6 between RFLP markers Aps1 and TG153 (Fig. 3b).
Map position of the Ol-4 locus
In order to verify whether Aps1 is linked to the Ol-4 locus, the RFLP marker Aps1 on chromosome 6 was converted into a CAPS marker (Table 3) that co-segregated with the original RFLP marker in the Ol-1 reference mapping population (see Materials and methods). The CAPS marker Aps1/TaqI revealed the expected multiple alleles in the F2 population. As shown in Fig. 2a, the e allele was from MM, two other alleles (p and p*) were from LA2172. By testing this marker on the F2 population, four marker phenotypes were observed (p/p*, e/p, e/p* and e/e, Table 4). Only the marker allele p was linked to the resistance allele of the Ol-4 locus. In addition, for the CAPS marker By-4 (Fig. 2b; Table 4), restriction enzyme HypCH4IV was specific for the p* allele (in equation p*≠p=e), and restriction enzyme ApoI was specific for the p allele (in equation p≠p*=e). Thus, when the results from these two restriction enzymes were combined, marker By-4 uncovered the four distinct marker genotypes of the F2 plants. The three marker alleles for CAPS markers Aps1 and By-4 were confirmed in the two BC1 families. The expected ratio for the four marker genotypes (p/p*, e/p, e/p* and e/e) in the F2 population was 1:1:1:1, while the observed frequencies were 74:15:80:17 for marker Aps1 and 73:16:80:17 for marker By-4, respectively (Table 4). For both CAPS markers By-4 and Aps1, distorted marker segregation was caused by preferential transmission of the L. peruvianum allele by the male parent (genotype e/p*, Appendix 3), with estimated transmission rates of about 18% and 82% for alleles e and p*, respectively.
Electrophoretic patterns of cleaved amplified polymorphic sequence (CAPS) markers (2% agarose gel) to show marker phenotypes in a F2 population from a cross of MM × LA2172. M DNA size marker of 1-kb ladder, lanes 1 and 2 resistant F2 plants, lanes 3 and 4 susceptible F2 plants. Lanes 1–4 show marker phenotypes of p/p*, e/p, e/p* and e/e, respectively; lane 5 MM with marker phenotype of e/e; lane 6 LA2172 (DNA is pooled from several plants) and shows marker phenotype of p/p*. a CAPS marker Aps1/TaqI (p≠p*≠e). The upper fragment in lane 6 (LA2172) is the p* allele and the lower fragment is the p allele; the single fragment in lane 5 (MM) is the e allele. b CAPS marker By-4. In lane 6 (LA2172) of the upper panel, HpyCH41V (p=e≠p*) reveals the upper fragment for p* allele; in lane 6 (LA2172) of the lower panel, ApoI (p≠p*=e) yields the upper fragment for p allele. c CAPS marker Aps1/Sau96I (p=p*≠e). The fragment in lane 6 (LA2172) is diagnostic for both p and p* alleles
All possible marker-allele configurations in the F2 population are shown in Table 4 and Fig. 2, and marker data for cross pollinator (CP) population type were prepared for JoinMap to construct a linkage map around the Ol-4 locus. As the JoinMap program assumes no distortion, parameters for both distortion and linkage between the Ol-4 and CAPS markers were also estimated using maximum likelihood estimations. A full account of the linkage analysis for various types of the CAPS markers is given in the Appendices. A linkage map was constructed for a part of tomato chromosome 6, where the Ol-4 locus was positioned above CAPS marker Aps1 (Fig. 3a). A comparison of the Ol-4 maps with the Ol-1 map showed that the order of the marker loci on the maps was identical. Therefore, we concluded that Ol-4 is on chromosome 6, but at a position different from the Ol-1 locus.
Genetic maps of part of chromosome 6 showing map positions of Ol-4 and Ol-1 loci, based on two different populations. a The Ol-4 locus map position in a F2 population derived from the cross of MM × LA2172, respectively. AFLP and CAPS markers were used. b Skeleton map showing positions of the Ol-1 locus and some restriction fragment length polymorphism markers on chromosome 6, which was retrieved from Huang et al. (2000)
Discussion
L. peruvianum is an out-crossing species and can harbour multi-alleles per locus. as illustrated in the present study and by others (Ganal and Tanksley 1996; Kaloshian et al. 1998; Miller and Tanksley 1990; van Ooijen et al. 1994). In several mapping studies, mapping strategies have been applied in order to deal with multi-allelic loci (Ganal and Tanksley 1996; van Ooijen et al. 1994). In the present study, it was illustrated that the information of multi-alleles at one locus can be fully exploited. Taking into account the high degree of heterogeneity in the donor parent L. peruvianum LA2172, we anticipated the possibility of more than two marker alleles per locus in the F2 population that was generated by mating F1 half-sibs. To test this hypothesis, multi-allelic CAPS markers for single loci were developed, which uncovered all marker-allele configurations (four marker genotypes, Fig. 2) in the F2 population. Consequently, markers could not be processed in the same way as a normal backcross or F2 population from inbreeding species, but rather as a population resulting from a cross between two heterogenously heterozygous diploid parents (coded as CP in JoinMap program).
The recognition of multi-alleles at a single locus in the F2 population by the multi-allelic markers has been of crucial importance. As shown in Table 4, CAPS markers uncovered two, three or four genotypes in the F2 population, depending on the restriction enzymes used (Fig. 2). For example, the CAPS marker Aps1/Sau96I could not differentiate between the alternative L. peruvianum alleles p and p* (Fig. 2c). Thus, this marker revealed only three marker phenotypes in the F2 population: homozygous as LA2172 (p/p*), homozygous as MM (e/e) and heterozygous (marker genotype of e/p and e/p*). F2 plants carrying the heterozygous marker phenotype segregated into classes: resistant (n=22) and susceptible (n=73) (Table 4). As explained in Appendix 4, incorrect estimates of recombination frequency between the Ol-4 locus and this marker could be obtained, if the distortion cannot unambiguously be attributed to either of the parents. Thus, the usage of bi-allelic markers of this type may complicate the mapping study because of the joint effect of linkage and preferential transmission of certain alleles. Similarly, bi-allelic markers like CAPS marker By-4/HypCH4IV (where p=e≠p*) were uninformative for mapping in this F2 population (Appendix 1), as this marker type uncovers two marker phenotypes (Fig. 2b; Table 4). One is heterozygous as LA2172 (marker genotypes of p/p* and e/p*) with 153 F2 plants (78 resistant and 75 susceptible), while the other is homozygous as MM (marker genotypes of e/p and e/e) with 33 F2 plants (15 resistant and 18 susceptible). Without discrimination of the p and e alleles, one would conclude that By-4/HypCH41V segregates independently from the Ol-4 gene. For a reliable genetic analysis, it is crucial to have markers that reveal different marker-allele configurations, like the multi-allelic, single-locus marker Aps1/TaqI that detected the three marker alleles (p, p* and e) and distinguished the four marker genotypes in the F2 population (Fig. 2a; Table 4).
By recognition of multi-alleles at a single locus in the F2 population, distorted segregation in the F2 population became uncovered that was caused by preferential transmission of L. peruvianum alleles by the male parent only. In this case, it was demonstrated that the selection acted against male gametes having L. esculentum alleles at the Ol-4 locus, which hardly influences the estimated recombination frequencies between the genetic loci (Appendix 3). In agreement with this, other authors (Ritter et al. 1990; van Ooijen et al. 1994) suggested that the estimated recombination frequency is unbiased when the segregation distortion is caused by selection at one locus per chromosome, but is biased when the selection acts on two (or more) loci on one chromosome.
In summary, we mapped the Ol-4 gene on the tomato chromosome 6, which governs complete resistance to O. neolycopersici and originates from L. peruvianum LA2172. It was illustrated that the hypothesis of close linkage between genetic markers can be erroneously rejected in case bi-allelic marker data are used for multi-allelic loci. The recognition of multiple alleles allowed accurate mapping of Ol-4 and revealed the large segregation distortion in the F2 population of MM × LA2172.
References
Ammiraju JSS, Veremis JC, Huang X, Roberts PA, Kaloshian I (2003) The heat-stable root-knot nematode resistance gene Mi-9 from Lycopersicon peruvianum is localized on the short arm of chromosome 6. Theor Appl Genet 106:478–484
Bai Y, Huang CC, Van der Hulst R, Meijer-Dekens F, Bonnema G, Lindhout P (2003) QTLs for tomato powdery mildew resistance (Oidium neolycopersici) in Lycopersicon parviflorum G1.1601 co-localize with two qualitative powdery mildew resistance genes. Mol Plant Microbe Interact 16:169–176
Baudry E, Kerdelhue C, Innan H, Stephan W (2001) Species and recombination effect on DNA variability in the tomato genus. Genetics 158:1725–1735
Brüggemann W, Linger P, Wenner A, Koornneef M (1996) Improvement of post-chilling photosynthesis in tomato by sexual hybridisation with a Lycopersicon peruvianum line from elevated altitude. Adv Hortic Sci 10:215–218
Brugmans B, Van der Hulst RGM, Visser RGF, Lindhout P, Van Eck HJ (2003) A rapid and reliable method for successful conversion of AFLP markers into simple single locus markers. Nucleic Acids Res 31:e53
Chetelat RT, De Verna JW (1991) Expression of unilateral incompatibility in pollen of Lycopersicon pennellii is determined by major loci on chromosomes 1, 6 and 10. Theor Appl Genet 81:704–712
Ganal MW, Tanksley SD (1996) Recombination around the Tm2a and Mi resistance genes in different crosses of Lycopersicon peruvianum. Theor Appl Genet 92:101–108
Haanstra JPW, Wye C, Verbakel H, Meijer-Dekens F, Van den Berg P, Odinot P, Van Heusden AW, Tanksley SD, Lindhout P, Peleman J (1999) An integrated high-density RFLP-AFLP map of tomato based on two Lycopersicon esculentum × L. pennellii F2 populations. Theor Appl Genet 99:254–271
Huang CC, Van de Putte PM, Haanstra-van der Meer JG, Meijer-Dekens F, Lindhout P (2000) Characterization and mapping of resistance to Oidium lycopersicum in two Lycopersicon hirsutum accessions: evidence for close linkage of two Ol-genes on chromosome 6. Heredity 85:511–520
Kaloshian I, Yaghoobi J, Liharska T, Hontelez J, Hanson D, Hogan P, Jesse T, Wijbrandi J, Simons G, Vos P, Zabel P, Williamson VM (1998) Genetic and physical location of the root-knot nematode resistance locus Mi in tomato. Mol Gen Genet 257:376–385
Kiss L, Cook RTA, Saenz GS, Cunnington JH, Takamatsu S, Pascoe I, Bardin M, Nicot PC, Sato Y, Rossman AY (2001) Identification of two powdery mildew fungi, Oidium neolycopersici sp. nov. and O. lycopersici, infecting tomato in different parts of the world. 2001. Mycol Res 105:684–697
Lindhout P, Pet G, van der Beek JG (1994) Screening wild Lycopersicon species for resistance to powdery mildew (Oidium lycopersicum). Euphytica 72:43–49
Miller JC, Tanksley SD (1990) RFLP analysis of phylogenetic relationships and genetic variation in the genus Lycopersicon. Theor Appl Genet 80:437–448
Ooijen JW van, Voorrips RE (2001) JoinMap 3.0, software for the calculation of genetic linkage maps. Plant Research International, Wageningen
Ooijen JW van, Sandbrink JM, Vrielink M, Verkerk R, Zabel P, Lindhout P (1994) An RFLP linkage map of Lycopersicon peruvianum. Theor Appl Genet 89:1007–1013
Poysa V (1990) The development of bridge lines for interspecific gene transfer between Lycopersicon esculentum and L. peruvianum. Theor Appl Genet 79:187–192
Rick CM (1982) Genetic relationships between self-incompatibility and flora traits in the tomato species. Biol Zentralbl 101:185–198
Ritter E, Gebhardt C, Salamini F (1990) Estimation of recombination frequencies and construction of RFLP linkage maps in plants from crosses between heterozygous parents. Genetics 125:645–654
Sandbrink JM, van Ooijen JW, Purimahua CC, Vrielink M, Verkerk R, Zabel P, Lindhout P (1995) Localization of genes for bacterial canker resistance in Lycopersicon peruvianum using RFLPs. Theor Appl Genet 90:444–450
Tanksley SD, Loaiza-Figueroa F (1985) Gametophytic self-incompatibility is controlled by a single major locus on chromosome 1 in Lycopersicon peruvianum. Proc Natl Acad Sci USA 82:5093–5096
Taylor IB (1986) Biosystematics of the tomato. In: Atherton JG, Rudich J (eds) The tomato crop—a scientific basis for improvement. Chapman and Hall, London, pp 1–34
Van der Beek JG, Pet G, Lindhout P (1994) Resistance to powdery mildew (Oidium lycopersicum) in Lycopersicon hirsutum is controlled by an incompletely dominant gene Ol-1 on chromosome 6. Theor Appl Genet 89:467–473
Van Heusden AW, Koornneef M, Voorrips RE, Brüggemann W, Pet P, Van Vrielink-Ginkel RV, Chen X, Lindhout P (1999) Three QTLs from Lycopersicon peruvianum confer a high level of resistance to Clavibacter michiganensis ssp. michiganensis. Theor Appl Genet 99:1068–1074
Veremis JC, Roberts PA (1996) Differentiation of Meloidogyne incognita and M. arenaria novel resistance phenotypes in Lycopersicon peruvianum and derived bridge-lines. Theor Appl Genet 93:960–967
Zamir D, Tadmor Y (1986) Unusual segregation of nuclear genes in plants. Bot Gaz 147:355–358
Acknowledgements
We thank Fien Meijer-Dekens and Thierry Marcel for assistance and the Unifarm of Wageningen University for nursing the plants. We thank Dr. Guusje Bonnema, Dr. Rients Niks and Dr. Sjaak van Heusden for critical reading of the manuscript and valuable comments. This work is sponsored by the Dutch Technology Foundation (STW, grant no. WBI 4835) and Dutch tomato-breeding companies.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by F. Salamini
Appendices
Appendix
Estimation of recombination frequency between Ol-4 and (multi-allelic) markers from the F2 population
Altogether, we used four types of markers (all possible marker-allele configurations in the F2 population), for each of which we discuss the linkage analysis below.
Appendix 1
Markers that distinguish between the alternative L. peruvianum alleles (p, p*), but only one of these alleles is distinct from the L. esculentum allele (p*≠p=e)
Marker By-4/HypCH4IV is of this type (Table 4; Fig. 2). The configuration of the cross between the two F1 genotypes (F1a and F1b, see Fig. 1) looks as follows:

Since neither of the parents is heterozygous at both loci, markers of this type are uninformative for linkage. In fact, the loci behave as if they segregate independently (see below). Not being aware of this non-informative nature, the joint segregation can be misinterpreted as ‘unlinked loci’.
Later on, we will have to introduce a parameter for preferential transmission of marker alleles. Let this parameter be β for allele transmission rate in the male parent (β=0.5 corresponds to a 1:1 Mendelian ratio). This leads to the following table of gamete combinations and their frequencies:
♀ | ♂ | |
|---|---|---|
β (ol-4 p*) | 1−β (ol-4 e) | |
(1/2) Ol-4 e | 1/2β | 1/2(1−β) |
(1/2) ol-4 e | 1/2β | 1/2(1−β) |
The following two-way table shows genotypes for the progeny (numbers in parentheses are the observed numbers for the marker By-4/HypCH4IV from Table 4).
| Ol-4/ol-4 | ol-4/ol-4 | Sum |
|---|---|---|---|
e/p* | 1/2β (78) | 1/2β (75) | β (153) |
e/e | 1/2(1−β) (15) | 1/2(1−β) (18) | (1−β) (33) |
Sum | 1/2 (93) | 1/2 (93) | 1 (186) |
Despite the distorted segregation, we indeed observe independent segregation between this marker and the Ol-4 locus. The distortion parameter β is estimated as \( \hat{\beta } = 153/186 = 0.8226. \)
Appendix 2
Markers that distinguish between the two alternative L. peruvianum alleles (p, p*), but only one of these is distinct from the L. esculentum allele (p≠p*=e)
This is the ‘mirror’ situation of (p*≠p=e), discussed above. Marker By-4/ApoI is of this type. The configuration of the cross looks like:

This represents the classical test cross configuration, from which recombination frequency (r), is readily estimated by counting the recombinant genotypes among the offspring. However, for the sake of completeness we introduce a parameter, α, for preferential transmission of marker alleles by the female parent (α=0.5 corresponds to a 1:1 ratio). This leads to the two-way table of genotype frequencies below (observed numbers are in parentheses, which are taken from Table 4 for marker By-4/ApoI).
| Ol-4/ol-4 | ol-4/ol-4 | Sum |
|---|---|---|---|
e/p | α(1−r) (85) | αr (4) | α (89) |
e/e | (1−α)r (8) | (1−α)(1−r) (89) | (1−α) (97) |
Sum | (93) | (93) | 1 (186) |
Notice that the boldfaced genotypes together occur with frequency r. So we estimate
We note that α is close to 0.5, indicating that no significant preferential transmission of alleles by the female parent occurs at this marker locus.
Appendix 3
Markers that distinguish between the two L. peruvianum alleles, as well as the L. esculentum allele (p≠p*≠e)
This represents the most informative class of markers that allow straightforward estimation of the two distortion parameters (α and β) as well as r. Marker By-4 and Aps1/TaqI are of this category. The configuration of the cross reads like:

Using the same parameter notation as above, we have the following table of gamete combinations
♀ | ♂ | |
|---|---|---|
β (ol-4 p*) | 1−β (ol-4 e) | |
α(1−r) (Ol-4 p) | α β(1−r) | α(1−β)(1−r) |
(1−α)r (Ol-4 e) | (1−α)βr | (1−α)(1−β)r |
αr (ol-4 p) | αβr | α(1−β)r |
(1−α)(1−r) (ol-4 e) | (1−α)β(1−r) | (1−α)(1−β)(1−r) |
Thus, the two-way table of genotypes reads:
| Ol-4/ol-4 | ol-4/ol-4 | Sum |
|---|---|---|---|
p/p* | αβ(1−r) | α β r | αβ |
e/p* | (1−α)βr | (1−α)β(1−r) | (1−α)β |
e/p | α(1−β)(1−r) | α(1−β)r | α(1−β) |
e/e | (1−α)(1−β)r | (1−α)(1−β)(1−r) | (1−α)(1−β) |
Notice that the boldfaced genotypes together represent a proportion, r (independent of α, β). So r is estimated by counting these genotypes. Likewise, α and β are estimated by adding the appropriate classes in the right margin of the above table (pp*+ep for α, pp*+ep* for β). Using the numbers given in Table 4 we obtain for
We observe that the estimates for α and β are the same as the ones obtained for markers By-4/HypCH4IV and By-4/ApoI, which is not surprising, since they represented the same locus (By-4).
Appendix 4
Markers that distinguish between the L. peruvianum and L. esculentum alleles, but not between the two L. peruvianum alleles (p*=p≠e)
Marker Aps1/Sau96I is of this type. The configuration of the cross reads like:

From this we see that, considering the marker locus only, any segregation distortion at the marker cannot be ascribed to either parent or to both parents. In other words, estimation of the distortion parameters must go along with estimation of r, and vice versa. In this sense, this marker category represents the most ‘difficult’ one: it requires simultaneous estimation of α, β and r.
Proceeding as before, using α and β for female and male transmission frequencies at the marker and r for recombination frequency, we obtain the following two-way table of genotypes (numbers in parentheses are observed numbers for marker Aps1/Sau96I in Table 4).
| Ol-4/ol-4 | ol-4/ol-4 | Sum |
|---|---|---|---|
p/p | αβ(1−r) (71) | αβr (3) | αβ (74) |
e/p | α(1−β)(1−r)+(1−α)βr (22) | α(1−β)r+(1−α)β(1−r) (73) | α(1−β)+(1−α)β (95) |
e/e | (1−α)(1−β)r (0) | (1−α)(1−β)(1−r) (17) | (1−α)(1−β) (17) |
Sum | (93) | (93) | 1 (186) |
For the simultaneous estimation of α, β, and r, we proceed as follows. As can be seen from the table above, α and β can be estimated from the observed frequencies at the marker locus (probabilities in the right margin). We also see that for the marker genotype frequencies, α and β are interchangeable, which means that, in case α and β are not equal, we cannot ascribe the estimate to either parent. However, we observe a 1:1 ratio at the Ol-4 locus, which means that for a marker closely linked to Ol-4, the value of α must be close to 0.5 (α is the female transmission rate: a clear deviation from 0.5 would ‘drag’ along the alleles at Ol-4). Estimates of α and β are obtained by solving the appropriate likelihood equations, using probabilities and observed frequencies at the marker locus. The resulting quadratic equation yields two equivalent solutions, i.e.
We accept the first one, since α should be close to 0.5; it also is in close agreement with the estimates obtained for the other markers. Next, we substitute (α, β)=(0.483,0.823) in the expressions for the probabilities in the body of the table and (numerically) solve the resulting likelihood equation for r. This yields \(\hat{r} = 0.0394.\) However, should we have used the ‘mirror’ estimates of\((\hat{\alpha },\hat{\beta }),\) i.e.\((\hat{\alpha },\hat{\beta }) = (0.823,0.483),\) the incorrect estimate of r would have been\(\hat{r} = 0.287.\) This, again, shows the necessity of carefully interpreting the joint segregation data in order to avoid wrong conclusions.
Using the obtained estimates, we have calculated the corresponding LOD values for linkage. The table below summarizes the results.
Marker | \({\hat \alpha }\) | \({\hat \beta }\) | \({\hat r}\) | LOD |
|---|---|---|---|---|
Aps1/TaqI | 0.479 | 0.828 | 0.054 | 39.1 |
Aps1/Sau96I | 0.483 | 0.823 | 0.039 | 27.7 |
By-4 | 0.479 | 0.823 | 0.065 | 36.7 |
By-4/ApoI | 0.479 | – | 0.065 | 36.7 |
By-4/HypCH4IV | – | 0.823 | – | – |
Rights and permissions
About this article
Cite this article
Bai, Y., van der Hulst, R., Huang, C.C. et al. Mapping Ol-4, a gene conferring resistance to Oidium neolycopersici and originating from Lycopersicon peruvianum LA2172, requires multi-allelic, single-locus markers. Theor Appl Genet 109, 1215–1223 (2004). https://doi.org/10.1007/s00122-004-1698-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-004-1698-5