Abstract
The Limnanthaceae (Order Brassicales) is a family of 18 taxa of Limnanthes (meadowfoam) native to California, Oregon, and British Columbia. Cultivated meadowfoam (L. alba Benth.), a recently domesticated plant, has been the focus of research and development as an industrial oilseed for three decades. The goal of the present research was to develop several hundred simple sequence repeat (SSR) markers for genetic mapping, molecular breeding, and genomics research in wild and cultivated meadowfoam taxa. We developed 389 SSR markers for cultivated meadowfoam by isolating and sequencing 1,596 clones from L. alba genomic DNA libraries enriched for AG n or AC n repeats, identifying one or more unique SSRs in 696 clone sequences, and designing and testing primers for 624 unique SSRs. The SSR markers were screened for cross- taxa utility and polymorphisms among ten of 17 taxa in the Limnanthaceae; 373 of these markers were polymorphic and 106 amplified loci from every taxon. Cross-taxa amplification percentages ranged from 37.3% in L. douglasii ssp. rosea (145/389) to 85.6% in L. montana (333/389). The SSR markers amplified 4,160 unique bands from 14 genotypes sampled from ten taxa (10.7 unique bands per SSR marker), of which 972 were genotype-specific. Mean and maximum haplotype heterozygosities were 0.71 and 0.90, respectively, among six L. alba genotypes and 0.63 and 0.93, respectively, among 14 genotypes (ten taxa). The SSR markers supply a critical mass of high-throughput DNA markers for biological and agricultural research across the Limnanthaceae and open the way to the development of a genetic linkage map for meadowfoam (x = 5).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Limanthaceae (Order Brassicales) is a family of 18 species and subspecies of Limnanthes native to Pacific Western North America (Mason 1952; Ornduff and Carvello 1968; Ornduff 1971) and one species, Floerkea proserpinacoides (false mermaid), native to the northeastern United States and Canada (Houle et al. 2000). Wild Limnanthes (meadowfoam) populations fluorish in temporally wet habitats (vernal pools) west of the Sierra Nevada mountain range in California and the Cascade mountain range in Oregon and British Columbia (Mason 1952). Several taxa in the family are threatened or endangered and native habitats are rapidly disappearing, particularly in urban growth boundaries. Twelve taxa in the family (L. bakeri, L. douglasii ssp. sulphurea, L. floccosa ssp. bellingeriana, L. floccosa ssp. californica, L. floccosa ssp. grandiflora, L. floccosa ssp. pumila, L. gracilis ssp. gracilis, L. gracilis ssp. parishii, L. macounii, L. montana, L. striata, and L. vinculans) are listed as rare, vulnerable, or endangered (Eastman 1990; Dole and Sun 1992; Walter and Gillett 1998). The greatest threat to the family is human activity, particularly agricultural and urban development. Cultivated meadowfoam (L. alba ssp. alba and L. alba ssp. versicolor) and three subspecies of L. douglasii do not seem to be seriously threatened or endangered (Walter and Gillett 1998). Nevertheless, several wild populations of L. alba, a native of California (Mason 1952), are known to be threatened (Knapp and Crane 1998).
Cultivated meadowfoam has been the focus of research and development as an industrial oilseed for three decades, particularly the last (Knapp and Crane 1998). The seed oil of meadowfoam is a rich source of novel very long-chain fatty acids (C20 and C22) (Smith et al. 1960; Bagby et al. 1961). The oil has unusually high concentrations of Δ5 unsaturated (typically 86%) and C20 and C22 (typically 96%) fatty acids and extraordinarily low concentrations of saturated fatty acids (typically less than 2%) (Knapp and Crane 1995). The novelty of the oil has been the primary factor behind the development of the meadowfoam oil industry and, more recently, the development of novel chemical derivatives for industrial markets (Isbell 1997).
The prospect for developing meadowfoam into an economically significant oilseed crop hinges on the development of genetically superior cultivars, and on conserving genetic diversity in the Limnanthaceae. Meadowfoam germplasm has not been collected from the wild on a significant scale for more than 20 years, and a recent census of wild populations has not been performed (Knapp and Crane 1998). The early wild collections and a few domesticated germplasm accessions are maintained by the United States Department of Agriculture National Plant Germplasm System (http://www.ars-grin.gov/npgs), and have supplied genetic diversity crucial for meadowfoam cultivar development and, consequently, for the development of the meadowfoam oil industry. Genetic diversity for several biologically and agriculturally important traits has been surveyed and cataloged in cultivated meadowfoam (Knapp and Crane 1995, 1997, 1998); however, molecular genetic diversity has not been surveyed, essentially because high-throughput, meadowfoam-specific DNA markers have not been developed.
Simple sequence repeats (SSRs) are logical targets for DNA marker development in species lacking high-throughput, sequence-tagged-site (STS) DNA markers (Weber and May 1989; Morgante and Olivieri 1993; Rafalski and Tingey 1993; Powell et al. 1996). SSRs can be rapidly isolated in significant numbers from SSR-enriched genomic DNA libraries (Rae et al. 2000; Ramsay et al. 2000; Tang et al. 2002), are often highly polymorphic, and can be rapidly and efficiently assayed. While our primary goal was to develop SSR markers for cultivated meadowfoam (L. alba ssp. alba and L. alba ssp. versicolor), the utility and polymorphisms of cultivated meadowfoam SSR markers across wild meadowfoam taxa was assessed. We developed 389 SSR markers for cultivated meadowfoam and identified 106 conserved SSR markers for the Limnanthaceae. The cross-taxa utility and polymorphisms of the SSR markers are described herein.
Materials and methods
Genomic DNA library development and screening
Genomic DNA was isolated from OMF40–11 (L. alba ssp. alba) and OMF64 (L. alba ssp. versicolor) (Crane and Knapp 2000) leaf tissue as described by Lodhi et al. (1994). The two DNA samples were pooled for library construction. Genomic DNA libraries enriched for AG n , AC n , AAT n , or ATC n repeats, where n is the number of repeats, were constructed by Genetic Identification Services (Chatsworth, Calif., USA), essentially as described by Karagyozov et al. (1993) and Edwards et al. (1996). DNA fragments in the 350 –800 bp range were cloned into pUC19 plasmids. The latter were transformed into the Escherichia. coli strain DH5α (yields ranged from 5,000 to 10,000 recombinant cells/1.4 ml).
Fifty to 100 μl of the original stock solutions of each of the four genomic DNA libraries were spread on LB-agar media with 75 μg/ml of ampicillin. Recombinant clones were isolated using the IPTG and X-Gal procedures. White colonies were transferred to ampicillin-LB agar medium in 96-well format plates. The plates were incubated at 37°C overnight. Clone scrapes were used as templates for colony PCRs. Meadowfoam genomic DNA inserts were amplified using universal M13 forward (5′-CGCCAGGGTTTTCCCAGTCACGAC-3′) and reverse (5′- TCACACAGGAAACAGCTATGAC-3′) primers. PCR reactions were performed with Perkin Elmer 9600 or MJ PTC 200 thermocyclers using reaction mixtures in a total volume of 35 μl containing 1x PCR buffer, 2 mM Mg2+, 0.2 μM each of dNTPs, 1% Tween 20, 1.5 mM Cresol Red, 3% sucrose, 0.5 μM per primer, and 0.75 U Taq polymerase (Qiagen, Valencia, California, USA). PCRs were performed by denaturing at 95°C for 3 min, followed by 35 cycles of 94°C for 30 s, 57°C for 30 s, 72°C for 1 min, and a final 10 min. extension. We estimated insert lengths on 1.5% agarose gels and selected inserts in the 350 – 800 bp range for DNA sequencing and SSR marker development. The selected PCR products were purified using QIAquick PCR purification kits. Half of the purifications were processed on a Qiagen BioRobot 3000. DNA sequencing was performed using 1.5–3.0 pmol of the M13 forward primer and 25 – 50 ng of purified PCR products on ABI 377 or 3700 automated sequencers (Perkin Elmer, Foster City, Calif., USA). Dimethyl sulfoxide (5%) was added as denaturant to increase DNA sequence quality, especially for clones harboring long repeats.
DNA sequence analyses and SSR marker development
DNA sequences were searched for the presence of dinucleotide, trinucleotide, and tetranucleotide repeat motifs. SSRs with five or more repeat units were selected for further analysis and SSR marker development. Sequence analyses were performed using the SEQLAB module of the Genetics Computing Group (GCG) Software (Madison, Wis., USA). Sequences harboring common repeat motifs were trimmed to remove the plasmid sequence and checked for redundancy using the GCG PILEUP function. Unique sequences were divided into perfect, imperfect, and compound repeat classes according to Weber (1990).
PRIMER 3.0 (http://www.genome.wi.mit.edu/cgi-bin/primer/primer3_www.cgi) was used to design primers complementary to DNA sequences flanking the SSRs. DNA sequences for reference alleles and primers for the SSR markers have been deposited in GenBank (http://www.ncbi.nlm.gov); BV007038-BV007426. We designed primers in the 20 – 27 bp range with G-C contents ranging from 35 to 60% and annealing temperatures (T m) ranging from 59 to 63°C. The other parameter settings for primer design were a T m variance of 2.0oC, maximum 3′ stability of 9.0, maximum self-complementarity of 6.0, maximum 3′ self-complementarity of 2.0, and max ploy-X of 4.0. To facilitate multiplexing, predicted PCR product lengths were systematically varied from 108 to 554 bp and the 5′ ends of forward primers were labeled with one of three fluorophores (6-FAM, HEX, or TET) in combinations to minimize overlap in multiplexed genotyping assays. Primers were produced and supplied by MWG Biotech (High Point, N.C., USA).
The SSR markers were screened for polymorphisms among 14 L. alba, L. floccosa, L. gracilis, L. montana, and L. douglasii germplasm accessions (Table 1). We screened four inbred lines of L. alba (OMF64, OMF40–11, LE76, and OMF109–2) and ten randomly selected individuals from open-pollinated populations of L. alba (Wheeler and OMF156), L. flocossa ssp. flocossa (LFF), L. flocossa ssp. grandiflora (LFG), L. gracilis ssp. gracilis (LGG), L. gracilis ssp. parishii (LGP), L. montana (LM), L. douglasii ssp. douglasii (LDD), L. douglasii ssp. nivea (LDN), and L. douglasii ssp. rosea (LDR) (Table 1) (http://www.ars-grin.gov/npgs; Crane and Knapp 2000, 2002). DNA samples for the inbred lines were produced from leaf samples collected from ten greenhouse grown plants per inbred, whereas DNA samples for open-pollinated populations were produced from leaf samples collected from a single greenhouse grown plant per population. Genomic DNA was isolated from the leaf tissue as described by Lodhi et al. (1994). PCRs were performed in a 20 μl volume containing 1x PCR buffer, 2.5 mM Mg2+, 0.2 μM each of dNTPs, 0.1% Tween 20, 5 – 7.5 pmol of each primer, 0.75 U Taq polymerase, and 15 – 20 ng of meadowfoam genomic DNA. We used a ‘touchdown’ PCR protocol (Don et al. 1991) to minimize non-specific PCR amplification. PCRs were performed using an initial denaturation step at 95°C for 3 min, followed by one cycle of 94°C for 30 s, 68°C for 30 s, and 72°C for 1 min. Subsequent cycles were performed by decreasing the annealing temperature by 1°C to a base temperature of 53 – 58°C (annealing temperatures varied from 53° to 58°C as a function of the T m of each primer pair). The annealing temperature for each SSR primer pair is listed in the SSR primer database supplied as supplemental data. Finally, PCRs were continued for 30 cycles at 94°C for 30 s, 58°C for 30 s, 72°C for 1 min, and a final 20 min extension.
SSR genotyping assays were performed on sequencing gels run on an ABI 377 equipped with the GeneScan and Genotyper software and filter set C (Applied Biosystems Incorporated, Foster City, Calif.) by post-PCR multiplexing three SSR markers per lane labeled with different fluorophores. The pooled amplicons were diluted 20-fold. Samples containing 0.5 μl of the diluted PCR products, 0.2 μl GeneScan 500 internal lane standard labeled with TAMRA, and 50% formamide were heated to 92°C for 5 min and chilled on ice for 5 min before gel loading. SSR allele calling was performed using GeneScan and allele lengths were estimated using Genotyper. We manually checked the output and corrected allele lengths as necessary.
Statistical analyses
Because more than half of the SSR markers amplified multiple loci and the allelism of bands amplified by multilocus SSR markers could not be ascertained, we calculated the number of alleles per locus, heterozygosities (H), and other statistics from a subset of 90 single-locus SSR markers, in addition to calculating the number of bands per marker and haplotype heterozygosities (H h ) from the genotypes of the complete set of 389 single-locus and multilocus SSR markers. H was estimated from SSR allele frequencies as described by Ott (1991), whereas H h was estimated by substituting haplotype frequencies for allele frequencies in H, where each haplotype is a unique SSR banding pattern. Hence, if each of 14 genotypes have unique haplotypes, then H h = 1 - [14 x (1/14)2] = 0.929. H estimates the probability of observing an allelic polymorphism between two individuals (genotypes) drawn at random, whereas H h estimates the probability of observing a minimum of one allelic polymorphism between two genotypes drawn at random. We estimated the number of loci amplified by each SSR marker from the genotypes of two F6 lines developed by single-seed descent (OMF40–11 and OMF64). Statistics were estimated using PROC FREQ, GLM, and CORR of the Statistical Analysis System (http://www.sas.com).
Results
SSR marker development
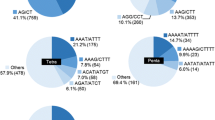
Thirty-six clones from the AG n , AC n , AAT n , and ATC n -enriched genomic DNA libraries, where n is the number of repeat units, were sequenced to check for SSR enrichment. Nine clones from each library were analyzed. SSRs were not found in DNA sequences from the AAT n and ATC n libraries, whereas SSRs were found in ten of the 18 DNA sequences from the AG n and AC n libraries, equally split between the two. We performed colony PCR on 3,264 recombinant clones from the AG n and AC n -enriched genomic DNA libraries and selected 1,596 clones for sequencing, 817 from the AC n and 779 from the AG n library. The selected clones had inserts ranging in length from 350 to 800 bp. Colony PCR products from the selected clones were purified, sequenced, and screened for all possible dinucleotide, trinucleotide, and tetranucleotide repeats. SSRs with a minimum of five repeat units (n>5) were identified in 1,237 clone sequences, 620 from the AC n and 617 from the AG n library. DNA sequences harboring common motifs were aligned and screened for redundancy. We identified 541 redundant and 696 unique SSR sequences, 281 from the AC n and 415 from the AG n library. The total and unique SSR enrichment percentages were 77.5% and 43.6%, respectively. The most common repeat motifs found in the selected sequences were AG n (328) and AC n (136), the repeats targeted for enrichment; however, 59 different perfect, imperfect, and compound repeat motifs were isolated and are listed in the SSR primer database (supplied as supplemental data).
We designed and tested primers for 624 unique SSRs (544 dinucleotide, 75 trinucleotide, and five tetranucleotide repeats), but could not design primers for the other 72 unique SSRs because of short flanking sequences. Of the 624 SSR primers, 389 amplified bands of the lengths predicted from OMF40–11 or OMF64 reference allele sequences (110 from the AC n and 279 from the AG n library), 24 produced multiple bands, and 211 failed to amplify bands from any of the 14 genotypes. Hence, 389 SSR markers were developed for cultivated meadowfoam from a starting point of 1,596 genomic DNA clone sequences and 696 unique SSR sequences. Fifteen of the SSR markers failed to amplify bands from OMF40–11 (L. alba ssp. alba) (Fig. 1).
Cross-taxa amplification percentages for 389 SSR markers genotyped in Limnanthes alba ssp. alba (LAA), L. alba ssp. versicolor (LAV), L. floccosa ssp. floccosa (LFF), L. floccosa ssp. grandiflora (LFG), L. gracilis ssp. gracilis (LGG), L. gracilis ssp. parishii (LGP), L. montana (LM), L. douglasii ssp. douglasii (LDD), L. douglasii ssp. nivea (LDN), and L. douglasii ssp. rosea (LDR)
Cross-taxa utility and polymorphisms
The SSR markers were screened for cross-taxa utility and intra- and interspecific polymorphisms among 14 genotypes from ten taxa (Table 1). Of 389 SSR markers, 106 amplified bands from every taxon, whereas 283 failed to amplify bands from one or more taxa (Fig. 1). Cross-taxa amplification percentages were greater for section Inflexae (82.0%) than section Reflexae (61.4%) taxa; however, bands were amplified from two out of three section Reflexae taxa (L. douglasii ssp. douglasii and L. douglasii ssp. nivea) as often as from wild section Inflexae taxa. The lowest cross-taxa amplification percentage was observed for L. douglasii ssp. rosea (37.3%). The cross-taxa amplification percentages for taxa other than L. douglasii ssp. rosea ranged from 68.1% for L. floccosa ssp. grandiflora to 85.6% for L. montana (Fig. 1). The utility of individual SSR markers within and between taxa can be ascertained from the SSR primer and polymorphism databases supplied as supplemental data.
One to 28 unique bands were amplified by each SSR marker among the 14 genotypes (Fig. 2). Sixteen of the SSR markers (4.1%) were monomorphic. One SSR marker produced two unique bands per genotype (14 unique haplotypes). The number of loci amplified from L. alba was estimated from the number of bands amplified from two inbred lines (OMF40–11 and OMF64); 171 SSR markers amplified a single locus (single band) and 218 SSR markers amplified two or more loci (two or more bands). Of the former, 90 amplified one or two bands from non-inbred, non-reference L. alba genotypes and were selected for calculating allele numbers, heterozygosities, and other single-locus SSR marker statistics (Table 2).
The number of loci amplified from OMF40–11 and OMF64 was estimated to be 421–431, respectively; 374 SSR markers amplified loci from OMF40–11, whereas 389 SSR markers amplified loci from OMF64. The number of bands amplified from non-reference L. alba genotypes ranged from 276 for LE76 to 347 for OMF156. SSR amplification rates dropped off for wild taxa in both sections (Fig. 1). The mean number of bands amplified from different germplasm or taxonomic groups were 353.3 for the six L. alba genotypes, 317.8 for the four non- reference L. alba genotypes, 267.8 for wild section Inflexae taxa (LFF, LFG, LGP, LGG, and LM), and 232.7 for wild section Reflexae taxa (LDD, LDN, and LDR). SSR markers failed to amplify bands from 0.74 out of six L. alba genotypes and 1.37 out of 14 genotypes across the ten taxa.
Of the 4,160 unique bands amplifed by the SSR markers from the 14 genotypes (10.7 unique bands per SSR marker), 972 were genotype-specific. We identified 410 genotype-specific bands among the six L. alba genotypes, 52 from LE76 to 130 from OMF40–11. The number of genotype-specific SSR bands amplified from wild meadowfoam taxa ranged from 37 for L. douglasii ssp. rosea to 89 for L. gracilis ssp. parishii. The number of genotype-specific bands was positively correlated (0.71) with the number of bands amplified from each genotype (the sum of shared and genotype-specific bands).
Ninety-six percent of the SSR markers (373 out of 389) were polymorphic, whereas 100% of the SSR markers in the selected single-locus subset (90 out of 389) were polymorphic (Fig. 3). Mean and maximum heterozygosities for the single-locus subset were 0.45 and 0.80, respectively, among the six L. alba germplasm accessions and 0.49 and 0.80, respectively, among the 14 genotypes (Fig. 3, Table 2). Similarly, mean and maximum H h for the complete set of SSR markers were 0.71 and 0.90, respectively, among the six L. alba germplasm accessions and 0.63 and 0.93, respectively, among the 14 genotypes. The H h mean was lower across taxa than within L. alba because null allele frequencies were greater for the former than the latter (thereby reducing allelic diversity) (Table 2). By contrast, the H mean was slightly greater across taxa than within L. alba because null allele frequencies were greatly reduced in the single-locus SSR marker subset.
The H distribution was platykurtic and approximately normal, whereas the H h distribution was right-skewed (Fig. 3). The skewness was produced by hypervariable, multilocus SSR markers. The heterozygosities for single-locus and haplotype heterozygosities for multilocus SSR markers are listed in the SSR primer database (supplied as supplemental data). Haplotype heterozygosities were calculated to supply a complete summary of the data and show that a many of the more complex, multilocus SSR markers were hypervariable. The probability of observing a minimum of one allelic difference between two individuals (H h ) was greater than 0.80 for 150 multilocus SSR markers (Fig. 3). Slightly more than half of the SSR markers (202/389) had haplotype heterozygosities ranging from 0.70 to 0.93 and 60% of the SSR markers uniquely identified a minimum of one out of 14 genotypes. Moreover, individual SSR markers uniquely identified up to 11 genotypes, e.g., LS196 distinguished 11 out of 14 and LS356 distinguished 10 out of 14 genotypes. When combined, LS196 and LS356 distinguished 14 out of 14 genotypes.
Discussion
The SSR markers described herein supply a critical mass of hypervariable, high-throughput DNA markers for biological and agricultural research in the Limnanthaceae, and are the first DNA markers developed specifically for meadowfoam. Heretofore, 18 allozyme markers had been described for meadowfoam (Brown and Jain 1979; Kesseli and Jain 1985; McNeill and Jain 1985; Ritland and Jain 1984). The meadowfoam SSR markers were designed for multiplex genotyping by systematically varying SSR allele lengths. Typically, 6–14 meadowfoam SSR markers can be multiplexed when the allele lengths of the genotypes are known a priori. The multiplexing density depends on the allelic diversity of the genotypes sampled, the genotyping or assay method, and number of loci amplified by the individual SSR markers. Because the allele lengths were not known in the initial screening, we multiplexed three SSR markers, each labelled with a different fluorophore, so that the bands amplified by each SSR marker could be unequivocally identified. The SSR allele length database has been supplied as supplemental data for planning new experiments and selecting SSR markers for multiplex genotyping.
The meadowfoam SSR markers were found to be extraordinarily polymorphic (Fig. 3, Table 2). Most importantly, 373 (96.1%) were polymorphic in L. alba, the domesticated species targeted for commercial development and genetic mapping (Knapp and Crane 1998; Katengam et al. 2002). The range of heterozygosities we observed were similar to the ranges reported for SSR markers isolated from unenriched and SSR-enriched genomic DNA libraries in other plant taxa (Akkaya et al. 1992; Morgante and Olivieri 1993; Wu and Tanksley 1993; Saghai Maroof et al. 1994; Kresovich et al. 1995; Plaschke et al. 1995; Röder et al. 1995; Rongwen et al. 1995; Powell et al. 1996; Taramino and Tingey 1996; Innan et al. 1997; Olufowote et al. 1997; Smulders et al. 1997; Loridon et al. 1998; Cho et al. 2000; Smith et al. 1997, 2000; Temnykh et al. 2000; Yu et al. 2002; Tang and Knapp 2003). Because wild and domesticated genotypes were sampled (Table 1), the haplotype heterozygosity distribution was right-skewed; nearly two-thirds of the SSR markers had haplotype heterozygosities greater than 0.6 (Fig. 3). By contrast, slightly less than one-third of the single-locus SSR markers (29/90) had heterozygosities greater than 0.6. H and H h was greater than the probability of observing a polymorphism between any of the loci amplified by a multilocus SSR marker, which was greater than the probability of observing a polymorphism between the locus amplified by a single-locus SSR marker. Because cultivated meadowfoam has not yet passed through severe domestication or breeding bottlenecks (Knapp and Crane 1998), SSRs seem to be as polymorphic among cultivated meadowfoam germplasm accessions as among wild taxa (Table 2).
Despite a propensity for redundantly amplifying DNA sequences, SSR-enrichment procedures yield genomic DNA libraries greatly enriched for SSR sequences and facilitate the development of SSR markers on a large scale (Ostrander et al. 1992; Karagyozov et al. 1993; Edwards et al. 1996; Rae et al. 2000; Ramsay et al. 2000; Tang et al. 2002). The meadowfoam genomic DNA libraries were greatly enriched for SSR sequences (SSRs were found in 77.5% of the genomic DNA clones); however, 43.7% of the SSR sequences were redundant (541/1,237), primers could not be designed for 10.3% of the unique SSR sequences (72/696), and 37.6% of the primers failed to amplify SSR alleles from any of the genotypes (235/624), primarily because of DNA sequence quality. Of the original 1,596 meadowfoam DNA sequences we produced, 6.3% yielded SSR markers that amplified loci from every taxon, whereas 25.3% yielded SSR markers that amplified loci from L. alba. Similarly, of 696 unique meadowfoam SSR sequences, 15.2% yielded SSR markers that amplified loci from all of the taxa, 20.8 to 47.8% yielded SSR markers that amplified loci from taxa other than L. alba, and 55.9% yielded SSR markers that amplified loci from L. alba. Hence, aside from DNA sequence redundancy, the attrition we observed was primarily caused by SSR primer failures, a common problem in SSR marker development (Kresovich et al. 1995; Liu et al. 1995; Plaschke et al. 1995; Röder et al. 1995; Rongwen et al. 1995; Szewc-Mcfadden et al. 1996; Olufowote et al. 1997; Kubik et al. 1999, Ramsay et al. 1999; Saal and Wricke et al. 1999; Cho et al. 2000; Temnykh et al. 2000; Tang et al. 2002; Yu et al. 2002).
Slightly more than one-fourth of the Limnanthes SSR markers (106/389) were conserved across the ten taxa (Figs. 1, 3). Predictably, fewer amplified alleles (bands) from every taxon (106) than from each of the wild taxa (145 to 333). While cross-taxa amplification rates generally decreased as taxonomic differences increased, null allele frequencies were greater for two non-reference L. alba genotypes (Wheeler and LE76) than for two wild species from section Inflexae (L. gracilis ssp. gracilis and L. montana) and one wild species from section Reflexae (L. douglasii ssp. douglasii) (Fig. 1). The failure of SSR markers to amplify loci, both within L. alba and across wild meadowfoam taxa, was undoubtedly caused by polymorphisms in the DNA sequences targeted by the SSR primers (Mogg et al. 2002). BLAST analyses of the Limnanthes DNA sequences identified no known gene sequences; hence, the SSRs undoubtedly reside in mutation-rich retrotransposon and dispersed repetitive element DNA (Ramsay et al. 1999, 2000).
SSRs generally seem to be less conserved among plants than animals (Schlötterer et al. 1991; Fitzsimmons et al. 1995; Garza et al. 1995; Brown et al. 1996; Garza and Freimer 1996; Primmer et al. 1996; Rico et al. 1996; Whitton et al. 1997; Peakall et al. 1998; Reed et al. 2000; Smulders et al. 2000; Cordiero et al. 2001; Kutil and Williams 2001). Schlötterer et al. (1991), Fitzsimmons et al. (1995), and Rico et al. (1996) identified SSRs that amplified loci across fish, turtle, or whale taxa sharing common ancestors 450, 300, or 40 million years before the present, respectively. Similarly, half of the SSRs tested by Primmer et al. (1996) amplified loci from birds sharing common ancestors ten million years before the present. SSRs frequently amplify loci from closely related plant taxa, but seldom from diverse plant taxa (Röder et al. 1995; Brown et al. 1996; Provan 1996; Whitton et al. 1997; Peakall et al. 1998; Smulders 2000; Kutil and Williams 2001). The interspecific SSR amplification rates we observed in Limnanthes (37.3–85.6%) (Fig. 1) were similar to interspecific SSR amplification rates in Glycine (48.4–61.3%) (Peakall et al. 1998) and intergeneric SSR amplification rates between tomato (Lycopersicon esculentum L.) and potato (Solanum tuberosum L.) (Provan et al. 1996; Smulders et al. 1997), among wheat (Triticum aestivum L.), barley (Hordeum vulgare L.), and rye (Secale cereale L.) (Röder et al. 1995), and between sorghum [Sorghum bicolor (L.) Moench.] and maize (Zea mays L.) (Brown et al. 1996).
We could not locate seed of and hence could not screen 9 of 19 taxa in the Limnanthaceae, specifically F. proserpinacoides and eight meadowfoam taxa, four from section Inflexae (L. bakeri, L. floccosa ssp. bellingeriana, L. flocossa ssp. californica, and L. flocossa ssp. pumila) and four from section Reflexae (L. douglasii ssp. sulphurea, L. macounii, L. striata, and L. vinculans). Because the tested and untested meadowfoam taxa are closely related (Mason 1952; Ornduff and Crovello 1968; Ornduff 1971) and 145 to 279 of the cultivated meadowfoam SSR markers amplified loci from L. douglasii (Fig. 1), the species most distantly related to L. alba, 100 or more should amplify loci from each of the untested meadowfoam taxa and supply a significant number of conserved SSR markers for genetic conservation, phylogenetic, phylogeographic, ecological, and evolutionary genetics research in the Limnanthaceae.
References
Akkaya MS, Bhagwat AA, Cregan PB (1992) Length polymorphisms of simple sequence repeat DNA in soybean. Genetics 132:1131–1139
Bagby MO, Smith CR, Miwa TK, Lohmar RL, Wolff LA (1961) A unique fatty acid from Limnanthes douglasii seed oil: the C22 diene. J Org Chem 26:1261–1265
Bowcock AM, Ruiz-Linares A, Tomfohrde J, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368:455–457
Brown CR, Jain SK (1979) Reproductive system and pattern of genetic variation in two Limnanthes species. Theor Appl Genet 54:181–190
Brown SM, Hopkins MS, Mitchell SE, Senior ML, Wang TL, Duncan RR, Gonzalez-Candelas F, Kresovich S (1996) Multiple methods for the identification of polymorphic simple sequence repeats (SSRs) in sorghum [Sorghum bicolor (L.) Moench.]. Theor Appl Genet 93:190–198
Cho YG, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch SR, Park, Ayres N, Cartinhour S (2000) Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100:713–722
Cordiero GM, Casu R, McIntyre, Manner JM, Henry RJ (2001) Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross-transferable to Erianthus and sorghum. Plant Sci 160:1115–1123
Crane JM, Knapp SJ (2000) Registration of the self-pollinated meadowfoam germplasm line OMF64. Crop Sci 40:1511–1512
Crane JM, Knapp SJ (2002) Registration of ‘Wheeler’ meadowfoam. Crop Sci 42:2208–2209
Dole JA, Sun M (1992) Field and genetic survey of the endangered Butte County meadowfoam Limnanthes floccossa subsp. californica (Limnanthaceae). Conserv Biol 6:549–558
Don RH, Cox PT, Wainwright BJ, Baker K, Mattick JS (1991) ‘Touchdown’ PCR to circumvent spurious priming during gene amplification. Nucleic Acid Res 19:4008
Eastman DC (1990) Rare and endangered plants of Oregon. Beautiful Amer, Wilsonville, Ore., pp1–194
Edwards KJ, Barker JHA, Daly A, Jones C, Karp A (1996) Microsatellite libraries enriched for several microsatellite sequences in plants. BioTechniques 20:8–760
FitzSimmons N, Moritz C, Moore S (1995) Conservation and dynamics of microsatellite loci over 300 million years of marine turtle evolution. Mol Biol Evol 12:1106–1113
Garza JC, Freimer NB (1996) Homoplasy for size at microsatellite loci in humans and chimpanzees. Genome Res 6:211–217
Garza JC, Slatkin M, Freimer NB (1995) Microsatellite allele frequencies in humans and chimpanzees, with implications for constraints on allele size. Mol Biol Evol 12:594–603
Houle G, McKenna MF, Lapointe L (2000) Spatiotemporal dynamics of Floerkea proserpinacoides (Limnanthaceae), an annual plant of the deciduous forest of eastern North America. Am J Bot 88:594–607.
Innan H, Terauchi R, Miyashita T (1997) Microsatellite polymorphism in natural populations of the wild plant Arabidopsis thaliana. Genetics 146:1441–1452
Isbell TA (1997) Development of meadowfoam as industrial crop through novel fatty acid derivatives. Lipid Tech 9:140–144
Karagyozov L, Kalcheva ID, Chapman VM (1993) Construction of random small-insert genomic libraries highly enriched for simple sequence repeats. Nucleic Acid Res 21:3911–3912
Katengam S, Crane JM, Knapp SJ (2002) The development of a genetic map for meadowfoam comprised of amplified fragment length polymorphisms. Theor Appl Genet 104:92–96
Kesseli RV, Jain SK (1985) Breeding systems and population structure in Limnanthes. Theor Appl Genet 71:292–299.
Knapp SJ, Crane JM (1995) Fatty acid diversity of Section Inflexae Limnanthes. Ind Crops Prod 4:219–227
Knapp SJ, Crane JM (1997) The development of self-pollinated inbred lines of meadowfoam by direct selection in open-pollinated populations. Crop Sci 37:1770–1775
Knapp SJ, Crane JM (1998) The development of meadowfoam as an industrial oilseed: breeding challenges and germplasm resources. In: Janick J (ed). Proc Fourth Natl Symp New Crops. Wiley, New York, pp 225–233
Kresovich S, Szewc-McFadden AK, Blick SM (1995) Abundance and characterization of simple sequence repeats (SSRs) isolated from a size-fractionated genomic library of Brassica napus L. (rapeseed). Theor Appl Genet 91:206–211
Kubik C, Meyer WA, Gaut BS (1999) Assessing the abundance and polymorphism of simple sequence repeats in perennial ryegrass. Crop Sci 39:1136–1141
Kutil BL, Williams CG (2001) Triplet-repeat microsatellites shared among hard and soft pines. J Hered 92:327–332
Liu ZW, Jarret RL, Kresovich S, Duncan RR (1995) Characterization and analysis of simple sequence repeat (SSR) loci in seashore paspalum (Paspalum vaginatum Swartz). Theor Appl Genet 91:47–52
Lodhi MA, Ye GN, Weeden NF, Resich BI (1994) A simple and efficient method for DNA extraction from grapevine cultivars and Vitis species. Plant Mol Biol Rep 12:6–13
Loridon K, Cournoyer B, Goubely C, Depeiges A, Picard G (1998) Length polymorphism and allele structure of trinucleotide microsatellites in natural accessions of Arabidopsis thaliana. Theor Appl Genet 97:591–604
Mason CT (1952) A systematic study of the genus Limnanthes. Botanical Public 25:455–512
McNeill CI, Jain SK (1985) Genetic differentiation studies and phylogenetic inference in plant genus Limnanthes (section Inflexae). Theor Appl Genet 66:257–269
Mogg R, Batley J, Hanley S, Edwards D, Sullivan HO, Edwards KJ (2002) Characterization of the flanking regions of Zea mays microsatellites reveals a large number of useful sequence polymorphisms. Theor Appl Genet 105:532–543
Morgante M, Olivieri AM (1993) PCR-amplified microsatellites as markers in plant genetics. Plant J 3:175–182
Olufowote JO, Xu Y, Chen X, Park WD, Beachell HM, Dilday RH, Goto M, McCouch SR (1997) Comparative evaluation of within-cultivar variation of rice (Oryza sativa L.) using microsatellite and RFLP markers. Genome 40:370–378
Ornduff R (1971) Systematic studies of Limnanthaceae. Madrona 21:103–111
Ornduff R, Crovello TJ (1968) Numerical taxonomy of Limnanthaceae. Amer J Bot 55:173–182
Ostrander EA, Jong PM, Rine J, Duyk G (1992) Construction of small-insert genomic DNA libraries highly enriched for microsatellite repeat sequences. Proc Natl Acad Sci USA 89:3419–3423
Ott J (1991) Analysis of human genetic linkage. John Hopkins Univ Press, Baltimore, Md.
Peakall R, Gilmore S, Keys W, Morgante M, Rafalski A (1998) Cross-species amplification of soybean (Glycine max) simple sequence repeats (SSRs) within the genus and other legume genera: implications for the transferability of SSRs in plants. Mol Biol Evol 15:1275–1287
Plaschke J, Ganal MW, Röder MS (1995) Detection of the genetic diversity in closely related bread wheats using microsatellite markers. Theor Appl Genet 91:1001–1007
Powell W, Morgante M, Andre C, Hanafey C, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Primmer CR, Moller AP, Ellegren H (1996) A wide-range survey of cross-species microsatellite amplification in birds. Mol Ecol 5:365–378
Provan J, Powell W, Waugh R (1996) Microsatellite analysis of relationships within cultivated potato (Solanum tuberosum). Theor Appl Genet 92:1078–1084
Rae SJ, Aldam C, Dominguez I, Hoebrechts M, Barnes SR, Edwards KJ (2000) Development and incorporation of microsatellite markers into the linkage map of sugar beet (Beta vulgaris ssp.). Theor Appl Genet 100:1240–1248
Rafalski A, Tingey S (1993) Genetic diagnostics in plant breeding: RAPDs, microsatellites, and machines. Trends Genet 9:275–280
Ramsay L, Macaulay M, Cardle L, Morangte M, Ivanissevich DS, Maestri E, Powell W, Waugh R (1999) Intimate association of microsatellite repeats with retrotransposons and other dispered repetitive elements in barley. Plant J 17:415–425
Ramsay L, Macaulay M, Degli Ivanissevich S, MacLean K, Cardle L, Fuller J, Edwards KJ, Tuvesson S, Morgante M, Massari A, Maestri E, Marmiroli N, Sjakste T, Ganal M, Powell W, Waugh R (2000) A simple sequence repeat-based linkage map of barley. Genetics 156:1997–2005
Reed KM, Mendoza KM, Beattie CW (2000) Comparative analysis of microsatellite loci in chicken and turkey. Genome 43:796–802
Rico C, Rico I, Hewitt G (1996) 470 million years of conservation of microsatellite loci among fish species. Proc R Soc Lond B 263:549–557
Ritland K, Jain SK (1984) A comparative study of floral and electrophoretic variation with life history variation in Limnanthes alba (Limnanthaceae). Theor Appl Genet 63:243–251
Röder MS, Plaschke J, König SU, Börner A, Sorrels ME, Tanksley SD, Ganal SW (1995) Abundance, variability and chromosomal location of microsatellites in wheat. Mol Gen Genet 246:327–333
Rongwen J, Akkaya MS, Bhagwat AA, Lavi V, Cregan PB (1995) The use of microsatellite DNA markers for soybean genotype identification. Theor Appl Genet 90:43–48
Saal B, Wricke G (1999) Development of simple sequence repeat markers in rye (Secale cereale L.) Genome 42:964–972
Saghai Maroof MA, Biyashev RM, Yang GP, Zhang Q, Allard RW (1994) Extraordinarily polymorphic microsatellite DNA in barley: species diversity, chromosomal locations, and population dynamics. Proc Nat Acad Sci 91:5466–5470
Schlötterer C, Amos B, Tautz D (1991) Conservation of polymorphic simple sequence repeat loci in cetacean species. Nature 354:63–65
Smith CR, Bagby MO, Miwa TK, Lohmar RL, Wolff IA (1960) Unique fatty acids from Limnanthes douglasii seed oil: The C20- and C22-monoenes. J Org Chem 25:1770–1774
Smith JSC, Chin ECL, Shu H, Smith OH, Wall SJ, Senior ML, Mitchell SE, Kresovich S, Ziegle J (1997) An evaluation of utility of SSR loci as molecular markers in maize (Zea mays L.): comparisons with data from RFLPs and pedigrees. Theor Appl Genet 95:163–173
Smith JSC, Kresovich S, Hopkins MS, Mitchell SE, Dean RE, Woodman WL, Lee M, Porter K (2000) Genetic diversity among elite sorghum inbred lines assessed with simple sequence repeats. Crop Sci 40:226–232
Smulders MJM, Bredemeijer G, Ruskortekaas W, Arens P, Vosman B (1997) Use of short microsatellites from database sequences to generate polymorphisms among Lycopersicon esculentum cultivars and accessions of other Lycopersicon species. Theor Appl Genet 94:264–272
Smulders MJM, Rus-Kortekaas W, Vosman B (2000) Microsatellite markers useful throughout the genus Dianthus. Genome 43:208–210
Szewc-McFadden AK, Kresovich S, Bliek SM, Mitchell SE, McFerson JR (1996) Identification of polymorphic, conserved simple sequence repeats (SSRs) in cultivated Brassica species. Theor Appl Genet 93:534–538
Tang S, Knapp SJ (2003) Microsatellites uncover extraordinary molecular genetic diversity in Native American land races and wild populations of cultivated sunflower. Theor Appl Genet 106:990–1003
Tang S, Yu JK, Slabaugh MB, Shintani DK, Knapp SJ (2002) Simple sequence repeat map of the sunflower genome. Theor Appl Genet 105:1124–1136
Taramino G, Tingey S (1996) Simple sequence repeats for germplasm analysis and mapping in maize. Genome 39:227–287
Temnykh S, Park WD, Ayres N, Cartinhour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Walter KS, Gillett, HJ (1998) IUCN red list of threatened plants. World Conservation Monitoring Centre. The World Conservation Union, Gland, Switzerland and Cambridge, UK, pp 1–862
Weber JL (1990) Informativeness of human (dC-dA)n(dG-dT)n polymorphisms. Genomics 7:524–530
Weber JL, May PE (1989) Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Amer J Human Genet 44:388–396
Whitton J, Rieseberg LH, Ungerer MC (1997) Microsatellite loci are not conserved across the Asteraceae. Mol Biol Evol 14:204–209
Wu KS, Tanksley SD (1993) Abundance, polymorphism and genetic mapping of microsatellies in rice. Mol Gen Genet 241:225–235
Yu JK, Mangor J, Thompson L, Edwards KJ, Slabaugh MB, Knapp SJ (2002) Allelic diversity of simple sequence repeat markers among elite inbred lines in cultivated sunflower. Genome 45:652–660
Acknowledgements
This research was funded by a grant from the USDA (99–34407–7509) to S.J.K. The DNA sequences described herein (GenBank BV007038-BV007426) were produced by the Nevada Genomics Center (NGC). NGC was established in 2000 with a National Science Foundation EPSCoR grant (P20 RR16464) to provide genomics services to the state of Nevada, is located in the Fleischmann Agricultural Building at the University of Nevada, Reno, and is a component of the Nevada Biomedical Research Infrastructure Resources (NBRR).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by O. Savolainen
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Kishore, V.K., Velasco, P., Shintani, D.K. et al. Conserved simple sequence repeats for the Limnanthaceae (Brassicales). Theor Appl Genet 108, 450–457 (2004). https://doi.org/10.1007/s00122-003-1447-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-003-1447-1