Abstract.
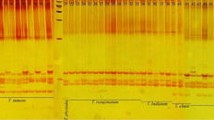
The development of microsatellite markers can be a time-consuming process, especially in species such as conifers where many microsatellites have been shown to be associated with the repetitive fraction of the genome and to produce complex banding patterns following electrophoresis. Therefore, procedures to eliminate this fraction from further processing are sought. In this paper, we report on the development of 53 dinucleotide SSR markers in Norway spruce, 35 of which (66%) produce simple, polymorphic patterns. This high efficiency is obtained by introducing a dot-blot selection against high copy number sequences, performed on the microsatellite-containing clones. The resulting markers turned out to be polymorphic and useful for population genetic studies and for linkage mapping. Seven additional markers that were not subject to the dot-blot selection are also presented.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Scotti, .I., Paglia, .G., Magni, .F. et al. Efficient development of dinucleotide microsatellite markers in Norway spruce (Picea abies Karst.) through dot-blot selection. Theor Appl Genet 104, 1035–1041 (2002). https://doi.org/10.1007/s00122-001-0843-7
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00122-001-0843-7