Abstract
Ischemia-induced metabolic remodeling plays a critical role in the pathogenesis of adverse cardiac remodeling and heart failure however, the underlying molecular mechanism is largely unknown. Here, we assess the potential roles of nicotinamide riboside kinase-2 (NRK-2), a muscle-specific protein, in ischemia-induced metabolic switch and heart failure through employing transcriptomic and metabolomic approaches in ischemic NRK-2 knockout mice. The investigations revealed NRK-2 as a novel regulator of several metabolic processes in the ischemic heart. Cardiac metabolism and mitochondrial function and fibrosis were identified as top dysregulated cellular processes in the KO hearts post-MI. Several genes linked to mitochondrial function, metabolism, and cardiomyocyte structural proteins were severely downregulated in the ischemic NRK-2 KO hearts. Analysis revealed significantly upregulated ECM-related pathways which was accompanied by the upregulation of several key cell signaling pathways including SMAD, MAPK, cGMP, integrin, and Akt in the KO heart post-MI. Metabolomic studies identified profound upregulation of metabolites mevalonic acid, 3,4-dihydroxyphenylglycol, 2-penylbutyric acid, and uridine. However, other metabolites stearic acid, 8,11,14-eicosatrienoic acid, and 2-pyrrolidinone were significantly downregulated in the ischemic KO hearts. Taken together, these findings suggest that NRK-2 promotes metabolic adaptation in the ischemic heart. The aberrant metabolism in the ischemic NRK-2 KO heart is largely driven by dysregulated cGMP and Akt and mitochondrial pathways.
Key messages
-
Post-myocardial infarction metabolic switch critically regulates the pathogenesis of adverse cardiac remodeling and heart failure. Here, we report NRK-2 as a novel regulator of several cellular processes including metabolism and mitochondrial function post-MI. NRK-2 deficiency leads to downregulation of genes important for mitochondrial pathway, metabolism, and cardiomyocyte structural proteins in the ischemic heart. It was accompanied by upregulation of several key cell signaling pathways including SMAD, MAPK, cGMP, integrin, and Akt and dysregulation of numerous metabolites essential for cardiac bioenergetics. Taken together, these findings suggest that NRK-2 is critical for metabolic adaptation of the ischemic heart.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiac diseases including myocardial infarction (MI)-induced heart failure account for a number of deaths larger than any other disease in humans [1]. The lifestyle and genetic makeup of an individual are among the largest risk factors of MI. The systemic hypoxia and metabolic alterations in MI switch the energy production towards anaerobic glycolysis at the expanse of oxidative metabolism [2]. Consequently, a metabolic switch between glycolysis and fatty acid usage occurs depending on the oxygen levels that determine mitochondrial bioenergetics. Ischemia-induced cardiac injury is primarily associated with the severity of cardiomyopathy, fibrosis, and metabolic conditions of the heart [3, 4]. During ischemia, oxygen depletion to the myocardium affects mitochondrial and metabolic functions. Overall, the metabolic response is important for cardiac homeostasis and successful adaptation to MI.
In accordance with the energy demand and oxygen supply of the heart for efficient pumping, oxidation of fatty acid occurs within myocardial cells based on the allosteric control of fatty acid uptake, esterification, and mitochondrial transport [5, 6]. Alterations in fatty acid metabolism, particularly abnormal fatty acid β-oxidation during and after ischemia, may contribute to heart failure [7]. Different fatty acids and their final metabolites have different effects on cardiomyocytes. Even though a reduced fatty acid metabolism eventually leads to increased glucose utilization [8, 9], it is unclear how well these metabolic pathways are regulated in the myocardium.
In ischemic conditions, aberrant metabolic response in the heart largely depends on mitochondrial function. Mitochondria generate ~ 90% of the ATPs for myocardial contraction, and mitochondrial oxidative phosphorylation (OXPHOS) is the primary source of energy in the heart [10]. Moreover, mitochondria maintain a homeostatic state by regulating the metabolic pathways responsible for balancing fatty acid and glucose oxidative phosphorylation [4]. In pathological conditions, this homeostatic state gets disturbed, which often causes mitochondrial dysfunction and, ultimately, cell death [11]. Dead myocardium is replaced by fibrotic tissues, characterized by the transformation of cardiac fibroblasts to myofibroblasts [3, 12]. Integrin signalings, such as the focal adhesion kinase (FAK), Rho-associated protein kinase (ROCK), and mitogen-activated protein kinase (MAPK), are well-established signaling pathways to regulate cardiac fibrosis under cardiac stress conditions [13]. Therefore, mitochondrial health and metabolic derangement play important roles in the pathogenesis of heart failure post-ischemia. Though many studies reported the role of mitochondrial dysfunction and metabolic switch in MI-induced cardiac remodeling, the underlying molecular mechanisms remain elusive.
Studies from our lab and other groups have characterized the role of a muscle integrin-binding protein nicotinamide riboside kinase-2 (NRK-2) in heart failure models including ischemia and pressure overload [14,15,16]. NRK-2 is highly expressed in skeletal muscle, and a trace amount is reported in healthy cardiac muscles [15]. We previously identified that NRK-2 specifically represses the P38α activity in mouse ischemic heart [15], and JNK in angiotensin II-treated human cardiomyocytes to limit apoptotic cardiac cell death, dilatative cardiac remodeling, and excessive fibrosis [16]. Despite these critical roles and being a top upregulated protein in dilated cardiomyopathy model [17, 18], the downstream targets of NRK-2 in cardiac pathogenesis are largely unknown.
Here, we performed transcriptomic and metabolomic studies in the ischemic NRK-2 KO hearts to identify the novel dysregulated genes/pathways and metabolites. We report, for the first time, that the loss of NRK-2 profoundly upregulates genes related to cardiac fibrosis and downregulates genes related to metabolic pathways and mitochondrial function. Moreover, metabolic studies suggest the dysregulation of important cardiac metabolites, including mevalonic acid, 3,4-dihydroxyphenylglycol (DHPG), 2-penylbutyric acid, stearic acid, 2-pyrrolidinone, and 8,14,11-eicosatrienoic acid. Hence, our findings strongly suggest that NRK-2 interplays roles between mitochondrial function and metabolic remodeling in the ischemic heart.
Method
NRK-2 knockout mice generation and myocardial infarction induction
NRK-2 knockout mouse was generated by the Jackson Laboratory (stock #018,683) using embryonic stem cells as described previously [15]. The experimental NRK-2 knockout (KO) mice were generated by crossing heterozygous males with heterozygous females under controlled conditions. The mouse strain was maintained on the C57BL/6 background. Eight- to ten-week old NRK-2 KO and littermate control male mice (n = 7–8) were anesthetized through an IP injection of ketamine (50 mg/kg) and xylazine (2.5 mg/kg) and subjected to permanent ligation of proximal left anterior descending (LAD) coronary artery (myocardial infarction), as described previously [19]. Since NRK-2 KO mice presented a severe phenotype two weeks following MI [15], we selected a time point of 1-week post-MI for our studies. The mice were euthanized by CO2 overdosing in a closed chamber, and cardiac tissue was harvested immediately for the studies. All animal procedures were carried out as per current NIH guidelines. The Institutional Animal Care and Use Committee (IACUC) of Vanderbilt University Medical Center approved all animal procedures and treatments, and animal-related experiments were performed at the Vanderbilt University Medical Center.
RNA extraction and cDNA library preparation
RNA isolation was performed as described previously [20]. Briefly, total RNA was isolated from remote LV tissues (after careful exclusion of the fibrotic scar and border area) (n = 5–4) using RNeasy kit (Qiagen) according to the manufacturer’s protocol. The quality of total RNA was assessed by the ratios OD260/OD280 and OD260/OD230 using Nano-drop 2000. Complimentary DNA (cDNA) was synthesized from 1 µg total RNA through SuperScript II reverse-transcriptase (Life Technologies). Prior to the next-generation sequencing (n = 3 each group), adapters were ligated onto both ends of the cDNA fragments. After amplifying fragments using PCR, fragments with insert sizes between 200 and 400 bp were selected for further processing. The libraries were pooled and sequenced on NovaSeq (paired-end sequencing).
Read processing and QC alignment
Library quality control was performed to analyze the quality of the overall raw reads and total bases, as well as the percentage of the GC content. The library was checked with Qubit and real-time PCR for quantification and size detection. The total number of bases, reads, GC (%), Q20 (%), and Q30 (%) were calculated for each sample. The quality of produced data is determined by the phred quality score (Q20) at each cycle referring to 99% accuracy and reads over the score of 20 are accepted as good quality. Phred quality score 20 Box plot containing the average quality at each cycle is created with FastQC wherein the x-axis shows the number of cycles and y-axis shows phred quality score.
Prior to sequencing, artifacts such as low-quality reads, adaptor sequence, contaminant genomic DNA, or PCR duplicates are removed. Trimmomatic program was used to remove adapter sequences and bases with base quality lower than three from the ends. Reads with a length shorter than 36 bp were removed during data trimming. Trimmed reads were then mapped to the reference genome using HISAT2, through Bowtie2 splice-aware aligner. UCSC mm10 was used as a reference genome.
Expression profiling analysis
Transcripts were assembled by StringTie with aligned reads (n = 3 each group) as described previously. StringTie is a highly efficient assembler of RNA-Seq alignments into potential transcripts. It uses a network flow algorithm as well as de novo assembly step to assemble and quantitate full-length transcripts representing multiple splice variants for each gene locus. Expression profiles are represented as read count and normalization value which are based on transcript length and depth of coverage. The FPKM (fragments per kilobase of transcript per million mapped reads) value is used as a normalization value. Differentially expressed genes were determined using a volcano plot and genes with log2 fold change of 2 or higher were taken.
Enrichment analysis
Gene Ontology (GO terms for biological process and molecular function) and KEGG pathways enrichment analysis were performed (n = 3 each group) to identify the differentially expressed genes as described previously [21]. R documentation and GO enrichment analysis for differentially expressed genes were carried out by clusterProfiler R as described before [22], and any GO term with corrected p < 0.05 was considered an enriched term. The clusterProfiler was also used to test the statistical enrichment of differential gene expression in KEGG pathways, and any term with an adjusted p < 0.05 was considered statistically significant.
Quantitative real-time PCR (qRT-PCR)
To validate the NGS data, a set of six top dysregulated genes were chosen for qRT-PCR (n = 5–4) to validate the transcript data. qRT-PCR was performed using the same procedure as described previously [23]. Briefly, total RNA was extracted using the RNeasy kit (Qiagen), and cDNA was synthesized using SuperScript II reverse-transcriptase (Life Technologies). qRT-PCR is performed using the SYBR Green PCR master mix using the primers provided in Suppl. Table 1. The calculations were performed using delta-delta-Ct method, and the results were normalized with 18 s rRNA.
LV lysate preparation
LV Lysates were prepared as described previously [24]. Briefly, heart tissues were harvested from anesthetized mice post 1 week of MI. Then, the atrium and right ventricle were excised carefully. The remote LV tissue (n = 3 each group) was homogenized in 1 × lysis buffer (Cell Signaling #9803) containing protease and phosphatase inhibitor cocktail. After homogenization, tissue lysates were spun down at 15,000 g for 15 min at 4 °C. The supernatant was collected in fresh sterile tubes, and protein concentration was quantified using bicinchoninic acid (BCA) protein assay (Pierce# 23,225). A total of 100 µg of the lysate were taken for further metabolomics study.
Tandem mass high-performance liquid chromatography mass spectrometry (HPLC–MS/MS)
For the separation of metabolites, the TimsTOF mass spectrometer and Elute UHPLC and autosampler (Bruker, Billerica, MA, USA) were employed as described previously [25]. Each metabolite was analyzed using electrospray ionization (ESI) with MS/MS-positive scan mode within the range of 20–1300 m/z. A Hamilton Intensity Solo 2 C18 column (100 mm × 2.1 mm, 1.8 µm beads) was maintained at 35 ℃ for metabolomics analyses. Ten microliters of the samples reconstituted with acetonitrile was injected twice consecutively and eluted using a gradient elution mode. Sodium formate was injected as an external calibrant in the first 0.3 min of each LC–MS/MS run.
Metabolite processing and integration of metabolomics with transcriptomics
MetaboAnalyst V5 workflow software [26] was used for metabolite processing and statistical analysis (n = 3 each group). Parameters set for peak detection are as follows: a minimum intensity threshold of 1000 counts as well as a minimum peak duration of 7 spectra, using peak area for feature quantification. The file masses were recalibrated based on the external calibrant injected between 0 and 0.3 min. To analyze the distribution of metabolite intensities among groups, independent, two-tailed Student’s t-test was performed using MetaboAnalyst V5.
Pathway enrichment and integration of metabolomics with transcriptomics were performed using MetaboAnalyst 5.0 and ImPala software. Enrichment was carried out with selected different metabolites (DM) with significant p value (0.0) versus top upregulated or downregulated DEG’s. The DMs with p < 0.1, combined with the DEGs with p < 0.05, and |log2FoldChange|> 1.5, were selected for integrative analysis.
Statistical analysis
The results are presented as mean ± SEM, and the comparisons among the groups were performed by the Mann–Whitney test unless it is mentioned. Data were analyzed using GraphPad Prism 6 (GraphPad Software, La Jolla, CA), and p < 0.05 was considered statistically significant.
Results
NRK-2 deficiency induces differential expression of genes post-MI
We recently reported that NRK-2 deficiency promotes rapid LV chamber dilatation, increased fibrosis and scar formation, LV wall thinning, and heart failure post-MI [15]. However, precise molecular mechanisms and downstream targets of NRK-2 are unknown. To identify the same, transcriptomic analysis was performed in remote LV harvested from NRK-2 KO mice post 1 week of MI. Following total RNA library construction, sequencing and transcriptome analysis were performed (Fig. 1A). The quality check of the transcript data was performed using phred quality score. The phred quality score of 20 was considered as 99% accuracy, and the reads over the score of 20 were accepted as good quality.
A Schematic diagram shows the experimental design. NRK-2 knockout (KO) and littermate control mice were subjected to left anterior descending (LAD) coronary artery ligation (myocardial infarction) followed by cardiac tissue harvesting and total RNA isolation. mRNA sequencing followed transcriptomic analysis and metabolomic studies were performed using remote LV region to identify the differentially expressed genes (DEGs). B MA plot shows log2 fold-changes in the gene expression versus the mean of normalized count between NRK-2 KO and control hearts. C The volcano plot shows DEGs from control NRK-2 KO hearts post-MI. The top left genes (red) show significantly downregulated genes while the top right panel (red) shows significantly upregulated genes. Only a few top dysregulated genes are labeled
The dispersion scatters and volcano plots based on differentially expressed genes (DEGs) revealed a total of 24,528 variables between the NRK-2 KO and control groups post-MI (Fig. 1B). The fibrotic genes such as Col8a1, Col12a1, Col5a2, Mfap5, and Fbn1 were among the most upregulated genes (Fig. 1C, Table 1). Collectively, these data suggest that loss of NRK-2 induces differential expression of several genes post-MI.
Uniquely and differentially regulated genes and pathways in the NRK-2 KO heart post-MI
The R package of clusterProfiler was performed to assess the functional insights of DEGs and strongly influenced KEGG pathways. The heatmap shows the group of DEGs of extremely affected pathways which includes mitochondrial function, metabolism, and extracellular matrix (ECM) (Fig. 2A). Most of the genes in the top downregulated group in the ischemic KO hearts were related to mitochondrial function and metabolism. Consequently, most of the genes in the top upregulated group were related to components of ECM. The Venn diagram showed a total of 4028 differentially expressed genes shared by both control and KO groups, while 39 genes in the KO and 77 genes in the control groups were uniquely expressed (Fig. 2B). Based on enriched ontology clustering, DEGs were significantly enriched in the pathways like nucleoside biphosphate metabolic process [Padj = −log10(3.5)], protein homooligomerization [Padj = −log10(3.2)], and drug metabolism [Padj = −log10(3)] and were enriched among the top downregulated pathways (Fig. 2C). However, the fibrotic pathways including ECM organization, assembly of collagen fibrils (Padj = −log10(12.5)), integrin cell surface interactions (Padj = −log10(12)), and cellular responses to TGF-β stimulus [Padj = −log10(12)] were among the top upregulated pathways (Fig. 2D). Analysis from the KEGG pathways indicated that pathways related to metabolic functions were exceptionally downregulated and tissue fibrosis regulatory pathways were among the top upregulated in the NRK-2 KO hearts post-MI.
NRK-2 deficiency upregulates ECM components genes and downregulates metabolic and mitochondrial pathways genes post-MI. A The heat map shows lower expression levels of many genes (right upper panel) related to mitochondrial dysfunction (MD) and metabolism in the NRK-2 KO while upregulation of many genes of extracellular matrix (ECM) components (right lower panel). B The circles show the number of unique and overlapping (4028) differentially expressed genes in control and NRK-2 KO hearts. KEGG pathway analysis shows C downregulated and D upregulated pathways in the KO heart post-MI
NRK-2 regulates fibrosis and metabolic derangement in the ischemic heart
Next, we investigated the top differentially expressed genes in the KO hearts. Most of the upregulated genes in the NRK-2-deficient heart were related to ECM deposition and fibrosis. A total of 29 genes related to ECM components were identified to be upregulated in the KO hearts post-MI (Fig. 3A). Other than the fibrotic genes, Lox, Thbs1, Nppa, Nppb, Aspn, Sfrp1, Myh7, Loxl1, and Fat1 were also among the top upregulated genes (Table 1). Interestingly, genes that play important roles in mitochondrial and metabolic functions were downregulated. A total of 34 genes related to metabolic processes and 40 genes related to mitochondrial function were found to be downregulated in the NRK-2 hearts post-MI (Suppl. Table 2A, B, C). The genes including Atp2a2, Slc25a4, Gbas, Nnt, Sdhb, Sod2, Eno3, Acsl1, Etfdh, Hadhb, Idh2, Hadha, Acadm, Acadvl, Acaa2, and Ech1 were among the top 50 downregulated genes. Further analysis revealed that the other gene categories including Smyd1, Fabp3, Pygm, Fyco1, Ldhb, Gsn, Pink1, Myl3, Myl2, Pln, Tcap, and Tnni3 were also included among the top downregulated genes (Fig. 3B, C). These findings suggest that NRK-2 plays important roles in the regulation of fibrosis, mitochondrial function, and metabolic remodeling in the ischemic heart.
Loss of NRK-2 modulates MI-induced expression of genes related to cardiac structural and Ca2+ handing proteins
Next, we validated a few genes from the top differentially expressed group of genes. A total of six from top upregulated and another six from top downregulated gene sets were assessed through qRT-PCR using RNA isolated from 1 week post-MI KO and control LV tissue. As expected, analyses revealed that NRK-2 deficiency significantly downregulated the mRNA transcript levels for the genes Tcap, Tnni3, Atp2a22, Acadm, Acaa2, and Hadha (Fig. 4A–F). Moreover, consistent with our transcriptomic data, qRT-PCR analysis for the genes Ltbp2, Col1a1, Col12a1, Col8a1, Mmp2, and Fn1 showed significantly increased mRNA expression in the NRK-2 KO hearts post-MI (Fig. 4G–L). The genes which were significantly upregulated are related to ECM structural components like LTBP2 which forms microfibrils and involves in cell adhesion. In this group, other genes such as Col1a1, Col12a1, and Col8a1 are collagen related, and FN1 is a fibronection coding gene that plays a role in cell migration. Many of the validated genes under the downregulated category like Acadm, Acaa2, and Hadha are related to mitochondrial enzymes, whereas Tcap and Tnni3 encode cardiac troponin and related proteins.
Validation of differentially expressed genes through qRT-PCR: A A few top downregulated (A–F) and upregulated genes (G–L) validation show the relative mRNA expressions in the control and NRK-2 KO hearts post-MI. Values are expressed as mean ± SEM, Mann–Whitney test was performed to compare the group. ns, not significant; *p < 0.05
NRK-2 deficiency dysregulates important metabolites in the ischemic heart
Our transcriptomic findings strongly suggest that NRK-2 may play critical roles in the derangement of metabolic activity in the heart post-ischemia. Therefore, we sought to assess the metabolites expression profile in the NRK-2 KO hearts post-MI. After metabolomic analysis using MetaboAnalyst, a total of 96 metabolites were confidently (p < 0.05) assigned to the HMDB 4.0 library (Suppl. Table 3). The principal component analysis (PCA) revealed incomplete separation between the control and NRK-2 groups. Thus, multivariate analysis was performed using a score plot for orthogonal projections to latent structures discriminant analysis (OPLSDA) providing a better separation and differences in key metabolites between the control and KO groups (Fig. 5A). Moreover, a volcano plot showed differential expression patterns of seven different metabolites (Fig. 5B).
NRK-2 deficiency induces differential regulation of cardiac metabolites post-MI. A Orthogonal partial least squares (OPLS) for metabolites in the ischemic NRK-2 KO (red) and control (green) hearts. Each point represents biological replicate. B The volcano plot shows significantly dysregulated metabolites. Further analysis shows profound upregulation of C mevalonic acid, D 3,4-dihydroxyphenylglycol, E 2-phenylbutyric acid, F uridine, and downregulation of G stearic acid, H 8,11,14-eicosatrienoic acid, and I 2-pyrrolidinone in the NRK-2 KO post-MI. n = 3 each group
Further analysis of these seven metabolites revealed that the relative concentration of four of them, including mevalonic acid (mevalonate), DHPG, 2-phenylbutyric acid, and uridine, were profoundly upregulated in the ischemic NRK-2 KO compared to control hearts (Fig. 5C–F). Interestingly, the relative concentration levels of the stearic acid, 8,11,14-eicosatrienoic acid and 2-pyrrolidinone were significantly lower in the ischemic KO vs. control hearts (Fig. 5G–I). Taken together, these findings suggest that NRK-2 is important for metabolic processes in the ischemic heart and deficiency leads to metabolic derangement and heart failure.
Network analysis of DEGs with targeted metabolomics
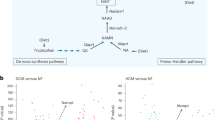
To illustrate the association between DEGs and metabolites, we conducted a pathway-based analysis using the online software MetaboAnalyst 5.0. Based on our inputs, three major metabolic pathways were identified which mainly include biosynthesis of unsaturated fatty acid, pyrimidine metabolism, and tyrosine metabolism. The metabolic pathways that significantly influenced metabolites with respect to NRK-2 are displayed as different colored circles (Fig. 6A), and the depth of the color indicates the significance of the altered pathways along with the distance from the origin circles. The size of the circle implies to the relative impact values (Fig. 6B). The bar chart summarizes the enrichment analysis based on differential metabolites with significant p-value (Fig. 6C).
To understand the alteration of metabolic responses at the transcriptome level, a comprehensive analysis was conducted with ImPala software which shows the relationships between DEGs and targeted metabolomics (Tables 2 and 3). With this software, the most probable molecular interactions among the DM and DEGs were identified and listed according to p value significance. The strongest coordinated responsiveness to NRK-2 was identified; with the most significant change related to ferroptosis, mitochondrial β-oxidation of long-chain saturated fatty acids, binding and uptake of ligands by scavenger receptors, and scavenging by class A pathways. The most significantly downregulated DEGs (Hadhb, Acaa2, Acsl1, Hadha) and downregulated DM stearic acid are involved in mitochondrial β-oxidation of long-chain saturated fatty acids. Another significantly upregulated DEGs, Col1a1, Col1a2, Col3a1, Col4a1, and Col4a2, along with guanosine monophosphate, were connected through a range of biological reactions that are involved in metabolism, signaling, and scavenging pathways. Collectively, enrichment and integration of pathways based on DM and DEGs suggest that NRK-2 knockout affects major pathways linked to metabolism along with many other biological pathways of minor concern.
Discussion
Cardiac fibrosis, mitochondrial dysfunction, and metabolic derangement are the major complications found robustly associated with MI [27, 28]. Previous studies from our and other groups revealed that NRK-2 plays a critical role in the pathogenesis of cardiac diseases. The expression of NRK-2 dramatically increases in ischemic and pressure-overloaded hearts, and deficiency induces dilated cardiomyopathies’ post-cardiac stressors [15, 16]. There is currently no treatment to limit or reverse dilatative cardiac remodeling and heart failure. Here, we identified NRK-2 as a critical regulator of cardiac fibrosis, mitochondrial function, and metabolic processes through altering myocardial transcriptomic and metabolomic profile post-MI. Our novel findings indicate that NRK-2 deficiency leads to an upregulation of fibrotic gene panels and downregulation of mitochondrial and metabolic genes post-MI. Moreover, loss of NRK-2 leads to profound upregulation of mevalonic acid, DHPG, 2-phenylbutyric acid, and uridine and downregulation of stearic acid, 8,11,14-eicosatrienoic acid, and 2-pyrrolidinone metabolites in the ischemic heart.
Post-MI transcriptomic analysis shows differential expression of many genes in the NRK-2 KO hearts, which largely play roles in fibrosis induction, mitochondrial dysfunction, and metabolic derangement post-MI. These findings are consistent with our previous report of increased fibrosis, extensive scar formation, dilated cardiac remodeling with profound LV dysfunction, and heart failure in NRK-2-deficient mice post-MI [15]. ECM organization and several other fibrosis-related pathways, including SMAD and MAP kinase, were among the top upregulated pathways in the KO post-MI. These observations strongly suggest the anti-fibrotic roles of NRK-2 in the ischemic heart though it is not clear whether NRK-2 directly modulates profibrotic pathways. There is also a possibility that the upregulation of profibrotic pathways in the KO hearts is secondary to deranged metabolism and mitochondrial pathways-induced cell death post-MI. Moreover, similar to NRK-2, other integrin-binding proteins, including FAK, integrin-linked kinase (ILK), Talin, Vinculin, and Melusin, are previously linked to cardiomyopathy and fibrosis in the stressed heart [29,30,31,32].
Mitochondrial bioenergetics and metabolism are critical for the cardiomyocytes to perform a healthy state of action. Upon MI onset, low availability of oxygen renders the cardiomyocytes and other cardiac cells in a hypoxic state, which induces a metabolic shift [33]. In this condition, mitochondrial function and turnover are also affected. We identified several downregulated genes related to mitochondrial ion channels and metabolism in the NRK-2 KO hearts post-MI. Among them, Acadm is primarily involved in fatty acid metabolism. Major studies in the mammalian heart suggest that fatty acid β-oxidation is the preferred pathway of energy production for maintaining efficient cardiac activity [34, 35]. We also found a downregulation of acetyl-CoenzymeA acyltransferase 2 (Acaa2) in the NRK-2 KO heart post-MI which is involved as a potential binding partner in medium-chain fatty acid metabolism and catalyzes the last step in the β-oxidation pathway [36, 37]. Though the role of the Acaa2 gene is poorly characterized in the cardiomyocytes, we propose that its suppression may affect the adaptation of the metabolic pathways during cardiac injury. Similar to Acadm and Acaa2, several other genes related to fatty acid oxidation were downregulated in the KO hearts. Enoyl coenzyme A hydratase 1 (ECH1) is another gene among downregulated group found in mitochondria [38] and helps in metabolism primarily through catabolizing the polyunsaturated fatty acids [39]. The shifting of the myocardial bioenergetic substrate from fatty acid to glucose leads to the downregulation of fatty acid β enzyme, which ultimately results in heart failure [40, 41]. Hence, NRK-2 regulates genes that are required for normal mitochondrial function and adequate metabolic activity of the ischemic heart.
Consistent with mitochondrial dysfunction and metabolic pathway defects, our findings further reveal the dysregulation of critical metabolites in the ischemic NRK-2 KO heart. Growing datasets suggest that the alterations in metabolites level may help better understand the pathogenesis of heart failure and early detection of heart failure [42, 43]. NRK-2 KO heart displayed an upregulation of metabolites mevalonate, DHPG, 2-phenylbutyric acid, and uridine post-MI. Mevalonate is synthesized from 3-hydroxy-3-methylglutaryl-coenzyme A (HMG-CoA) [44] and is metabolized into isoprenoids, which is crucial for a variety of cellular functions, including cholesterol synthesis, cell growth, proliferation, and death [45, 46]. Studies have shown that targeting the mevalonate pathway limits myocardial fibrosis, adverse cardiac remodeling, and dysfunction post-MI [47]. These studies suggest the detrimental effects of elevated mevalonate in the pathological phenotype of the ischemic NRK-2 KO animals. Another dysregulated metabolite in NRK-2 KO heart was DHPG which is a noradrenaline metabolite and is elevated in the infarcted heart of rats. The treatment with an angiotensin-converting enzyme (ACE) inhibitor reverses the level of DHPG and reduces heart failure [48]. Therefore, induction of DHPG in the NRK-2 KO hearts seems to occur due to increased LV dysfunction and heart failure.
The precise role of metabolite 2-phenylbutyric acid in cardiac pathophysiology is unknown. The 2-phenylbutyric acid is a secondary metabolite, which is generally non-essential and may play roles in cell signaling. Such metabolites are often produced due to the incomplete metabolism of secondary metabolites [49]. Uridine was another metabolite with elevated expression in the NRK-2 KO hearts, which has also been reported to be induced in human ischemic hearts [50]. Interestingly, a recent study demonstrated the protective roles of uridine in rats challenged with myocardial ischemia [51]. The study concluded that uridine protects the heart against ischemia potentially through the activation of mitochondrial ATP-dependent potassium channels.
In stark contrast, 8,11,14-eicosatrienoic acid, stearic acid, and 2-pyrolidinone were robustly downregulated in the KOs hearts compared to control heart pots-MI. The 8,11,14-eicosatrienoic acid is a metabolite of alpha-linolenic acid and linoleic acid metabolism pathway. A higher plasma level of 8,11,14-eicosatrienoic acid was recently identified in patients with acute myocardial infarction (AMI) [52]. These observations suggest that the induction of 8,11,14-eicosatrienoic acid in AMI patients is compensatory, and NRK-2 likely helps in the upregulation of such metabolites upon ischemia for metabolic adaptation. Stearic acid is another non-essential saturated fatty acid chain [53] found severely attenuated in the NRK-2 KO vs. control hearts post-MI. The loss of NRK-2 in the KO may limit the induction of these fatty acids, and their metabolites consequently lead to detrimental cardiac phenotypes.
In the same line, our transcriptomic analysis showed significant downregulation of fatty acid-binding protein-3 (FABP3). FABP3 is a small protein in the heart that plays a crucial role in cardiomyocyte metabolism and limiting lipid toxicity by binding and transporting the free long-chain fatty acids [54,55,56]. A study reported a positive correlation between circulating FABP3 and cardiac hypertrophy and heart failure in pressure-overloaded mice and humans [57]. These observations suggest that NRK-2 deficiency in the heart downregulates cardiac FABP3 which, in part, perturbs the cardiomyocyte metabolism post-MI.
An aberrant metabolism in cardiac pathogenesis is linked with mitochondrial dysfunction which is regulated by a variety of signaling pathways [58]. Activation of MAPK pathways triggers several downstream signaling molecules which ultimately aggravates mitochondrial dysfunction and cellular injury under cardiac stressors [59]. Our current and previous studies [16] show that NRK-2 is one of the potential mediators of MAPK activation-induced mitochondrial dysfunction which has ultimate larger effects on the cellular metabolism in the ischemic heart.
Besides these, NRK-2 is a muscle-specific β1 integrin-binding protein that selectively reduces cell adhesion to laminin and plays a crucial role in terminal myogenesis [60]. However, it is largely unknown if NRK-2 regulates the structural organization of cardiac muscle under pathogenic conditions. Myl3, Myl2, Pln, Tcap, and Tnni3 are cardiac muscle-related genes that were among the most downregulated genes in the cellular components of the post-MI KO hearts. A variety of mutations in these genes cause dilated cardiomyopathy in humans [61]. It is not clear how NRK-2 potentially regulates these cardiac muscle structural genes, and further studies are warranted to understand the precise mechanism. Other genes such as Fyco1 and Pink1 regulate autophagy and mitophagy [62, 63], and their downregulation in the post-MI NRK-2 KO heart indicates aberrant autophagy and mitophagy. Consistent with these observations, our previous study reported that NRK-2 overexpression attenuates angiotensin II-induced mitochondrial membrane depolarization in human cardiomyocytes [16]. These observations emphasized NRK-2 as a regulator of mitochondrial function in the injured heart.
There are certain limitations of the current study. We cannot exclude the possibility that some observed differences may be due to undesired compensatory effects to the permanent deletion of the Nrk2 gene, where genes potentially become differently regulated during the growth and development of the animal. Though this possibility is less likely as NRK-2 is minimally expressed in healthy cardiac tissue, expression profoundly upregulates only upon cardiac stressors [15, 16]. Moreover, there is also a possibility that the metabolic alterations may partly be due to the activation of fibrosis in the KO heart post-MI. However, we selected the minimally fibrotic remote area of LV tissues for all analyses, which reduces the potential contribution of fibrosis to the metabolic remodeling of ischemic NRK-2 KO heart.
Collectively we show, for the first time, that NRK-2 facilitates cardiac metabolic switch in ischemic conditions. Our transcriptomic and metabolomic studies show that the loss of NRK-2 promotes cardiac fibrosis, mitochondrial dysfunction, and aberrant cardiac metabolism in the ischemic heart. The NRK-2 deficiency leads to the MI-induced upregulation of genes related to the ECM component; however, the downregulation of several genes linked to mitochondrial dysfunction, metabolic derangements, and impaired contraction. Importantly, we observed upregulations of mevalonic acid, DHPG, 2-penylbutyric acid, and uridine and downregulations of stearic acid, 8,11,14-eicosatrienoic acid, and 2-pyrrolidinone in the ischemic KO hearts. Therefore, NRK-2 lies at the nexus of mitochondrial function and metabolic remodeling in the ischemic heart and potentially be manipulated for the treatment of ischemia-induced heart failure.
Data availability
Data is included in the manuscript and supplementary files. All the raw data is deposited in the NCBI database with the accession number GSE223868.
References
Virani SS, Alonso A, Benjamin EJ, Bittencourt MS, Callaway CW, Carson AP, Chamberlain AM, Chang AR, Cheng S, Delling FN et al (2020) Heart disease and stroke statistics-2020 update: a report from the American Heart Association. Circulation 141:e139–e596. https://doi.org/10.1161/CIR.0000000000000757
Tran DH, Wang ZV (2019) Glucose metabolism in cardiac hypertrophy and heart failure. J Am Heart Assoc 8:e012673. https://doi.org/10.1161/JAHA.119.012673
Lal H, Ahmad F, Woodgett J, Force T (2015) The GSK-3 family as therapeutic target for myocardial diseases. Circ Res 116:138–149. https://doi.org/10.1161/CIRCRESAHA.116.303613
Lee L, Horowitz J, Frenneaux M (2004) Metabolic manipulation in ischaemic heart disease, a novel approach to treatment. Eur Heart J 25:634–641. https://doi.org/10.1016/j.ehj.2004.02.018
Kudo N, Barr AJ, Barr RL, Desai S, Lopaschuk GD (1995) High rates of fatty acid oxidation during reperfusion of ischemic hearts are associated with a decrease in malonyl-CoA levels due to an increase in 5’-AMP-activated protein kinase inhibition of acetyl-CoA carboxylase. J Biol Chem 270:17513–17520. https://doi.org/10.1074/jbc.270.29.17513
Lopaschuk GD, Belke DD, Gamble J, Itoi T, Schonekess BO (1994) Regulation of fatty acid oxidation in the mammalian heart in health and disease. Biochim Biophys Acta 1213:263–276. https://doi.org/10.1016/0005-2760(94)00082-4
Lopaschuk GD, Ussher JR, Folmes CD, Jaswal JS, Stanley WC (2010) Myocardial fatty acid metabolism in health and disease. Physiol Rev 90:207–258. https://doi.org/10.1152/physrev.00015.2009
Ahmad F, Woodgett JR (2020) Emerging roles of GSK-3alpha in pathophysiology: emphasis on cardio-metabolic disorders. Biochim Biophys Acta Mol Cell Res 1867:118616. S0167–4889(19)30224–1. https://doi.org/10.1016/j.bbamcr.2019.118616
Yamamoto T, Sano M (2022) Deranged myocardial fatty acid metabolism in heart failure. Int J Mol Sci 23. https://doi.org/10.3390/ijms23020996
Jiang M, Xie X, Cao F, Wang Y (2021) Mitochondrial metabolism in myocardial remodeling and mechanical unloading: implications for ischemic heart disease. Front Cardiovasc Med 8:789267. https://doi.org/10.3389/fcvm.2021.789267
Ahmad F, Singh AP, Tomar D, Rahmani M, Zhang Q, Woodgett JR, Tilley DG, Lal H, Force T (2019) Cardiomyocyte-GSK-3alpha promotes mPTP opening and heart failure in mice with chronic pressure overload. J Mol Cell Cardiol 130:65–75. https://doi.org/10.1016/j.yjmcc.2019.03.020
Lal H, Ahmad F, Zhou J, Yu JE, Vagnozzi RJ, Guo Y, Yu D, Tsai EJ, Woodgett J, Gao E et al (2014) Cardiac fibroblast glycogen synthase kinase-3beta regulates ventricular remodeling and dysfunction in ischemic heart. Circulation 130:419–430. https://doi.org/10.1161/CIRCULATIONAHA.113.008364
Israeli-Rosenberg S, Manso AM, Okada H, Ross RS (2014) Integrins and integrin-associated proteins in the cardiac myocyte. Circ Res 114:572–586. https://doi.org/10.1161/CIRCRESAHA.114.301275
Diguet N, Trammell SAJ, Tannous C, Deloux R, Piquereau J, Mougenot N, Gouge A, Gressette M, Manoury B, Blanc J et al (2018) Nicotinamide riboside preserves cardiac function in a mouse model of dilated cardiomyopathy. Circulation 137:2256–2273. https://doi.org/10.1161/CIRCULATIONAHA.116.026099
Ahmad F, Tomar D, Aryal ACS, Elmoselhi AB, Thomas M, Elrod JW, Tilley DG, Force T (2020) Nicotinamide riboside kinase-2 alleviates ischemia-induced heart failure through P38 signaling. Biochim Biophys Acta Mol Basis Dis 1866:165609. https://doi.org/10.1016/j.bbadis.2019.165609
Shahzadi SK, Marzook H, Qaisar R, Ahmad F (2022) Nicotinamide riboside kinase-2 inhibits JNK pathway and limits dilated cardiomyopathy in mice with chronic pressure overload. Clin Sci (Lond) 136:181–196. https://doi.org/10.1042/CS20210964
Zhou J, Ahmad F, Parikh S, Hoffman NE, Rajan S, Verma VK, Song J, Yuan A, Shanmughapriya S, Guo Y et al (2016) Loss of adult cardiac myocyte GSK-3 leads to mitotic catastrophe resulting in fatal dilated cardiomyopathy. Circ Res 118:1208–1222. https://doi.org/10.1161/CIRCRESAHA.116.308544
Zhou J, Ahmad F, Lal H, Force T (2016) Response by Zhou et al to letter regarding article, “loss of adult cardiac myocyte gsk-3 leads to mitotic catastrophe resulting in fatal dilated cardiomyopathy.” Circ Res 119:e29–e30. https://doi.org/10.1161/CIRCRESAHA.116.309093
Lal H, Zhou J, Ahmad F, Zaka R, Vagnozzi RJ, Decaul M, Woodgett J, Gao E, Force T (2012) Glycogen synthase kinase-3alpha limits ischemic injury, cardiac rupture, post-myocardial infarction remodeling and death. Circulation 125:65–75. https://doi.org/10.1161/CIRCULATIONAHA.111.050666
Ahmad F, Lal H, Zhou J, Vagnozzi RJ, Yu JE, Shang X, Woodgett JR, Gao E, Force T (2014) Cardiomyocyte-specific deletion of gsk3alpha mitigates post-myocardial infarction remodeling, contractile dysfunction, and heart failure. J Am Coll Cardiol 64:696–706. https://doi.org/10.1016/j.jacc.2014.04.068
Khan AA, Gul MT, Karim A, Ranade A, Azeem M, Ibrahim Z, Ramachandran G, Nair VA, Ahmad F, Elmoselhi A et al (2022) Mitigating sarcoplasmic reticulum stress limits disuse-induced muscle loss in hindlimb unloaded mice. NPJ Microgravity 8:24. https://doi.org/10.1038/s41526-022-00211-w
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287. https://doi.org/10.1089/omi.2011.0118
Gupte M, Lal H, Ahmad F, Sawyer DB, Hill MF (2017) Chronic neuregulin-1beta treatment mitigates the progression of postmyocardial infarction heart failure in the setting of type 1 diabetes mellitus by suppressing myocardial apoptosis, fibrosis, and key oxidant-producing enzymes. J Card Fail 23:887–899. https://doi.org/10.1016/j.cardfail.2017.08.456
Yusuf AM, Qaisar R, Al-Tamimi AO, Jayakumar MN, Woodgett JR, Koch WJ, Ahmad F (2022) Cardiomyocyte-GSK-3beta deficiency induces cardiac progenitor cell proliferation in the ischemic heart through paracrine mechanisms. J Cell Physiol 237:1804–1817. https://doi.org/10.1002/jcp.30644
Sharaf BM, Giddey AD, Alniss H, Al-Hroub HM, El-Awady R, Mousa M, Almehdi A, Soares NC, Semreen MH (2022) Untargeted metabolomics of breast cancer cells MCF-7 and SkBr 3 treated with tamoxifen/trastuzumab. Cancer Genomics Proteomics 19:79–93. https://doi.org/10.21873/cgp.20305
Pang Z, Chong J, Zhou G, de Lima Morais DA, Chang L, Barrette M, Gauthier C, Jacques PE, Li S, Xia J (2021) MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res 49:W388–W396. https://doi.org/10.1093/nar/gkab382
Talman V, Ruskoaho H (2016) Cardiac fibrosis in myocardial infarction-from repair and remodeling to regeneration. Cell Tissue Res 365:563–581. https://doi.org/10.1007/s00441-016-2431-9
Zuurbier CJ, Bertrand L, Beauloye CR, Andreadou I, Ruiz-Meana M, Jespersen NR, Kula-Alwar D, Prag HA, Eric Botker H, Dambrova M et al (2020) Cardiac metabolism as a driver and therapeutic target of myocardial infarction. J Cell Mol Med 24:5937–5954. https://doi.org/10.1111/jcmm.15180
Chen C, Li R, Ross RS, Manso AM (2016) Integrins and integrin-related proteins in cardiac fibrosis. J Mol Cell Cardiol 93:162-174. S0022-2828(15)30115-2. https://doi.org/10.1016/j.yjmcc.2015.11.010
Schwartz MA, Ginsberg MH (2002) Networks and crosstalk: integrin signalling spreads. Nat Cell Biol 4:E65-68. https://doi.org/10.1038/ncb0402-e65
Schroer AK, Merryman WD (2015) Mechanobiology of myofibroblast adhesion in fibrotic cardiac disease. J Cell Sci 128:1865–1875. https://doi.org/10.1242/jcs.162891
Brancaccio M, Fratta L, Notte A, Hirsch E, Poulet R, Guazzone S, De Acetis M, Vecchione C, Marino G, Altruda F et al (2003) Melusin, a muscle-specific integrin β1-interacting protein, is required to prevent cardiac failure in response to chronic pressure overload. Nat Med 9:68–75
Su Z, Liu Y, Zhang H (2021) Adaptive cardiac metabolism under chronic hypoxia: mechanism and clinical implications. Front Cell Dev Biol 9:625524. https://doi.org/10.3389/fcell.2021.625524
Mozaffarian D, Katan MB, Ascherio A, Stampfer MJ, Willett WC (2006) Trans fatty acids and cardiovascular disease. N Engl J Med 354:1601–1613. https://doi.org/10.1056/NEJMra054035
Marin-Garcia J, Goldenthal MJ (2002) Fatty acid metabolism in cardiac failure: biochemical, genetic and cellular analysis. Cardiovasc Res 54:516–527. https://doi.org/10.1016/s0008-6363(01)00552-1
Wang W, Ledee D (2021) ACAA2 is a ligand-dependent coactivator for thyroid hormone receptor beta1. Biochem Biophys Res Commun 576:15–21. https://doi.org/10.1016/j.bbrc.2021.08.073
Martines AMF, van Eunen K, Reijngoud DJ, Bakker BM (2017) The promiscuous enzyme medium-chain 3-keto-acyl-CoA thiolase triggers a vicious cycle in fatty-acid beta-oxidation. PLoS Comput Biol 13:e1005461. https://doi.org/10.1371/journal.pcbi.1005461
Liang X, Zhu D, Schulz H (1999) Delta 3,5,7, Delta2,4,6-trienoyl-CoA isomerase, a novel enzyme that functions in the beta-oxidation of polyunsaturated fatty acids with conjugated double bonds. J Biol Chem 274:13830–13835. https://doi.org/10.1074/jbc.274.20.13830
Mao X, Huang D, Rao C, Du M, Liang M, Li F, Liu B, Huang K (2020) Enoyl coenzyme A hydratase 1 combats obesity and related metabolic disorders by promoting adipose tissue browning. Am J Physiol Endocrinol Metab 318:E318–E329. https://doi.org/10.1152/ajpendo.00424.2019
Rai A, Kumar V, Jerath G, Kartha CC, Ramakrishnan V (2021) Mapping drug-target interactions and synergy in multi-molecular therapeutics for pressure-overload cardiac hypertrophy. NPJ Syst Biol Appl 7:11. https://doi.org/10.1038/s41540-021-00171-z
Abdulhag UN, Soiferman D, Schueler-Furman O, Miller C, Shaag A, Elpeleg O, Edvardson S, Saada A (2015) Mitochondrial complex IV deficiency, caused by mutated COX6B1, is associated with encephalomyopathy, hydrocephalus and cardiomyopathy. Eur J Hum Genet 23:159–164. https://doi.org/10.1038/ejhg.2014.85
McGarrah RW, Crown SB, Zhang GF, Shah SH, Newgard CB (2018) Cardiovascular metabolomics. Circ Res 122:1238–1258. https://doi.org/10.1161/CIRCRESAHA.117.311002
de Couto G, Ouzounian M, Liu PP (2010) Early detection of myocardial dysfunction and heart failure. Nat Rev Cardiol 7:334–344. https://doi.org/10.1038/nrcardio.2010.51
Bathaie SZ, Ashrafi M, Azizian M, Tamanoi F (2017) Mevalonate pathway and human cancers. Curr Mol Pharmacol 10:77–85. https://doi.org/10.2174/1874467209666160112123205
Xu H, Shen Y, Liang C, Wang H, Huang J, Xue P, Luo M (2021) Inhibition of the mevalonate pathway improves myocardial fibrosis. Exp Ther Med 21:224. https://doi.org/10.3892/etm.2021.9655
Goldstein JL, Brown MS (1990) Regulation of the mevalonate pathway. Nature 343:425–430. https://doi.org/10.1038/343425a0
Yang Y, Rong X, Lv X, Jiang W, Lai D, Xu S, Fu G (2017) Inhibition of mevalonate pathway prevents ischemia-induced cardiac dysfunction in rats via RhoA-independent signaling pathway. Cardiovasc Ther 35. https://doi.org/10.1111/1755-5922.12285
Howes LG, Hodsman GP, Rowe PR, Sumithran E, Johnston CI (1989) Cardiac 3,4-dihydroxyphenylethylene glycol (DHPG) and catecholamine levels during perindopril therapy of chronic left ventricular failure in rats. J Auton Pharmacol 9:15–21. https://doi.org/10.1111/j.1474-8673.1989.tb00192.x
Bohner H, Janiak PS, Nitsche V, Eichinger A, Schutz H (1997) Relative bioavailability of different butamirate citrate preparations after single dose oral administration to 18 healthy volunteers. Int J Clin Pharmacol Ther 35:117–122
Cao JY, Parker B, Koay YC, Lal S, O’Sullivan JF (2018) Identification of new pathways in human ischemic myocardium. Circulation 138:A12816
Krylova IB, Selina EN, Bulion VV, Rodionova OM, Evdokimova NR, Belosludtseva NV, Shigaeva MI, Mironova GD (2021) Uridine treatment prevents myocardial injury in rat models of acute ischemia and ischemia/reperfusion by activating the mitochondrial ATP-dependent potassium channel. Sci Rep 11:16999. https://doi.org/10.1038/s41598-021-96562-7
Guo M, Fan X, Tuerhongjiang G, Wang C, Wu H, Lou B, Wu Y, Yuan Z, She J (2021) Targeted metabolomic analysis of plasma fatty acids in acute myocardial infarction in young adults. Nutr Metab Cardiovasc Dis 31:3131–3141. https://doi.org/10.1016/j.numecd.2021.06.024
Renaud SC (1992) What is the epidemiologic evidence for the thrombogenic potential of dietary long-chain fatty acids? Am J Clin Nutr 56:823S-824S. https://doi.org/10.1093/ajcn/56.4.823s
Zhuang L, Mao Y, Liu Z, Li C, Jin Q, Lu L, Tao R, Yan X, Chen K (2021) FABP3 deficiency exacerbates metabolic derangement in cardiac hypertrophy and heart failure via pparalpha pathway. Front Cardiovasc Med 8:722908. https://doi.org/10.3389/fcvm.2021.722908
Furuhashi M, Hotamisligil GS (2008) Fatty acid-binding proteins: role in metabolic diseases and potential as drug targets. Nat Rev Drug Discov 7:489–503. https://doi.org/10.1038/nrd2589
Lee SM, Lee SH, Jung Y, Lee Y, Yoon JH, Choi JY, Hwang CY, Son YH, Park SS, Hwang GS et al (2020) FABP3-mediated membrane lipid saturation alters fluidity and induces ER stress in skeletal muscle with aging. Nat Commun 11:5661. https://doi.org/10.1038/s41467-020-19501-6
Varrone F, Gargano B, Carullo P, Di Silvestre D, De Palma A, Grasso L, Di Somma C, Mauri P, Benazzi L, Franzone A et al (2013) The circulating level of FABP3 is an indirect biomarker of microRNA-1. J Am Coll Cardiol 61:88–95. https://doi.org/10.1016/j.jacc.2012.08.1003
Li A, Zheng N, Ding X (2022) Mitochondrial abnormalities: a hub in metabolic syndrome-related cardiac dysfunction caused by oxidative stress. Heart Fail Rev 27:1387–1394. https://doi.org/10.1007/s10741-021-10109-6
Rose BA, Force T, Wang Y (2010) Mitogen-activated protein kinase signaling in the heart: angels versus demons in a heart-breaking tale. Physiol Rev 90:1507–1546. https://doi.org/10.1152/physrev.00054.2009
Li J, Rao H, Burkin D, Kaufman SJ, Wu C (2003) The muscle integrin binding protein (MIBP) interacts with alpha7beta1 integrin and regulates cell adhesion and laminin matrix deposition. Dev Biol 261:209–219
McNally EM, Mestroni L (2017) Dilated Cardiomyopathy: genetic determinants and mechanisms. Circ Res 121:731–748. https://doi.org/10.1161/CIRCRESAHA.116.309396
Gomez-Sanchez R, Yakhine-Diop SM, Bravo-San Pedro JM, Pizarro-Estrella E, Rodriguez-Arribas M, Climent V, Martin-Cano FE, Gonzalez-Soltero ME, Tandon A, Fuentes JM et al (2016) PINK1 deficiency enhances autophagy and mitophagy induction. Mol Cell Oncol 3:e1046579. https://doi.org/10.1080/23723556.2015.1046579
Kuhn C, Menke M, Senger F, Mack C, Dierck F, Hille S, Schmidt I, Brunke G, Bunger P, Schmiedel N et al (2021) FYCO1 Regulates Cardiomyocyte Autophagy and Prevents Heart Failure Due to Pressure Overload In Vivo. JACC Basic Transl Sci 6:365–380. https://doi.org/10.1016/j.jacbts.2021.01.001
Funding
The work was supported by Collaborative (22010901112) and Targeted (1801090144), research grants from the University of Sharjah to Firdos Ahmad. R.H. is funded by ASPIRE “Abu Dhabi Precision Medicine ARI” (VRI-20-10) grant.
Author information
Authors and Affiliations
Contributions
HM performed experiments, collected data, and helped with manuscript writing, AG performed experiments and collected data, MAS, KP, MHS, and RH helped with data analysis and interpretation, DT and NCS helped with experiments and data analysis, RQ helped with data analysis and interpretation and manuscript writing, and FA designed the study, acquired funding, supervised the project, performed data analysis and interpretation, and wrote first draft and revised the manuscript.
Corresponding author
Ethics declarations
Ethical approval
The Institutional Animal Care and Use Committee (IACUC) of Vanderbilt University Medical Center approved all animal procedures and treatments.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Marzook, H., Gupta, A., Tomar, D. et al. Nicotinamide riboside kinase-2 regulates metabolic adaptation in the ischemic heart. J Mol Med 101, 311–326 (2023). https://doi.org/10.1007/s00109-023-02296-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-023-02296-6