Abstract
Amphiphysin 2, also named bridging integrator-1 (BIN1) or SH3P9, has been recently implicated in rare and common diseases affecting different tissues and physiological functions. BIN1 downregulation is linked to cancer progression and also correlates with ventricular cardiomyopathy and arrhythmia preceding heart failure. Increased BIN1 expression is linked to increased susceptibility for late-onset Alzheimer’s disease. In addition, altered splicing may account for the muscle component of myotonic dystrophies, while recessive germinal mutations cause centronuclear myopathy. Despite undoubtedly underlining the relevance of BIN1 in human diseases, the molecular and cellular bases leading to such different diseases are unclear at present. BIN1 is a key regulator of endocytosis and membrane recycling, cytoskeleton regulation, DNA repair, cell cycle progression, and apoptosis. In light of the recent findings on the molecular, cellular, and physiological roles of BIN1, we discuss potential pathological mechanisms and highlight common disease pathways and also tissue-specific regulation. Next challenges will be to validate BIN1 both as a prognostic marker for the related diseases and as a potential therapeutic target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amphiphysin 2, also named BIN1 (MYC box-dependent interacting protein-1 or bridging integrator-1) or SH3P9, is an ubiquitously expressed nucleocytoplasmic protein that was identified as a binding partner of the transcription factor MYC and in parallel as an interactor of a tyrosine kinase SRC SH3 ligand peptide.
BIN1 is a key regulator of different cellular functions, including endocytosis and membrane recycling, cytoskeleton regulation, DNA repair, cell cycle progression, and apoptosis. BIN1 alteration is of strong medical relevance as it was linked to cancer progression, several myopathies, heart failure, and late-onset Alzheimer’s disease. Here, we focus on the different roles of BIN1 in specific tissues, in relation to BIN1-associated human diseases.
BIN1 gene organization and expression
BIN1 is located on human chromosome 2q14 and mouse 18q32. The human and mouse coding sequences have high DNA and protein sequence homology (89 and 95 %, respectively) [1]. Human Bin1/amphiphysin 2 and amphiphysin 1 are members of the amphiphysin family. Amphiphysin 1 and 2 share 49 % amino acid sequence homology [2, 3]. Amphiphysin 1 is a neuronal protein involved in synaptic vesicle endocytosis [4]. Orthologues have been studied in Caenorhabditis elegans, drosophila, and zebrafish. The amphiphysins are structurally related to yeast RVS 167, which is involved in endocytosis [5] and is a negative regulator of the cell cycle [6].
BIN1 is ubiquitously expressed with highest expression in skeletal muscle and brain [7, 8]. The regulatory sequence in the promoter region of human BIN1 encompasses predicted myogenin/myoD, Sp1, and serum response factor (SRF) binding sites, providing a molecular explanation for the high expression of BIN1 in striated muscle [1]. Sp1 and SRF sites are conserved in mouse, whereas the myogenin/myoD sites are substituted by Mef2 and Tef sites [1]. Moreover, in both human and mouse, a strong NF-kB binding site is conserved [1]. Additionally, E2F1 and c-MYC were found to modulate transcription of BIN1 [9, 10].
The BIN1 gene has 20 exons which can be spliced into multiple isoforms (Fig. 1). Exons 7, 13, 14, 15, and 16 are brain specific [8, 11, 12]; whereas, exon 11 is a muscle-specific exon [1, 13, 14]. Mis-splicing of exons 7 and 11 was identified in muscles from myotonic dystrophy patients [15], while exon 13 is often mis-spliced in cancer resulting in lack of c-MYC binding [16]. Isoforms 1–7 are expressed in brain. The muscle-specific isoform is isoform 8; whereas, isoforms 9 and 10 are ubiquitously expressed.
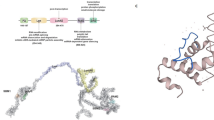
BIN1 functional domains and tissue-specific isoforms. a Protein domains of BIN1: The N-BAR domain is involved in membrane binding and tubulation; The PI motif is skeletal muscle-specific, and it is involved in phosphoinositides binding; The PS motif is a Proline-Serine rich region encoded by exon 12 and present in all isoforms; The CLAP domain is involved in clathrin and AP2 binding and is only present in neuronal isoforms; The MBD domain binds c-Myc, and it is involved in its regulation; and the SH3 domain interacts with Proline-rich domains found in proteins such as dynamin 2. b Gene organization of BIN1. The main nomenclature used is from NCBI and in italic is the nomenclature by Weschler-Reya et al.1997. Transcript isoforms found in brain (c), skeletal muscle (d), most of other tissues (e), and in melanoma (f). Alternative exons are indicated
BIN1 domain organization and protein regulation
BIN1 has several distinctive protein domains (Fig. 1). The N-BAR (BIN-amphiphysin/Rvs) domain encoded by exons 1 to 10 is ubiquitously expressed, binds lipid membranes in a dimer conformation, and has a role in generating and sensing membrane curvature [17, 18]. The N-terminal amphipathic helix inserts into the membrane and promotes curvature, while the positively charged surface of the BAR domain interacts with the negatively charged membrane to sense and maintain the curvature [18]. Dimerization of the BAR domain is a prerequisite for this function. The BAR domain is involved in homo or hetero dimerization (with amphiphysin 1) [19, 20], and the deletion of AMPH1 lead to loss of both AMPH1 and BIN1 in brain [21].
The phosphoinositide (PI) binding motif is a short polybasic sequence encoded by the muscle exon 11 [8, 13, 22]. The PI motif increases BIN1 affinity towards the negatively charged lipids PtdIns (4, 5) P2 [23] and/or PtdIns3P and PtdIns5P [15]. This motif may potentially target BIN1 to membrane compartments such as T-tubules, muscle-specific plasma membrane invaginations. Additionally, it could provide a muscle-specific regulation of BIN1 conformation through binding to the SH3 domain and phosphoinositides [24].
The clathrin and AP2 (CLAP) binding domain encoded by exons 13–16 is found only in brain isoforms [8, 11, 25], and it is responsible for binding to endocytic proteins clathrin and AP2 [19].
The Myc-binding domain (MBD) is encoded by exons 17 and 18 [7]. Alternative splicing of exon 17 leads to the loss of this interaction [26] [13].
The Src homology 3 (SH3) domain encoded by the last two exons (19 and 20) is present in all isoforms. SH3 domains bind proline-rich motifs [27]. The SH3 domains of amphiphysin 1 and 2 differ from other SH3 domains due to a large patch of negative electrostatic potential and an unusually extended n-Src loop [3].
BIN1 functions are regulated through specific exon splicing. Open and closed conformations of BIN1 have been suggested to be regulated by inclusion of the PI motif and integrity of the SH3 domain [24, 28]. In isoform 8, the BAR + PI domains bind the SH3 domain intramolecularly and compete with SH3 interactors, and PtdIns (4, 5) P2 binding to the PI motif induces the open conformation and interaction with protein effectors as dynamin on PtdIns (4, 5) P2 enriched membranes [24]. BIN1 mutations causing centronuclear myopathy and found in the SH3 domain affect BIN1 conformation and binding to dynamin [22, 28]. Differential splicing of exon 7 in the BAR domain also modulates protein-protein interaction with dynamin [12]. In addition, BIN1 may also be regulated by phosphorylation [29, 30].
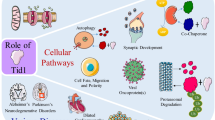
BIN1 cellular functions
Membrane trafficking and membrane remodeling
The most characterized function of amphiphysins is in endocytosis. Amphiphysins are involved in endocytosis in neuronal and non-neuronal cells [4, 31–33] through interaction with several proteins associated with clathrin-coated pits: dynamin [31], AP2 adaptor complexes [31, 33], clathrin [25, 34], synaptojanin [35], and endophilin/SH3GL2 [36, 37] (Table 1). Noteworthy, dephosphorylation of amphiphysin 1 and BIN1 by calcineurin is necessary to induce endocytosis [29]. Moreover, exogenous expression of the amphiphysin SH3 domain alone inhibits dynamin-dependent transferrin uptake [3]. The mutual exclusivity for clathrin or dynamin binding may be correlated to their non-overlapping sites at the clathrin-coated pit [34]. Thus, it is believed that amphiphysins orchestrate the sequential recruitment and action of effectors in parallel to membrane deformation, at sites of membrane fission [38].
However, knockdown of BIN1 in HeLa cells or knockout (KO) in MEF cells did not alter significantly transferrin uptake, but increased the intracellular transferrin levels due to defects in transferrin receptor recycling [39, 40]. Similarly, the only C. elegans orthologue named AMPH-1 colocalizes and directly interacts with RME-1 (dynamin-like protein and EHD orthologue) to initiate endosome tubulation and membrane recycling [39]. Thus, depending on the context, amphiphysins control endocytosis and/or recycling (Fig. 2). Functional interaction of EHD1 and BIN1 is further sustained by the finding that EHD1 mediates vesicle trafficking and development or maintenance of T-tubules in muscle; EHD1 loss impairs T-tubule structure and BIN1 localization [41].
BIN1 cellular roles and proposed mechanisms of linked diseases. BIN1 is implicated in membrane remodeling for intracellular trafficking (top left), or for T-tubule maturation or maintenance (top right). BIN1 also directly regulates c-MYC nuclear function and proteins implicated in DNA repair (bottom), leading to a tight balance between proliferation and apoptosis
The role of BIN1 in membrane trafficking reflects its ability to regulate membrane remodeling. In vitro studies have shown that the N-BAR domain of amphiphysin and endophilin binds and evaginates lipid membranes into narrow tubules [42–44]. Exogenously expressed amphiphysin 1 or the brain isoform of BIN1 in Chinese hamster ovary (CHO) cells exhibit diffused and cytosolic distribution. Conversely, overexpression of BIN1 muscle isoform 8 in CHO and C2C12 muscle cells leads to its strong concentration at the cell surface and formation of numerous narrow tubules that are prevented by cholesterol depletion. Electron microscopy (EM) and correlative microscopy identified that these membrane tubules are connected to the plasma membrane [23, 45], resembling the T-tubules found in skeletal muscle. Deletion of the PI motif strongly decreases membrane tubulation [15, 46, 47], implying that the PI motif promotes BIN1-dependent membrane tubulation. Overall, a tight regulation of BIN1 controls the balance between membrane fission and trafficking versus membrane tubulation, most probably in a tissue-specific manner.
Cytoskeleton network
Several studies have suggested a potential role for BIN1 in regulating the actin cytoskeleton. In particular, amphiphysin 1 directly binds N-WASP [48] and regulates actin polymerization during phagocytosis [49]. Moreover, the yeast amphiphysin orthologue RVS167 is involved in actin dynamics regulation in response to environmental signals [6, 50].
BIN1 may also modulate the microtubule (MT) network thought binding via its BAR domain to the coiled-coil region of CLIP170, a plus-end protein involved in MT stability [46]. BIN1 membrane tubulation in HeLa cells is sensitive to nocodazole (MT depolymerizing agent), and depletion of CLIP170 decreased BIN1 tubulation capacity [46]. Moreover, in human neuroblastoma cells and in mouse brain, BIN1 colocalizes and interacts with TAU, a microtubule-associated protein promoting microtubule assembly and stabilization and implicated in Alzheimer’s disease (AD) [51]. While mechanistic data are needed, these studies support the hypothesis that BIN1 coordinates membrane and cytoskeleton remodeling.
DNA repair, cell cycle, and apoptosis
In addition to cytosolic functions, BIN1 has several roles in the nucleus (Fig. 2). BIN1 isoform 9 that lacks the PI motif and maintains a functional MYC-binding domain (without exon 11 and with exon 17) was reported to localize mainly in the nucleus of proliferating cultured cells [26, 52]. Indeed, the MBD of BIN1 interacts with the MB1 domain of c-MYC, a transcription factor with the basic helix-loop-helix-leucine zipper (bHLH-LZ) structure that plays a central role in cell growth, apoptosis, and malignancy [7]. Only isoforms of BIN1 which localize in the nucleus can activate programmed cell death [16, 53].
Interestingly, overexpression of BIN1 in malignant but not in non-malignant cells induced apoptosis [53, 54]. BIN1 can inhibit cell transformation in a caspase-independent cell death process through both Myc-dependent and Myc-independent mechanisms [55]. The cell death program engaged by BIN1 is susceptible to the serine protease inhibitor AEBSF and to inhibition by the SV40 large T antigen [53]. In addition, BIN1 interacts with the proto-oncogene c-ABL, and BIN1 overexpression results in morphological transformation of NIH 3T3 fibroblasts in a c-ABL-dependent manner [56]. Moreover, BIN1 is part of the transforming pathway induced by the adenovirus gene product E1A. E1A can inactivate the tumor suppressor retinoblastoma protein (RB), leading to the decreased release of the DNA-binding factor E2F1 and the subsequent failure in activation of BIN1 transcription [9, 57], which again leads to the decreased inhibition of c-MYC and potentially impaired cell cycle control. Interestingly, while BIN1 interacts with and inhibits c-MYC [10], BIN1 may also impact directly on the activation of c-MYC target genes. Additionally, in transformed cells, c-MYC can repress BIN1 transcription which further facilitates cell transformation [10].
Other studies suggested that BIN1 plays an important role in DNA repair by binding and inhibiting poly (ADP-ribose) polymerase (PARP1), a key component of the base excision repair pathway, via the BAR domain [10, 58]. Depletion of endogenous BIN1 abolishes cisplatin-induced cell death via activation of c-MYC and PARP1 [10]. BIN1 downregulation also reduces cell sensitivity to the DNA-damaging chemotherapeutic agents etoposide and doxorubicin [10]. BIN1-dependent decrease of intrinsic PARP1 activity may sensitize cancer cells to DNA damage [10]. In addition, BIN1 interacts with two other proteins important for DNA repair in the non-homologous end-joining (NHEJ) pathway, Ku and XRCC4 [59, 60], strengthening a role for BIN1 as a regulator of DNA repair.
Additional evidence of a role for BIN1 in cell cycle regulation came from studies of the fission yeast homologue hob1 (homologue of Bin1). Hob1p was shown to be dispensable for actin organization and endocytosis but mutants were hypersensitive to starvation or genotoxic stress [61]. In particular, hob1 mutants were more susceptible to DNA damage and failed to undergo G1 arrest after DNA damage [61]. BIN1 but not amphiphysin 1 rescued this phenotype, confirming an evolutionary conserved role for BIN1 in cell cycle regulation [61].
Physiological functions of BIN1 and physiopathology of related diseases
BIN1 and cancer
BIN1 expression is reduced or altered in several cancer types including breast, colon, prostate and lung cancers, hepatocarcinoma, and neuroblastoma [62–65]. Loss of heterozygosity or mis-splicing of BIN1 also correlates with cancer prognosis and increased metastasis [16, 65–67].
In a mosaic Bin1-null mouse, mammary gland specific Bin1 deletion delays the differentiation of the glandular ductal network during pregnancy but does not increase tumor susceptibility [62]. Nevertheless, when tumor formation is initiated by the carcinogen DMBA, mice lacking Bin1 developed more aggressive tumors which are characterized by an increase in proliferation, survival, and motility [62].
When present at a normal level, BIN1 can inhibit MYC-dependent transformation and tumor growth [7, 68]. BIN1 also regulates another member of MYC family of oncogenes, N-Myc [69], which contains the N-terminal domain and MYC box 1 needed for the interaction with BIN1 [7]. N-Myc is highly expressed in neurons and overexpressed in aggressive neuroblastoma [70, 71]. In 6/8 neuroblastoma cell lines overexpressing N-Myc, BIN1 is strongly downregulated, while its overexpression in these cells leads to apoptosis [72].
Decrease of BIN1 may contribute to malignant development via the loss of processes required for terminal differentiation (through downregulation of p21 and p53), and by contributing to Myc deregulation and loss of control over NHEJ repair. Cisplatin-based chemotherapy is a commonly used approach for the various human malignancies; however, many cancer patients respond poorly to this treatment [73]. BIN1 downregulation was shown to increase resistance to this therapy, most likely through loss of control over the DNA repair mechanism [10]. Quantification of BIN1 expression may thus be an important marker to predict cisplatin sensitivity. Additionally, increasing the expression of BIN1 may be a novel therapeutic strategy for treatment of cisplatin-resistant cancers [74].
In addition to the nuclear functions discussed above, BIN1 could be implicated in tumor cell migration and invasion through cytoskeleton and membrane remodeling. Altogether, it appears that BIN1 decrease does not strongly increase the risk of tumor formation but favors tumor progression [75].
BIN1 in skeletal muscle and myopathies
BIN1 is implicated in two myopathies: centronuclear myopathy (CNM) and myotonic dystrophy (DM). Germline mutations cause autosomal recessive centronuclear myopathy (ARCNM), a rare congenital disease associated with non-progressive muscle weakness with onset at birth or infancy [22, 47, 76]. Patient muscle presents histologically with increased centralized nuclei and fiber atrophy not linked to excessive muscle regeneration [47, 77]. No history of cancer has been noted in patients, and potential cardiac involvement was reported in only one patient [22, 47, 76, 78]. To date, the following homozygous mutations have been reported: missense changes in the amphipathic helix and BAR domain (K35N, D151N, and R154C) [22, 76], stop codon mutations in the last exon truncating the SH3 domain [22, 78], and a mutation in the exon 11 donor splice site leading to exon 11 skipping [47]. While BIN1 expression appears not altered, mutations in N-BAR or skipping of the PI motif strongly decreases the membrane tubulation properties of BIN1, while the SH3 truncations impair both intramolecular binding and binding to dynamin 2. Noteworthy, dynamin 2 is a GTPase implicated in cytoskeleton and membrane remodeling and mutated in autosomal dominant CNM [79].
Myotonic dystrophies are multisystemic diseases characterized by progressive muscle wasting, myotonia, cataracts, and heart conduction defects. Congenital myotonic dystrophy (CDM1) and myotonic dystrophy of type 1 (DM1) or of type 2 (DM2) are caused by the expression of mutant RNAs containing expanded CUG or CCUG repeats that sequester splicing factors as muscleblind-like-1 (MBNL1), resulting in aberrant splicing of other mRNAs, including BIN1. In DM patient muscles, the BIN1 embryonic isoform containing exon 7 and lacking exon 11 is persistently expressed [15]. As centronuclear myopathy and myotonic dystrophy share several clinical and histopathological features such as muscle weakness and centralized myonuclei, alteration of BIN1 represents a potential common molecular mechanism.
BIN1 is highly expressed in skeletal muscles, where expression increases during in vitro differentiation of muscle cells, and Bin1 knockdown inhibits myoblast fusion and differentiation [7, 8, 23, 26]. The inclusion of the muscle-specific PI motif during muscle differentiation and its importance for membrane tubulation also indicate a significant role for BIN1 in membrane remodeling in muscles [8, 26]. In addition, BIN1 localizes to the triad, the membrane structure sustaining the excitation-contraction (EC) coupling machinery [8].
Several studies in animal models revealed the physiological importance of BIN1 in skeletal muscles. Drosophila with a deletion of the unique amphiphysin orthologue (amph) is viable but flightless and generally sluggish [44]. Drosophila AMPH is localized on the postsynaptic membrane of the neuromuscular junction and on the T-tubule network, and adult amph mutants have severe defects in T-tubule organization and EC coupling, without any strong defects in synaptic vesicle endocytosis or recycling, suggesting Drosophila AMPH is functionally closer to mammalian BIN1 than to AMPH1 [44]. Concordantly, knockdown of Bin1 in adult mouse flexor digitorum brevis (FDB) muscle leads to alteration of T-tubule structure and Ca2+ levels [80]. Similarly, U7-antisense-induced exon 11 skipping in mouse tibialis anterior induces a defect in T-tubule organization and DHPR distribution and a decrease in muscle strength while muscle mass and fiber size are unaffected [15]. Recently, a BIN1 canine model was reported with a spontaneous mutation affecting exon 11 splicing, reducing the overall BIN1 expression and resulting in the inherited myopathy of Great Danes, a late onset and very progressive myopathy [47]. A high number of centrally located nuclei was detected, reminiscent of CNM, associated with abnormal sarcolemmal invaginations and structural defects in T-tubules [47]. T-tubule defects and altered localization of T-tubule markers (RYR1, DHPR, and CAV3) are also noted in patients with CNM [14, 47]. Taking together the molecular data indicating BIN1 is a membrane tubulating protein and the consistent data from animal models and patients, it appears BIN1 is essential in the positioning and remodeling of T-tubules. BIN1 may also have additional roles in skeletal muscle, for example in sarcomere organization, as suggested through transgenic overexpression of the BIN1 SH3 domain in mice [81], or in autophagosome maturation [47].
BIN1 in heart and cardiac failure
Decreased BIN1 expression was observed in patients with ventricular arrhythmia. Plasma BIN1 level appears correlated with disease progression and advanced heart failure in patients, although these recent findings await confirmation [82, 83]. Moreover, BIN1 mRNA and protein expression was significantly decreased in failing cardiomyocytes [84]. BIN1 also localizes to T-tubules in cardiac muscle and knockdown in cardiomyocytes reduces the distribution of DHPR (CAV1.2) to the T-tubules and delays calcium influx [85]. Thus, in addition to a structural role at the T-tubules, BIN1 may regulate DHPR transport to the T-tubules.
Preliminary data sustained a physiological role for BIN1 in heart function. Injection of a bin1 antisense morpholino in zebrafish induces a strong cardiac phenotype, with altered calcium transient and contractility and a significant decrease in heart rate [84]. Complete KO of Bin1 in mice is perinatally lethal [40]. An increased thickness and occlusion of both ventricular chambers was identified, suggesting mice die from ventricular cardiomyopathy. As genetic decrease of Bin1 specifically in mouse skeletal muscle also leads to perinatal death (Prokic et al, unpublished), the contribution of the cardiac phenotype to survival remains to be investigated.
BIN1 in brain and Alzheimer’s disease
Recently, large scale genome-wide association studies (GWAS) linked BIN1 to late-onset Alzheimer’s disease (LOAD) [86–93]. Alzheimer’s disease (AD) is the most common cause of dementia, affecting 13 % of the population older than 65 years of age and 30–50 % of the population older than 80 years. LOAD is characterized by a large number of senile plaques and neurofibrillary tangles in the brain [94]. BIN1 is the second most important susceptibility locus for LOAD after apolipoprotein E (APOE) (http://www.alzgene.org/) [95]. SNPs upstream of BIN1 correlate with a higher risk to develop LOAD, and to increased BIN1 transcript levels [51, 95]. Altered expression of BIN1 was demonstrated in aging mice, transgenic AD models, and AD brains. Whether BIN1 may be a biomarker or a therapeutic target for Alzheimer’s disease remains to be investigated. Mammalian models to confirm the implication of BIN1 in Alzheimer’s disease and investigate the normal role of BIN1 in mammalian brain are required.
BIN1 was shown to interact with the microtubule-associated protein TAU (MAPT) in drosophila, and while altered amph expression did not modify the Aβ42-induced rough eye and neurodegeneration phenotype, loss of amph was able to suppress TAU-induced neurotoxicity [51]. The authors hypothesized BIN1 overexpression could disrupt vesicle transport at the synapse, promoting TAU sequestration at the membrane. Importantly, these results associate BIN1 modulation of the AD pathogenesis with TAU signaling. BIN1 and several proteins recently linked to AD appear interconnected through multiple interacting proteins and share related functions on membrane trafficking, sustaining a potential central role of BIN1 in AD [96]. A recent study linked PtdIns3P, a regulator of membrane trafficking, with AD [97]. Interestingly, PtdIns3P is also implicated in CNM as the PtdIns3P phosphatase myotubularin is mutated in X-linked CNM [98]. These findings suggest a PtdIns3P-BIN1 signaling pathway is common to both CNM (muscle) and AD (brain).
Conclusion and pending questions
BIN1 is linked to several human diseases affecting different tissues. Downregulation is linked to increased cancer progression and may correlate with ventricular cardiomyopathy and arrhythmia preceding heart failure, while increased expression increases susceptibility to develop late-onset Alzheimer’s disease. Altered splicing may account for the muscle compound of the multisystemic phenotypes in myotonic dystrophies, and germline mutations cause congenital centronuclear myopathy. While it undoubtedly underlines the medical importance of BIN1 in human diseases, the molecular and cellular bases leading to such different diseases linked to alterations in BIN1 are unclear at present and represent an exciting area of future research. A more precise clarification of BIN1 isoforms, tissue-specific regulation, and interactors will be needed, together with the generation of mammalian models to test the pathological hypotheses summarized in this review. Future translational research should aim to validate BIN1 as a prognostic marker for the related diseases and as a potential therapeutic target.
References
Mao NC, Steingrimsson E, DuHadaway J, Wasserman W, Ruiz JC, Copeland NG, Jenkins NA, Prendergast GC (1999) The murine Bin1 gene functions early in myogenesis and defines a new region of synteny between mouse chromosome 18 and human chromosome 2. Genomics 56:51–58
Leprince C, Romero F, Cussac D, Vayssiere B, Berger R, Tavitian A, Camonis J (1997) A new member of the amphiphysin family connecting endocytosis and signal transduction pathways. FASEB J 11:A928–A928
Owen DJ, Wigge P, Vallis Y, Moore JDA, Evans PR, McMahon HT (1998) Crystal structure of the amphiphysin-2 SH3 domain and its role in the prevention of dynamin ring formation. EMBO J 17:5273–5285
Bauerfeind R, Takei K, De Camilli P (1997) Amphiphysin I is associated with coated endocytic intermediates and undergoes stimulation-dependent dephosphorylation in nerve terminals. J Biol Chem 272:30984–30992
Munn AL, Stevenson BJ, Geli MI, Riezman H (1995) End5, End6, and End7—mutations that cause actin delocalization and block the internalization step of endocytosis in Saccharomyces cerevisiae. Mol Biol Cell 6:1721–1742
Bauer F, Urdaci M, Aigle M, Crouzet M (1993) Alteration of a yeast SH3 protein leads to conditional viability with defects in cytoskeletal and budding patterns. Mol Cell Biol 13:5070–5084
Sakamuro D, Elliott KJ, WechslerReya R, Prendergast GC (1996) BIN1 is a novel MYC-interacting protein with features of a tumour suppressor. Nat Genet 14:69–77
Butler MH, David C, Ochoa GC, Freyberg Z, Daniell L, Grabs D, Cremona O, De Camilli P (1997) Amphiphysin II (SH3P9; BIN1), a member of the amphiphysin/Rvs family, is concentrated in the cortical cytomatrix of axon initial segments and nodes of Ranvier in brain and around T tubules in skeletal muscle. J Cell Biol 137:1355–1367
Cassimere EK, Pyndiah S, Sakamuro D (2009) The c-MYC-interacting proapoptotic tumor suppressor BIN1 is a transcriptional target for E2F1 in response to DNA damage. Cell Death Differ 16:1641–1653
Pyndiah S, Tanida S, Ahmed KM, Cassimere EK, Choe C, Sakamuro D (2011) c-MYC Suppresses BIN1 to Release Poly (ADP-Ribose) Polymerase 1: A Mechanism by Which Cancer Cells Acquire Cisplatin Resistance. Sci Signal 4. doi:10.1126/scisignal.2001556
Tsutsui K, Maeda Y, Tsutsui K, Seki S, Tokunaga A (1997) cDNA cloning of a novel amphiphysin isoform and tissue-specific expression of its multiple splice variants. Biochem Biophys Res Commun 236:178–183
Ellis JD, Barrios-Rodiles M, Colak R, Irimia M, Kim T, Calarco JA, Wang XC, Pan Q, O’Hanlon D, Kim PM et al (2012) Tissue-specific alternative splicing remodels protein-protein interaction networks. Mol Cell 46:884–892
WechslerReya R, Sakamuro D, Zhang J, Duhadaway J, Prendergast GC (1997) Structural analysis of the human BIN1 gene. Evidence for tissue-specific transcriptional regulation and alternate RNA splicing. J Biol Chem 272:31453–31458
Toussaint A, Cowling BS, Hnia K, Mohr M, Oldfors A, Schwab Y, Yis U, Maisonobe T, Stojkovic T, Wallgren-Pettersson C et al (2011) Defects in amphiphysin 2 (BIN1) and triads in several forms of centronuclear myopathies. Acta Neuropathol 121:253–266
Fugier C, Klein AF, Hammer C, Vassilopoulos S, Ivarsson Y, Toussaint A, Tosch V, Vignaud A, Ferry A, Messaddeq N et al (2011) Misregulated alternative splicing of BIN1 is associated with T tubule alterations and muscle weakness in myotonic dystrophy. Nat Med 17:720–725
Ge K, DuHadaway J, Du W, Herlyn M, Rodeck U, Prendergast GC (1999) Mechanism for elimination of a tumor suppressor: aberrant splicing of a brain-specific exon causes loss of function of Bin1 in melanoma. Proc Natl Acad Sci U S A 96:9689–9694
Frost A, Unger VM, De Camilli P (2009) The BAR domain superfamily: membrane-molding macromolecules. Cell 137:191–196
Peter BJ, Kent HM, Mills IG, Vallis Y, Butler PJ, Evans PR, McMahon HT (2004) BAR domains as sensors of membrane curvature: the amphiphysin BAR structure. Science 303:495–499
Ramjaun AR, McPherson PS (1998) Multiple amphiphysin II splice variants display differential clathrin binding: identification of two distinct clathrin-binding sites. J Neurochem 70:2369–2376
Slepnev VI, Ochoa GC, Butler MH, Grabs D, De Camilli P (1998) Role of phosphorylation in regulation of the assembly of endocytic coat complexes. Science 281:821–824
Di Paolo G, Sankaranarayanan S, Wenk MR, Daniell L, Perucco E, Caldarone BJ, Flavell R, Picciotto MR, Ryan TA, Cremona O et al (2002) Decreased synaptic vesicle recycling efficiency and cognitive deficits in amphiphysin 1 knockout mice. Neuron 33:789–804
Nicot AS, Toussaint A, Tosch V, Kretz C, Wallgren-Pettersson C, Iwarsson E, Kingston H, Garnier JM, Biancalana V, Oldfors A et al (2007) Mutations in amphiphysin 2 (BIN1) disrupt interaction with dynamin 2 and cause autosomal recessive centronuclear myopathy. Nat Genet 39:1134–1139
Lee E, Marcucci M, Daniell L, Pypaert M, Weisz OA, Ochoa GC, Farsad K, Wenk MR, De Camilli P (2002) Amphiphysin 2 (Bin1) and T-tubule biogenesis in muscle. Science 297:1193–1196
Kojima C, Hashimoto A, Yabuta I, Hirose M, Hashimoto S, Kanaho Y, Sumimoto H, Ikegami T, Sabe H (2004) Regulation of Bin1 SH3 domain binding by phosphoinositides. Embo J 23:4413–4422
Ramjaun AR, Micheva KD, Bouchelet I, McPherson PS (1997) Identification and characterization of a nerve terminal-enriched amphiphysin isoform. J Biol Chem 272:16700–16706
Wechsler-Reya RJ, Elliott KJ, Prendergast GC (1998) A role for the putative tumor suppressor Bin1 in muscle cell differentiation. Mol Cell Biol 18:566–575
Yu HT, Chen JK, Feng SB, Dalgarno DC, Brauer AW, Schreiber SL (1994) Structural basis for the binding of proline-rich peptides to SH3 domains. Cell 76:933–945
Royer B, Hnia K, Gavriilidis C, Tronchere H, Tosch V, Laporte J (2013) The myotubularin-amphiphysin 2 complex in membrane tubulation and centronuclear myopathies. EMBO Rep 14:907–915
Marks B, McMahon HT (1998) Calcium triggers calcineurin-dependent synaptic vesicle recycling in mammalian nerve terminals. Curr Bio 8:740–749
Hojlund K, Bowen BP, Hwang H, Flynn CR, Madireddy L, Geetha T, Langlais P, Meyer C, Mandarino LJ, Yi Z (2009) In vivo phosphoproteome of human skeletal muscle revealed by phosphopeptide enrichment and HPLC-ESI-MS/MS. J Proteome Res 8:4954–4965
David C, McPherson PS, Mundigl O, de Camilli P (1996) A role of amphiphysin in synaptic vesicle endocytosis suggested by its binding to dynamin in nerve terminals. Proc Natl Acad Sci USA 93:331–335
Shupliakov O, Low P, Grabs D, Gad H, Chen H, David C, Takei K, De Camilli P, Brodin L (1997) Synaptic vesicle endocytosis impaired by disruption of dynamin-SH3 domain interactions. Science 276:259–263
Wigge P, Kohler K, Vallis Y, Doyle CA, Owen D, Hunt SP, McMahon HT (1997) Amphiphysin heterodimers: potential role in clathrin-mediated endocytosis. Mol Biol Cell 8:2003–2015
McMahon HT, Wigge P, Smith C (1997) Clathrin interacts specifically with amphiphysin and is displaced by dynamin. FEBS letters 413:319–322
McPherson PS, Garcia EP, Slepnev VI, David C, Zhang X, Grabs D, Sossin WS, Bauerfeind R, Nemoto Y, De Camilli P (1996) A presynaptic inositol-5-phosphatase. Nature 379:353–357
Micheva KD, Ramjaun AR, Kay BK, McPherson PS (1997) SH3 domain dependent interactions of endophilin with amphiphysin (vol 414, pg 308, 1997). FEBS letters 419:150
Ringstad N, Nemoto Y, DeCamilli P (1997) The SH3p4/Sh3p8/SH3p13 protein family: binding partners for synaptojanin and dynamin via a Grb2-like Src homology 3 domain. P Natl Acad Sci USA 94:8569–8574
Taylor MJ, Perrais D, Merrifield CJ (2011) A High Precision Survey of the Molecular Dynamics of Mammalian Clathrin-Mediated Endocytosis. Plos Biol 9. doi:10.1371/journal.pbio.1000604
Pant S, Sharma M, Patel K, Caplan S, Carr CM, Grant BD (2009) AMPH-1/Amphiphysin/Bin1 functions with RME-1/Ehd1 in endocytic recycling. Nat Cell Biol 11:1399–1410
Muller AJ, Baker JF, DuHadaway JB, Ge K, Farmer G, Donover PS, Meade R, Reid C, Grzanna R, Roach AH et al (2003) Targeted disruption of the murine Bin1/Amphiphysin II gene does not disable endocytosis but results in embryonic cardiomyopathy with aberrant myofibril formation. Mol Cell Biol 23:4295–4306
Posey AD, Jr., Swanson KE, Alvarez MG, Krishnan S, Earley JU, Band H, Pytel P, McNally EM, Demonbreun AR (2014) EHD1 mediates vesicle trafficking required for normal muscle growth and transverse tubule development. Dev Biol. doi:10.1016/j.ydbio.2014.01.004
Takei K, Slepnev VI, Haucke V, De Camilli P (1999) Functional partnership between amphiphysin and dynamin in clathrin-mediated endocytosis. Nat Cell Biol 1:33–39
Farsad K, Ringstad N, Takei K, Floyd SR, Rose K, De Camilli P (2001) Generation of high curvature membranes mediated by direct endophilin bilayer interactions. J Cell Biol 155:193–200
Razzaq A, Robinson IM, McMahon HT, Skepper JN, Su Y, Zelhof AC, Jackson AP, Gay NJ, O’Kane CJ (2001) Amphiphysin is necessary for organization of the excitation-contraction coupling machinery of muscles, but not for synaptic vesicle endocytosis in Drosophila. Genes Dev 15:2967–2979
Spiegelhalter C, Tosch V, Hentsch D, Koch M, Kessler P, Schwab Y, Laporte J (2010) From dynamic live cell imaging to 3D ultrastructure: novel integrated methods for high pressure freezing and correlative light-electron microscopy. PLoS One 5. doi:10.1371/journal.pone.0009014
Meunier B, Quaranta M, Daviet L, Hatzoglou A, Leprince C (2009) The membrane-tubulating potential of amphiphysin 2/BIN1 is dependent on the microtubule-binding cytoplasmic linker protein 170 (CLIP-170). Eur J Cell Biol 88:91–102
Bohm J, Vasli N, Maurer M, Cowling B, Shelton GD, Kress W, Toussaint A, Prokic I, Schara U, Anderson TJ et al (2013) Altered splicing of the bin1 muscle-specific exon in humans and dogs with highly progressive centronuclear myopathy. PLoS Genet 9. doi:10.1371/journal.pgen.1003430
Yamada H, Padilla-Parra S, Park SJ, Itoh T, Chaineau M, Monaldi I, Cremona O, Benfenati F, De Camilli P, Coppey-Moisan M et al (2009) Dynamic interaction of amphiphysin with N-WASP regulates actin assembly. J Biol Chem 284:34244–34256
Yamada H, Ohashi E, Abe T, Kusumi N, Li SA, Yoshida Y, Watanabe M, Tomizawa K, Kashiwakura Y, Kumon H et al (2007) Amphiphysin 1 Is Important for actin polymerization during phagocytosis. Mol Biol Cell 18:4669–4680
Colwill K, Field D, Moore L, Friesen J, Andrews B (1999) In vivo analysis of the domains of yeast Rvs167p suggests Rvs167p function is mediated through multiple protein interactions. Genetics 152:881–893
Chapuis J, Hansmannel F, Gistelinck M, Mounier A, Van Cauwenberghe C, Kolen KV, Geller F, Sottejeau Y, Harold D, Dourlen P et al (2013) Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Mol Psychiatry 18:1225–1234
WechslerReya R, Elliott K, Herlyn M, Prendergast GC (1997) The putative tumor suppressor BIN1 is a short-lived nuclear phosphoprotein, the localization of which is altered in malignant cells. Cancer Res 57:3258–3263
Elliott K, Ge K, Du W, Prendergast GC (2000) The c-Myc-interacting adaptor protein Bin1 activates a caspase-independent cell death program. Oncogene 19:4669–4684
DuHadaway JB, Sakamuro D, Ewert DL, Prendergast GC (2001) Bin1 mediates apoptosis by c-Myc in transformed primary cells. Cancer Res 61:3151–3156
Elliott K, Sakamuro D, Basu A, Du W, Wunner W, Staller P, Gaubatz S, Zhang H, Prochownik E, Eilers M et al (1999) Bin1 functionally interacts with Myc and inhibits cell proliferation via multiple mechanisms. Oncogene 18:3564–3573
Kadlec L, Pendergast AM (1997) The amphiphysin-like protein 1 (ALP1) interacts functionally with the cABL tyrosine kinase and may play a role in cytoskeletal regulation. P Natl Acad Sci USA 94:12390–12395
Kinney EL, Tanida S, Rodrigue AA, Johnson JK, Tompkins VS, Sakamuro D (2008) Adenovirus E1A oncoprotein liberates c-Myc activity to promote cell proliferation through abating Bin1 expression via an Rb/E2F1-dependent mechanism. J Cell Physiol 216:621–631
Meyer-Ficca ML, Meyer RG, Jacobson EL, Jacobson MK (2005) Poly (ADP-ribose) polymerases: managing genome stability. Int J Biochem Cell Biol 37:920–926
Ramalingam A, Farmer GE, Stamato TD, Prendergast GC (2007) Bin1 interacts with and restrains the DNA end-binding protein complex Ku. Cell Cycle 6:1914–1918
Grelle G, Kostka S, Otto A, Kersten B, Genser KF, Muller EC, Walter S, Boddrich A, Stelzl U, Hanig C et al (2006) Identification of VCP/p97, carboxyl terminus of Hsp70-interacting protein (CHIP), and amphiphysin II interaction partners using membrane-based human proteome arrays. Mol Cell Proteomics 5:234–244
Routhier EL, Donover PS, Prendergast GC (2003) hob1+, the fission yeast homolog of Bin1, is dispensable for endocytosis or actin organization, but required for the response to starvation or genotoxic stress. Oncogene 22:637–648
Chang MY, Boulden J, Katz JB, Wang L, Meyer TJ, Soler AP, Muller AJ, Prendergast GC (2007) Bin1 ablation increases susceptibility to cancer during aging, particularly lung cancer. Cancer Res 67:7605–7612
Zhong XL, Hoelz DJ, Kumar HR, Sandoval JA, Rescorla FJ, Hickey RJ, Malkas LH (2009) Bin1 is linked to metastatic potential and chemosensitivity in neuroblastoma. Pediatr Blood Cancer 53:332–337
Pan K, Liang XT, Zhang HK, Zhao JJ, Wang DD, Li JJ, Lian QZ, Chang AE, Li Q, Xia JC (2012) Characterization of bridging integrator 1 (bin1) as a potential tumor suppressor and prognostic marker in hepatocellular carcinoma. Mol Med 18:507–518
Ghaneie A, Zemba-Palko V, Itoh H, Itoh K, Sakamuro D, Nakamura S, Soler AP, Prendergast GC (2007) Bin1 attenuation in breast cancer is correlated to nodal metastasis and reduced survival. Cancer Biol Ther 6:192–194
Ge K, Duhadaway J, Sakamuro D, Wechsler-Reya R, Reynolds C, Prendergast GC (2000) Losses of the tumor suppressor BIN1 in breast carcinoma are frequent and reflect deficits in programmed cell death capacity. Int J Cancer 85:376–383
Ge K, Minhas F, Duhadaway J, Mao NC, Wilson D, Buccafusca R, Sakamuro D, Nelson P, Malkowicz SB, Tomaszewski J et al (2000) Loss of heterozygosity and tumor suppressor activity of Bin1 in prostate carcinoma. Int J Cancer 86:155–161
Kennah E, Ringrose A, Zhou LL, Esmailzadeh S, Qian H, Su MW, Zhou YW, Jiang XY (2009) Identification of tyrosine kinase, HCK, and tumor suppressor, BIN1, as potential mediators of AHI-1 oncogene in primary and transformed CTCL cells. Blood 113:4646–4655
Hogarty MD, Liu X, Thompson PM, White PS, Sulman EP, Maris JM, Brodeur GM (2000) BIN1 inhibits colony formation and induces apoptosis in neuroblastoma cell lines with MYCN amplification. Med Pediatr Oncol 35:559–562
Seeger RC, Brodeur GM, Sather H, Dalton A, Siegel SE, Wong KY, Hammond D (1985) Association of multiple copies of the N-Myc oncogene with rapid progression of neuroblastomas. New Engl J Med 313:1111–1116
Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM (1984) Amplification of N-Myc in untreated human neuroblastomas correlates with advanced disease stage. Science 224:1121–1124
Tajiri T, Liu XY, Thompson PM, Tanaka S, Suita S, Zhao HQ, Maris JM, Prendergast GC, Hogarty MD (2003) Expression of a MYCN-interacting isoform of the tumor suppressor BIN1 is reduced in neuroblastomas with unfavorable biological features. Clin Cancer Res 9:3345–3355
Brabec V, Kasparkova J (2005) Modifications of DNA by platinum complexes. Relation to resistance of tumors to platinum antitumor drugs. Drug Resist Updat 8:131–146
Tanida S, Mizoshita T, Ozeki K, Tsukamoto H, Kamiya T, Kataoka H, Sakamuro D, Joh T (2012) Mechanisms of Cisplatin-Induced Apoptosis and of Cisplatin Sensitivity: Potential of BIN1 to Act as a Potent Predictor of Cisplatin Sensitivity in Gastric Cancer Treatment. Int J Surg Oncol 2012: 862879. doi:10.1155/2012/862879
Prendergast G, MAJ C, Ramalingam A, Chang MY (2009) Bar the door: cancer suppression by amphiphysin-like genes. Biochemica et Biophysica Acta 1795:25–36
Claeys KG, Maisonobe T, Bohm J, Laporte J, Hezode M, Romero NB, Brochier G, Bitoun M, Carlier RY, Stojkovic T (2010) Phenotype of a patient with recessive centronuclear myopathy and a novel BIN1 mutation. Neurology 74:519–521
Romero NB, Bitoun M (2011) Centronuclear myopathies. Semin Pediatr Neurol 18:250–256
Bohm J, Yis U, Ortac R, Cakmakci H, Kurul SH, Dirik E, Laporte J (2010) Case report of intrafamilial variability in autosomal recessive centronuclear myopathy associated to a novel BIN1 stop mutation. Orphanet J Rare Dis 5: 35. doi:10.1186/1750-1172-5-35
Bitoun M, Maugenre S, Jeannet PY, Lacene E, Ferrer X, Laforet P, Martin JJ, Laporte J, Lochmuller H, Beggs AH et al (2005) Mutations in dynamin 2 cause dominant centronuclear myopathy. Nat Genet 37:1207–1209
Tjondrokoesoemo A, Park KH, Ferrante C, Komazaki S, Lesniak S, Brotto M, Ko JK, Zhou J, Weisleder N, Ma J (2011) Disrupted membrane structure and intracellular Ca (2) (+) signaling in adult skeletal muscle with acute knockdown of Bin1. PLoS One 6: e25740. doi:10.1371/journal.pone.0025740
Fernando P, Sandoz JS, Ding W, de Repentigny Y, Brunette S, Kelly JF, Kothary R, Megeney LA (2009) Bin1 SRC homology 3 domain acts as a scaffold for myofiber sarcomere assembly. J Biol Chem 284:27674–27686
Hong TT, Cogswell R, James CA, Kang G, Pullinger CR, Malloy MJ, Kane JP, Wojciak J, Calkins H, Scheinman MM et al (2012) Plasma BIN1 correlates with heart failure and predicts arrhythmia in patients with arrhythmogenic right ventricular cardiomyopathy. Heart Rhythm 9:961–967
Asimaki A (2012) BIN1: a new biomarker to track ARVC? Heart rhythm 9:968–969
Hong TT, Smyth JW, Chu KY, Vogan JM, Fong TS, Jensen BC, Fang K, Halushka MK, Russell SD, Colecraft H et al (2012) BIN1 is reduced and Cav1.2 trafficking is impaired in human failing cardiomyocytes. Heart Rhythm 9:812–820
Hong TT, Smyth JW, Gao D, Chu KY, Vogan JM, Fong TS, Jensen BC, Colecraft HM, Shaw RM (2010) BIN1 localizes the L-type calcium channel to cardiac T-tubules. PLoS Biol 8. doi:10.1371/journal.pbio.1000312
Seshadri S, Fitzpatrick AL, Ikram MA, DeStefano AL, Gudnason V, Boada M, Bis JC, Smith AV, Carassquillo MM, Lambert JC et al (2010) Genome-wide analysis of genetic loci associated with Alzheimer disease. Jama 303:1832–1840
Masoodi TA, Al Shammari SA, Al-Muammar MN, Alhamdan AA, Talluri VR (2013) Exploration of deleterious single nucleotide polymorphisms in late-onset Alzheimer disease susceptibility genes. Gene 512:429–437
Hu XL, Pickering E, Liu YC, Hall S, Fournier H, Katz E, Dechairo B, John S, Van Eerdewegh P, Soares H et al. (2011) Meta-analysis for genome-wide association study identifies multiple variants at the BIN1 locus associated with late-onset Alzheimer's disease. PLoS One 6. doi:10.1371/journal.pone.0016616
Wijsman EM, Pankratz ND, Choi Y, Rothstein JH, Faber KM, Cheng R, Lee JH, Bird TD, Bennett DA, Diaz-Arrastia R et al (2011) Genome-wide association of familial late-onset Alzheimer’s disease replicates BIN1 and CLU and nominates CUGBP2 in interaction with APOE. Plos Genet 7. doi:10.1371/journal.pgen.1001308
Carrasquillo MM, Belbin O, Hunter TA, Ma L, Bisceglio GD, Zou FG, Crook JE, Pankratz VS, Sando SB, Aasly JO et al (2011) Replication of BIN1 association with Alzheimer’s disease and evaluation of genetic interactions. J Alzheimers Dis 24:751–758
Lambert JC, Zelenika D, Hiltunen M, Chouraki V, Combarros O, Bullido MJ, Tognoni G, Fievet N, Boland A, Arosio B et al (2011) Evidence of the association of BIN1 and PICALM with the AD risk in contrasting European populations. Neurobiol Aging 32. doi:10.1016/j.neurobiolaging.2010.11.022
Lee JH, Cheng R, Barral S, Reitz C, Medrano M, Lantigua R, Jimenez-Velazquez IZ, Rogaeva E, George-Hyslop PHS, Mayeux R (2011) Identification of novel loci for Alzheimer disease and replication of CLU, PICALM, and BIN1 in Caribbean Hispanic individuals. Arch Neurol-Chicago 68:320–328
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, Jun G, Destefano AL, Bis JC, Beecham GW et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45:1452–1458
Schaeffer EL, Figueiro M, Gattaz WF (2011) Insights into Alzheimer disease pathogenesis from studies in transgenic animal models. Clinics 66:45–54
Tan MS, Yu JT, Tan L (2013) Bridging integrator 1 (BIN1): form, function, and Alzheimer’s disease. Trends Mol Med 10:594–603
Raj T, Shulman JM, Keenan BT, Chibnik LB, Evans DA, Bennett DA, Stranger BE, De Jager PL (2012) Alzheimer disease susceptibility loci: evidence for a protein network under natural selection. Am J Hum Genet 90:720–726
Morel E, Chamoun Z, Lasiecka ZM, Chan RB, Williamson RL, Vetanovetz C, Dall’Armi C, Simoes S, Du Jour KSP, McCabe BD et al (2013) Phosphatidylinositol-3-phosphate regulates sorting and processing of amyloid precursor protein through the endosomal system. Nat Commun 4:2250
Hnia K, Vaccari I, Bolino A, Laporte J (2012) Myotubularin phosphoinositide phosphatases: cellular functions and disease pathophysiology. Trends Mol Med 18:317–327
Ramjaun AR, Philie J, de Heuvel E, McPherson PS (1999) The N terminus of amphiphysin II mediates dimerization and plasma membrane targeting. J Biol Chem 274:19785–19791
Huser S, Suri G, Crottet P, Spiess M (2013) Interaction of amphiphysins with AP-1 clathrin adaptors at the membrane. Biochem J 450:73–83
Grabs D, Slepnev VI, Zhou SY, David C, Lynch M, Cantley LC, DeCamilli P (1997) The SH3 domain of amphiphysin binds the proline-rich domain of dynamin at a single site that defines a new SH3 binding consensus sequence. J Biol Chem 272:13419–13425
Dong J, Misselwitz R, Welfle H, Westermann P (2000) Expression and purification of dynamin II domains and initial studies on structure and function. Protein Expr Purif 20:314–323
Kajiho H, Saito K, Tsujita K, Kontani K, Araki Y, Kurosu H, Katada T (2003) RIN3: a novel Rab5 GEF interacting with amphiphysin II involved in the early endocytic pathway. J Cell Sci 116:4159–4168
Leprince C, Le Scolan E, Meunier B, Fraisier V, Brandon N, De Gunzburg J, Camonis J (2003) Sorting nexin 4 and amphiphysin 2, a new partnership between endocytosis and intracellular trafficking. J Cell Sci 116:1937–1948
Lee C, Kim SR, Chung JK, Frohman MA, Kilimann MW, Rhee SG (2000) Inhibition of phospholipase D by amphiphysins. J Biol Chem 275:18751–18758
Cousin MA, Tan TC, Robinson PJ (2001) Protein phosphorylation is required for endocytosis in nerve terminals: potential role for the dephosphins dynamin I and synaptojanin, but not AP180 or amphiphysin. J Neurochem 76:105–116
Wixler V, Laplantine E, Geerts D, Sonnenberg A, Petersohn D, Eckes B, Paulsson M, Aumailley M (1999) Identification of novel interaction partners for the conserved membrane proximal region of alpha-integrin cytoplasmic domains. FEBS Lett 445:351–355
Messina S, Onofri F, Bongiorno-Borbone L, Giovedi S, Valtorta F, Girault JA, Benfenati F (2003) Specific interactions of neuronal focal adhesion kinase isoforms with Src kinases and amphiphysin. J Neurochem 84:253–265
Acknowledgements
We apologize to colleagues whose work was not cited because of space limitations. A.P. is supported by Association Francaise contre les Myopathies, B.S.C. by a fellowship from Fondation Recherche Médicale and J.L. by the Institut National de la Santé et de la Recherche Médicale and by a translational research stipend with Assistance Publique–Hôpitaux de Paris.
Conflict of interest
The authors declare that they have no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Prokic, I., Cowling, B.S. & Laporte, J. Amphiphysin 2 (BIN1) in physiology and diseases. J Mol Med 92, 453–463 (2014). https://doi.org/10.1007/s00109-014-1138-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-014-1138-1