Abstract
Behçet's disease (BD) is a complex disease with genetic and environmental risk factors implicated in its etiology; however, its pathophysiology is poorly understood. To decipher BD's genetic underpinnings, we combined gene expression profiling with pathway analysis and association studies. We compared the gene expression profiles in peripheral blood mononuclear cells (PBMCs) of 15 patients and 14 matched controls using Affymetrix microarrays and found that the neuregulin signaling pathway was over-represented among the differentially expressed genes. The Epiregulin (EREG), Amphiregulin (AREG), and Neuregulin-1 (NRG1) genes of this pathway stand out as they are also among the top differentially expressed genes. Twelve haplotype tagging SNPs at the EREG-AREG locus and 15 SNPs in NRG1 found associated in at least one published BD genome-wide association study were tested for association with BD in a dataset of 976 Iranian patients and 839 controls. We found a novel association with BD for the rs6845297 SNP located downstream of EREG, and replicated three associations at NRG1 (rs4489285, rs383632, and rs1462891). Multifactor dimensionality reduction analysis indicated the existence of epistatic interactions between EREG and NRG1 variants. EREG-AREG and NRG1, which are members of the epidermal growth factor (EGF) family, seem to modulate BD susceptibility through main effects and gene–gene interactions. These association findings support a role for the EGF/ErbB signaling pathway in BD pathogenesis that warrants further investigation and highlight the importance of combining genetic and genomic approaches to dissect the genetic architecture of complex diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Behçet's disease (BD) is a systemic immuno-inflammatory disease characterized by the peculiar form it affects the organism, especially in the mucocutaneous and ocular structures. Its most common clinical manifestations are recurrent orogenital ulcerations and inflammatory eye lesions. Although its etiopathogenesis remains to be elucidated, genetic (e.g., HLA-B51) and environmental factors as well as immune-mediated mechanisms have been pointed out as important factors [1].
Genome-wide association studies (GWAS) for BD recently revealed two novel susceptibility loci at a genome-wide significant level: IL23R-IL12RB2 and IL10 [2, 3]. However, the genetic susceptibility to BD is governed by additional genes that cannot be pinpointed using this approach. To identify additional genetic risk factors for BD, we hereby pursued another strategy which intersects data from a genome-wide expression study with an in silico pathway analysis and association studies.
Gene expression profiling allows the simultaneous monitoring of the transcriptional behavior of thousands of genes, and genes differentially expressed between patients and controls in a relevant tissue are likely to be involved at some stage of the pathogenic process. Gene profiling experiments have allowed the identification of novel and unsuspected pathways involved in disease etiology, including in rheumatic and immune diseases [4–6]. Pathway analysis of differentially expressed genes is a useful approach to highlight undetected patterns in the data and to determine which differentially expressed genes to follow-up in independent assays such as association studies. This convergence of genomic and genetic approaches enables data mining beyond the top findings. With this strategy, we were able to highlight the role of the neuregulin signaling pathway in BD pathogenesis and strengthen the previously observed association of variants in NRG1 with BD.
Materials and methods
Gene profiling study subjects
Fourteen BD patients and 15 healthy controls were enrolled in the gene expression study through the Hospital Infante D. Pedro, Hospital de Sousa Martins, Hospital Pediátrico de Coimbra and Instituto Português de Reumatologia in Portugal. These participants are of Portuguese Caucasian origin. All patients satisfied the International Criteria for Behçet's disease (ICBD) [7]. Patients were considered immunosuppressed if medicated with azathioprine, cyclosporine, deflazacort, or thalidomide. BD patients with age-at-diagnosis after 60 years were excluded. Controls were evaluated using the same evaluation procedures as the cases and selected when negative for BD and any other rheumatologic or autoimmune disorder. This research was approved by the ethics committee at Hospital Infante D. Pedro and Instituto Português de Reumatologia where the samples were collected. All participants were informed of the study and provided informed written consent.
Total RNA isolation and microarray hybridization
Whole blood samples were obtained by venipuncture and collected in BD Vacutainer CPT tubes (BD, Franklin Lakes, NJ, USA). These samples were centrifuged to isolate PBMCs, which were then washed twice. Total RNA was extracted from PBMCs using RNeasy Mini kit (Qiagen, Hilden, Germany) according to the protocol recommended by the manufacturer. Total RNA (3.5 μg) from each individual was hybridized to GeneChip Human Genome U133 Plus 2.0 microarrays (Affymetrix, Santa Clara, CA, USA) at the Instituto Gulbenkian de Ciência's Affymetrix Core Facility following manufacturer's protocols. Extensive quality control checks were performed in all steps of the process.
Gene expression data normalization and statistical analysis
The intensity array data were analyzed with their respective CDF file from Affymetrix on the Partek software (Partek Incorporated, St Louis, MO, USA). Background correction, normalization, and summarization of the CEL files were performed using the robust multichip average algorithm. Differentially expressed genes among cases and controls were identified with analysis of variance (ANOVA), adjusting for known experimental (immunosuppression status) and study design (geographic origin and scan date) co-variates. The genes with more than 1.20-fold-change and P ≤ 0.05 were considered differentially expressed. To account for multiple testing, we calculated false discovery rates (FDR) using the Partek Q value method [8].
The gene expression profiling was conducted and reported in accordance with the minimum information about microarray experiment (MIAME) guidelines [9]. The microarray data were deposited on Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/projects/geo/) with the accession number GSE17114.
Hierarchical clustering (HC) analysis was performed using the Partek software. The effects of the geographic origin of the participants, the scan date of the microarrays and the immunosuppression status of the participants were removed using the batch-remove tool implemented in Partek prior to visualization, since these visualization tools cannot correct for study design batch effects.
Quantitative real-time polymerase chain reaction confirmation of some of the microarray results was not performed because of an insufficient amount of RNA available for several of the controls and patients used in this study.
Pathway analyses
Ingenuity Pathway Analysis 9.0 (IPA; Ingenuity Systems, http://www.ingenuity.com) was used to functionally annotate genes according to molecular networks, biological function, and canonical pathways. An IPA list is built upon a very large manually curated and up-to-date database of genes, proteins, functions, interactions/networks, and pathways. Functional analysis was performed on the 621 probe IDs differentially expressed (P ≤ 0.05 and absolute fold-change > 1.20). The IPA “Core Analysis” was run using the Human Genome U133 Plus 2.0 array as the reference set, using direct and indirect relationships for network analysis, including endogenous chemicals, data from Human species only, experimental observed confidence, and all tissues, cell lines, and data sources. The significance (P values) of the association between a dataset and a canonical pathway was determined by comparing the number of genes in a dataset that participate in a given pathway to the total number of occurrences of these genes in all pathway annotations. A Fisher's exact test was used to calculate the P value to determine the probability that the association between the genes in the dataset and the canonical pathway is explained only by chance. The level of statistical significance was set to P ≤ 0.05.
Association study subjects
Patients (976) and 839 controls, ascertained and collected as described previously [10], were included in the association study. Diagnosis of BD was made according to the ICBD criteria [7]. This research was approved by the ethics committee at the Tehran University for Medical Sciences, Iran. All participants were informed of the study and provided informed written consent.
Genotyping
Thirteen haplotype tagging SNPs (htSNP) at the EREG-AREG locus (chr. 4: 75449724–75545341 bp) were identified in Haploview 4.1 [11] using genotypes of 30 European (CEU) trios downloaded from the HapMap Release 24/phase II Nov08 (NCBI B36 assembly, dbSNP b126), and with the following options: pairwise mode, r 2 > 0.80, and minor allele frequency (MAF) > 0.1. Fifteen SNPs with a r 2 < 0.8 in NRG1 (chr. 8: 31517663–32066776 bp) found associated with BD in either one of two published GWAS (personal communications by Dr. Elaine Remmers and Dr. Nobuhisa Mizuki) were genotyped.
Genomic DNA was extracted from whole blood samples using a salting out procedure. SNPs were genotyped using Sequenom's (San Diego, USA) iPlex assay (primer extension of multiplex products with detection by matrix assisted laser desorption/ionization time-of-flight mass spectrometry) following the manufacturer's protocol and detected in a Sequenom MassArray K2 platform. The primer sequences are available in Supplementary Table 1 and were designed using Sequenom's MassARRAY ® Assay Design 3.0 software. The genotyping was performed at the Instituto Gulbenkian de Ciência's Genomics Unit.
Extensive quality control was performed using eight HapMap (http://hapmap.ncbi.nlm.nih.gov/) controls of diverse ethnic affiliation, sample duplication within and across plates, Mendelian inheritance check in three large pedigrees, Hardy–Weinberg equilibrium (HWE) in the control group (P > 0.01), and a minimum of 95 % call rate for each SNP. Genotype determinations were performed blinded to affection status.
Association analyses
Student's unpaired t tests and chi-square tests were used to compare quantitative and qualitative clinical and demographic data, respectively, between BD cases and controls. χ 2 tests for HWE in controls and crude allelic association of SNPs with BD risk were performed using Haploview 4.1. The SNPassoc v.1.4-9 package [12] implemented in the R freeware (http://cran.r-project.org/) was used to assess the associations with BD adjusted for gender. Odds ratios (ORs) and their associated 95 % confidence intervals (CIs) were calculated for SNPs with significant allelic associations with BD. Results were considered significant below the conventional level of 0.05.
Meta-analyses
GWAS data for EREG-AREG and NRG1 SNPs (e.g., allele counts, OR, CI) was obtained by personal communications from Drs. Ahmet Gul and Elaine F. Remmers for the Turkish dataset [2], and from Drs. Shigeaki Ohno, Nobuhisa Mizuki and Akira Meguro for the Japanese dataset [3]. Fixed effects (Mantel-Haenszel) meta-analyses were performed using the rmeta package in R.2.7.2 and PLINK v1.047.
Gene–gene interactions
Epistasis between EREG-AREG and NRG1 variants was detected and characterized using the multifactor dimensionality reduction (MDR) method (v2.0 beta 8.3) (www.epistasis.org) [13]. Missing genotypes were first imputed for each SNP using the PLINK database (http://pngu.mgh.harvard.edu/purcell/plink/) [14] with genotypes from individuals in the CEU HapMap population as a reference. MDR reduces high-dimensional data into a single dimension data by pooling multilocus genotypes into “high- risk” and “low-risk” groups according to ratio of affected and unaffected individuals within each genotype combination. This new multilocus variable is then tested for its ability to predict disease status. Cross validation and testing balance accuracy (TBA) are used to select the best models [13, 15]. The statistical significances of the best models were calculated after 1,000 permutations using the MDR Permutation Tool (v1.0, beta 2). Entropy-based interaction dendrograms were drawn to interpret epistasis models.
Results
Gene expression study
The principal clinical and demographic characteristics of the 29 individuals used in the gene profiling study are shown in Table 1. Cases and controls were matched for age-at-examination (AEE) (mean AAE ± SD of 37.1 ± 11.0 years in cases and 36.7 ± 13.5 years in controls, P = 9.39E-01). All of the microarrays performed were of good quality (average ± SD of present calls and of background were 45.0 ± 3.0 % and 42.0 ± 6.1 %, respectively).
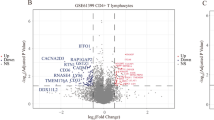
Using analysis of variance on the normalized expression data, 621 probe sets representing 508 genes (Supplementary Table 2) were found differentially expressed among BD cases and controls with a threshold of 1.20-fold-change, and P ≤ 0.05. 373 of these probe sets (representing 314 genes) were down-regulated. All the probe sets with P ≤ 0.05 also had a Q value ≤ 0.05. The false discovery rate was determined based on Q values, as these have a higher apparent power when compared to other standard methods [8]. The hierarchical cluster diagram shows a distinct gene expression profile between patients and controls that clearly separates the two groups of samples (Fig. 1). The top 30 genes with larger fold-change differences between cases and controls are shown in Table 2. EREG, S100B (S100 calcium binding protein B), NAMPT (nicotinamide phosphoribosyltransferase), AREG, and NRG1 were the top five genes under-expressed in patients, and HBG1/HBG2 (hemoglobin subunit gamma-1/gamma-2), SRSF6 (serine/arginine-rich splicing factor 6), LGALS2 (lectin, galactoside-binding, soluble, 2), NEBL (nebulette), and TRD@ (T cell receptor delta locus) the top five over-expressed genes in BD patients.
Illustration of the expression pattern differences among BD cases and controls. Hierarchical clustering analysis of the 29 samples using the 621 probe sets differentially expressed between BD cases and controls, with a threshold of a 1.2-fold-change and P ≤ 0.05. Each column represents an individual, and each row a probe set. Higher expression levels are dark red and lower levels are dark blue. Refer to Table 1 for sample identification. This figure is produced with the Partek software
Pathway analysis
Pathway analysis of the differentially expressed genes was performed using the Ingenuity Pathway Analysis (IPA). This software enables the discovery and analyses of functional relationships between differentially expressed genes and gives information about the molecular networks, biological functions, and canonical pathways over-represented among those genes.
Fourteen canonical pathways (Supplementary Table 3) were found to be significantly over-represented among the differentially expressed genes, including the lipid antigen presentation by CD1 (4/22 genes, P = 4.34E-04) and antigen presentation (4/40 genes, P = 9.12E-03) pathways which have functions related to cell-mediated immunity. IPA analysis also highlighted several pathways related to cell signaling such as the RhoA signaling (9/22 genes, P = 1.72E-03), Cdc42 signaling (9/143 genes, P = 3.51E-03), JAK2 in hormone-like cytokine signaling (4/34 genes, P = 8.17E-03), caveolar-mediated endocytosis signaling (6/81 genes, P = 8.88E-03), and neuregulin signaling (6/95 genes, P = 2.04E-02). Interestingly, three genes of the neuregulin signaling pathway—EREG (−2.39 fold-change, P = 2.99E-02), AREG (−1.80-fold-change, P = 4.90E-02), and NRG1 (−1.76-fold-change, P = 3.12E-02)—were among the most differentially expressed genes between cases and controls, and therefore these three genes were selected for follow up in an association study.
Association study
To evaluate the association between genes of the neuregulin signaling pathway (EREG-AREG and NRG1) and susceptibility to BD, we performed a case–control association study. The general characteristics of the association study dataset are shown in Table 3. The distribution of clinical symptoms in the Iranian BD patients (e.g., 98.7 % with oral aphthosis, 62.3 % with genital aphthosis, 55.2 % with skin lesions, 59.7 % with ocular lesions) is in line with what has been observed for larger datasets [16], suggesting that this is a representative group. Cases and controls were matched for age-at-examination (mean AAE ± SD of 39.1 ± 11.0 years in BD and 40.0 ± 12.3 years in controls, P = 1.93E-01).
Since no SNPs in the EREG-AREG locus showed evidence of association in the two published BD GWAS (personal communications by Dr. Elaine Remmers and Dr. Nobuhisa Mizuki), we followed a haplotype tagging SNP approach in order to explore the association of this locus with BD. In an extended region of chromosome 8 (chr. 8: 31517663–32713793 bp), including NRG1, 17 and 22 SNPs have been associated with BD in the Japanese and Turkish populations (P < 0.05), respectively (personal communications by Dr. Elaine Remmers and Dr. Nobuhisa Mizuki). Since NRG1 is a very large gene spanning over 1.1 Mb, we focused on 15 independent SNPs located within the smaller region showing stronger evidence of association (chr. 8: 31517663–32066776 bp). In the EREG-AREG region, one SNP failed genotyping (rs2132065) and was not included in the analysis. All the remaining SNPs were successfully genotyped (Supplementary Table 4), and the association results are shown in Supplementary Table 5.
In the EREG-AREG locus, we found an allelic association at rs6845297 (P = 2.51E-02, ORA [95%CI] = 0.86 [0.75–0.98]), located downstream of EREG. In NRG1, three markers demonstrated an association with BD risk in the allelic test: rs4489285 (P = 2.01E-02, ORG [95%CI] = 0.85 [0.73–0.97]) located upstream NRG1, and rs383632 (P = 2.78E-02, ORT [95%CI] = 1.22 [1.02–1.46]) and rs1462891 (P = 3.92E-02, ORT [95 % CI] = 1.18 [1.01–1.37]) located in NRG1. rs4489285 was also associated with BD in a Japanese sample, and rs383632 and rs1462891 were associated with BD in a Turkish dataset (personal communications by Dr. Elaine Remmers and Dr. Nobuhisa Mizuki). Since male to female ratio was significantly higher in the BD group than in the control group (52.5 % and 41.5 %, respectively, P = 3.76E-06), we tested the association of these four SNPs adjusted for gender. They remained significantly associated (P ≤ 0.05), albeit less so due to the concurrent loss of power associated with co-variate adjustment. None of these SNPs survive Bonferroni multiple testing correction. Haplotype associations did not provide additional insights over single marker associations (data not shown).
Meta-analysis of associated SNPs in NRG1
A combined analysis of the three associated SNPs at NRG1 with data from two published GWAS [2, 3] reveals that the G allele of rs4489285 confers protection to BD (P = 2.10E-03, ORG [95 % CI] = 0.83 [0.74–0.93]) and that the T alleles of rs383632 (P = 6.61E-04, ORT [95 % CI] = 1.23 [1.09–1.38]) and rs1462891 (P = 7.88E-04, ORT[95 % CI] = 1.19[1.07–1.32]) confer risk to BD (Table 4). The association consistency of these NRG1 polymorphisms is reinforced by this meta-analysis and further supports their role in BD susceptibility.
Epistatic interaction
Given that BD has clearly a multigenic inheritance and that EREG, AREG, and NRG1 belong to the same cellular pathway, we tested for the existence of non-additive gene–gene interactions using the multifactor-dimensionality reduction method. Supplementary Table 6 summarizes the best interaction models among all possible 2-, 3-, and 4-marker models tested. The best 4-marker interaction model was a significant predictor of Behçet's disease. The interaction between rs6845297 and rs9992496 (both located downstream of EREG) and rs2345991 and rs956203 (located in NRG1) had a statistically significant testing balanced accuracy of 0.566 (thus correctly classifying 56.6 % of the individuals tested), and a cross-validation consistency of 5/10, indicating that the model was selected five times out of ten cross-validation subsets (P = 8.00E-03 after 1,000 permutations). The interaction dendogram depicted in Fig. 2 shows a strong non-linear (epistatic) synergistic interaction between rs2345991 and rs956203 (located in NRG1) with rs9992496 (located downstream of EREG). The dot-dash line indicates an independent effect of rs6845297 (located downstream of EREG) over the three other SNPs. These results suggest that BD susceptibility can be modulated by epistatic interactions between epidermal growth factor receptor genes.
Discussion
Using a genomic and genetic convergence strategy for the first time in the BD genetics field, the neuregulin signaling pathway emerged as a novel player. Several genes in this pathway (EREG, AREG, NRG1) were under-expressed in BD cases when compared to controls, and genetic markers in these loci were associated with BD susceptibility.
To the best of our knowledge, we performed the first genome-wide study comparing the expression profiles of BD patients and controls. PBMCs were considered a relevant tissue for expression profiling since BD is a generalized vasculitis characterized by a profound inflammatory and immune dysregulation [1]. Furthermore, given that BD is a complex disease with multiple genes and environmental factors implicated in its etiology, we did not expect to find genes with major expression differences. Instead, the cumulative effect of small changes in expression levels of a large number of genes, over time, will result in the phenotype. Supporting this notion is the finding that HLA-DQA1, a well-established locus for BD, demonstrated a moderate differential expression in our study (1.56-fold-change, P = 3.19E-02). Therefore, we selected genes to follow up based on a conservative threshold of 1.20-fold-change.
Pathway analysis of differentially expressed genes drew our attention to the neuregulin signaling pathway and to some of its constituents, in particular to EREG, AREG, and NRG1, which were among the most under-expressed genes in BD patients. Association studies of these genes revealed one polymorphism in the EREG-AREG locus (rs6845297) and three SNPs in NRG1 (rs4489285, rs383632, and rs1462891) associated with BD. While the association of rs6845297 with BD has not been previously tested, meta-analyses of our associations in NRG1 with those from previous reports in Turkish and Japanese [2, 3] indicate that the susceptibility conferred by these variants is not restricted to the Iranian population and strengthen the role of NRG1 in BD susceptibility. The effect size of these NRG1 variants in BD susceptibility (ORs for the risk alleles of approximately 1.20) is much smaller than the effect of the well-established HLA-B51 allele (OR of approximately 3.50) [2, 3] but is of the same magnitude as reported non-HLA allelic associations (ORs typically between 1.20 and 1.60) in genome-wide associations studies for BD [2, 3, 17, 18]. Moreover, these three genetic markers have minor allele frequencies ranging from 0.122 to 0.473 in HapMap Caucasian (CEU) and Asian (CHB and JPT) populations, suggesting that these common variants may influence risk for BD in a substantial proportion of the population.
Concerns with multiple testing arise when screening a large number of genes or genetic variants. The results of the gene expression profiling were assessed for false discovery rate using the Q value, but no correction for the number of SNPs tested was performed. However, the NRG1 meta-analyses and the observed epistatic interactions between genetic markers in NRG1 and EREG-AREG reinforce the role of this pathway in BD pathogenesis and warrant further validation in independent population samples.
To date, these four genetic markers have not been associated with other diseases (NHGRI catalogue of GWAS), and no functions have been linked to these variants. However, NRG1 is encoded by 21 alternatively spliced exons spanning more than 1.1 Mb which, through alternative promoter usage and splicing, produce a variety of isoforms with different affinities to their receptors [19]. Interestingly, rs4489285 is in complete LD with rs1476540 (r 2 = 1, CEU population) that is located in a conserved transcription factor binding site for GATA-1, GATA-2, and GATA-3 in humans, and rs383632 is in complete LD with rs967205 (r 2 = 1, CEU population) which localizes to a conserved transcription factor binding site for FOXO3a, FOXO3b, and FOXD1 in humans (http://snp-nexus.org). The associated SNPs may therefore work as proxies for variants in intronic enhancers.
Epiregulin, the most under-expressed gene in BD patients, plays an essential role in immune/inflammatory-related responses in keratinocytes and macrophages in the epidermal layer [20] and has a pivotal role in peptidoglycan-induced proinflammatory cytokine production by antigen presenting cells [21]. Indeed, deficiency of epiregulin in mice results in chronic dermatitis and is correlated with an enhanced expression of the pro-inflammatory cytokine IL-18 by keratinocytes, supporting the role of this molecule in inflammatory diseases. An in silico analysis of the functional relevance of rs6845297 (located downstream of EREG) using SNPnexus (http://snp-nexus.org/) did not reveal previous associations of this polymorphism with other phenotypes or potential effects on regulatory elements or conserved sequences.
Amphiregulin binds exclusively to ErbB1 [22], and its overexpression has been associated with several autoimmune disorders such as systemic lupus erythematosus, psoriasis, and rheumatoid arthritis, and to synovial membrane inflammation [23–25]. Studies in transgenic mice demonstrate that amphiregulin overexpression in both the epidermis basal and suprabasal layer leads to a severe psoriasis-like phenotype and skin inflammation with a rich dermal and epidermal infiltration of neutrophils and lymphocytes [26]. Epidermal AREG expression is a possible mediator of innate cutaneous immunity and epidermal proliferation and a potential trigger of both cutaneous psoriasis and psoriatic arthritis [26].
Because of its multiple isoforms and various functional roles, NRG1 has been implicated in the pathophysiology of numerous disorders, including breast cancer, schizophrenia, and bipolar disorder in human studies, atherosclerosis, myocardial dysfunction, and multiple sclerosis in animal models [27]. Interestingly, a schizophrenia-associated missense mutation in NRG1 has been associated with immune system deregulation [27], establishing a link between this gene and immunological disorders. NRG1 can bind either to ErbB3 (that has no active kinase domain) and/or ErbB4. ErbB4, as well as two other genes of further downstream in the pathway (DCN, SOS) (Supplementary Table 1), were differentially expressed but not among the top genes, and thus were not selected for follow-up in the association study. On the other hand, ErbB1 was not differentially expressed, suggesting that the intracellular signaling pathway affected in BD may be via ErbB4 and not ErbB1. To further establish the role of this pathway in BD pathogenesis, comprehensive immunogenetic/cellular studies may be conducted.
We cannot formally exclude the possibility that the relatively small number of samples and the different immunossupressants (IS) taken by some patients had an impact on our findings in the expression study, and may prevented the detection of some important BD genes. Nevertheless, to minimize the effect of this potential confounding factor, we included the immunosuppression status as a co-variate in the analysis. The rationale behind including immunosuppressed patients in the expression profiling was that they are expected to be in a physiological state that more closely resembles the inactive phase of BD patients not taking immunosuppressants drugs. Our group of cases is therefore thought to be more homogeneous than a group of patients with varying degrees of disease activity. Furthermore, current treatments for BD target mostly environmental triggers (e.g., antibiotics to treat bacterial infections) or self-amplifying disease mechanisms (e.g., immunosuppressants to block auto-immunity, NSAIDs and corticosteroids to reduce inflammation), but are not specific for this disorder and are unlikely to affect the unique initial etiopathogenic mechanism(s) that we are searching for. Also, unlike canonical pathways related to immunity and inflammation, the neuroregulin signaling pathway is not as likely to be influenced by immunomodulatory drugs.
In summary, we have shown the power of combining genome-wide expression studies with in silico pathways analysis and association studies in the identification of novel genetic risk factors for BD. We have also yielded a series of candidate genes, blood biomarkers, pathways, and mechanisms that are prime targets for follow-up in hypothesis driven studies. Future research towards understanding the role of these genes and pathways in BD pathogenesis include candidate gene association studies, deep re-sequencing of selected genes, biochemical, cellular, and animal work.
References
Kapsimali VD, Kanakis MA, Vaiopoulos GA, Kaklamanis PG (2010) Etiopathogenesis of Behçet's disease with emphasis on the role of immunological aberrations. Clin Rheumatol 29:1211–1216
Remmers EF, Cosan F, Kirino Y, Ombrello MJ, Abaci N, Satorius C, Le JM, Yang B, Korman BD, Cakiris A et al (2010) Genome-wide association study identifies variants in the MHC class I, IL10, and IL23R-IL12RB2 regions associated with Behçet's disease. Nat Genet 42:698–702
Mizuki N, Meguro A, Ota M, Ohno S, Shiota T, Kawagoe T, Ito N, Kera J, Okada E, Yatsu K et al (2010) Genome-wide association studies identify IL23R-IL12RB2 and IL10 as Behçet's disease susceptibility loci. Nat Genet 42:703–706
Olsen N, Sokka T, Seehorn CL, Kraft B, Maas K, Moore J, Aune TM (2004) A gene expression signature for recent onset rheumatoid arthritis in peripheral blood mononuclear cells. Ann Rheum Dis 63:1387–1392
Achiron A, Gurevich M, Friedman N, Kaminski N, Mandel M (2004) Blood transcriptional signatures of multiple sclerosis: unique gene expression of disease activity. Ann Neurol 55:410–417
Mandel M, Gurevich M, Pauzner R, Kaminski N, Achiron A (2004) Autoimmunity gene expression portrait: specific signature that intersects or differentiates between multiple sclerosis and systemic lupus erythematosus. Clin Exp Immunol 138:164–170
International Team for the Revision of the International Criteria for Behçet's Disease (ITR-ICBD) (2006) Revision of the International Criteria for Behçet's Disease (ICBD). Clin Exp Rheumatol 24(Suppl 42):S14–S15
Quian HR, Huang S (2005) Comparison of false discovery rate methods in identifying genes with differential expression. Genomics 86:495–503
Brazma A, Hingamp P, Quackenbush J, Sherlock G, Spellman P, Stoeckert C, Aach J, Ansorge W, Ball CA, Causton HC et al (2001) Minimum information about a microarray experiment (MIAME): toward standards for microarray data. Nat Genet 29:365–371
Xavier JM, Shahram F, Davatchi F, Rosa A, Crespo J, Abdollahi BS, Nadji A, Jesus G, Barcelos F, Vaz Patto J et al (2012) Association study of IL10 and IL23R-IL12RB2 in Iranian Behçet's disease patients. Arthritis Rheum 64:2761–2772
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
González JR, Armengol L, Solé X, Guinó E, Mercader JM, Estivill X, Moreno V (2007) SNPassoc: an R package to perform whole genome association studies. Bioinformatics 23:644–645
Ritchie MD, Hahn LW, Roodi N, Bailey LR, Dupont WD, Parl FF, Moore JH (2001) Multifactor-dimensionality reduction reveals high-order interactions among estrogen-metabolism genes in sporadic breast cancer. Am J Hum Genet 69:138–147
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Moore JH (2004) Computational analysis of gene–gene interactions using multifactor dimensionality reduction. Expert Rev Mol Diagn 4:795–803
Davatchi F, Shahram F, Chams-Davatchi C, Shams H, Nadji A, Akhlaghi M, Faezi T, Ghodsi Z, Faridar A, Ashofteh F et al (2010) Behçet's disease: from east to west. Clin Rheumatol 29:823–833
Hou S, Yang Z, Du L, Jiang Z, Shu Q, Chen Y, Li F, Zhou Q, Ohno S, Chen R et al (2012) Identification of a susceptibility locus in STAT4 for Behçet's disease in Han Chinese in a genome-wide association study. Arthritis Rheum 64:4104–4013
Lee YJ, Horie Y, Wallace GR, Choi YS, Park JA, Song R, Kang YM, Kang SW, Baek HJ, Kitaichi N et al (2013) Genome-wide association study identifies GIMAP as a novel susceptibility locus for Behcet's disease. Ann Rheum Dis. doi:10.1136/annrheumdis-2011-200288
Falls DL (2003) Neuregulins: functions, forms, and signaling strategies. Exp Cell Res 284:14–30
Shirasawa S, Sugiyama S, Baba I, Inokuchi J, Sekine S, Ogino K, Kawamura Y, Dohi T, Fujimoto M, Sasazuki T (2004) Dermatitis due to epiregulin deficiency and a critical role of epiregulin in immune-related responses of keratinocyte and macrophage. Proc Natl Acad Sci U S A 101:13921–13926
Sugiyama S, Nakabayashi K, Baba I, Sasazuki T, Shirasawa S (2005) Role of epiregulin in peptidoglycan-induced proinflammatory cytokine production by antigen presenting cells. Biochem Biophys Res Commun 337:271–274
Shoyab M, Plowman GD, McDonald VL, Bradley JG, Todaro GJ (1989) Structure and function of human amphiregulin: a member of the epidermal growth factor family. Science 243:1074–1076
Ishii T, Onda H, Tanigawa A, Ohshima S, Fujiwara H, Mima T, Katada Y, Deguchi H, Suemura M, Miyake T et al (2005) Isolation and expression profiling of genes upregulated in the peripheral blood cells of systemic lupus erythematosus patients. DNA Res 12:429–439
Cook PW, Pittelkow MR, Keeble WW, Graves-Deal R, Coffey RJ Jr, Shipley GD (1992) Amphiregulin messenger RNA is elevated in psoriatic epidermis and gastrointestinal carcinomas. Cancer Res 52:3224–3227
Davies MR, Harding CJ, Raines S, Tolley K, Parker AE, Downey-Jones M, Needham MR (2005) Nurr1 dependent regulation of pro-inflammatory mediators in immortalised synovial fibroblasts. J Inflamm 2:15
Cook PW, Brown JR, Cornell KA, Pittelkow MR (2004) Suprabasal expression of human amphiregulin in the epidermis of transgenic mice induces a severe, earlyonset, psoriasis-like skin pathology: expression of amphiregulin in the basal epidermis is also associated with synovitis. Exp Dermatol 13:347–356
Marballi K, Quinones MP, Jimenez F, Escamilla MA, Raventós H, Soto-Bernardini MC, Ahuja SS, Walss-Bass C (2010) In vivo and in vitro genetic evidence of involvement of neuregulin 1 in immune system dysregulation. J Mol Med 88:1133–1141
Acknowledgments
We thank Drs. Ahmet Gul, Elaine F. Remmers, Shigeaki Ohno, Nobuhisa Mizuki and Akira Meguro for sharing their GWAS data. We thank to Dr. Majid Zeidi (Iranian Blood Transfusion Organization) for his excellent support. We are thankful to Doctor Sirous Zeinali (Department of Molecular Medicine, Biotechnology Research Center, Pasteur Institute of Iran) and Doctor Kayvan Saeedfar for their valuable help. We are also deeply grateful to all study participants and to the genotyping unit at the Instituto Gulbenkian de Ciência.
Funding
This research was supported by the Research Committee of the Tehran University of Medical Sciences (grant 132/714), the Portuguese Fundação para a Ciência e a Tecnologia (grant PTDC/SAU-GMG/098937/2008, doctoral fellowship SFRH/BD/43895/2008 to JMX, and a Ciência contract to SAO), and the Portuguese Instituto do Emprego e Formação Profissional (fellowship to JMX, TK, BVF).
Disclosure of potential conflict of interest
There is no conflict of interest to disclose.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 378 kb)
Rights and permissions
About this article
Cite this article
Xavier, J.M., Krug, T., Davatchi, F. et al. Gene expression profiling and association studies implicate the neuregulin signaling pathway in Behçet's disease susceptibility. J Mol Med 91, 1013–1023 (2013). https://doi.org/10.1007/s00109-013-1022-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-013-1022-4