Abstract
Hepatocellular carcinoma (HCC) is one of the most common malignancies in the world, and there is an urgent need to discover novel factors that can act as biomarkers for prognostic assessment and therapeutic targets of HCC. In this study, highly purified plasma membrane proteins from clinical tissue samples were obtained using a strategy combining sucrose density gradient centrifugation and subsequent phase partition. Using a two-dimensional gel electrophoresis and MALDI-Q-TOF MS/MS-based proteomics approach, we identified 13 plasma membrane-associated proteins that were differentially expressed in HCC and normal liver tissues. Of those, RhoA was one of the most significantly upregulated proteins in HCC, and its overexpression was confirmed using Western blotting. Immunohistochemistry suggested a link between RhoA expression and poor differentiation and clinicopathologic stage. Suppression of RhoA expression in HepG2 and Hep3B cells by RNA interference led to significant inhibition of cell growth, induction of apoptosis, and a decrease in migration. Our data suggest that RhoA may serve as a potential biomarker and an attractive therapeutic target for HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Primary liver cancer, also known as hepatocellular carcinoma (HCC), is the fifth most common cancer, with approximately one million new diagnoses annually, and it is one of the most deadly cancers, with approximately 600,000 yearly deaths attributed to this disease [1, 2]. HCC accounts for 80–90% of all cases of liver cancer, and its prevalence is highest in China, Southeast Asia, and sub-Saharan Africa [3, 4]. HCC begins in the main cell type in the liver and most frequently occurs in those people with liver disease and scarring called cirrhosis [5, 6]. As a precancerous lesion, cirrhosis typically occurs in patients who suffer from chronic hepatitis B or C infection or who have a long history of alcohol abuse [7, 8].
Proteomics is a promising approach that may overcome some of the limitations of previous approaches for the elucidation of the molecular mechanisms underlying many biological processes [9, 10]. Analysis of proteomic data has led to the identification of a number of novel, noteworthy biomarkers and signaling pathways [11, 12]. In recent decades, proteomic approaches have been successfully used in the investigation of many types of cancer, including breast, lung, colon, and stomach carcinomas [13, 14].
Nevertheless, few proteomic studies focusing on subcellular fractions such as the plasma membrane have been performed as yet because of the difficulty of obtaining purified plasma membrane proteins [15, 16]. To date, there are very few proteomic reports concerning the plasma membrane of HCC cells, especially at the level of clinical tissue samples from patients with HCC. Some comparative proteomic studies investigating HCC and normal cell lines or tissues have revealed significant and reproducible changes in the expression level of a number of proteins including metabolic enzymes, signal transduction factors, and oncoproteins; however, few of these proteins were found to change their expression levels in concert, reflecting regional variability or tissue heterogeneity [17, 18]. Also, few of these proteins have been subjected to further functional analysis concerning their roles in HCC carcinogenesis, and therefore we lack the complete understanding of the underlying biological processes that is required for clinical applications.
In this study, a strategy combining sucrose density gradient centrifugation and subsequent phase partition was applied to obtain plasma membrane proteins of high purity. The plasma membrane-associated proteins that were differentially expressed in cancer and in the corresponding normal tissues were profiled. One of these proteins, RhoA, was chosen for further clinical validation and functional investigation. The data resulting from this work are expected to bring about an improved understanding of the potential biomarker RhoA in the initiation and progression of HCC and thereby facilitate the translation of experimental findings into clinical applications.
Materials and methods
Clinical specimens
All tissue specimens were collected at the West China Hospital of Sichuan University. For two-dimensional electrophoresis (2-DE), fresh HCC and pair-matched adjacent normal tissues were collected from 12 HCC patients who underwent surgical resection. For immunohistochemical (IHC) analyses, paraffin-embedded HCC specimens were collected from 72 patients who underwent surgical resection. A summary of the clinical and pathologic profiles for these patients is shown in Table 1. The project was approved by the institutional ethics committee of Sichuan University. Informed consent for research was received from all patients prior to analysis.
Preparation of plasma membrane-associated proteins
The tissues were dissolved in homogenization buffer (50 mM HEPES, pH 7.4, 1 mM CaCl2, 1 mM EDTA, 1 mM vanadate, and 1 mM phenylmethylsulfonyl fluoride) and subjected to sucrose density gradient centrifugation (40%, 35%, 30%, 25%, 20%, 15%, 10%, and 5%) as previously described [19]. Solutions containing 40% (w/w) polyethylene glycol 3350 and 20% (w/w) Dextran T-500 were freshly prepared for subsequent phase partition [20]. The samples were subjected to Western blotting for evaluation of purification.
Proteomics
Approximately 300 μl of sample containing 1 mg of protein was applied to standard 2-DE gels [21]. The gels were stained in Coomassie Brilliant Blue R-250 (Bio-Rad) and then analyzed using PDQuest-7.1 software (Bio-Rad). The spots that showed a greater than 3.0-fold difference between conditions were chosen for in-gel digestion using Trypsin Gold (Promega) according to the manufacturer's instructions. MALDI-Q-TOF mass spectrometry was performed on a Q-Tof Premier mass spectrometer (Waters). The mass spectrometry data were acquired as peak list (PKL) files using MassLynx Version 4.0 software (Waters) and subsequently processed using the public MASCOT program (Matrix Science) [22].
Immunohistochemistry
The sections were incubated with a mouse monoclonal antibody against RhoA (ab54835, Abcam) and stained with Envision System horseradish peroxidase (DakoCytomation, Inc.) according to the manufacturer's instructions. For statistical analysis, total staining of RhoA was scored as the product of the staining intensity (on a scale of 0–3: negative 0, mild 1, moderate 2, and strong 3) × the percentage of positive cells (recorded on an ordered categorical scale: 0, zero; 1, 1–25%; 2, 26–50%; 3, 51–75%; and 4, 76–100%), resulting in a scale of 0–12 [23].
Cell culture
The human HCC cell lines HepG2 and Hep3B were grown in Dulbecco's modified Eagle's medium (Invitrogen) supplemented with 10% fetal calf serum (Invitrogen) and maintained at 37°C in a humidified atmosphere of 95% air and 5% CO2.
RNA interference
Four pairs of RhoA-specific small interfering RNA (siRNA), a scrambled siRNA oligonucleotide (used as a negative control; siNC), and a siRNA oligonucleotide targeting glyceraldehyde-3-phosphate dehydrogenase (used as a positive control) were synthesized by Ambion, Inc. The siRNA transfections were performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions. A pilot experiment was carried out to identify two RhoA-specific siRNAs (siRhoA-1 and siRhoA-2) that were shown to inhibit RhoA expression by Western blotting.
Western blotting
The protein extracts were subjected to 12% SDS-PAGE, transferred to PVDF membranes (Millipore), and incubated with primary antibodies against α-Na,K-ATPase (ab2871, Abcam), mitochondrial complex IV subunit I (ab14705, Abcam), calnexin (ab2798, Abcam), and RhoA (ab54835, Abcam). The blots were probed with secondary antibodies conjugated with horseradish peroxidase and visualized using an enhanced chemiluminescence system (Pierce).
MTT and colony formation assay
MTT assay was performed using an MTT reagent (Roche). The cell vitality index was calculated using the following formula: vitality index = OD treated wells/OD control wells × 100. For colony formation assay, cells were seeded in six-well plates at 3 × 102 cells per well and incubated overnight. After transfection of siRNA, the cells were allowed to grow continuously for an additional 10 days. The cell were fixed with methanol and stained with crystal violet (Sigma).
Flow cytometry, TUNEL, and Hoechst staining
Cells were seeded in six-well plates at 2 × 105 cells per well and harvested at 72 h post-transfection. After washing with PBS, the cells were then resuspended and incubated in a propidium iodide/Annexin V solution (R&D Systems). Flow cytometry was carried out on a FACSAria flow cytometer (BD Biosciences). TUNEL and Hoechst staining were performed according to the instructions of the DeadEndTM Fluorometric TUNEL System (Promega) and Hoechst 33258 (Sigma), respectively.
Cell migration and wound healing assays
Cell migration assay was performed on 8.0-μm pore, 24-well transwell plates (Millipore). Cells were seeded into the upper chamber, which contained serum-free medium, while the lower chamber was filled with complete medium that acted as a chemoattractant. Cells attached to the lower side were fixed in 4% paraformaldehyde in PBS for 20 min and stained for 10 min with 0.5% crystal violet. The number of migrated cells on the lower side of the membrane was counted in five microscopic fields. For wound healing assay, cells were seeded in six-well plates and cultured overnight. Adherent cells were scratched with a 100-μl micropipette tip and monitored for migration into the wound [24].
Statistical analysis
All quantitative data were recorded as means ± SD. The one-way analysis of variance was used to analyze differences among multiple groups. Statistical significance was defined as P < 0.05 for all analyses. Computations were performed using the statistical package SPSS 11.5 (SPSS).
Results
Preparation of plasma membrane-associated proteins
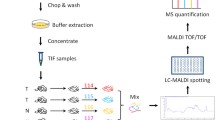
In this study, a strategy combining sucrose density gradient centrifugation and subsequent phase partition was used to obtain plasma membrane proteins of high purity (Fig. 1a). Western blotting showed that the level of α-Na,K-ATPase (a plasma membrane marker) was significantly enriched after our purification strategy, while the levels of mitochondrial complex IV subunit I (MTCO1, a mitochondrial membrane marker) and calnexin (an endoplasmic reticulum membrane marker) were reduced (Fig. 1b). These results demonstrate that the plasma membrane, but not the mitochondrial or endoplasmic reticulum membranes, was greatly enriched with our extraction approach.
Strategy combining sucrose density gradient centrifugation and subsequent phase partition to obtain plasma membrane-associated proteins. a The post-nuclear fractions were subjected to sucrose density gradient centrifugation (40%, 35%, 30%, 25%, 20%, 15%, 10%, and 5%). The crude membrane fractions were collected and mixed with a solution of 40% (w/w) polyethylene glycol 3350 and 20% (w/w) Dextran T-500 for further phase partition at 4°C. b Western blotting (25 μg protein per lane) showed that the level of ATPase was greatly increased after purification, indicating that plasma membrane proteins were enriched
Plasma membrane-associated proteins that differed between HCC and corresponding normal tissue samples
Coomassie-stained gels containing HCC and control normal tissue lysates had averages of 758 and 713 spots, respectively (Fig. 2). Thirteen plasma membrane-associated proteins were successfully identified; most of them have been described to be involved in cellular functions during carcinogenesis, including adhesion, proliferation, apoptosis, signal transduction, and cytoskeletal remodeling (Table 2). Of these, RhoA was one of the top hits, displaying a more than fivefold increase in HCC compared with normal liver tissue (Fig. 3a, b). Therefore, we hypothesized that RhoA could play important roles in HCC and therefore focused our attention on this protein. Western blotting showed that RhoA was overexpressed in HCC tissues compared to normal tissues (Fig. 3c), consistent with the results from the 2-DE analysis.
Profile of plasma membrane-associated proteins that were differentially expressed in HCC and normal liver tissues. Averages of 758 and 713 spots were visualized in 2-DE gels for HCC and normal liver tissues, respectively. The identified differentially expressed plasma membrane-associated proteins are indicated with arrows
Identification of RhoA overexpression in HCC using proteomics. a Cropped images from a 2-DE gel (upper panel) and three-dimensional images (lower panel, PDQuest) showing overexpression of RhoA in HCC tissues. b Tandem mass spectrometry map for RhoA identification. c RhoA overexpression in HCC was validated by Western blotting
Overexpression of RhoA was associated with poor differentiation and metastasis
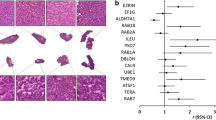
To further evaluate the potential diagnostic value of RhoA and investigate its oncogenic properties in HCC, immunohistochemistry was carried out to examine the pattern of RhoA expression pattern in clinical tissue samples. In normal liver tissues, weak positive staining of RhoA was detected at the plasma membrane and in the cytoplasm. In 72 HCC specimens, weak (29.2%), moderate (40.3%), and strong (30.5%) positive staining was detected in well, moderately, and poorly differentiated cancer tissues, respectively (Fig. 4, Table 1).
Immunohistochemical analysis of RhoA expression in clinical HCC tissues. The staining intensities of RhoA increase markedly as tissue differentiation decreases. a Normal liver, weak positive staining; b well-differentiated HCC, weak positive staining; c moderately differentiated HCC, moderate positive staining; d poorly differentiated HCC, strong positive staining
The clinicopathological correlation of RhoA staining with various HCC parameters was also evaluated, which included gender, age, differentiation, tumor nodules, lymph node involvement, distant metastasis, and clinical stage (Table 1). The quantity and extent of RhoA staining were significantly increased as tissue differentiation decreased (p = 0.03). Additionally, RhoA staining was stronger in stage III and IV tissues than in stage I and II tissues (p < 0.01). It is worth noting that there was no significant difference in RhoA staining among the four T stage groups, though it was significantly different in both the N stage (p < 0.01) and M stage (p < 0.01) groups. There was no significant difference between the male and female groups. However, RhoA staining had a negative correlation with age. In view of the overexpression of RhoA in poorly differentiated and metastatic HCC tissues, we next investigated its oncogenic properties in vitro using siRNA.
Suppression of RhoA in HepG2 cells decreased proliferation and induced apoptosis
SiRhoA-1, which targets RhoA expression, was first applied to HepG2 cells. Western blotting revealed that RhoA expression was reduced by 11%, 62%, and 96% at 72 h post-transfection with 1, 10, and 100 nM siRhoA-1, respectively (Fig. 5a). To observe changes in cellular characteristics, 10 and 100 nM siRhoA-1 were used for further experiments. MTT assay showed that cell proliferation was suppressed by siRhoA-1 (10 nM) in a time-dependent manner and that cell viability was decreased to approximately 50% of the siNC control level at 72 h after transfection (Fig. 5b). Colony formation assay showed that the colony number was reduced by 71.6% in the siRhoA-1 group versus the siNC group (Fig. 5c). These results demonstrate that RhoA suppression leads to a significant decrease in proliferation in HepG2 cells, reflecting the oncogenic capability of RhoA in HCC.
Suppression of RhoA is correlated with inhibition of proliferation and induction of apoptosis in HepG2 cells. a Western blotting confirmed that RhoA expression was inhibited in cells treated with siRhoA-1 (10 and 100 nM). b MTT assay showed that siRhoA-1 (10 nM) decreased cell growth in a time-dependent manner. c Colony formation assay showed that the number of colonies was decreased with siRhoA-1 (10 nM) after 14 days in culture. d TUNEL assay (×100 magnification) showed that the number of apoptotic cells in the siRhoA-1 (100 nM) treatment group was significantly increased. e Hoechst staining (×400) revealed evident chromatin condensation and segregation following siRhoA-1 (100 nM) treatment
We further investigate changes in cellular characteristics at a higher dose of siRhoA-1 (100 nM). TUNEL assay showed a significant increase in apoptosis in the siRhoA-1 group (Fig. 5d). Hoechst staining clearly showed that the nuclear morphology had apoptotic characteristics such as chromatin condensation and segregation (Fig. 5e). These data demonstrated that apoptosis could be specifically induced by oversuppression of RhoA in HepG2 cells.
Suppression of RhoA in HepG2 decreased cell migration
Since our IHC analysis suggested a close association between RhoA overexpression and HCC metastasis, we investigated cell migration following siRhoA-1 treatment (10 nM). Transwell assay showed that the number of the cells migrating from the upper to the lower well was significantly decreased compared to control (Fig. 6a). Wound healing assay showed that siRhoA-1 group had delayed migration, with cells loosely associating (Fig. 6b). These data demonstrate that suppression of RhoA did indeed decrease cell migration in vitro.
Suppression of RhoA is correlated with migration in HepG2 cells. a Transwell assay showed that cell migration was decreased upon siRhoA-1 (10 nM) treatment (*p < 0.01). b Scratch assay showed that cells treated with siRhoA-1 (10 nM) had a delay in migration, with cells loosely associating compared with the three control groups (*p < 0.01)
Suppression of RhoA in Hep3B cells is associated with growth inhibition and a reduction in cellular migration
To verify the effects caused by RhoA suppression in HepG2, we further performed a set of experiments in Hep3B, a human hepatoma cell line expressing hepatitis B surface antigen. Another siRNA construct against RhoA, termed siRhoA-2, was also used to eliminate the potential off-target effects of siRhoA-1. Western blotting showed that the RhoA expression was reduced by 68% (10 nM siRNA treatment) and 97% (10 nM siRNA treatment) at 72 h post-transfection, respectively (Fig. 7a). MTT assay showed that cell proliferation was suppressed by siRhoA-2 (10 nM) in a time-dependent manner and that cell viability was reduced by at least 40% at 72 h after transfection (Fig. 7b). Flow cytometry showed that the number of apoptotic cells in the siRhoA-2 (100 nM) group was significantly higher (24.3%) than that in siNC control group (Fig. 7c). Transwell assay showed that cell migration was reduced by more than 80% after treatment with siRhoA-2 (10 nM). These results in Hep3B cells are similar to those seen in HepG2 cells, indicating that RhoA plays similar roles in both hepatitis B virus (HBV)-negative (HepG2) and HBV-positive (Hep3B) cells.
RhoA suppression in Hep3B cells is associated with inhibition of growth and migration. a Western blotting confirmed that RhoA expression was inhibited in cells treated with siRhoA-2 (10 and 100 nM). b MTT assay showed inhibition of cell growth in a time-dependent manner after treatment with siRhoA-2 (10 nM). c Flow cytometric analysis showed an obvious increase in apoptosis induced by siRhoA-2 (100 nM). d Transwell assay showed the migration ability of cells was decreased following siRhoA-2 (10 nM) treatment (*p < 0.01)
Discussion
Identification of pathology-specific plasma membrane-associated proteins is a key step towards the discovery of potential biomarkers and therapeutic targets for cancer. However, plasma membrane-associated proteins are hydrophobic and expressed less abundantly than cytoplasmic proteins, presenting difficulties for the satisfactory preparation of plasma membrane-associated proteins. In the present study, we prepared plasma membrane-associated proteins using a strategy combining sucrose density gradient centrifugation and subsequent phase partition and compared the proteomic profiles of HCC and normal liver tissues by means of a 2-DE and mass spectrometry-based proteomics approach.
We focused on the subset of plasma membrane-associated proteins instead of the complete proteome because many plasma membrane-associated proteins play important roles in tumor development and progression [25, 26]. The enrichment of plasma membrane-associated proteins that is possible with our purification strategy increases the likelihood of finding valuable biomarkers on the cell surface. Although sucrose density gradient centrifugation is a classic method for enriching plasma membrane proteins, obtaining reproducible, satisfactory purification using this method is often troublesome, likely because of the similar densities of the plasma membrane and endomembranes such as the mitochondrial membrane and endoplasmic reticulum membrane [27, 28]. Therefore, we further adopted the use of phase partition after gradient centrifugation to enrich plasma membrane-associated proteins. This method of phase partition is based mainly on the principle that plasma membrane and endomembranes possess different charges, which leads to their different distribution coefficients in two-phase solution [20]. In order to assess the quality of the purification, we investigated the levels of marker proteins for the plasma membrane, mitochondrial membrane, and endoplasmic reticulum membrane. Immunoblotting revealed that the plasma membrane-associated proteins were greatly enriched with our combined purification strategy; thus, our high-quality approach provides the best chance to identify plasma membrane-associated proteins of interest.
A total of 13 differentially expressed plasma membrane-associated proteins were identified, all of which were involved in various biological processes related to carcinogenesis such as adhesion, proliferation, signal transduction, and cytoskeletal remodeling. Comprehensive analysis of the differences in the plasma membrane levels of these proteins in the HCC versus normal liver tissues could yield interesting data regarding HCC initiation and development. Of the 13 proteins identified, RhoA aroused our interest because it is highly expressed in HCC and plays interesting roles. RhoA belongs to a small GTPase superfamily that includes the Ras, Rho, Rab, Arf, and Ran subfamilies [29, 30]. As a member of the Rho subfamily, RhoA is mainly involved in the formation of stress fibers and focal adhesions and is essential to the maintenance of cell polarity as well as to intercellular junctions, cell movement, and migration [31, 32]. Recent studies have shown that RhoA can regulate cell proliferation and apoptosis and may be involved in the carcinogenesis of multiple malignant tumors including lung, breast, colon, ovarian, and prostate cancer [33, 34].
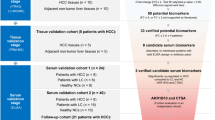
To the best of our knowledge, this study is the first proteomic report showing association of RhoA with HCC. In this work, proteomics was used as a preliminary screening tool for the identification of interesting proteins, and our final objective was to discover potential biomarkers with clinical significance. Consequently, it is necessary to translate these laboratory findings to clinical applications such as diagnosis and treatment. Therefore, a library composed of 72 clinical tissue samples was studied using immunohistochemistry to further assess the clinical significance of RhoA expression. The RhoA staining intensities in normal liver tissues and well, moderately, and poorly differentiated HCC tissues increased in a marked manner that paralleled the loss of differentiation. The close relation between the increased expression of RhoA and poor differentiation and metastasis indicated that RhoA could be used as an independent or supplementary biomarker for predicting HCC prognosis.
The biological functions of RhoA in cell proliferation and migration were investigated in vitro using RNA interference. Taking into account the fact that some cases of HCC are accompanied by infection with HBV, two cell lines, the HBV-negative HepG2 and the HBV-positive Hep3B, were used in this study. The similar results we obtained in the two different cell lines indicate that RhoA play similar roles regardless of HBV infection status. The fact that suppression of RhoA was sufficient to inhibit cell growth reflects the oncogenic properties of RhoA in HCC. In addition, the decrease in migration caused by RhoA suppression suggests that RhoA is involved in HCC migration, which correlates with the observation that higher RhoA expression was found in tissues from patients with lymph node and distant metastasis. These in vitro findings suggest that RhoA could be a potential therapeutic target in HCC and could be inhibited via treatments such as gene, antibody, and chemical therapies.
In summary, our data present a role for RhoA in HCC tumorigenesis through a combination of proteomic analysis, pathological validation, and cell function analysis. We demonstrated that RhoA overexpression was associated with poor differentiation and metastasis in HCC. We also found that RhoA suppression inhibited cell growth and migration in vitro. Our findings strongly suggest that RhoA is a promising biomarker for HCC and possibly a therapeutic target for HCC treatment.
References
El-Serag HB, Mason AC (1999) Rising incidence of hepatocellular carcinoma in the United States. N Engl J Med 340:745–750
Skolnick AA (1996) Armed with epidemiologic research, China launches programs to prevent liver cancer. J Am Med Assoc 276:1458–1459
Levin B, Amos C (1995) Therapy of unresectable hepatocellular carcinoma. N Engl J Med 332:1294–1296
Bruix J, Llovet JM (2002) Prognostic prediction and treatment strategy in hepatocellular carcinoma. Hepatology 35:519–524
Kim KR, Moon HE, Kim KW (2002) Hypoxia-induced angiogenesis in human hepatocellular carcinoma. J Mol Med 80:703–714
Aravalli RN, Steer CJ, Cressman EN (2008) Molecular mechanisms of hepatocellular carcinoma. Hepatology 48:2047–2063
Branda M, Wands JR (2006) Signal transduction cascades and hepatitis B and C related hepatocellular carcinoma. Hepatology 43:891–902
Sutton A, Nahon P, Pessayre D, Rufat P, Poiré A, Ziol M, Vidaud D, Barget N, Ganne-Carrié N, Charnaux N et al (2006) Genetic polymorphisms in antioxidant enzymes modulate hepatic iron accumulation and hepatocellular carcinoma development in patients with alcohol-induced cirrhosis. Cancer Res 66:2844–2852
Du XL, Hu H, Lin DC, Xia SH, Shen XM, Zhang Y, Luo ML, Feng YB, Cai Y, Xu X et al (2007) Proteomic profiling of proteins dysregulated in Chinese esophageal squamous cell carcinoma. J Mol Med 85:863–875
Cravatt BF, Simon GM, Yates JR 3rd (2007) The biological impact of mass-spectrometry-based proteomics. Nature 450:991–1000
Kondo T, Hirohashi S (2006) Application of highly sensitive fluorescent dyes (CyDye DIGE Fluor saturation dyes) to laser microdissection and two-dimensional difference gel electrophoresis (2D-DIGE) for cancer proteomics. Nat Protoc 1:2940–2956
Shields DJ, Niessen S, Murphy EA, Mielgo A, Desgrosellier JS, Lau SK, Barnes LA, Lesperance J, Bouvet M, Tarin D et al (2010) RBBP9: a tumor-associated serine hydrolase activity required for pancreatic neoplasia. Proc Natl Acad Sci USA 107:2189–2194
Wang L, Zhu YF, Guo XJ, Huo R, Ma X, Lin M, Zhou ZM, Sha JH (2005) A two-dimensional electrophoresis reference map of human ovary. J Mol Med 83:812–821
Han MH, Hwang SI, Roy DB, Lundgren DH, Price JV, Ousman SS, Fernald GH, Gerlitz B, Robinson WH, Baranzini SE et al (2008) Proteomic analysis of active multiple sclerosis lesions reveals therapeutic targets. Nature 451:1076–1081
Zhou J, Zhou T, Cao R, Liu Z, Shen J, Chen P, Wang X, Liang S (2006) Evaluation of the application of sodium deoxycholate to proteomic analysis of rat hippocampal plasma membrane. J Proteome Res 5:2547–2553
Jayanthi LD, Samuvel DJ, Ramamoorthy S (2004) Regulated internalization and phosphorylation of the native norepinephrine transporter in response to phorbol esters. Evidence for localization in lipid rafts and lipid raft-mediated internalization. J Biol Chem 279:19315–19326
Chignard N, Beretta L (2004) Proteomics for hepatocellular carcinoma marker discovery. Gastroenterology 127:S120–S125
Seow TK, Liang RC, Leow CK, Chung MC (2001) Hepatocellular carcinoma: from bedside to proteomics. Proteomics 1:1249–1263
Cao R, Li X, Liu Z, Peng X, Hu W, Wang X, Chen P, Xie J, Liang S (2006) Integration of a two-phase partition method into proteomics research on rat liver plasma membrane proteins. J Proteome Res 5:634–642
Norling B, Zak E, Andersson B, Pakrasi H (1998) 2D-isolation of pure plasma and thylakoid membranes from the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett 436:189–192
Peirce MJ, Wait R, Begum S, Saklatvala J, Cope AP (2004) Expression profiling of lymphocyte plasma membrane proteins. Mol Cell Proteomics 3:56–65
Perkins DN, Pappin DJ, Creasy DM, Cottrell JS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20:3551–3567
Kreisberg JI, Malik SN, Prihoda TJ, Bedolla RG, Troyer DA, Kreisberg S, Ghosh PM (2004) Phosphorylation of Akt (Ser473) is an excellent predictor of poor clinical outcome in prostate cancer. Cancer Res 64:5232–5236
Finlayson AE, Freeman KW (2009) A cell motility screen reveals role for MARCKS-related protein in adherens junction formation and tumorigenesis. PLoS One 4:e7833
Lajoie P, Goetz JG, Dennis JW, Nabi IR (2009) Lattices, rafts, and scaffolds: domain regulation of receptor signaling at the plasma membrane. J Cell Biol 185:381–385
Maxfield FR (2002) Plasma membrane microdomains. Curr Opin Cell Biol 14:483–487
McNiven MA, Thompson HM (2006) Vesicle formation at the plasma membrane and trans-Golgi network: the same but different. Science 313:1591–1594
Saraste J, Goud B (2007) Functional symmetry of endomembranes. Mol Biol Cell 18:1430–1436
Olson MF, Ashworth A, Hall A (1995) An essential role for Rho, Rac, and Cdc42 GTPases in cell cycle progression through G1. Science 269:1270–1272
Wei Y, Zhang Y, Derewenda U, Liu X, Minor W, Nakamoto RK, Somlyo AV, Somlyo AP, Derewenda ZS (1997) Crystal structure of RhoA-GDP and its functional implications. Nat Struct Biol 4:699–703
Chan CH, Lee SW, Li CF, Wang J, Yang WL, Wu CY, Wu J, Nakayama KI, Kang HY, Huang HY et al (2010) Deciphering the transcriptional complex critical for RhoA gene expression and cancer metastasis. Nat Cell Biol 12:457–467
Hoshino D, Tomari T, Nagano M, Koshikawa N, Seiki M (2009) A novel protein associated with membrane-type 1 matrix metalloproteinase binds p27(kip1) and regulates RhoA activation, actin remodeling, and matrigel invasion. J Biol Chem 284:27315–27326
Suzuki C, Daigo Y, Ishikawa N, Kato T, Hayama S, Ito T, Tsuchiya E, Nakamura Y (2005) ANLN plays a critical role in human lung carcinogenesis through the activation of RHOA and by involvement in the phosphoinositide 3-kinase/AKT pathway. Cancer Res 65:11314–11325
Zhao X, Lu L, Pokhriyal N, Ma H, Duan L, Lin S, Jafari N, Band H, Band V (2009) Overexpression of RhoA induces preneoplastic transformation of primary mammary epithelial cells. Cancer Res 69:483–491
Acknowledgments
This study was supported by the National Natural Science Foundation of China (No. 30872742).
Disclosure of potential conflict of interests
The authors declare no conflict of interests related to this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gou, L., Wang, W., Tong, A. et al. Proteomic identification of RhoA as a potential biomarker for proliferation and metastasis in hepatocellular carcinoma. J Mol Med 89, 817–827 (2011). https://doi.org/10.1007/s00109-011-0753-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-011-0753-3