Abstract
The low-density lipoprotein (LDL) receptor is a transmembrane glycoprotein that mediates the binding and endocytosis of lipoproteins containing apolipoprotein B and E, especially the cholesterol-rich LDL. Mutations in the LDL receptor gene can produce dysfunctional LDL receptors and cause familial hypercholesterolemia. The expression of the LDL receptor gene is under an intriguing regulation by sterol and nonsterol mediators either at the transcriptional level or at the posttranscriptional level, both of which are linked to cell signaling pathways. Upregulation of liver LDL receptor expression is effective in treating hypercholesterolemia. In this review, we focus on the latest progress on the mechanisms and regulation of the LDL receptor gene expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The low-density lipoprotein (LDL) receptor was first discovered by Goldstein and Brown in 1974 on cultured human skin fibroblasts. Since then, its structure, function, mutation, and physiological as well as pharmacological modulations have been extensively studied. The LDL receptor is a membrane-spanning glycoprotein with a highly conserved structure in human and other animal species [1]. LDL receptor is synthesized in the rough endoplasmic reticulum (ER) as a precursor protein, followed by a maturation process during the transportation to the Golgi apparatus, including elongation of carbohydrate chains, removal of signal peptide, and conformational change [2]. The mature form of the human LDL receptor protein contains 839 amino acid residues with a molecular mass of 160 kDa [3], and has five functional domains: the ligand-binding domain, the epidermal growth factor (EGF) precursor homology domain, the O-linked polysaccharide domain, the membrane-spanning domain, and the cytoplasmic domain. Structures and functions of these domains have been reviewed in detail elsewhere [4].
The matured LDL receptor proteins are guided to the cell surface, where they cluster into the coated pits on the cell membrane [5]. LDL receptors on the cell surface bind and uptake apolipoprotein B- and apolipoprotein E-containing lipoproteins (especially LDL) from the circulation. After endocytosis, the LDL receptor uncouples from its ligand and returns to the cell surface for recycling, while the LDL undergoes further metabolism [6]. The LDL is the most cholesterol-rich lipoprotein in the plasma, and elevation of plasma LDL cholesterol (LDL-c) is a major risk factor for atherosclerosis and coronary heart disease (CHD). LDL receptor plays a pivotal role in the clearance and metabolism of LDL. In the human body, the liver is the most LDL-receptor-abundant organ and accounts for more than 70% of the total LDL clearance in plasma [7]. Thus, LDL receptor gene mutations often result in familial hypercholesterolemia (FH), either heterozygous or homozygous, depending on the genetic background of the parents. The heterozygote of FH is one of the most common autosomal dominant genetic diseases in humans, whereas the homozygote is rare but more severe [8, 9]. Numerous LDL receptor gene mutations have been identified, and the mutations have been divided into several classes based on their phenotypic effects on the protein [4, 10].
At present, upregulation of liver LDL receptor expression has been proven to be one of the most effective means to lower plasma cholesterol level [11]. In this review, we focus on the latest progression on the regulation of the LDL receptor gene expression.
Human LDL receptor gene
The human LDL receptor structural gene is located in the short arm of chromosome 19. It spans approximately 45 kb and consists of 18 exons, each coding for a different protein domain and 17 introns [12]. The promoter is located on the 5′-flanking region, within which the majority of cis-acting DNA elements are found between base pairs (bp) −58 and −234, with the A of the initiator methionine codon as +1. The promoter region spans 177 bp, including three imperfect direct repeats with 16 bp of each, two TATA-like sequences, and several transcription initiation sites, all of which are essential for gene expression and regulation (Fig. 1, [13]). Among the three direct repeats, repeats 1 and 3 contain sequences that can be recognized by the general transcription factor Sp1. Their role is to maintain the basic transcription level of the LDL receptor gene regardless of the presence or absence of sterols [13, 14]. However, they are not sufficient for a high-level expression of the LDL receptor gene in the absence of sterols, in which the contribution from repeat 2 is needed. Repeat 2 contains a 10-bp DNA element termed sterol regulatory element (SRE) (Fig. 1, [15]), and its function will be discussed in detail.
The human LDL receptor mRNA has a 5.3-kb sequence in length, which contains an unusually 2.5-kb-long 3′ untranslated region (UTR) [3]. There are three AU-rich elements (AREs) in the 5′ proximal region and three copies of Alu-like repeat in the 3′ distal region of the 3′ UTR. These structures play a key role for the stability of the LDL receptor mRNA and serve as cis-acting elements for the posttranscriptional regulation of the LDL receptor gene expression [3, 16].
Like other eukaryotic genes, the expression of the LDL receptor gene is under a complex regulation, either at the transcriptional or posttranscriptional level, mediated through intriguing signaling pathways. Cholesterol and derivatives, and nonsterol mediators, like cytokines, growth factors, and some hormones, are able to regulate LDL receptor expression, many of which are of important significance in the clinic [17, 18].
Transcriptional regulation of the LDL receptor gene
Sterols
The sterol regulatory element-binding protein (SREBP) pathway is crucial in the transcriptional regulation of the LDL receptor gene expression by cholesterol and its derivatives [19]. The SREBPs are transcription factors belonging to the basic-helix–loop–helix–leucine zipper (bHLH-Zip) family [20]. They were identified in Brown and Goldstein’s laboratory to bind to SRE, which is not only present in the promoter of LDL receptor gene but also in promoters of other genes that code for enzymes participating in cholesterol or fatty acids biosynthesis, such as the 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase gene and the acetyl coenzyme A synthetase gene [19–22]. There are three members of the SREBPs in mammalian cells, SREBP-1a, SREBP-1c, and SREBP-2. They have different selectivity for different target genes, with SREBP-2 as a major activator of the LDL receptor gene [23]. The SREBPs are synthesized as inactive precursors embedded in the ER membrane with a molecular mass of about 125 kDa. They consist of approximately 1,150 amino acid residues and can be divided into three functional domains forming a hairpin structure, within which the amino terminal domain contains the bHLH-Zip transcription activator (Fig. 2) [24].
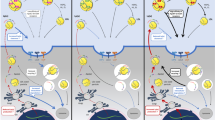
Intracellular regulation of LDL receptor gene expression. This figure illustrates how LDL receptor gene expression is regulated in the cells. As indicated in the text and shown in this figure, the SREBP pathway plays an important role for the transcriptional regulation, while the 3′ UTR of LDL receptor mRNA is a key factor for the posttranscriptional regulation. Targets and/or pathways for the clinical agents with cholesterol-lowering effect are demonstrated in the figure. bHLH-Zip, basic-helix–loop–helix–leucine zipper; Ch, cholesterol; ER, endoplasmic reticulum; ERK, extracellular signal-regulated kinase; HMG-CoA, 3-hydroxy-3-methylglutaryl coenzyme A; Insig, insulin-induced gene; LDL, low density lipoprotein; nSREBP, nuclear SREBP; S1P, site-1 protease; S2P, site-2 protease; SCAP, SREBP cleavage-activating protein; SRE, sterol regulatory element; SREBP, sterol regulatory element-binding protein; SSD, sterol sensing domain; UTR, untranslated region
To enter the nucleus and activate the transcription of target genes, the SREBP precursors must travel to the Golgi apparatus where the amino terminal domain is released [25]. The transfer of SREBPs needs the escort of another ER membrane protein termed SREBP cleavage-activating protein (SCAP). SCAP consists of 1,276 amino acids and can be divided into two functional domains (Fig. 2) [26]. The amino terminal forms eight membrane-spanning helices with short loops between them; five of the helices (2–6) serve as sterol sensors and are designated as sterol sensing domain (SSD) [26]. The carboxy terminal domain of SCAP mediates protein–protein interaction with the SREBP precursor and forms SREBP/SCAP complex [26, 27].
When cholesterol or its derivatives are abundant in cells, the SREBP pathway is suppressed, and the transcription of the LDL receptor gene or other genes required for lipids synthesis are turned off. Cholesterol can bind directly to the SSD of SCAP, causing a conformational change of SCAP, which permits it to bind to a pair of ER membrane proteins named insulin-induced genes (Insig) 1 and 2 then forms SREBP/SCAP/Insig ternary complex (Fig. 2) [28–32]. The binding of Insig proteins traps SREBP/SCAP in ER membrane so that the SREBPs are not able to get to the Golgi apparatus for cleavage, and the expression levels of their target genes decrease accordingly. As a result, the uptake and synthesis of cholesterol are inhibited, and the cells reach a cholesterol homeostasis [28–32].
On the contrary, when the cells are absent of sterols, SCAP does not interact with the Insig proteins. In this case, the SREBP/SCAP complex is free to leave the ER [33]. After getting into the Golgi apparatus, the transcriptional active domain of the SREBP precursor will be released by two sequential proteolytic cleavages catalyzed by two proteases residing in the Golgi membrane, while SCAP will return to the ER for recycling (Fig. 2) [34, 35]. The two proteases are site-1 protease (S1P) and site-2 protease (S2P), representing a serine protease and a zinc metalloprotease, respectively [36, 37]. The cleavage of SREBP precursor results in the release of a fragment containing the bHLH-Zip domain; its molecular weight is about 68 kDa and termed as nuclear SREBP (nSREBP), or the mature form of SREBP. The nSREBP enters into the nucleus and activates the transcription of target genes [34–37]. As a result, the cells uptake more cholesterol-containing lipoproteins and increase the cholesterol production to reach a new level of cholesterol homeostasis. The nSREBP is not stable and is polyubiquitinated and rapidly degraded by the proteasome with an estimated half-life of 3 h [38].
Pharmacological modulation of the LDL receptor gene expression through the SREBP pathway has been proven to be effective in treating hypercholesterolemia. Among the currently available lipid-altering drugs, cholesterol biosynthesis inhibitors, bile acid sequestrants, and cholesterol absorption inhibitors are effective in decreasing the plasma cholesterol and subsequently upregulate liver LDL receptor expression. These agents have been used in monotherapy and in combination [11], and they all reduce the plasma LDL-c level to a certain extent in the clinic, with statins being the most promising agents among them [11, 39]. Statins are currently the most prescribed lipid-lowering drugs around the world. Statins competitively inhibit HMG-CoA reductase and block cholesterol biosynthesis in the liver. Secondary to this action, the expression of LDL receptor is upregulated (Fig. 2). Statins are very effective in lowering LDL-c level and have been proven beneficial in preventing and ameliorating atherosclerosis and CHD in large-scale clinical trials [39].
Nonsterol mediators
In addition to cholesterol and derivatives, a number of nonsterol mediators such as hormones, cytokines, growth factors, and second messengers also regulate the transcription of the LDL receptor gene [17, 18, 40]. Some of them are of physiological or pharmacological significance. Their effects on the LDL receptor gene expression are summarized in Table 1.
Among hormones, estrogens such as 17 beta-estradiol dramatically increase liver LDL receptor expression [41, 42]. Therefore, estrogens lower plasma LDL-c and have atheroprotective effects both in animal models and in clinical studies [42, 43]. The estrogen-responsive element in the LDL receptor promoter has been located to repeat 3, which contains a consensus Sp1 binding site [44]. Estrogen receptor-alpha, a member of the family of nuclear hormone receptors, is necessary for the LDL receptor expression induced by estrogens. It interacts with Sp1 and forms a protein complex to activate the transcription of the LDL receptor gene [44–46]. Another hormone that acts through nuclear receptor and increases the LDL receptor transcription is triiodothyronine [47]. Through sequential deletion analysis, a potential thyroid hormone responsive element was found in the promoter region of the LDL receptor gene [48]. An obvious clinical feature of hypothyroidism in animals and man is hypercholesterolemia, mainly because of a reduced expression of the LDL receptor in the liver. Therefore, thyroid hormone therapy can improve the lipids’ profile in the circulation by increasing the hepatic LDL receptor expression [49, 50]. Insulin is also able to upregulate the expression of the LDL receptor gene both in cultured hepatoma cells and in mononuclear cells of type 2 diabetic patients [51, 52]. The induction of the LDL receptor gene transcription by insulin needs an intact SRE and the participation of SREBPs [53]. Moreover, activation of the extracellular signal-regulated kinase (ERK) in cells is essential for insulin’s induction because inhibition of the ERK pathway completely abolishes the effect of insulin on the LDL receptor gene promoter [54, 55]. Evidences have shown that SREBP-1a and SREBP-2 are direct substrates of ERK [56, 57]. Insulin-activated ERK can phosphorylate specific serine residues in the amino terminal domain of SREBP-1a and SREBP-2, followed by an enhanced transcriptional activity of the mature form of these SREBPs without affecting their nuclear abundance. Then, the transcription of the LDL receptor gene is activated accordingly. Mutations of the serine residues at the phosphorylation sites in SREBPs abolish the effect of insulin [54–57]. These evidences suggest that besides sterol-mediated proteolytic cleavage, SBEBPs are also regulated by the phosphorylation process.
A number of cytokines such as tumor necrosis factor (TNF) alpha, interleukin (IL) 1, IL-6, and oncostatin M (OM) are able to activate the transcription of the LDL receptor gene in hepatocytes [58–60] (Table 1). The plasma levels of these proinflammatory cytokines are often elevated in inflammation, infection, or trauma, during which hypocholesterolemia status is frequently observed [61]. Systemic infusion of cytokines also lowers plasma cholesterol level in animals, and this effect is partially explained by their induction of the liver LDL receptor expression [61]. Cytokines activate the transcription of the LDL receptor gene through different mechanisms. TNF-alpha and IL-1 are capable of regulating the LDL receptor gene transcription only when cells are cultured in sterol-free media, and their induction is repressed after sterols or LDL is added [58]. In contrast, OM or IL-6 upregulates the LDL receptor gene expression in a sterol-independent manner, similar to that by insulin and some growth factors [59, 60]. OM has been shown to increase the LDL receptor gene transcription by recruiting transcription factors early growth response gene 1 (Egr1) and CCAAT/enhancer binding protein beta (c/EBP beta) to bind to a DNA motif termed sterol-independent regulatory element (SIRE), which overlaps the TATA-like sequences in the promoter region of the LDL receptor gene (Fig. 1) [62, 63], whereas IL-6 needs SRE and the repeat 3 Sp1 binding site for mediating its transcriptional activation effect [59]. The cis-acting DNA elements and transcription factors involved in the induction of LDL receptor expression by TNF-alpha and IL-1 remain to be identified. Although cytokines regulate the LDL receptor gene expression through diverse mechanisms, they all act through the ERK pathway [64, 65].
Growth factors, including the platelet-derived growth factor (PDGF), EGF, and the fibroblast growth factor (FGF) also upregulate LDL receptor gene expression [66–68]. It was observed previously that the induction of the LDL receptor gene transcription by growth factors was related to the effects of their stimulation on cell growth, as upregulation of LDL receptor can provide proliferating cells with more cholesterol for biomembrane synthesis [69]. However, recent data suggests that LDL receptor gene expression and cell growth can be regulated independently [70]. The stimulation effect of growth factors on the LDL receptor gene promoter requires SRE and the Sp1 binding sites as cis-acting elements and is related to the ERK-mediated phosphorylation and activation of SREBPs, as growth factors potently activate this signaling pathway just like insulin [54, 56, 66]. Second messenger analog phorbol esters regulate the LDL receptor gene expression as well. For example, 12-O-tetradecanoylphorbol-13-acetate (TPA) increases LDL receptor gene transcription in a protein kinase C (PKC) dependent manner in hepatocytes. TPA-activated PKC induces hyperphosphorylation of histone H3 at the LDL receptor gene promoter region, thereby increasing its transcription [71]. On the other hand, phorbol-12-myristate-13-acetate (PMA) increases LDL receptor gene expression both at the transcriptional level and at the posttranscriptional level by stabilizing its mRNA [16].
While a variety of abovementioned extracellular stimuli upregulate the transcription of the LDL receptor gene, the activation of the ERK signaling cascade is crucial for these activities because blocking of this pathway closes down their effects [64, 65, 72]. ERK belongs to the subfamilies of the mitogen-activated protein kinases (MAPK), the activation of which by successive phosphorylation is secondary to the extracellular stimuli binding to their receptors on cell surface. Those receptors either have intrinsic tyrosine kinase activity themselves (like growth factor receptors and insulin receptor) or are coupled to another protein–tyrosine kinase (like receptors for cytokines) [73]. Upon activation, ERK phosphorylates and activates numerous cytoplasmic or nuclear protein factors and mediates multiple biological responses, including those control cell growth and differentiation [73]. But how the ERK pathway links to the promoter of the LDL receptor gene and increases its transcription through different mechanisms is not fully elucidated. It is speculated that transcription factors or coactivators participating in LDL receptor gene expression may be modulated by ERK or its downstream signals; therefore, increasing the LDL receptor transcription through a mechanism similar to that of insulin and some of the growth factors [72, 74].
In contrast to the ERK pathway, p38 decreases the transcription of the LDL receptor gene [64, 75]. There is a cross-talk mechanism between the p38 and ERK pathways, in which p38 exerts its inhibitory effect on the LDL receptor gene promoter by suppressing ERK activity. Blocking solely the p38 pathway activates ERK and induces LDL receptor gene transcription in hepatocytes [75]. Inhibiting p38 in IL-1 treated liver cells causes the superinduction of ERK activity and LDL receptor gene transcription [64, 72]. These data suggest that the transcription of the LDL receptor gene can be regulated through an interlinked signal network in response to various extracellular stimuli.
Posttranscriptional regulation of the LDL receptor gene
In addition to transcriptional regulation by sterols and various nonsterol mediators, the LDL receptor expression can be regulated at posttranscriptional level as well, although the mechanism and significance requires further investigation and appreciation. Current data suggest that the mRNA stability is the major mechanism for the posttranscriptional regulation of the LDL receptor gene expression [16]. The LDL receptor mRNA has a constitutively short half-life of about 45 min in HepG2 cells [76]. The stability of a specific mRNA is largely determined by the structure of its 3′ UTR. There are three AREs being identified in the 5′ proximal region of the LDL receptor 3′ UTR [16]. AREs have also been found in 3′ UTRs of many other short-lived mRNA species, like the mRNAs for cytokines. These elements confer their destabilizing effects on mRNAs, including the LDL receptor mRNA [16, 77]. Evidence has shown that after fusing the 5′-most ARE of the LDL receptor 3′ UTR to the coding region of the beta-globin cDNA, the degradation rate of the fusion transcript increases threefold as compared to the wild-type beta-globin mRNA. Fusion of LDL receptor 3′ UTR fragment containing all three AREs with the beta-globin gene increases the degradation rate of the fusion transcript by more than tenfold [16]. The destabilizing effect of AREs has also been demonstrated in animals in vivo. Transgenic mice expressing human LDL receptor gene, with two AREs in the 3′ UTR being truncated, can increase its mRNA stability up to threefold as compared to the native mice LDL receptor mRNA, resulting in a total of 2.5-fold increase in the LDL receptor expression in the mice liver [78].
The stability of the LDL receptor mRNA can be regulated by several mediators, including phorbol ester PMA, chenodeoxycholic acid (CDCA), berberine, and a fibrate drug, gemfibrozil [16, 76, 79–81]. PMA prolongs the half-life of the LDL receptor mRNA by more than twofold in HepG2 cells. The cis-acting element of PMA in the 3′ UTR of LDL receptor mRNA has been located to the 3′ distal region which contains three Alu-like repeats, and the stabilization effect of PMA also associates with the actin cytoskeleton in HepG2 cells [16, 76]. Berberine is a natural compound isolated from herbs such as Coptis chinensis. It was recently shown to be capable of upregulating LDL receptor expression through a posttranscriptional and sterol-independent mechanism in hepatocytes (Fig. 2). The 5′ proximal region of the LDL receptor 3′ UTR containing three AREs is indispensable for berberine to stabilize the receptor mRNA. Berberine has also shown a promising LDL-c-lowering effect and safety both in an animal model and in hypercholesterolemic patients [80]. The trans-acting protein factors used by these mediators to stabilize LDL receptor mRNA have not been identified yet and need extensive investigation. Interestingly, the stabilization of LDL receptor mRNA by berberine and CDCA also need the activation of the ERK signaling pathway in cells. Blocking this pathway abolishes their stabilization effects [79, 80]. But how the ERK pathway, activated by these mediators, links to the LDL receptor 3′ UTR to stabilize the receptor’s mRNA remains to be clarified.
Taken together, significant progression has been made to elucidate the mechanisms underlying the regulation of the LDL receptor gene expression since its discovery. Concomitantly, lipid-lowering agents working through upregulation of liver LDL receptor expression have been used in clinical practice [11, 80] and/or are in development in laboratories [82]. However, several key questions remain unresolved, including the ones regarding cross talks among the signal molecules or pathways, the other cytoplasmic factors that might participate in regulating LDL receptor mRNA stability, and whether or not other posttranscriptional mechanisms exist. Answers to these questions may lead to the discovery of new treatments for dyslipidemia.
Abbreviations
- ARE:
-
AU-rich element
- bHLH-Zip:
-
Basic-helix–loop–helix–leucine zipper
- bp:
-
Base pair(s)
- CDCA:
-
Chenodeoxycholic acid
- c/EBP:
-
CCAAT/enhancer binding protein
- CHD:
-
Coronary heart disease
- EGF:
-
Epidermal growth factor
- Egr1:
-
Early growth response gene 1
- ER:
-
Endoplasmic reticulum
- ERK:
-
Extracellular signal-regulated kinase
- FGF:
-
Fibroblast growth factor
- FH:
-
Familial hypercholesterolemia
- HMG-CoA:
-
3-hydroxy-3-methylglutaryl coenzyme A
- IL:
-
Interleukin
- Insig:
-
Insulin-induced gene
- kb:
-
Kilobases
- LDL:
-
Low-density lipoprotein
- LDL-c:
-
LDL-cholesterol
- MAPK:
-
Mitogen-activated protein kinase
- nSREBP:
-
Nuclear SREBP
- OM:
-
Oncostatin M
- PDGF:
-
Platelet-derived growth factor
- PKC:
-
Protein kinase C
- PMA:
-
Phorbol-12-myristate-13-acetate
- S1P:
-
Site-1 protease
- S2P:
-
Site-2 protease
- SCAP:
-
SREBP cleavage-activating protein
- SIRE:
-
Sterol-independent regulatory element
- SRE:
-
Sterol regulatory element
- SREBP:
-
Sterol regulatory element-binding protein
- SSD:
-
Sterol sensing domain
- TNF:
-
Tumor necrosis factor
- TPA:
-
12-O-tetradecanoylphorbol-13-acetate
- UTR:
-
Untranslated region
References
Südhoff TC, Goldstein JL, Brown MS, Russell DW (1985) The LDL-receptor gene. A mosaic of exons shared with different proteins. Science 228:815–822
Tolleshaug H, Goldstein JL, Schneider WJ, Brown MS (1982) Posttranslational processing of the LDL receptor and its genetic disruption in familial hypercholesterolemia. Cell 30:715–724
Yamamoto T, Davis CG, Brown MS, Schneider WJ, Casey ML, Goldstein JL, Russell DW (1984) The human LDL receptor: a cysteine-rich protein with multiple Alu sequences in its mRNA. Cell 39:27–38
Defesche JC (2004) Low-density lipoprotein receptor—its structure, function, and mutations. Semin Vasc Med 4:5–11
Davis CG, Van Driel IR, Russell DW, Brown MS, Goldstein JL (1987) The LDL-receptor: identification of aminoacids in cytoplasmic domain required for rapid endocytosis. J Biol Chem 262:4075–4079
Soutar AK (1996) Intracellular transport of the low-density lipoprotein receptor. Biochem Soc Trans 24:547–552
Spady DK (1992) Hepatic clearance of plasma low-density lipoproteins. Semin Liver Dis 12:373–385
Brown MS, Goldstein JL (1986) Receptor-mediated pathway for cholesterol homeostasis. Science 232:34–47
Hobbs HH, Russell DW, Brown MS, Goldstein JL (1990) The LDL receptor locus in familial hypercholesterolemia: mutation analysis of a membrane protein. Annu Rev Genet 24:133–170
Austin MA, Hutter CM, Zimmern RL, Humphries SE (2004) Genetic causes of monogenic heterozygous familial hypercholesterolemia: a HuGE prevalence review. Am J Epidemiol 160:407–420
Bays H, Stein EA (2003) Pharmacotherapy for dyslipidaemia—current therapies and future agents. Expert Opin Pharmacother 4:1901–1938
Lindgren V, Luskey KL, Russell DW, Francke U (1985) Human genes involved in cholesterol metabolism: chromosomal mapping of the loci for the low-density lipoprotein receptor and 3-hydroxy-3-methylglutaryl-coenzyme A reductase with cDNA probe. Proc Natl Acad Sci U S A 82:8567–8571
Südhoff TC, Van Der Westhuyzen DR, Goldstein JL, Brown MS, Russell DW (1987) Three direct repeats and a TATA-like sequence are required for regulated expression of the human low density lipoprotein receptor gene. J Biol Chem 262:10773–10779
Chang R, Yang E, Chamblis D, Kumar A, Wise J, Mehta KD (1996) In vivo role of the Sp1 site neighboring sterol-responsive element-1 in controlling low-density lipoprotein receptor gene expression. Biochem Biophys Res Commun 218:733–739
Smith JR, Osborne TF, Goldstein JL, Brown MS (1990) Identification of nucleotides responsible for enhancer activity of sterol regulatory element in low density lipoprotein receptor gene. J Biol Chem 265:2306–2310
Wilson GM, Vasa MZ, Deeley RG (1998) Stabilization and cytoskeletal-association of LDL receptor mRNA are mediated by distinct domains in its 3′ untranslated region. J Lipid Res 39:1025–1032
Soutar AK, Knight BL (1990) Structure and regulation of the LDL-receptor and its gene. Br Med Bull 46:891–916
Catapano AL (1989) The low density lipoprotein receptor: structure, function and pharmacological modulation. Pharmacol Ther 43:187–219
Rawson RB (2003) The SREBP pathway—insights from Insigs and insects. Nat Rev Mol Cell Biol 4:631–640
Yokoyama C, Wang X, Briggs MR, Admon A, Wu J, Hua X, Goldstein JL, Brown MS (1993) SREBP-1, a basic-helix–loop–helix–leucine zipper protein that controls transcription of the low density lipoprotein receptor gene. Cell 75:187–197
Briggs MR, Yokoyama C, Wang X, Brown MS, Goldstein JL (1993) Nuclear protein that binds sterol regulatory element of low density lipoprotein receptor promoter. I. Identification of the protein and delineation of its target nucleotide sequence. J Biol Chem 268:14490–14496
Wang X, Briggs MR, Hua X, Yokoyama C, Goldstein JL, Brown MS (1993) Nuclear protein that binds sterol regulatory element of low density lipoprotein receptor promoter. II. Purification and characterization. J Biol Chem 268:14497–14504
Horton JD, Goldstein JL, Brown MS (2002) SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest 109:1125–1131
Hua X, Sakai J, Ho YK, Goldstein JL, Brown MS (1995) Hairpin orientation of sterol regulatory element-binding protein-2 in cell membranes as determined by protease protection. J Biol Chem 270:29422–29427
Wang X, Sato R, Brown MS, Hua X, Goldstein JL (1994) SREBP-1, a membrane-bound transcription factor released by sterol-regulated proteolysis. Cell 77:53–62
Nohturfft A, Brown MS, Goldstein JL (1998) Topology of SREBP cleavage activating protein, a polytopic membrane protein with a sterol sensing domain. J Biol Chem 273:17243–17250
Hua X, Nohturfft A, Goldstein JL, Brown MS (1996) Sterol resistance in CHO cells traced to point mutations in SREBP cleavage activating protein (SCAP). Cell 87:415–426
Yang T, Espenshade PJ, Wright ME, Yabe D, Gong Y, Aebersold R, Goldstein JL, Brown MS (2002) Crucial step in cholesterol homeostasis: sterols promote binding of SCAP to INSIG-1, a membrane protein that facilitates retention of SREBPs in ER. Cell 110:489–500
Yabe D, Brown MS, Goldstein JL (2002) Insig-2, a second endoplasmic reticulum protein that binds SCAP and blocks export of sterol regulatory element-binding proteins. Proc Natl Acad Sci U S A 99:12753–12758
Radhakrishnan A, Sun LP, Kwon HJ, Brown MS, Goldstein JL (2004) Direct binding of cholesterol to the purified membrane region of SCAP: mechanism for a sterol-sensing domain. Mol Cell 15:259–268
Brown AJ, Sun L, Feramisco JD, Brown MS, Goldstein JL (2002) Cholesterol addition to ER membranes alters conformation of SCAP, the SREBP escort protein that regulates cholesterol metabolism. Mol Cell 10:237–245
Adams CM, Goldstein JL, Brown MS (2003) Cholesterol-induced conformational change in SCAP enhanced by Insig proteins and mimicked by cationic amphiphiles. Proc Natl Acad Sci U S A 100:10647–10652
Espenshade PJ, Li WP, Yabe D (2002) Sterols block binding of COPII proteins to SCAP, thereby controlling SCAP sorting in ER. Proc Natl Acad Sci U S A 99:11694–11699
Brown MS, Goldstein JL (1999) A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood. Proc Natl Acad Sci U S A 96:11041–11048
Nohturfft A, DeBose-Boyd RA, Scheek S, Goldstein JL, Brown MS (1999) Sterols regulate cycling of SREBP cleavage-activating protein (SCAP) between endoplasmic reticulum and Golgi. Proc Natl Acad Sci U S A 96:11235–11240
Duncan EA, Brown MS, Goldstein JL, Sakai J (1997) Cleavage site for sterol-regulated protease located to a Leu–Ser bond in the lumenal loop of sterol regulatory element-binding protein-2. J Biol Chem 272:12778–12785
Duncan EA, Dave UP, Sakai J, Goldstein JL, Brown MS (1998). Second-site cleavage in sterol regulatory element-binding protein occurs at transmembrane junction as determined by cysteine panning. J Biol Chem 273:17801–17809
Hirano Y, Yoshida M, Shimizu M, Sato R (2001) Direct demonstration of rapid degradation of nuclear sterol regulatory element-binding proteins by the ubiquitin-proteasome pathway. J Biol Chem 276:36431–36437
Corsini A, Bellosta S, Baetta R, Fumagalli R, Paoletti R, Bernini F (1999) New insights into the pharmacodynamic and pharmacokinetic properties of statins. Pharmacol Ther 84:413–428
Auwerx JH, Chait A, Wolfbauer G, Deeb SS (1989) Involvement of second messengers in regulation of the low-density lipoprotein receptor gene. Mol Cell Biol 9:2298–2302
Windler EET, Kovanen PT, Chao Y, Brown MS, Havel RJ, Goldstein JL (1980) The estradiol-stimulated lipoprotein receptor of rat liver. J Biol Chem 255:10464–10471
Hodgin JB, Maeda N (2002) Minireview: estrogen and mouse models of atherosclerosis. Endocrinology 143:4495–4501
Inukai T, Takanashi K, Takebayashi K, Tayama K, Aso Y, Takiguchi Y, Takemura Y (2000) Estrogen markedly increases LDL-receptor activity in hypercholesterolemic patients. J Med 31:247–261
Brüning JC, Lingohr P, Gillette J, Hanstein B, Avci H, Krone W, Müller-Wieland D, Kotzka J (2003) Estrogen receptor-α and Sp1 interact in the induction of the low density lipoprotein-receptor. J Steroid Biochem Mol Biol 86:113–121
Li C, Briggs MR, Ahlborn TE, Kraemer FB, Liu J (2001) Requirement of Sp1 and estrogen receptor α interaction in 17β-estradiol-mediated transcriptional activation of the low density lipoprotein receptor gene expression. Endocrinology 142:1546–1553
Parini P, Angelin B, Rudling M (1997) Importance of estrogen receptors in hepatic LDL receptor regulation. Arterioscler Thromb Vasc Biol 17:1800–1805
Ness GC (1991) Thyroid hormone. Basis for its hypocholesterolemic effect. J Fla Med Assoc 78:383–385
Bakker O, Hudig F, Meijssen S, Wiersinga WM (1998) Effects of triiodothyronine and amiodarone on the promoter of the human LDL receptor gene. Biochem Biophys Res Commun 249:517–521
Ness GC, Lopez D (1995) Transcriptional regulation of rat hepatic low-density lipoprotein receptor and cholesterol 7-α-hydroxylase by thyroid hormone. Arch Biochem Biophys 323:404–408
Duntas LH (2002) Thyroid disease and lipids. Thyroid 12:287–293
Wade DP, Knight BL, Soutar AK (1989) Regulation of low-density-lipoprotein-receptor mRNA by insulin in human hepatoma HepG2 cells. Eur J Biochem 181:727–731
Duvillard L, Florentin E, Lizard G, Petit JM, Galland F, Monier S, Gambert P, Verges B (2003) Cell surface expression of LDL receptor is decreased in type 2 diabetic patients and is normalized by insulin therapy. Diabetes Care 26:1540–1544
Streicher R, Kotzka J, Müller-Wieland D, Siemeister G, Munck M, Avci H, Krone W (1996) SREBP-1 mediates activation of the low density lipoprotein receptor promoter by insulin and insulin-like growth factor-I. J Biol Chem 271:7128–7133
Kotzka J, Müller-Wieland D, Roth G, Kremer L, Munck M, Schürmann S, Knebel B, Krone W (2000) Sterol regulatory element binding proteins (SREBP)-1a and SREBP-2 are linked to the MAP-kinase cascade. J Lipid Res 41:99–108
Kotzka J, Müller-Wieland D, Koponen A, Njamen D, Kremer L, Roth G, Munck M, Knebel B, Krone W (1998) ADD1/SREBP-1c mediates insulin-induced gene expression linked to the MAP kinase pathway. Biochem Biophys Res Commun 249:375–379
Roth G, Kotzka J, Kremer L, Lehr S, Lohaus C, Meyer HE, Krone W, Müller-Wieland D (2000) MAP kinases Erk1/2 phosphorylate sterol regulatory element-binding protein (SREBP)-1a at serine 117 in vitro. J Biol Chem 275:33302–33307
Kotzka J, Lehr S, Roth G, Avci H, Knebel B, Müller-Wieland D (2004) Insulin-activated Erk-mitogen-activated protein kinases phosphorylate sterol regulatory element-binding protein-2 at serine residues 432 and 455 in vivo. J Biol Chem 279:22404–22411
Stopeck AT, Nicholson AC, Mancini FP, Hajjar DP (1993) Cytokine regulation of low density lipoprotein receptor gene transcription in HepG2 cells. J Biol Chem 268:17489–17494
Gierens H, Nauck M, Roth M, Schinker R, Schürmann C, Scharnagl H, Neuhaus G, Wieland H, März W (2000) Interleukin-6 stimulates LDL receptor gene expression via activation of sterol-responsive and Sp1 binding elements. Arterioscler Thromb Vasc Biol 20:1777–1783
Liu J, Streiff R, Zhang YL, Vestal RE, Spence MJ, Briggs MR (1997) Novel mechanism of transcriptional activation of hepatic LDL receptor by oncostatin M. J Lipid Res 38:2035–2048
Hardardottir I, Grunfeld C, Feingold KR (1994) Effects of endotoxin and cytokines on lipid metabolism. Curr Opin Lipidol 5:207–215
Zhang F, Lin M, Abidi P, Thiel G, Liu J (2003) Specific interaction of Egr1 and c/EBPβ leads to the transcriptional activation of the human low density lipoprotein receptor gene. J Biol Chem 278:44246–44254
Liu J, Ahlborn TE, Briggs MR, Kraemer FB (2000) Identification of a novel sterol-independent regulatory element in the human low density lipoprotein receptor promoter. J Biol Chem 275:5214–5221
Kumar A, Middleton A, Chambers TC, Mehta KD (1998) Differential roles of extracellular signal-regulated kinase-1/2 and p38MAPK in interleukin-1β- and tumor necrosis factor-α-induced low density lipoprotein receptor expression in HepG2 cells. J Biol Chem 273:15742–15748
Li C, Kraemer FB, Ahlborn TE, Liu J (1999) Induction of low density lipoprotein receptor (LDLR) transcription by oncostatin M is mediated by the extracellular signal-regulated kinase signaling pathway and the repeat 3 element of the LDLR promoter. J Biol Chem 274:6747–6753
Basheeruddin K, Li X, Rechtoris C, Mazzone T (1995) Platelet-derived growth factor enhances Sp1 binding to the LDL receptor gene. Arterioscler Thromb Vasc Biol 15:1248–1254
Graham A, Russell LJ (1994) Stimulation of low-density lipoprotein uptake in HepG2 cells by epidermal growth factor via a tyrosine kinase-dependent, but protein kinase C-independent mechanism. Biochem J 298:579–584
Hsu HY, Nicholson AC, Hajjar DP (1994) Basic fibroblast growth factor-induced low density lipoprotein receptor transcription and surface expression. Signal transduction pathways mediated by the bFGF receptor tyrosine kinase. J Biol Chem 269:9213–9220
Mazzone T, Basheeruddin K, Ping L, Frazer S, Getz GS (1989) Mechanism of the growth-related activation of the low density lipoprotein receptor pathway. J Biol Chem 264:1787–1792
Kapoor GS, Atkins BA, Mehta KD (2002) Activation of Raf-1/MEK-1/2/p42/44MAPK cascade alone is sufficient to uncouple LDL receptor expression from cell growth. Mol Cell Biochem 236:13–22
Huang W, Mishra V, Batra S, Dillon I, Mehta KD (2004) Phorbol ester promotes histone H3-Ser10 phosphorylation at the LDL receptor promoter in a protein kinase C-dependent manner. J Lipid Res 45:1519–1527
Mehta KD (2002) Role of mitogen-activated protein kinases and protein kinase C in regulating low-density lipoprotein receptor expression. Gene Expr 10:153–164
Robinson MJ, Cobb MH (1997) Mitogen-activated protein kinase pathways. Curr Opin Cell Biol 9:180–186
Kapoor GS, Golden C, Atkins B, Mehta KD (2003) pp90RSK-and protein kinase C-dependent pathway regulates p42/44MAPK-induced LDL receptor transcription in HepG2 cells. J Lipid Res 44:584–593
Singh RP, Dhawan P, Golden C, Kapoor GS, Mehta KD (1999) One-way cross-talk between p38MAPK and p42/44MAPK. Inhibition of p38MAPK induces low density lipoprotein receptor expression through activation of p42/44 MAPK cascade. J Biol Chem 274:19593–19600
Wilson GM, Roberts EA, Deeley RG (1997) Modulation of LDL receptor mRNA stability by phorbol esters in human liver cell culture models. J Lipid Res 38:437–446
Zubiaga AM, Belasco JG, Greenberg ME (1995) The nonamer UUAUUUAUU is the key AU-rich sequence motif that mediates mRNA degradation. Mol Cell Biol 15:2219–2230
Knouff C, Malloy S, Wilder J, Altenburg MK, Maeda N (2001) Doubling expression of the low density lipoprotein receptor by truncation of the 3′-untranslated region sequence ameliorates type III hyperlipoproteinemia in mice expressing the human ApoE2 isoform. J Biol Chem 276:3856–3862
Nakahara M, Fujii H, Maloney PR, Shimizu M, Sato R (2002) Bile acids enhance low density lipoprotein receptor gene expression via a MAPK cascade-mediated stabilization of mRNA. J Biol Chem 277:37229–37234
Kong W, Wei J, Abidi P, Lin M, Inaba S, Li C, Wang Y, Wang Z, Si S, Pan H, Wang S, Wu J, Wang Y, Li Z, Liu J, Jiang JD (2004) Berberine is a novel cholesterol-lowering drug working through a unique mechanism distinct from statins. Nat Med 10:1344–1351
Goto D, Okimoto T, Ono M, Shimotsu H, Abe K, Tsujita Y, Kuwano M (1997) Upregulation of low density lipoprotein receptor by gemfibrozil, a hypolipidemic agent, in human hepatoma cells through stabilization of mRNA transcripts. Arterioscler Thromb Vasc Biol 17:2707–2712
Kong W, Abidi P, Kraemer FB, Jiang JD, Liu J (2005) In vivo activities of cytokine oncostatin M in regulation of plasma lipid levels. J Lipid Res 46:1163–1171
Acknowledgements
The article was supported by the “10th 5-year Plan” drug discovery program and “973” program of the Ministry of Sciences and Technology, People’s Republic of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kong, WJ., Liu, J. & Jiang, JD. Human low-density lipoprotein receptor gene and its regulation. J Mol Med 84, 29–36 (2006). https://doi.org/10.1007/s00109-005-0717-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-005-0717-6