Abstract
Hepatitis C viral infection is a cause of chronic liver disease, and current therapies are only effective in 50 % of patients. Serine proteases, which are present in both hepatitis C virus (HCV) and the dengue virus, are the most studied class of proteolytic enzymes and are the primary targets for drug development in this field. In this paper, we describe the synthesis of a novel class of isomannide-based peptide mimetic compounds based on a tartaric acid backbone. Our data showed that substitutions at position 168 (D168A) and 170 (V170A) conferred low-level resistance against compound 5a3, whereas substitutions at position 155 (R155K) and 156 (A156V) conferred no resistance. These data suggest that even though compound 5a3 is a noncompetitive inhibitor; it is able to interact with important residues located near the catalytic site. In addition, this novel compound class exhibits potent antiviral activity against variants carrying resistance mutations to boceprevir and telaprevir. Our docking studies showed important interactions, including hydrogen bonds and a π–π interaction, between compound 5a3 and residues of the allosteric site of NS3/4A. Biological and theoretical results indicate that 5a3 is a promising lead compound for the development of new drugs targeting HCV infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Millions of people worldwide are affected by Flaviviruses from the Flaviviridae family, hepatitis C virus (HCV), West Nile virus, and dengue virus (DV). HCV results in acute clinical hepatitis in approximately 20 % of infected individuals. It is the leading cause of chronic liver diseases, including cirrhosis, carcinoma, and liver failure (Cohen, 1999; Feld and Hoofnagle, 2005). Therapies based on alpha interferon and the nucleoside analog ribavirin have adverse effects and are only partially effective against HCV (Yusof et al., 2000; Wright et al., 2001).
HCV and DV NS3 protease (NS3 pro) is a serine protease that possesses a characteristic catalytic triad composed of histidine (His), aspartic acid (Asp), and serine (Ser), which is conserved across all flaviviruses. NS3 pro activity is essential for viral replication and the maturation of infectious virions; therefore, NS3 pro is a suitable target for the development of chemotherapeutic approaches to treat hepatitis C and dengue infections (Bazan and Fletterick, 1989; Gorbalenya et al., 1989; Melino and Paci, 2007). In this context, some natural and synthetic NS3 pro inhibitors have been described (Sidique et al., 2009; Chee et al., 2010).
Peptides that interact with NS3 pro have garnered significant interest due to their activity and specificity (Ingallinella et al., 1998). However, there are some limitations associated with peptide drugs. For example, peptide drugs have low bioavailabilities and can be hydrolyzed by proteases (Frokjaer and Otzen, 2005). Studies on the development of peptide mimetic inhibitors have demonstrated the ability of this compound class to potentially overcome these obstacles (Lampa et al., 2010). The FDA approved boceprevir and telaprevir as the first direct-acting antiviral agents for chronic hepatitis C treatment. Both are NS3/4A pro peptide mimetic inhibitors and belong to a class of α-ketoamide derivatives that have remarkable antiviral activity against HCV genotype 1 (Lange and Sarrazin, 2002). In addition, a variety of peptide mimetic-based NS3 pro inhibitors has been synthesized over the past decade. Notably, these inhibitors possess several distinct backbones with moderate to good activity against HCV (Maryanoff and Costanzo, 2008; Örtqvist et al., 2010; Lampa et al., 2010).
Mutations in the NS3/4A protease are associated with drug resistance, which is the major cause of antiviral therapy failure (Sarrazin and Zeuzem, 2010; Halfon and Locarnin, 2011; Lim et al., 2012). Indeed, the V170A mutation has been shown to confer resistance against boceprevir and narlaprevir (Vermehren and Sarrazin, 2011).
Over the past few years, we have shown that some peptide mimetics derived from isomannide (1) (Fig. 1) exhibit activity against both HCV and DV. Computational studies suggest that these mimetics interact with NS3/4A pro (Muri et al., 2004, 2005; Barros et al., 2009, 2010, 2012).
Herein, we describe new isomannide-based peptide mimetic compounds with a tartaric acid backbone. Notably, 5a3 is a noncompetitive inhibitor of NS3 pro in HCV and possesses potent antiviral activity against HCV variants with resistance mutations to boceprevir and telaprevir.
Results and discussion
Compound design and synthesis
Previous reports have shown that compounds containing the tartaric acid moiety possess inhibitory activity against HIV-1 protease (Peçanha et al., 2003; Barros et al., 2006; Resende et al., 2007; Shriner and Furow, 1963). These results encouraged us to explore this core for the design of new peptide mimetic inhibitors of NS3 pro. Compounds possessing a fused-bicyclic structure, such as darunavir and telaprevir, represent an effort toward the design and development of new antiviral drugs (Lin et al., 2004; Ghosh et al., 2007).
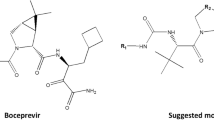
Commercially available isomannide (1) possesses a U-shaped structure, and inverting its hydroxyl group configuration leads to a W-shaped conformation which is a quasi linear structure. These structural manipulations are of great interest in the design of new compounds, particularly peptide mimetic compounds (Westermann et al., 2004; Ghosh et al., 2007). Based on our earlier results concerning the design and synthesis of peptide derivatives from 1 (see infra), the isomannide-tartaric acid scaffold was selected for new peptide mimetics (Fig. 1).
Peptide mimetics derived from l-tartaric acid have already been synthesized either via the classical EDC/HOBt protocol from the free acid or through the corresponding acid chloride, yielding the corresponding pseudo-peptide with C 2 symmetry (Shriner and Furow, 1963; Resende et al., 2007).
The proposed synthetic route for the new isomannide-based peptide mimetics with a tartaric acid core devoid of C 2 symmetry makes use of di-O-acetyl-tartaric anhydrides 6a,b (Lin et al., 2004; Ghosh et al., 2007). The amine 3 is a known compound that our group prepared in four steps from commercially available 1 (Peçanha et al., 2003; Barros et al., 2006). The condensations of 3 and carboxylic acids 4a1–4a4 or 4b2–4b4 are key steps in the syntheses of peptide mimetics 5a1–5a4 and 5b2–5b4. Notably, all of the carboxylic acids (4a1–4a4 and 4b2–4b4) used in the condensation reaction possess side chains derived from amino acids.
Short and efficient synthetic routes to peptide mimetics 5a1–5a4 employed classical procedures, as shown in Scheme 1. Crystalline di-O-acetyl-d-tartaric anhydride (6a) was prepared by the dehydration of d-tartaric acid (2a) with Ac2O/H2SO4 at reflux, as previously described in the literature (Lin et al., 2004; Ghosh et al., 2007). Ring opening of compound 6a was accomplished by reacting the compound with l-glycine ethyl ester hydrochloride (l-Gly-OEt.HCl), l-phenylalanine methyl ester hydrochloride (l-Phe-OMe.HCl), or l-valine methyl ester hydrochloride (l-Val-OMe.HCl) to produce pseudo-peptides 4a1, 4a2, and 4a3, respectively. These ring-opening reactions were carried out in the presence of N-methylmorpholine (NMM) at low temperatures. The ring-opening reactions afforded the expected product as viscous oils in moderate yield (Scheme 1). Similarly, compound 4a4 was obtained in good yield using l-proline methyl ester (l-Pro-OMe). Subsequent condensation reactions of acids 4a1–4a4 and amine 3 utilizing the classical EDC/HOBt/NMM protocol afforded the corresponding peptide mimetics 5a1–5a4 in moderate yields. All compounds were characterized spectroscopically.
Synthesis of the peptide mimetics 4a1–4a4 and 5a1–5a4. i l-Gly-OEt.HCl or l-Phe-OMe.HCl or l-Val-OMe.HCl or l-Pro-OMe, NMM, CH2Cl2, 0 °C, 30 min., 40 % (4a1), 55 % (4a2), 50 % (4a3), 84 % (4a4). ii Compound 3, EDC.HCl, HOBt, NMM, CH2Cl2, 0 °C, 1 h, then r.t., overnight, 40 % (5a1), 40 % (5a2), 42 % (5a3), 50 % (5a4)
Crystalline di-O-acetyl-l-tartaric anhydride (6b) was prepared in the same manner as 6a, by dehydration of the l-tartaric acid (2b) with Ac2O/H2SO4 at reflux (Lin et al., 2004; Ghosh et al., 2007).
Ring opening of commercially available di-O-acetyl-l-tartaric anhydride (6b) was accomplished by reacting the compound with Phe-OMe.HCl and l-Val-OMe.HCl at low temperatures. The reaction afforded good yields of the corresponding acids, 4b2 and 4b3. Notably, compound 4b4 was obtained in excellent yield using l-Pro-OMe (Scheme 2). The subsequent condensation reactions of acids 4b2–4b4 with amine 3 employed the classical EDC/HOBt/NMM protocol. This protocol afforded the corresponding peptide mimetics 5b2–5b4 in moderate to good yields. All compounds were characterized spectroscopically. To evaluate the contribution of the acetyl group to the biological activity of the compound, the diacetyl esters 5b2–5b4 were selectively hydrolyzed under basic conditions with K2CO3 and methanol to afford diols 7b2–7b4 in good yields (Barros et al., 2006; Resende et al., 2007).
Synthesis of peptide mimetics 4b2–4b4, 5b2–5b4 and 7b2–7b4 i l-Phe-OMe.HCl or l-Val-OMe.HCl or l-Pro-OMe, NMM, CH2Cl2, 0 °C, 30 min., 83 % (4b2), 78 % (4b3), 95 % (4b4). ii Compound 3, EDC.HCl, HOBt, NMM, CH2Cl2, 0 °C, 1 h, then r.t., overnight, 50 % (5b2), 75 % (5b3), 51 % (5b4). iii MeOH, 0 °C, K2CO3, r.t. 30 min., 92 % (7b2), 74 % (7b3), 90 % (7b4)
Biology
To express and purify the recombinant NS3/4A pro protein complex, a plasmid encoding the NS4A cofactor (21–32aa) was covalently joined to NS3 (1–182aa) via GlySerGlySer (GSGS), a flexible amino acid linker with the NS3 (1–182aa) N-terminal protease domain. The construct was inserted into a pET21dHT expression vector that added a His-tag followed by a TEV cleavage site (ENLYFQGS) at the N-terminus of the NS4A cofactor to facilitate purification. Protein expression is inducible with IPTG. Overall, the construct results in the production of a 6×His-TEVsite-NS3/4A pro protein with a molecular weight of 22.4 kDa. The soluble recombinant protein was purified via nickel affinity column chromatography. The 6×His-TEVsite tag was excised with TEV protease to yield NS3/4A pro. NS3/4 pro has a predicted molecular weight of 20.6 kDa, which was confirmed by SDS-PAGE (Fig. 2a). Western blot using an anti-6xHis tag monoclonal antibody was performed to confirm the absence of the 6×His tag on purified NS3/4A pro (Fig. 2b). As expected, approximately 1 mg of protein per liter of culture was obtained.
a SDS-PAGE analysis of NS3/4A pro protein purity after different purification steps. The products in the soluble fraction of the cell lysates were purified by metal affinity chromatography (His-Trap column) and stained with Coomassie brilliant blue G. Lane 1, molecular weight marker; lane 2, 6×His-TEV-NS3/4A pro soluble fraction eluted from the His-Trap column over an increasing imidazole gradient; lane 3, NS3/4A pro after 6×His-TEVsite tag excision with TEV pro at a ratio of 1:1; and lane 4, untagged NS3/4A pro (lane 3) was applied to the His-TRAP column, which retained both the 6×His-TEV site tag and the TEV protease and allowed flow-through of only NS3/4A pro. b Western blot of the same samples in A reacted with the anti-6×HisTag antibody to confirm the absence of the 6×His tag on the purified NS3/4A pro protein shown lane 4
To verify the proteolytic activity of the recombinant protein, kinetic parameters were determined using a fluorogenic peptide substrate based on the NS4A–NS4B cleavage site sequence of the HCV polyprotein. Regarding the activity of NS3/4A pro, data showed hyperbolic kinetic curves as a function of substrate concentration. The k cat was found to be 0.046 ± 0.008 s−1; the KM was 2.67 ± 0.64 µM.
In vitro inhibitory effects of peptide mimetics
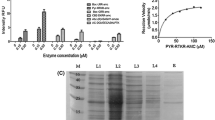
A medium-throughput NS3/4A pro inhibition-based screening assay (MTS) was used to evaluate the inhibitory activity of isomannide-based peptide mimetic derivatives against HCV-1b. Compounds 4a1 and 5a3 showed NS3/4A pro inhibitory activities of 49 ± 17 and 63 ± 13 %, respectively (Fig. 3).
Peptide mimetics derived from isomannide were tested for their activity against the HCV-1b NS3 protease at 100 μM using an MTS inhibition assay. The compounds were pre-incubated in individual wells with 30 ng of NS3/4A pro for 10 min at 25 °C. Subsequently, the peptide substrate 5-FAM/QXL™ 520-FRET was added, and the solution was incubated for 60 min at 25 °C. The reactions were performed in a black 96-well plate with final volumes of 50 μL; the reactions were performed following the manufacturer’s protocol. The percent inhibition was calculated from control reactions carried out without inhibitors (NI). Error bars represent the standard deviation. SI: reference inhibitor Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH; NI: no inhibitor
Structure–activity relationship (SAR) study
The isomannide ring of 5a3 shows increased activity compared to compound 4a3. Isomannide-based compounds have already been reported as potential HCV serine protease inhibitors (Barros et al., 2009). In the 4a series, 4a1 showed the greatest activity, most likely due to the l-glycine ethyl ester substituent. However, inserting the isomannide ring, as shown in 5a1, was not favorable, indicating a possible rotation of the binding site due to steric hindrance. Furthermore, the valine methyl ester substituent, as in 5a3, showed the greatest activity among the tested compounds.
Furthermore, the dose–response curve of the most active compound, 5a3, was compared with that of the reference inhibitor Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH (Cha = β-cyclohexylalanine; Dif = 3,3-diphenylalanine) (AnaSpec, USA). Compound 5a3 and Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH have IC50 values of 76 ± 14 and 0.60 ± 0.09 μM, respectively (Fig. 4) (Sarrazin and Zeuzem, 2010).
Inhibition of the proteolytic activity of NS3/4A by compound 5a3 and the reference inhibitor (Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH). Each experimental point in the graph represents the average of two independent replicates tested at different concentrations. The bars for each point represent the standard deviations
Substrate titration experiments were performed at different concentrations of 5a3 or Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH to evaluate whether the inhibitory mechanism of compound 5a3 is similar to that of the reference inhibitor (Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH). By fitting the experimental data to the Michaelis–Menten equation, we determined the K i values for 5a3 and Ac–Asp–Glu–Dif–Glu–Cha–Cys–OH, which were 116.5 ± 5.7 and 1.90 ± 0.27 µM, respectively. Moreover, the Lineweaver–Burk plot indicated that 5a3 inhibited the proteolytic activity of NS3/4A noncompetitively (Fig. 5).
Lineweaver-Burk (double reciprocal) plot of NS3/4A pro inhibition with compound 5a3. The concentrations of compound 5a3 used were 0, 84, 112, and 150 μM; assays were performed in duplicate. Initial rate data were fitted by linear regression to the standard noncompetitive inhibition equation using GraphPad Prism 5
In the literature, mutations to NS3/4A protease are associated with drug resistance, which is the major cause of antiviral therapy failure (Halfon and Locarnin, 2011; Sarrazin and Zeuzem, 2010; Lim et al., 2012). In addition, the A156T mutant gradually disappeared from populations treated with boceprevir and was replaced with the V170A mutant (Tong et al., 2006). Recently, the D168A mutant has become resistant to the second and third generation protease inhibitors, whereas the V170A mutant confers resistance to boceprevir and narlaprevir (Vermehren and Sarrazin, 2011).
Thus, compound 5a3 was also evaluated against the following four NS3/4A protease mutants: R155K, A156V, D168A, and V170A (AnaSpec, USA). The D168A and V170A mutants conferred low-level resistance (3.6 and 2.4-fold increase in IC50, respectively) to 5a3, whereas the R155K and A156V mutants conferred no resistance (Table 1).
Molecular docking
In order to better understand how 5a3 interacts with NS3/4A protease, we performed molecular docking studies. The docking results of compound 5a3 were generated via the best docking-scoring combinations from Autodock4.2 for Windows-based PCs. Biliverdin is the only noncompetitive NS3/4A inhibitor described in the literature (Zhu et al., 2010). Unfortunately, no crystallographic structure has been solved until now.
Shiryaev and co-workers described three potential allosteric sites in the NS3 domain (Shiryaev et al., 2012). Based on their findings and the experimental data of the mutant enzymes, docking studies were confined within a 10 Å sphere centered at V167.
To evaluate the accuracy of this methodology, re-docking was performed with the co-crystallized form. The docked ligand revealed a conformation similar to the crystallized conformation and had a root-mean-square deviation (RMSD) of 1.09 Å, indicating that the scoring function was successful.
Analysis of the binding mode showed that the peptide mimetic moiety of compound 5a3 forms hydrogen bonds with residues R123, G124, R155, and D168 in the NS3/4A protease, and these bonds have distances of 2.4, 1.6, 3.5, and 1.6 Å, respectively (Fig. 6). Some of these interactions were also described by Shiryaev and co-workers. Interestingly, a π–π interaction was observed between the phenyl moiety of compound 5a3 and residue F154. These interactions may contribute to the stabilization and maintenance of 5a3 in the allosteric site of the NS3/4A protease.
a A visual inspection of compound 5a3 and important residues for HCV NS3/4A protease (2OC0) binding and inhibition (blue); the figure was constructed using Maestro Version 9.2 (Schrodinger Inc). Weak interactions were omitted to allow improved visualization. The π–π interaction between 5a3 and F154 is shown as a green line; a strong hydrogen bond between 5a3 and R123 is shown as a pink dashed line. b Three-dimensional view of 5a3 in 2OC0 (image shown as pink cartoon). Atoms are represented with different colors: red (oxygen), blue (nitrogen), white (hydrogen), purple (carbon in 5a3), and green (carbon in important residues). Principal hydrogen bonds are shown as black dashed lines (R123, V168, and R155). PyMOL was used to construct the images (Schrodinger Inc) (Color figure online)
In silico pharmacokinetic and toxicity analyses
The advent of predictive tools to screen the pharmacokinetic and toxicological properties of drugs has revolutionized the drug discovery process. This technology enables the filtering of weak drug candidates. Eliminating drugs early in the discovery process helps to decrease the number of failures in clinical trials. Traditionally, these predictive tools were applied only at the end of the drug discovery process, but they are now utilized during initial phases of drug development. Removing molecules with poor pharmacokinetic properties at an early stage leads to significant cost savings (Kumar et al., 2011). Therefore, 5a3 was submitted for in silico pharmacokinetics evaluation. Because good absorption is necessary for oral administration, we analyzed this derivative according to the “rule-of-five” developed by Lipinski and co-workers (Lipinski, 2004). The rule-of-five defines the theoretical parameters for a chemical compound to have good oral bioavailability. The rule states that the most “druglike” molecules possess a clogP of less than or equal to 5, a molecular weight (MW) of less than or equal to 500, 10 or fewer hydrogen bond acceptors, and 5 or fewer hydrogen bond donors. Molecules violating more than one of these rules may suffer from bioavailability problems. The results showed that compound 5a3 fulfilled the Lipinski “rule-of-five”. Importantly, according to a theoretical analysis of lipophilicity (clog P), the most active inhibitor possesses sufficient hydrophobicity to penetrate biological membranes.
We also analyzed the druglikeness and drug score values of compound 5a3 and compared them with the corresponding values for boceprevir and telaprevir. In both evaluations, 5a3 showed better results than both of the marketed drugs (Fig. 7).
Comparison of the druglikeness (a) and drug score (b) values of compound 5a3, boceprevir, and telaprevir. These parameters were calculated using Osiris Property Explorer (Sander et al., 2009)
Furthermore, the theoretical toxicity risks obtained from the Osiris program (Sander et al., 2009) for compound 5a3 were low with respect to the mutagenic, tumorigenic, irritant, and reproductive risks (data not shown).
Although these are not conclusive results, the molecular modeling data obtained in this work show the potential for compound 5a3, which could be a lead candidate for the design of safe derivatives with activity against HCV.
Conclusions
Novel peptide mimetic compounds derived from an isomannide-tartaric acid scaffold were synthesized. The synthesized compounds were assayed using HCV protease genotype 1b. Compound 5a3 was found to be a noncompetitive inhibitor (K i/K M = 43.6) of NS3/4A protease. Docking studies showed important interactions between compound 5a3 and residues R123, G124, R155, and D168 in NS3/4A pro. These interactions included hydrogen bonds and a π–π interaction with residues in the allosteric site of NS3/4A pro. This structural information may be helpful for the design of more potent noncompetitive inhibitors. The biological and theoretical results indicate that 5a3 is a promising lead compound for the development of new drugs to combat HCV infection.
Experimental section
Isomannide and both acids (2a–2b) were purchased from Aldrich Chem. Co. and used without further purification.
Spectroscopic data for all synthesized compounds for formation of the 3 (Barros et al., 2009)
1,4:3,6-Dianhydro-2-O-tosyl-d-mannitol
Yield: 40 %; white solid; mp 103–104 °C.
IR (KBr, ν cm−1): 3526, 2933, 2866, 1596, 1359, 1189, 1173, 1050, 1019, 818, 663.
1H NMR (300 MHz, CDCl3): δ = 7.84 (2H, d, J = 8.1 Hz, Ar–H), 7.34 (2H d, J = 8.1 Hz, Ar–H), 4.91 (1H dd, J = 6.6, 5.5 Hz, H6′), 4.49 (1H, t, J = 4.8 Hz, H3′a), 4.42 (1H, t, J = 5.1 Hz, H1′), 4.29–4.25 (1H, t, m, H1′), 4.04–3.95 (2H, m, H4′), 3.79 (1H, t, J = 7.8 Hz, H3′), 3.55 (1H, dd, J = 7.2, 1.8 Hz, H6′a), 2.46 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 145.2 (Ar-Tosyl), 133.0 (Ar-Tosyl), 129.8 (Ar-Tosyl), 127.9 (Ar-Tosyl); 81.3 (C5′), 80.0 (C2′), 78.3 (C3′), 73.9 (C1′), 72.2 (C6′), 70.0 (C4′), 21.6 (CH3).
1,4:3,6-Dianhydro-2-O-benzyl-5-O-tosyl-d-mannitol
Yield: 75 %; colorless oil.
[α] 20D +98 (c 0.10, DMSO).
IR (KBr, ν cm−1): 3063, 2977, 2950, 2879, 1598, 1454, 1366, 1190, 1178, 1141, 1027, 853.
1H NMR (300 MHz, CDCl3): δ = 7.81 (2H, d, J = 8.4 Hz, Ar–H), 7.35–7.32 (7H, m, Ar–H), 4.91–4.89 (1H, m, H3′), 4.69 (1H, d, J = 12.0 Hz, CH2), 4.50 (1H, d, J = 12.0 Hz, CH2), 4.47 (2H, t, J = 2.1 Hz, H3′a e H4′), 4.04–3.79 (4H, m, H1′, H4′ e H6′), 3.62 (1H, t, J = 8.4 Hz, H6′a), 2.44 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 145.0 (Ar-Tosyl), 137.4 (Ar-Tosyl), 133.2 (Ar-Benzyl), 129.7 (Ar-Tosyl), 128.4 (Ar-Tosyl), 127.9 (Ar-Benzyl), 127.8 (Ar-Benzyl), 80.1 (C2′), 80.0 (C5′), 78.8 (C6′), 78.4 (C3′), 72.5 (CH2-Benzyl), 71.0 (C4′), 70.1 (C1′), 21.6 (CH3).
1,4:3,6-Dianhydro-2-azido-5-O-benzyl-2-deoxy-d-glucitol
Yield: 73 %; pale yellow oil;
[α] 20D +92 (c 0.10, DMSO).
IR (KBr, ν cm−1): 3063, 3031, 2946, 2878, 2102, 1455, 1320, 1256, 1135, 1100, 1083, 1021, 739.
1H NMR (300 MHz, CDCl3): δ = 7.36–7.26 (5H, m, Ar–H), 4.75 (1H, d, J = 11.7 Hz, CH2), 4.66 (1H, t, J = 4.5 Hz, H3′), 4.54 (1H, d, J = 11.7 Hz, CH2), 4.48 (1H, d, J = 4.2 Hz, H3′a), 4,15–4,00 (4H, m, H1′ e H4′), 3.86 (1H, dd, J = 6.3 e 2.7 Hz, H6′a), 3.66 (1H, t, J = 7.8 Hz, H6′).
13C NMR (75 MHz, CDCl3): δ = 137.4 (Ar-Benzyl), 128.3 (Ar-Benzyl), 127.8 (Ar-Benzyl), 127.7 (Ar-Benzyl), 86.4 (C2′), 80.5 (C5′), 78. 8 (C6′), 72.6 (C3′), 72.4 (CH2-Benzyl), 70.7 (C4′), 66.2 (C1′).
(3 S,6R)-6-(benzyloxy)hexahydrofuro[3,2-b]furan-3-amine (3)
Colorless oil in quantitative yield.
[α] 20D +104 (c 0.10, DMSO).
IR (KBr, ν cm−1): 3360, 2876, 1605, 1455, 1369, 1209, 1065, 1017, 751, 700.
1H NMR (300 MHz, CDCl3): δ = 7.37–7.35 (5H, m, Ar–H), 4.76 (2H, d, J = 11.4 Hz, CH2–Ar), 4.68 (1H, t, J = 4.2 Hz, H4′), 4.54 (2H, d, J = 12.0 Hz, CH2–Ar), 4.27 (1H, d, J = 4.2 Hz, H1′ or H4′), 4.06–4.02 (1H, m, H1′ or H4′), 3.85 (1H, dd, J = 6.3 e 2.4 Hz, H1′), 3.76 (1H, d, J = 9.3 Hz, H6′), 3.64 (1H, t, J = 7.8 Hz, H3′a), 3.53 (1H, d, J = 4.2 Hz, H6′a), 3.45 (1H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 137.9 (Ar–H), 128.6 (Ar–H), 128.0 (Ar–H), 127.8 (Ar–H), 90.0 (C2′), 80.1, 79.4 (C6′), 76.3 (CH2–Ar), 72.6 (C3′), 70.5 (C4′), 58.8 (C1′).
HRMS (FAB): m/z [M + H]+ calcd for C13H18NO3: 236.1287; found: 236.1284.
General procedure for the formation of intermediates 4
To a 0 °C cooled solution of anhydride 6a,b (4.02 mmol, 0.868 g) in dry dichloromethane (5 mL) was added dropwise a solution of the appropriate amino acid ester hydrochloride (a1–a4) (4.82 mmol) and 4-methylmorpholine (4.82 mmol, 0.53 mL) in dry dichloromethane (2.5 mL). The mixture was stirred overnight, and the volatiles were removed. The residue was dissolved in ethyl acetate (or dichloromethane) and washed successively with 10 % HCl and brine. The organic layer was dried over Na2SO4 and concentrated to furnish 4a1–4a3 and 4b2–4b3 as viscous oils, which were purified by recrystallization (CH2Cl2/MeOH) to afford the hygroscopic solids 4a1 (40 %), 4a2, (55 %), 4a3, (50 %), 4b2, (84 %), and 4b3 (78 %).
The free amine of proline methyl ester hydrochloride was obtained by dissolving the salt (7.87 mmol, 1.3 g) in a minimum volume of 20 % K2CO3 and extracting eight times with diethyl ether. Concentrating the pooled organic phases afforded the free aminoester (4.02 mmol, 0.519 g) as an oil; this oil was directly dissolved in dry dichloromethane (2.5 mL) and added to a solution of anhydride 3 (4.02 mmol, 0.868 g) in dry dichloromethane (5 mL). The mixture was stirred overnight, and the solvent was removed. The product was purified by recrystallization (CH2Cl2/MeOH) to afford hygroscopic solids 4a4 (84 %) and 4b4 (95 %).
a1–a4 Substituents
Ethyl 2-aminoacetate (a1)
Yield: 99 %; white solid; mp 140–142 °C
1H NMR (300 MHz, CDCl3): δ = 4.30 (2H, s, H5); 4.00–3.91 (2H, q, H7, J = 7,5 Hz); 1.32 (3H, t, H8, J = 7,5 Hz).
13C NMR (75 MHz, CDCl3): δ = 167.2 (C6), 61.1 (C7), 41.8 (C5), 14.4 (C8).
(S)-1-Methoxy-1-oxo-3-phenylpropan-2-ammonium chloride (a2)
Yield: 99 %; white solid; mp 159–161 °C.
IR (KBr, ν cm−1): 2840, 2702, 2625, 2358, 2014, 1744, 1583, 1495, 1447, 1399, 1291, 1238, 1146, 1118, 1083, 1061, 989, 934, 902, 864, 809, 741, 701, 626.
1H NMR (300 MHz, CDCl3): δ = 7.40–7.29 (5H, m, Ar–H), 4.79 (1H, dd. J = 7.4, 6.0 Hz, H5), 3.66 (3H, s, CH3), 3.52 (1H, dd. J = 10.5, 4.0 Hz, H8), 3.25 (1H, dd. J = 10.5, 4.0 Hz, H8).
13C NMR (75 MHz, CDCl3): δ = 130.2 (C9), 128.1 (C10), 127.6 (C11), 125.9 (C12), 52.3(C5), 50.7 (C8), 52.6 (C5), 50.3 (C7).
(S)-1-Methoxy-3-methyl-1-oxobutan-2-ammonium chloride (a3)
Yield: 99 %; white solid; mp 171 °C.
IR (KBr, ν cm−1): 3391, 2915, 2741, 2565, 2361, 1741, 1633, 1442, 1358, 1238, 1043, 1005, 919, 860.
1H NMR (300 MHz, CDCl3): δ = 3.87 (3H, s, CH3), 3.50–3.36 (2H, m, H2 e H5), 2.50–2.39 (1H, m, H5), 2.23–2.06 (4H, m, H3 e H4).
13C NMR (75 MHz, CDCl3): δ = 169.9 (C6), 62.9 (C5), 56.3(C7), 26.1 (C8), 18.4 (C9), 18.4 (C10).
(S)-2-(Methoxycarbonyl)pyrrolidinium chloride (a4)
Yield: 99 %; white solid; mp 69 °C.
IR (KBr, ν cm−1): 3437; 3400; 2970; 2882; 1732; 1594; 1571; 1434; 1379; 1354; 1286; 1219; 1170; 1103; 1064; 1037; 972; 928; 877; 831; 771; 734
1H NMR (300 MHz, CDCl3): δ = 4.04 (1H, d, J = 4.7 Hz, H5), 3.87 (3H, s, CH3), 2.41–2.32 (1H, m, H8), 1.04 (6H, m, H9 and H10).
13C NMR (75 MHz, CDCl3): δ = 169.9 (COOCH3), 62.9 (C5), 56.3(COOCH3), 26.1 (CH(CH3)2), 18.4 (CH3), 18.4 (CH3).
(2S,3S)-2,3-Diacetoxy-4-(2-ethoxy-2-oxoethylamino)-4-oxobutanoic acid (4a1)
[α] 20D +20 (c 0.10, CH2Cl2).
IR (KBr, ν cm−1): 3851, 3626, 3369, 2926, 2853, 2479, 1746, 1731, 1694, 1681, 1652, 1634, 1538, 1532, 1463, 1455, 1377, 1271, 1212, 1116, 1073, 1032, 992, 896, 866, 799, 738, 702.
1H NMR (300 MHz, CDCl3): δ = 5.83 (1H, d, J = 2.4 Hz, H2 or H3), 5.64 (1H, d, J = 2.4 Hz, H2 or H3), 4.23 (2H, q, J = 7.2 Hz, H7), 4.13 (1H, dd, J = 18.3, 5.1 Hz, H5), 3.99 (1H, dd, J = 18.3, 5.1 Hz, H5), 2.22 (3H, s, CH3), 2.17 (3H, s, CH3), 1.30 (3H, t, J = 7.2 Hz, H8).
13C NMR (75 MHz, CDCl3): δ = 169.8 (C1), 169.6 (C4), 169.3 (C6), 169.1(C=O), 166.2 (C=O), 71.7 (C2), 70.7 (C3), 61.8 (C7), 41.2 (C5), 20.4 (CH3), 20.2 (CH3), 14.0 (C8).
HRMS-FAB: m/z [M + 1] Calcd for C12H17NO9: 319.0903; found: 320.0611.
(2S,3S)-2,3-Diacethoxy-4-((S)-1-methoxy-1-oxo-3-phenylpropan-2-ylamino)-4-oxobutanoic acid (4a2)
[α] 20D +26.2 (c 1.0, CH2Cl2).
IR (KBr, ν cm−1): 3627, 3585, 3389, 2955, 1732, 1716, 1681, 1652, 1538, 1497, 1455, 1436, 1374, 1220, 1132, 1069, 946, 877, 738, 702.
1H NMR (300 MHz, CDCl3): δ = 7.32–7.18 (5H, m, H10–12), 7.06 (1H, d, J = 5.4 Hz, NH), 5.74 (1H,sl, H2 or H3), 5.34 (1H, sl, H2 or H3), 4.79 (1H, dd, J = 8.4, 3.9 Hz, H5), 3.66 (3H, s, H7), 3.07 (2H, m, CH2PHe), 2.05 (3H, s, CH3), 2.04 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 171.9 (C1), 171.2 (C4), 170.2 (C6), 169.4 (C=O), 166.4 (C=O), 135.2 (C9), 129.1 (C10), 128.6 (C11), 127.2 (C12), 72.3(C2), 70.7 (C3), 52.6 (C5), 52.3 (C7), 37.3 (C8), 20.4 (CH3), 20.3 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C18H21NO9: 395.1216; found: 396.6837.
(2S,3S)-2,3-Diacethoxy-4-(S)-1-methoxy-3-methyl-1-oxobutan-2-ylamino)-4-oxobutanoic acid (4a3)
[α] 20D +12.6 (c 1.0, CH2Cl2).
IR (KBr, ν cm−1): 3854, 3752, 3736, 3650, 3399, 2966, 2936, 2850, 2615, 1737, 1662, 1540, 1439, 1376, 1221, 1151, 1129, 1073, 1017, 945, 892, 823.
1H NMR (300 MHz, CDCl3): δ = 6.68 (1H, d, J = 7.6 Hz, NH), 5.76 (1H, d, J = 2.4 Hz, H2 or H3), 5.64 (1H, d, J = 2.7 Hz, H2 or H3), 4.13 (1H, dd, J = 14.4, 7.2 Hz, H5), 3.76 (3H, s, H7), 2.22 (3H, s, CH3), 2.17 (3H, s, CH3), 1.28–1.24 (1H, m, H8), 0.91 (3H, d, J = 6.9 Hz, H9), 0.86 (3H, d, J = 6.9 Hz, H10).
13C NMR (75 MHz, CDCl3): δ = 172.0 (C1), 169.8 (C4), 169.6 (C6), 169.2 (C=O), 166.1 (C=O), 72.1 (C2), 70.5 (C3), 56.9 (C5), 52.3(C7), 31.0 (C8), 20.4 (CH3), 20.1 (CH3), 18.8 (C9), 17.4 (C10).
HRMS-FAB: m/z [M + 1] Calcd for C14H21NO9: 347.1216; found: 348.6792.
(2S,3S)-2,3-Diacetoxy-4-(2-(methoxycarbonyl)pyrrolidin-1-yl)-4-oxobutanoic acid (4a4)
[α] 20D −47.0 (c 1.0, CH2Cl2).
IR (KBr, ν cm−1): 3315, 2965, 2882, 1753, 1669, 1538, 1453, 1368, 1212, 1136, 1096, 1078, 961, 881, 740.
1H NMR (300 MHz, CDCl3): δ = 5.74 (1H, d, J = 4.5 Hz, H2 or H3), 5.59 (1H, d, J = 4.5 Hz, H2 or H3), 4.43 (1H, dd, J = 8.2, 3.0 Hz, H5), 4.00–3.94 (1H, m, C10), 3.69 (3H, s, C7), 3.60–3.51 (1H, m, C10), 2.16 (3H, s, CH3), 2.15 (3H, s, CH3), 2.11–2.09 (2H, m, C8), 2.07–1.98 (2H, m, C9).
13C NMR (75 MHz, CDCl3): δ = 171.8 (C1), 169.9 (C4), 169.7 (C6), 169.1 (C=O), 165.1 (C=O), 70.9 (C2), 70.3 (C3), 59.7 (C5), 52.2 (C7), 46.9 (C10), 28.5 (C8), 24.7 (C9), 20.4 (CH3), 20.3 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C14H19NO9: 345.1060; found: 346.1280.
(2R,3R)-2,3-Diacetoxy-4-((S)-1-methoxy-1-oxo-3-phenylpropan-2-ylamino)-4- oxobutanoicacid (4b2)
[α] 20D +32.5 (c 0.8, CH2Cl2).
IR (KBr, ν cm−1): 3500, 2957, 1750, 1677, 1534, 1455, 1374, 1214, 1115, 1063, 972, 703.
1H NMR (300 MHz, CDCl3): δ = 7.32–7.22 (5H, m, H9–14), 6.60 (1H, d, J = 7.9 Hz, NH), 5.73 (1H, sl, H2 or H3), 5.50 (1H, sl, H2 or H3), 4.88 (1H, q, J = 5.7 Hz, H5), 3.73 (3H, s, H7), 3.15–3.06 (2H, m, H8), 2.07 (3H, s, CH3), 2.05 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 171.2 (C1), 171.1 (C4), 169.9 (C6), 169.1 (C=O), 166.1 (C=O), 135.2 (C9), 129.1 (C10), 128.6 (C11), 127.3 (C12), 72.3 (C2), 70.6 (C3), 52.6 (C5), 52.4 (C7), 37.4 3 (C8), 20.4 (CH3), 20.3 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C18H21NO9: 395.1216; found: 396.1488.
(2R,3R)-2,3-Diacetoxy-4-((S)-1-methoxy-3-methyl-1-oxobutan-2-ylamino)-4-oxobutanoic acid (4b3)
[α] 20D +26.7 (c 0.9, CH2Cl2).
IR (KBr, ν cm−1): 3344, 2966, 1745, 1535, 1438, 1375, 1214, 1151, 1120, 1066, 973, 875, 736.
1H NMR (300 MHz, CDCl3): δ = 6.77 (1H, d, J = 8.9 Hz, NH), 5.73 (1H, d, J = 2.4 Hz, H2 or H3), 5.58 (1H, d, J = 2.4 Hz, H2 or H3), 4.53 (1H, dd, J = 8.8, 4.8 Hz, H5), 3.74 (3H, s, H7), 2.20 (3H, s, CH3), 2.21 (3H, s, CH3), 2.15 (1H, m, H8), 0.90 (3H, d, J = 6.8 Hz, H9), 0.86 (3H, d, J = 6.8 Hz, H10).
13C NMR (75 MHz, CDCl3): δ = 172.0 (C1), 170.2 (C4), 169.9 (C6), 169.4 (C=O), 166.5 (C=O), 72.4 (C2), 71.0 (C3), 56.9 (C5), 52.3 (C7), 31.1 (C8), 20.4 (CH3), 20.1 (CH3), 18.9 (C9), 17.5 (C10).
HRMS-FAB: m/z [M + 1] Calcd for C14H21NO9: 347.1216; found: 348.1465.
(2R,3R)-2,3-Diacetoxy-4-((S)-(2-(methoxycarbonyl)pyrrolidin-1-yl)-4-oxobutanoic acid (4b4)
[α] 20D −42.5 (c 0.8, CH2Cl2).
IR (KBr, ν cm−1): 3449, 2957, 1751, 1655, 1637, 1618, 1459, 1438, 1375, 1220, 1120, 1073, 874, 733.
1H NMR (300 MHz, CDCl3): δ = 5.74 (1H, d, J = 4.3 Hz, H2 or H3), 5.63 (1H, d, J = 4.3 Hz, H2 or H3), 4.44 (1H, dd, J = 8.4, 3.1 Hz, H5), 4.00–3.95 (1H, m, C10), 3.69 (3H, s, C7), 3.57–3.52 (1H, m, C10), 2.17 (3H, s, CH3), 2.16 (3H, s, CH3), 2.11–2.07 (2H, m, C8), 2.04–1.95 (2H, m, C9).
13C NMR (75 MHz, CDCl3): δ = 171.8 (C1), 169.9 (C4), 169.7 (C6), 168.9 (C=O), 164.9 (C=O), 70.7 (C2), 69.9 (C3), 59.7 (C5), 52.3 (C7), 47.0 (C10), 28.6 (C8), 24.7 (C9), 20.4 (CH3), 20.3 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C14H19NO9: 345.1060; found: 346.1271.
General procedure for the formation of final products 5
To a 0 °C cooled suspension of the appropriate carboxylic acid 4 (0.515 mmol) and amine 3 (0.618 mmol, 0.145 g) in dry dichloromethane (10 mL) under argon atmosphere was added to EDC.HCl (0.790 mmol, 0.125 g), HOBt (0.790 mmol, 0.106 g), and NMM (1.614 mmol, 0.17 mL). The mixture was stirred at 0 °C for 1 h under an argon atmosphere and then stirred overnight at room temperature. The volatiles were removed, and the resulting residue was dissolved in ethyl acetate and washed successively with saturated NaHCO3, 10 % HCl, and brine. The organic phase was dried over Na2SO4 and concentrated to yield crude 5.
(2S,3S)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-(2-ethoxy-2-oxoethylamino)-1,4-dioxobutane-2,3-diyl diacetate (5a1)
Yield: 40 %; pale yellow oil (after purification by chromatographic column; eluent: hexane/ethyl acetate).
[α] 20D +28.0 (c 0.10, CH2Cl2).
IR (neat, cm−1): 3311, 3066, 3032, 2959, 2929, 2875, 1746, 1669, 1602, 1546, 1496, 1455, 1374, 1344, 1274, 1225, 1134, 1094, 1085, 1070, 1017, 962, 912, 880, 845, 798, 780, 746, 698, 662, 638, 603.
1H NMR (300 MHz, CDCl3): δ = 7.37–7.35 (5H, m, Ar–H), 6.75 (1H, t, J = 4.8 Hz, NH); 6.42 (1H, d, J = 8.7 Hz, NH), 5.65 (1H, d, J = 3.6 Hz, H2 or H3), 5.53 (1H, d, J = 3.3 Hz, H2 or H3), 4.76 (1H, d, J = 11.7 Hz, CH2-Ar), 4.62 (1H, t, J = 4.2 Hz, H4′), 4.55 (1H, d, J = 12.0 Hz, CH2-Ar), 4.42 (1H, d, J = 4.2 Hz, H1′ or H4′), 4.39–4.34 (1H, m, H3′), 4.22 (2H, q, J = 7.2 Hz, H7), 4.10–4.05 (4H, m; H1′, H6′ e H3′a), 4.00 (1H, d, J = 5.1 Hz, H6′a) 3.86 (1H, dd, J = 9.0, 7.5 Hz, H5), 3.67 (1H, dd, J = 18.3, 5.1 Hz, H5), 2.18 (3H, s, CH3), 2.17 (3H, s, CH3), 1.29 (3H, t, J = 7.2 Hz, H8).
13C NMR (75 MHz, CDCl3): δ = 169.3 (C1), 169.0 (C4), 168.9 (C6), 166.3 (C=O), 165.7 (C=O), 137.6 (Ar–H), 128.4 (Ar–H), 127.8 (Ar–H), 127.7 (Ar–H), 86.6 (C3′a and C6′a), 80.3 (C2), 78.9 (C3), 72.9 (CH2-Ar), 72.5 (C4′), 71.9 (C6′), 70.8 (C1′), 61.7 (C7), 57.0 (C3′), 41.1 (C5), 20.5 (CH3), 20.4 (CH3), 14.0 (C8).
HRMS-FAB: m/z [M + 1] Calcd for C25H32N2O11: 536.5284; found: 537.73676.
(2S,3S)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-((S)-1-methoxy-1-oxo-3-phenylpropan-2-ylamino)-1, 4-dioxobutane-2,3-dihyl diacetate (5a2)
Yield: 40 %; pale yellow solid (after recrystallization from dichloromethane/ethyl ether); mp 122–123 °C.
[α] 20D +50.0 (c 0.10, CH2Cl2).
IR (KBr, ν cm−1): 3286, 3064, 3031, 2952, 2880, 2359, 1755, 1657, 1540, 1497, 1455, 1436, 1373, 1343, 1374, 1256, 1206, 1136, 1098, 1084, 1058, 1029, 963, 912, 879, 749, 700.
1H NMR (300 MHz, CDCl3): δ = 7.35–7.26 (8H, m, H10-12, Ar–H), 7.05 (2H, d, J = 4.2 Hz, Ar–H), 6.55 (1H, d, J = 4.8 Hz, NH), 6.46 (1H, d, J = 4.5 Hz, NH), 5.66 (1H, d, J = 1.5 Hz, H2 or H3), 5.54 (1H, d, J = 1.5 Hz, H2 or H3), 5.30 (2H, s, CH2Phe), 4.62 (1H, dt, J = 11.4, 2.7 Hz, H5), 4.55 (1H, d, J = 6.9 Hz, H4′), 4.41 (1H, dd, J = 10.1, 2.4 Hz, H3′a), 4.35–4.33 (1H, m, H3′), 4.08–4.03 (2H, m, H1′), 3.85–3.82 (2H, m, H6′ or H6′a), 3.75 (3H, s, H7), 3.66 (1H, t, J = 5.0 Hz, H4′), 3.13 (1H, dd, J = 11.6 e 8.1 Hz, H8), 3.08 (1H, dd, J = 11.6 e 8.1 Hz, H8), 2.13 (3H, s, H13 or H14), 2.06 (3H, s, H13 or H14).
13C NMR (75 MHz, CDCl3): δ = 169.3 (C1), 169.2 (C4), 168.7 (C6), 165.8 (C=O), 165.7 (C=O), 137.6 (C9), 135.2 (Ar–H), 129.2 (C10), 128.7 (C11), 128.5 (C12), 127.9 (Ar–H), 127.8 (Ar–H), 127.4 (Ar–H), 86.8 (C3′a or C6′a), 86.7 (C3′a or C6′a), 80.3 (C2 or C3), 79.0 (C2 or C3), 73.0 (CH2Phe), 72.5 (C1′ or C4′), 72.3 (C6′), 70.8 (C1′ or C4′), 57.1 (C3′), 52.6 (C5), 52.4 (C7), 37.5 (C8), 20.5 (CH3), 20.4 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C31H36N2O11: 612.2319; found: 613.7522.
(2S,3S)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-((S)-1-methoxy-3-methyl-1-oxobutan-2-ylamino)-1, 4-dioxobutane-2,3-dihyl diacetate (5a3)
Yield: 42 %; pale yellow solid (after recrystallization from dichloromethane/ethyl ether); mp 112–113 °C.
[α] 20D +24.0 (c 0.10, CH2Cl2)
IR (KBr, ν cm−1): 3851, 3742, 3646, 3287, 3065, 2962, 2879, 1754, 1656, 1539, 1455, 1436, 1373, 1274, 1207, 1141, 1098, 1059, 1027, 964, 876, 845, 750, 699, 651.
1H NMR (300 MHz, CDCl3): δ = 7.35–7.33 (5H, m, Ar–H), 6.68 (1H, d, J = 5.1 Hz, NH), 6.58 (1H, d, J = 4.5 Hz, NH), 5.66 (1H, d, J = 2.1 Hz, H2 or H3), 5.56 (1H, d, J = 1.8 Hz, H2 or H3), 5.51 (2H, s, CH2Phe), 4.74 (1H, d, J = 7.2 Hz, H5), 4.64 (1H, t, J = 3.0 Hz, H4′), 4.55 (1H, t, J = 3.0 Hz, H3′a), 4.35–4.33 (1H, m, H3′), 4.06–4.03 (2H, m, H1′), 3.87–3.80 (3H, m, H6′ or H6′a or H4′), 3.73 (3H, s, H7), 2.17–2.15 (1H, m, H8), 2.17 (3H, s, CH3), 2.16 (3H, s, CH3), 0.91 (3H, d, J = 3.9 Hz, H9 or H10), 0.87 (3H, d, J = 4.2 Hz, H9 or H10).
13C NMR (75 MHz, CDCl3): δ = 169.3 (C1), 169.2 (C4), 168.9 (C6), 166.2 (C=O), 166.0 (C=O), 137.6 (Ar–H), 128.3 (Ar–H), 127.8 (Ar–H), 127.7 (Ar–H), 86.6 (C3′a or C6′a), 86.5 (C3′a or C6′a), 80.2 (C2 or C3), 78.9 (C2 or C3), 72.7 (CH2Phe), 72.4 (C1′ or C4′), 72.1 (C6′), 70.8 (C1′ or C4′), 57.0 (C3′), 56.9 (C5), 52.2 (C7), 30.9 (C8), 20.5 (CH3), 20.4 (CH3), 18.7 (C9 and C10).
HRMS-FAB: m/z [M + 1] Calcd for C27H36N2O11: 564.5815; found: 565.76803.
(2S,3S)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-(R)-(2-(methoxycarbonyl)pyrrolidin-1-yl)-1,4-dioxobutane-2,3-diyl diacetate (5a4)
Yield: 50 %; hygroscopic solid (after purification by chromatographic column; eluent: hexane/ethyl acetate).
[α] 20D +13.0 (c 0.8, CH2Cl2).
IR (KBr, ν cm−1): 3414, 2953, 2882, 1751, 1665, 1535, 1451, 1438, 1372, 1213, 1135, 1071, 960, 700.
1H NMR (300 MHz, CDCl3): δ = 7.36–7.28 (5H, m, Ar–H), 6.48 (1H, br s, NH), 5.68 (1H, d, J = 4.9 Hz, H2 or H3), 5.55 (1H, d, J = 4.9 Hz, H2 or H3), 4.78 (1H, d, J = 11.7 Hz, H6′a), 4.64–4.61 (1H, m, H5), 4.58 (1H, d, J = 11.7 Hz, H3′a), 4.43–4.32 (3H, m, CH2Phe, H3′), 4.12–4.03 (2H, m, H4′), 3.96–3.90 (1H, m, H6′), 3.90–3.83 (3H, m, H1′, H10), 3.78 (1H, s, H10), 3.69 (3H, s, H7), 2.21–2.18 (2H, m, H8), 2.15 (3H, s, CH3), 2.10 (3H, s, CH3), 2.08–2.00 (2H, m, H9).
13C NMR (75 MHz, CDCl3): δ = 171.7 (C1), 169.5 (C4), 169.1(C6), 165.7 (C=O), 164.7 (C=O), 137.6 (Ar–H), 128.4 (Ar–H), 127.9 (Ar–H), 127.8 (Ar–H), 86.7 (C3′a or C6′a), 86.5 (C3′a or C6′a), 80.3 (C2 or C3), 78.9 (C2 or C3), 72.8 (CH2Phe), 72.5 (C1′ or C4′), 71.2 (C6′), 70.9 (C1′ or C4′), 59.3 (C5), 57.0 (C3′), 52.1 (C7), 46.8 (C10), 28.8 (C8), 24.6 (C9), 20.6 (CH3), 20.3 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C27H34N2O11: 562.2163; found: 563.2430.
(2R,3R)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-((S)-(1-methoxy-1-oxo-3-phenylpropan-2-ylamino)-1,4-dioxobutane-2,3-diyl diacetate (5b2)
Yield: 50 %; white solid (after purification by chromatographic column; eluent: hexane/ethyl acetate); mp 178–180 °C.
[α] 20D +68.2 (c 0.8, CH2Cl2).
IR (KBr, ν cm−1): 3287, 3066, 3030, 2953, 2879, 1755, 1657, 1540, 1497, 1436, 1372, 1257, 1204, 1134, 1083, 1057, 962, 747, 699.
1H NMR (300 MHz, CDCl3): δ = 7.35–7.24 (8H, m, H10, H12, Ar–H), 7.04 (2H, d, J = 6.2 Hz, H11), 6.59 (1H, d, J = 2.8 Hz, NH), 5.66 (1H. dd, J = 13.7, 5.5 Hz, H2 or H3), 5.54 (1H, d, J = 11.8 Hz, H2 or H3), 4.89 (1H, t, J = 4.6 Hz, H5), 4.75 (1H, d, J = 11.8 Hz, H6′a), 4.60–4.59 (2H, m, CH2Phe), 4.56 (1H, d, J = 11.8 Hz, H3′a), 4.40 (1H, d, J = 4.3 Hz, H6′), 4.34–4.32 (1H, m, H3′), 4.06–4.02 (2H, m, H1′), 3.85–3.82 (2H, m, H4′), 3.74 (3H, s, H7), 3.15 (1H, dd, J = 11.6, 8.1 Hz, H8), 3.08 (1H, dd, J = 11.6, 8.1 Hz, H8), 2.12 (3H, s, CH3), 2.05 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 171.2 (C1), 169.3 (C4), 168.7 (C6), 165.9 (C=O), 165.7 (C=O), 137.6 (C9), 135.2 (Ar–C), 129.2 (C10), 128.6 (C11), 128.4 (C12), 127.9 (Ar–C), 127.8 (Ar–C), 127.3 (Ar–C), 86.7 (C3′a or C6′a), 80.3 (C3′a or C6′a), 78.9 (C2 or C3), 72.9 (CH2Phe), 72.5 (C1′ or C4′), 72.2 (C2 or C3), 71.8 (C6′), 70.8 (C1′ or C4′), 57.1 (C3′), 52.6 (C5), 52.4 (C7), 37.5 (C8), 20.5 (CH3), 20.4 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C31H36N2O11: 612.2319; found: 613.2689.
(2R,3R)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-((S)-1-methoxy-3-methyl-1-oxobutan-2-ylamino)-1,4-dioxobutane-2,3-diyl diacetate (5b3)
Yield: 75 %; white solid (after purification by chromatographic column; eluent: hexane/ethyl acetate); mp 84–85 °C.
[α] 20D +71.2 (c 0.8, CH2Cl2).
IR (KBr, ν cm−1): 3279, 2963, 2876, 1753, 1654, 1541, 1468, 1373, 1259, 1208, 1144, 1061, 963, 874, 737.
1H NMR (300 MHz, CDCl3): δ = 7.34–7.27 (5H, m, Ar–H), 6.70 (1H, d, J = 8.7 Hz, NH), 6.61 (1H, d, J = 7.4 Hz, NH), 5.66 (1H, d, J = 3.1 Hz, H2 or H3), 5.56 (1H, d, J = 3.1 Hz, H2 or H3), 4.75 (d, 1H, d, J = 11.8 Hz, H6′a), 4.64–4.61 (1H, m, H5), 4.54 (1H, t, J = 11.8 Hz, H3′a), 4.51 (1H, dd, J = 8.8, 4.8 Hz, H6′), 4.42–4.40 (1H, m, H3′), 4.30–4.33 (2H, m, CH2Phe), 4.06–4.03 (2H, m, H1′), 3.87–3.82 (2H, m, H4′), 3.73 (3H, s, H7), 3.65 (1H, dd, J = 9.0, 7.3 Hz, H8), 2.17 (3H, s, CH3), 2.15 (3H, s, CH3), 0.89 (3H, d, J = 3.9 Hz, H9 or H10), 0.85 (3H, d, J = 6.8 Hz, H9 or H10).
13C NMR (75 MHz, CDCl3): δ = 171.7 (C1), 169.4 (C4), 169.0 (C6), 166.2 (C=O), 165.8 (C=O), 137.6 (Ar–C), 128.4 (Ar–C), 127.9 (Ar–C), 127.8 (Ar–C), 86.7 (C3′a or C6′a), 80.3 (C3′a or C6′a), 78.9 (C2 or C3), 72.9 (CH2Phe), 72.5 (C1′ or C4′), 72.4 (C2 or C3), 71.8 (C6′), 70.8 (C1′ or C4′), 57.1 (C3′), 56.9 (C5), 52.2 (C7), 31.0 (C8), 20.6 (CH3), 20.5 (CH3), 18.2 (C9), 17.5 (C10).
HRMS-FAB: m/z [M + 1] Calcd for C27H36N2O11: 564.2319; found: 565.2684.
(2R,3R)-1-((3S,6R)-6-(Benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-4-((R)-2(methoxycarbonyl)pyrrolidin-1-yl)-1,4-dioxobutane-2,3-diyl diacetate (5b4)
Yield: 51 %; white solid (after purification by chromatographic column; eluent: hexane/ethyl acetate); mp 53–54 °C.
[α] 20D +20.5 (c 0.8, CH2Cl2).
IR (KBr, ν cm−1): 3314, 2955, 2880, 1751, 1668, 1538, 1453, 1372, 1212, 1135, 1096, 1070, 961, 881, 742.
1H NMR (300 MHz, CDCl3): δ = 7.36–7.28 (5H, m, Ar–H), 6.61 (1H, d, J = 7.5 Hz, NH), 5.68 (1H, d, J = 5.0 Hz, H2 or H3), 5.54 (1H, d, J = 5.0 Hz, H2 or H3), 4.74 (1H, d, J = 11.7 Hz, H6′a), 4.64 (1H, t, J = 4.7 Hz, H5), 4.57 (1H, d, J = 11.7 Hz, H3′a), 4.42–4.40 (3H, m, H3′), 4.35–4.33 (2H, m, CH2Phe), 4.08–4.03 (3H, m, H4′, H6′), 3.96–3.91 (1H, s, H10), 3.87–3.84 (3H, m, H1′, H10), 3.69 (3H, s, H7), 2.23–2.19 (2H, m, H8), 2.15 (3H, s, CH3), 2.10 (3H, s, CH3), 2.04–1.97 (2H, m, H9).
13C NMR (75 MHz, CDCl3): δ = 171.7 (C1), 169.5 (C4), 169.2 (C6), 165.8 (C=O), 164.7 (C=O), 137.6 (Ar–C), 128.4 (Ar–C), 127.9 (Ar–C), 127.8 (Ar–C), 86.7 (C3′a or C6′a), 80.3 (C3′a or C6′a), 78.9 (C2 or C3), 72.9 (CH2Phe), 72.5 (C1′ or C4′), 71.2 (C2 or C3), 70.9 (C1′ or C4′), 70.3 (C6′), 59.3 (C5), 57.1 (C3′), 52.2 (C7), 46.8 (C10), 28.8 (C8), 24.6 (C9), 20.7 (CH3), 20.3 (CH3).
HRMS-FAB: m/z [M + 1] Calcd for C27H34N2O11: 562.2163; found: 563.2421.
(3S,4S)-2,5-Dioxotetrahydrofuran-3,4-diyl diacetate (6a)
Yield: 82 %; hygroscopic crystals; mp 130 °C (lit. 133–134 °C; Shriner and Furow, 1963)
1H NMR (300 MHz, CD3OD): δ = 5.77 (1H, s, CH), 5.69 (1H, s, CH), 2.23 (3H, s, CH3), 2.20 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 171.2 (COCH3), 167.0 (C=O), 82.2 (CH), 17.7 (COCH3).
(3R,4R)-2,5-Dioxotetrahydrofuran-3,4-diyl diacetate (6b)
Yield: 82 %; hygroscopic crystals; mp 130 °C (lit. 133–134 °C; Shriner and Furow, 1963)
1H NMR (300 MHz, CD3OD): δ = 5.77 (1H, s, CH), 5.77 (1H, s, CH), 2.21 (3H, s, CH3), 1.19 (3H, s, CH3).
13C NMR (75 MHz, CDCl3): δ = 171.0 (COCH3), 167.0 (C=O), 82.2 (CH), 17.0 (COCH3).
(2S)-Methyl-2-((2R,3R)-4-((3S,6R)-6-(benzyloxy)hexahydrofuro[3,2-b]furan-3-ylamino)-2,3-dihydroxy-4-oxobutanamido)-3-phenylpropanoate (7b2)
Yield: 92 %; yellow solid (after purification by chromatographic column; eluent: hexane/ethyl acetate); mp 41–42 °C.
[α] 20D +124.3(c 0.6, MeOH).
IR (KBr, ν cm−1): 3393, 30663, 3030, 2950, 2879, 1736, 1670, 1648, 1541, 1497, 1454, 1368, 1215, 1139, 1081, 911, 737, 700.
1H NMR (300 MHz, CD3OD): δ = 7.37–7.18 (10H, m, H10, H11, H12, Ar–H), 4.76–4.75 (1H, m, H2 or H3), 4.71 (1H, d, J = 11.4 Hz, NH), 4.70 (1H, t, J = 4.7 Hz, H2 or H3), 4.53 (1H, d, J = 11.4 Hz, H4′), 4.44 (d, 2H, J = 4.4 Hz, CH2Phe), 4.41 (1H, d, J = 1.8 Hz, H5), 4.39 (1H, d, J = 1.8 Hz, H3′a), 4.34–4.31 (1H, m, H3′), 4.13 (1H, dd, J = 11.6, 6.6 Hz, H1′), 4.04 (1H, dd, J = 9.6, 4.8 Hz, H1′), 3.87–3.85 (1H, m, H6′ or H6′a), 3.84–3.82 (1H, m, H6′ or H6′a), 3.70 (3H, s, H7), 3.64 (1H, t, J = 5.0 Hz, H4′), 3.18 (1H, dd, J = 13.8 e 5.6 Hz, H8), 3.08 (1H, dd, J = 13.8 e 5.6 Hz, H8).
13C NMR (75 MHz, CD3OD): δ = 174.3 (C1), 174.0 (C4), 173.1 (C6), 139.4 (Ar–C), 137.6 (C9), 130.4 (Ar–C), 129.6 (C10), 129.4 (Ar–C), 129.1 (C11), 128.9 (C12), 128.0 (Ar–C), 88.5 (C3′a or C6′a), 81.5 (C3′a or C6′a), 80.7 (C2 or C3), 74.1 (C2 or C3), 74.0 (CH2Phe), 73.5 (C6′), 71.8 (C1′ or C4′), 71.7 (C1′ or C4′), 58.2 (C3′), 54.8 (C5), 52.8 (C7), 38.6 (C8).
HRMS-FAB: m/z [M + 1] Calcd for C27H32N2O9: 528.2108; found: 529.2430.
(2S)-Methyl-2-((2R,3R)-4-((3S,6R)-6-(benzyloxy)hexahydrofuro[3,2b]furan-3-ylamino)-2,3-dihydroxy-4-oxobutanamido)-3-methyl butanoate (7b3)
Yield: 74 %; yellow solid (after purification by chromatographic column; eluent: hexane/ethyl acetate); mp 44–45 °C.
[α] 20D +96.0 (c 0.9, MeOH).
IR (KBr, ν cm−1): 3387, 2961, 2877, 1751, 1655, 1638, 1541, 1438, 1371, 1272, 1210, 1141, 1023, 735, 699.
1H NMR (300 MHz, CD3OD): δ = 7.38–7.26 (5H, m, Ar–H), 4.70–4.69 (2H, m, CH2Phe), 4.54 (1H, d, J = 11.5 Hz, H2 or H3), 4.49 (1H, d, J = 1.7 Hz, H2 or H3), 4.47 (1H, d, J = 4.4 Hz, H5), 4.43 (2H, d, J = 1.8 Hz, H4′), 4.41 (1H, d, J = 5.6 Hz, H3′a), 4.14 (1H, dd, J = 11.6, 6.6 Hz, H3′), 4.05 (2H, dd, J = 9.6, 4.8 Hz, H1′), 3.89–3.86 (1H, m, H6′ or H6′a), 3.85–3.82 (1H, m, H6′ or H6′a), 3.73 (3H, s, H7), 2.21–2.14 (1H, m, C8), 0.96 (3H, d, J = 4.0 Hz, H9 or H10), 0.94 (3H, d, J = 4.0 Hz, H9 or H10).
13C NMR (75 MHz, CD3OD): δ = 174.3 (C1), 174.0 (C4), 173.2 (C6), 139.4 (Ar–H), 129.4 (Ar–H), 129.1 (Ar–H), 128.9 (Ar–H), 88.5 (C3′a or C6′a), 81.8 (C3′a or C6′a), 80.7 (C2 or C3), 74.1 (C2 or C3), 74.0 (CH2PHe), 73.9 (C1′ or C4′), 73.5 (C6′), 71.8 (C1′ or C4′), 58.7 (C3′), 58.2 (C5), 52.7 (C7), 32.4 (C8), 19.3 (CH3), 18.3 (C9 and C10).
HRMS-FAB: m/z [M + 1] Calcd for C23H32N2O9: 480.2108; found: 481.2320.
(2R)-Methyl-1-((2R,3R)-4-((3S,6R)-6-(benzyloxy)hexahydrofuro[3,2b]furan-3-ylamino)-2,3-dihydroxy-4-oxobutanoyl)pyrrolidine-2-carboxylate (7b4)
Yield: 90 %; white solid (after purification by chromatographic column; eluent: hexane/ethyl acetate); mp 47–48 °C.
[α] 20D +79.6 (c 0.6, MeOH).
IR (KBr, ν cm−1): 3356, 2880, 1714, 1681, 1651, 1614, 1538, 1455, 1312, 1227, 1198, 1138, 1025, 911, 883, 742, 699.
1H NMR (300 MHz, CD3OD): δ = 7.36–7.28 (5H, m, Ar–H), 4.74 (1H, d, J = 11.7 Hz, H6′a), 4.71–4.69 (2H, m, H2, H3), 4.54 (1H, d, J = 11.7 Hz, H3′a), 4.51–4.43 (1H, m, H5), 4.42 (d, 1H, J = 6.0 Hz, H3′), 4.39–4.28 (2H, m, CH2Phe), 4.14 (1H, dd, J = 13.5, 7.0 Hz, H6′), 4.07–4.03 (1H, m, H4′), 3.91–3.89 (1H, m, H4′), 3.86–3.82 (2H, m, H1′), 3.71 (3H, s, H7), 3.66–3.63 (2H, m, H10), 2.27–2.17 (2H, m, H8), 2.07–1.91 (2H, m, H9).
13C NMR (75 MHz, CD3OD): δ = 174.3 (C1), 172.4 (C4), 172.2 (C6), 139.4 (Ar–C), 129.4 (Ar–C), 129.1 (Ar–C), 128.9 (Ar–C), 88.5 (C3′a or C6′a), 81.8 (C3′a or C6′a), 80.7 (C2 or C3), 74.1 (C2 or C3), 73.9 (CH2PHe), 73.5 (C1′ or C4′), 73.3 (C6′), 71.7 (C1′ or C4′), 60.9 (C5), 58.3 (C3′), 52.8 (C7), 46.6 (C10), 30.4 (C8), 25.8 (C9).
HRMS-FAB: m/z [M + 1] Calcd for C23H30N2O9: 478.1951; found: 479.2291.
Construction of NS3/4A pro expression plasmids
Con1/SG-Neo (I) was used as template in the generation of constructs fusing the NS3 protease and cofactor NS4A to form NS3/4A pro. Con1/SG-Neo (I) contains the cDNA of the HCV 1b subgenomic replicon, which codes for the S1179I mutant in the NS5a sequence. The fragment corresponding to amino acid residues 1–182 of NS3 pro was amplified using forward primer NS3 pro-GSGS-F (5′-ggtagtggtagtATggCgCCTATTACggCCTAC-3′, which incorporates the nucleotide sequence encoding the GSGS linker at the 5´ terminus) and reverse primer NS3 pro-HindIII-R (5′-TATTAAgCTTTTAggACCgCATAgTggTTTC-3′, which incorporates a Hind III restriction site and a stop codon at the 3´terminus). To amplify the NS4A cofactor (residues 21–32), we used primer NS4A-BamHI-F (5′-AATAggATCCggCAgCgTggTCATTgTg-3′, which incorporates a BamHI restriction site at the 5′ terminus) and primer NS4A-GSGS-R (5′-actaccactaccggACAAgATgATCCTgCC-3′, which incorporates the nucleotide sequence encoding the GSGS linker at the 3´ terminus). PCR reactions were performed in 1X PCR buffer (Invitrogen), 1.5 mM MgSO4 (Invitrogen), 250 µM of each dNTP, 50 pmol of each primer, and 1.25 U of Pfx DNA polymerase (Invitrogen) with 100–200 ng of genomic DNA in a final volume of 50 µL. The thermocycle program to amplify the construct contained an initial denaturing step of 94 °C for 4 min, followed by 35 cycles of 94 °C (1 min), 60 °C (1 min), and 68 °C (2 min). To construct the NS3 protease domain fused to cofactor NS4A (NS3/4Apro), the PCR products were then joined by overlap PCR using the outer primers NS4A-BamHI-F and NS3 pro-HindIII-R. The resultant PCR product contained the NS4A cofactor domain attached via a flexible GSGS linker to NS3 pro. The PCR product was purified with a GFX PCR DNA and gel band purification kit (GE-Healthcare, USA). The resultant fragment was digested with BamHI and HindIII and subsequently inserted into the pET21dHT vector modified for expression of a 6×His tag and tev protease-site (6×HT) at the N-terminus (homemade construction, IBCCF/UFRJ). The construct was validated by DNA sequencing using a ABI3730 sequencer (Applied Biosystems) that employs the PDTIS/FIOCRUZ platform.
Expression and purification of NS3/4A pro
The recombinant plasmid pET21dHT-HCVNS3/4A pro was inserted into E. coli strain BL21 [λDE3] (Novagen) by heat shock. Transformed cells were grown at 37 °C in 2 L of LB medium containing 100 μg/mL ampicillin (USB) until an OD600 of 0.8 was reached. The culture was then cooled down to 25 °C, and 0.5 mM isopropyl-β-d-thiogalactopyranoside (IPTG) was added to induce 6×His-NS3/4A pro (with 6×His-TEVsite) protein expression. After 3 h, cells were harvested by centrifugation and stored at −80 °C until used.
To purify the protein, cell pellets were thawed and resuspended in 25 mL buffer A [50 mM Tris–HCl (pH 7.5), 300 mM NaCl, 2 mM β-Mercaptoethanol, 0.5 % Triton X-100, and 10 % glycerol] per L of original culture. Resuspensions were lysed with the addition of 5 mg/mL lysozyme (Sigma), 20 µg/mL DNase, and 2 mM MgCl2. To prevent the proteolytic cleavage of protein during cell lysis, all procedures were performed on ice. The cell lysate was centrifuged at 30,000×g and 4 °C for 40 min. The supernatant was applied to a His-Trap column (GE-Healthcare) which was washed with buffer A. The 6xHT-NS3/4Apro protein bound to the column was eluted with an imidazole gradient from 0 to 500 mM. Fractions containing the purified protein were dialyzed for 16 h at 4 °C in buffer B [50 mM Tris–HCl (pH 7.5), 300 mM NaCl, 2 mM β-Mercaptoethanol, 0.5 % Triton X-100, and 30 % glycerol]. The 6×His-TEV site of the protein was excised with TEV protease [40]. The untagged NS3/4A pro was again applied to the His-TRAP column, which retained both 6×His-TEV and the TEV protease. Only NS3/4A pro was eluted. NS3/4A pro was dialyzed against buffer B and concentrated in a Stirred Ultrafiltration Cell with a 3000-MWCO membrane (Millipore-Amicon). The NS3/4A pro concentration was measured using a Coomasie Kit (Bradford) Protein Assay Kit (Pierce) following the manufacturer´s protocol.
In vitro inhibition measurements
To evaluate the inhibitory activity of peptide mimetic compounds against the HCV NS3/4A protease, the SensoLyte®520 HCV Protease Assay Kit *Fluorimetric* (AnaSpec, CA, USA) was used following the manufacturer’s protocol. The compound screening assay was performed in a black 96-well plate. Each well contained 30 ng NS3/4A pro that was pre-incubated for 10 min at 25 °C with 100 µM (or varying concentrations) of the tested compound. Subsequently, the 5-FAM/QXL™ 520-FRET peptide substrate was added and incubated for 60 min at 25 °C. Fluorescent signal substrate cleavage was monitored at the excitation and emission wavelengths of 490 and 520 nm, respectively, using a SpectraMax M5 fluorimeter (Molecular Devices). Compounds were diluted into assay buffer from 20 mM or 40 mM stock solutions prepared in 100 % DMSO. These compounds were diluted to a final concentration of 10 % DMSO in the assay buffer. The initial reaction velocities (Vi) of product formation were determined from progress curves using the linear regression method (less than 10 % peptide cleavage). SigmaPlot v.10.0 and GraphPad Prism v.5 software were used to calculate IC50 or Ki values assuming Michaelis–Menten kinetics.
Molecular docking studies
Docking studies were performed using AutoDock 4.2 for Windows. AutoDock Tools (ADT) was used to set up both crystal and ligand structure parameters. The tridimensional structure of 5a3 was built and minimized to the PM6 level in the molecular modeling program Spartan’10 (Wavefunction Inc.) and then exported to ADT. The coordinates of the wild-type NS3/4A crystal structure were obtained from the Protein Data Bank (PDB code 2OC0) (Prongay et al., 2007). The ligand (i.e., HU1) and solvent molecules were removed. Hydrogen atoms were added, and non-hydrogen atoms were merged with the corresponding carbon atoms.
First, the grid center was established by centering the grid box on the allosteric site with 0.375 Å spacing and identical 60 × 60 × 60 points. Docking studies were carried out using the empirical free energy function and the Lamarckian genetic algorithm applying a standard protocol. An initial population of 150 randomly placed individuals and a maximum number of 2.5 × 106 energy evaluations were employed. A total of 50 independent docking runs were carried out. Structures differing by less than 2.0 Å in positional RMSD were clustered. The results of the most favorable free energies of binding were selected as the resultant complex structure.
In silico pharmacokinetic and toxicity analyses
Herein, we used Osiris Property Explorer (http://www.organic-chemistry.org/prog/peo/) (Thomas Sander, Actelion Pharmaceuticals Ltd., Gewerbestrasse 16, 4123 Allschwil, Switzerland) to predict the physicochemical properties (i.e., cLogP and solubility), toxicity, druglikeness, and drug score of the most active compound. Values of druglikeness are based on the occurrence frequencies of each fragment of the molecule in commercial drugs, while the drug score evaluates the compound’s potential to qualify as a drug. The drug score is also related to topological descriptors, fingerprints of molecular druglikeness, structural keys, and other properties such as cLog P, log S, and molecular weight.
References
Barros JC, da Silva JFM, Calazans A, Tanuri A, Brindeiro RM, Williamson JS, Antunes OAC (2006) Synthesis of pseudopeptides derived from (R, R)-tartaric acid as potential inhibitors of HIV-protease. Lett Org Chem 3:882–886
Barros TG, Pinheiro S, Williamson JS, Tanuri A, Pereira HS, Brindeiro RM, Alonso Neto JB, Antunes OAC, Muri EMF (2009) Novel peptide mimetic inhibitors of hepatitis C serine protease derived from isomannide. Synthesis 4:620–626
Barros TG, Pinheiro S, Williamson JS, Tanuri A, Pereira HS, Brindeiro RM, Alonso Neto JB, Antunes OAC, Muri EMF (2010) Pseudo-peptides derived from isomannide: inhibitors of serine proteases. Amino Acids 38:701–709
Barros TG, Zorzanelli BC, Pinheiro S, Brito MA, Tanuri A, Costa ECB, Mohana-Borges RS, Rodrigues CR, Souza AMT, Ferreira VF, Muri EMF (2012) Novel peptide mimetics based on N-protected amino acids derived from isomannide as potential inhibitors of NS3 serine protease of hepatitis C virus. Lett Org Chem 9:239–249
Bazan JF, Fletterick RJ (1989) Detection of a trypsin-like serine protease domain in flaviviruses and pestviruses. Virology 171:637
Chee CF, Abdullah I, Buckle MJC, Rahman NA (2010) An efficient synthesis of (±)-panduratin A and (±)-isopanduratin A, inhibitors of dengue-2 viral activity. Tetrahedron Lett 51:495–498
Cohen J (1999) The scientific challenge of hepatitis C. Science 285:26–30
Feld JJ, Hoofnagle JH (2005) Mechanism of action of interferon and ribavirin in treatment of hepatitis C. Nature 436:967–972
Frokjaer S, Otzen DE (2005) Protein drug stability: a formulation challenge. Nat Rev Drug Discov 4:298–306
Ghosh AKZL, Dawson ZL, Mitsuya H (2007) Darunavir, a conceptually new HIV-1 protease inhibitor for the treatment of drug-resistant HIV. Bioorg Med Chem 15:7576–7580
Gorbalenya AE, Donchenko AP, Kunin EV, Blinov VM (1989) N-terminal domains of putative helicases of flavi- and pestiviruses may be serine proteases. Nucleic Acids Res 17:3889–3897
Halfon P, Locarnin S (2011) Hepatitis C virus resistance to protease inhibitors. J Hepatol 55:192–206
Ingallinella P, Altamura S, Bianchi E, Taliani M, Ingenito R, Cortese R, De Francesco R, Steinkühler C, Pessi A (1998) Potent peptide inhibitors of human hepatitis C virus NS3 protease are obtained by optimizing the cleavage products. Biochemistry 37:8906–8914
Kumar M, Sharma S, Srinivasan A, Singh T, Kaur B (2011) Structure-based in silico rational design of a selective peptide inhibitor for thymidine monophosphate kinase of mycobacterium tuberculosis. J Mol Model 17:1173–1182
Lampa A, Ehrenberg AE, Gustafsson SS, Vema A, Åkerblom E, Lindeberg G, Karlén A, Danielson UH, Sandström A (2010) Improved P2 phenylglycine-based hepatitis C virus NS3 protease inhibitors with alkenylic prime-side substituents. Bioorg Med Chem 18:5413–5424
Lange C, Sarrazin C (2002) In hepatitis C: new drugs in hepatology. A clinical textbook, 3rd edn. Flying Publisher, Germany
Lim SR, Qin X, Susser S, Nicholas JB, Lange C, Herrmann E, Hong J, Arfsten A, Hooi L, Bradford W, Nájera I, Smith P, Zeuzem S, Kossen K, Sarrazin C (2012) Virologic escape during danoprevir (ITMN-191/RG7227) monotherapy is hepatitis C virus subtype dependent and associated with R155K substitution. Antimicrob Agents Chemother 56:271–279
Lin C, Lin K, Luong YP, Rao BG, Wei YY, Brennan DL, Fulghum JR, Hsiao HM, Ma S, Maxwell JP, Cottrell KM, Perni RB, Gates CA, Kwong ADJ (2004) In vitro resistance studies of hepatitis C virus serine protease inhibitors, VX-950 and BILN 2061: structural analysis indicates different resistance mechanisms. J Biol Chem 279:17508–17514
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341
Maryanoff BE, Costanzo MJ (2008) Inhibitors of proteases and amide hydrolases that employ an alpha-ketoheterocycle as a key enabling functionality. Bioorg Med Chem 16:1562–1595
Melino S, Paci M (2007) Progress for dengue virus diseases. Towards the NS2B-NS3pro inhibition for a therapeutic-based approach. FEBS J 274:2986
Muri EMF, Gomes M Jr, Alencar FL, Sales A Jr, Bastos ML, Hernandez-Valdes R, Albuquerque MG, da Cunha EFF, de Alencastro RB, Williamson JS, Antunes OAC (2004) N-t-Boc-amino acid esters of isomannide—Potential inhibitors of serine proteases. Amino Acids 27:153–159
Muri EMF, Gomes MJr, Albuquerque MG, de Alencastro RB, Williamson JS, Antunes OAC, Williamson JS, Antunes OAC (2005) Pseudo-peptides derived from isomannide as potential inhibitors of serine proteases. Amino Acids 28:413–419
Örtqvist P, Gising J, Ehrenberg AE, Vema A, Borg A, Karlén A, Larhed M, Danielson UH, Sandström A (2010) Discovery of achiral inhibitors of the hepatitis C virus NS3 protease based on 2(1H)-pyrazinones. Bioorg Med Chem 18:6512–6525
Peçanha EP, Figueiredo LJO, Brindeiro RM, Tanuri A, Calazans AR, Antunes OAC (2003) Synthesis and anti-HIV activity of new C2 symmetric derivatives designed as HIV-1 protease inhibitors. Il Farmaco 58:149–157
Prongay AJ, Guo Z, Yao N, Pichardo J, Fischmann T, Strickland C, Myers JJr, Weber PC, Beyer BM, Ingram R, Hong Z, Prosise WW, Ramanathan L, Taremi SS, Yarosh-Tomaine T, Zhang R, Senior M, Yang RS, Malcolm B, Arasappan A, Bennett F, Bogen SL, Chen K, Jao E, Liu YT, Lovey RG, Saksena AK, Venkatraman S, Girijavallabhan V, Njoroge FG, Madison V (2007) Discovery of the HCV NS3/4A protease inhibitor (1R,5S)-N-[3-amino-1-(cyclobutylmethyl)-2,3-dioxopropyl]-3-[2(S)-[[[(1,1-dimethylethyl)amino]carbonyl]amino]-3,3-dimethyl-1-oxobutyl]- 6,6-dimethyl-3-azabicyclo[3.1.0]hexan-2(S)-carboxamide (Sch 503034) II. Key steps in structure-based optimization. J Med Chem 50:2310–2318
Resende GO, Aguiar LCS, Cotrim BA, da Silva JFC, Antunes OAC (2007) Synthesis of asymmetric peptide mimetic compounds containing tartaric acid core. potential inhibitors of HIV-1 protease. Lett Org Chem 4:168–171
Sander T, Freyss J, von Korff M, Reich JR, Rufener C (2009) OSIRIS, an entirely in-house developed drug discovery informatics system. J Chem inf Model. 49(2):232–246
Sarrazin C, Zeuzem S (2010) Resistance to direct antiviral agents in patients with hepatitis C virus infection. Gastroenterol 138:447–462
Shiryaev SA, Cheltsov AV, Strongin AY (2012) Probing of exosites leads to novel inhibitor Scaffolds of HCV NS3/4A proteinase. PLoS One 7:7–10
Shriner RL, Furow CL (1963) Diacetyl–d-tartaric-acid. Org Synth Coll 4:242
Sidique S, Shiryaev SA, Ratnikov BI, Herath A, Su Y, Strongin AY, Cosford NDP (2009) Structure-activity relationship and improved hydrolytic stability of pyrazole derivatives that are allosteric inhibitors of West Nile Virus NS2B-NS3 proteinase. Bioorg Med Chem Lett 19:5773
Tong X, Chase R, Skelton A, Che T, Wright-Minogue J, Malcolm BA (2006) Activity of a potent hepatitis C virus polymerase inhibitor in the chimpanzee model. Antivir Res 70:28–38
Vermehren J, Sarrazin C (2011) New HCV therapies on the horizon. Clin Microbiol Infect 17:122–134
Westermann B, Diedrichs N, Krelaus R, Walter A, Gedrath I (2004) Diastereoselective synthesis of homologous bicyclic lactams-potential building blocks for peptide mimics. Tetrahedron Lett 45:5983–5986
Wright M, Main J, Thomas HC (2001) Treatment of chronic viral Hepatitis. Antiv Chem Chemother 12:201–212
Yusof R, Clum S, Wetzel M, Murthy HM, Padmanabhan R (2000) Purified NS2B/NS3 serine protease of dengue virus type 2 exhibits cofactor NS2B dependence for cleavage of substrates with dibasic amino acids in vitro. J Biol Chem 275:9963–9969
Zhu Z, Wilson AT, Luxon BA, Brown KE, Mathahs MM, Bandyopadhyay S, McCaffrey AP, Schmidt WN (2010) Biliverdin inhibits hepatitis C virus nonstructural 3/4A protease activity: mechanism for the antiviral effects of heme oxygenase? Hepatology 52:1897–1905
Acknowledgments
This work was supported by Coordination for the Improvement of Higher Education Personnel (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—CAPES), National Council for Scientific and Technological Development (Conselho Nacional de Desenvolvimento Científico e Tecnológico—CNPq) and the Carlos Chagas Filho Foundation for Support of Research of the State of Rio de Janeiro (Fundação Carlos Chagas Filho de Amparo a Pesquisa do Estado do Rio de Janeiro—FAPERJ).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abrahim-Vieira, B., da Costa, E.C.B., de A. Azevedo, P.H.R. et al. Novel isomannide-based peptide mimetics containing a tartaric acid backbone as serine protease inhibitors. Med Chem Res 23, 5305–5320 (2014). https://doi.org/10.1007/s00044-014-1058-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-014-1058-1