Abstract
Thriving populations of invasive species often decline, but the mechanisms behind such declines are rarely elucidated. The microbial community living within insects can have significant impacts on host health and population dynamics and may be one such mechanism. Here, we use Illumina RNAseq and 16S amplicon sequencing to compare the bacterial communities, and investigate the presence of viruses, in queens of the invasive yellow crazy ant, Anoplolepis gracilipes, from populations in various stages of decline or expansion in Australia. Black queen cell virus and sequences with homology to the Dicistroviridae family of viruses were detected in queens from declining ant populations. This is the first instance of sequences with homology to a virus being found in A. gracilipes and may indicate a new virus. The overall bacterial communities were not statistically different between the population types and a number of potential mutualists were discovered. Several putative pathogens were also detected, including Candidatus Rhabdochlamydia, the entomopathogen Serratia marcescens and the reproductive parasite Candidatus Cardinium. These taxa are known to significantly affect host biology in other organisms. Our survey has identified several candidates that may be responsible for population declines of this invasive ant, and therefore may have potential as biological control agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Social insects have been shown to suffer from a wide range of pathogens (Schmid-Hempel 1998; Rose et al. 1999; Evans and Schwarz 2011). The honey bee, Apis mellifera, is particularly well studied and may carry many pathogens, including, but not limited to, bacteria, such as Paenibacillus larvae (American foulbrood), microsporidia, such as Nosema, and a number of viruses in the Dicistroviridae and Iflaviridae families (Evans and Schwarz 2011). All of these pathogens have been associated with significant declines of A. mellifera populations (Evans and Schwarz 2011). Pathogens can influence the community and population dynamics of other social insects, including the South American bumble bee, Bombis dahlbomii (Schmid-Hempel et al. 2013).
Pathogens, including viruses, are being increasingly discovered in ants. Three dicistroviruses have been found in Solenopsis invicta (Valles et al. 2009; Valles 2012). The first virus, SINV-1, was only recently discovered (Valles 2012) and since then several viruses have also been identified in the invasive ants Linepithema humile, Nylanderia pubens and the non-invasive Formica exsecta (Valles 2012; Johansson et al. 2013; Sébastien et al. 2015). Some of these viruses may be asymptomatic, at least under normal, non-stress conditions, but other viruses may be strongly pathogenic (de Miranda et al. 2010; Porter et al. 2013). It seems likely that many viruses infect ant species, but before the advent of next-generation sequencing (NGS) they were simply too difficult to identify (Valles et al. 2012). NGS techniques, such as Roche 454 and Illumina sequencing have revolutionized metagenomic studies and the discovery of microbes such as viruses (Caporaso et al. 2012). Sequencing of the universal 16S rRNA gene in bacteria has brought new appreciation for the complexity of the bacterial community within organisms (Bartram et al. 2011). This technology enables high-throughput, swift, relatively inexpensive sequencing for the discovery of microorganisms and understanding their population dynamics (Johansson et al. 2013). One such use is the identification of potential biocontrol agents for invasive pests (Liu et al. 2011; Valles et al. 2012).
The microbiota of an insect can have both positive and negative effects on health, reproduction and longevity (Cox-Foster et al. 2007; Russell et al. 2009; Vasquez et al. 2012). Mutualistic bacteria play a myriad of roles in host physiology, such as providing essential nutrients that are lacking in the host’s diet, mediating host thermal tolerance, or enhancing pathogen and parasitoid resistance (Russell et al. 2009; Feldhaar 2011). For example, beneficial taxa such as lactic acid bacteria have been implicated in facilitating positive honey bee health by inhibiting the bacterial brood pathogen, Paenibacillus larva (Vasquez et al. 2012).
The yellow crazy ant, Anoplolepis gracilipes, is one of the most widespread, ecologically and economically damaging invasive ant species (Holway et al. 2002). Populations of this ant were first detected in northeast Arnhem Land in the Northern Territory of Australia in 1982 (Majer 1984), though it likely had arrived several decades prior (Young et al. 2001). Several A. gracilipes populations have been observed to naturally decline in this region (Gruber et al. 2012; Cooling and Hoffmann 2015) and elsewhere (Haines et al. 1994). The proximate cause of the declines remains unclear, but it has been hypothesized that pathogens have played a role (Gruber 2012; Cooling 2016; Lester and Gruber 2016). Here, we compare the microbial fauna of A. gracilipes queens from populations in northeast Arnhem Land exhibiting either population decline or expansion. We focused on pathogens in queens because of the vital role queens play in reproduction and population dynamics of ant populations. Specifically, we (1) determine pathogen and mutualist presence in A. gracilipes and (2) test whether declining and expanding populations differ in their bacterial and viral communities.
Methods
Population selection
Population selection was determined by two factors: each population’s history of decline or expansion and A. gracilipes worker abundance in each population at the time of sampling (B. Hoffmann, personal observation; Cooling and Hoffmann 2015). Importantly, populations that displayed relatively static population abundances over time were not used in this study. Abundance was determined using a combination of (1) qualitative visual presence/absence surveys that were predominantly used to determine the spatial extent of each population and (2) pitfall traps which provide a quantified measure of A. gracilipes abundance.
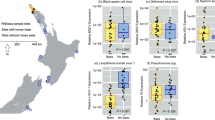
We selected six populations, four where A. gracilipes was present in various stages of decline without human intervention (Fig. 1) and two that were expanding. The two low-abundance populations (L1, L2) were the remainder of much larger populations that had declined substantially in the previous 5 years (B. Hoffmann, unpublished data). The two medium-abundance populations (M1, M2) covered a much wider area and had higher abundances than the low-abundance populations. These populations were stable, but previously belonged to larger populations in which some decline had been observed (B. Hoffmann, unpublished data). The two high-abundance populations (H1, H2) were expanding, covered a large area and had abundances several orders of magnitude higher than the low- and medium-abundance populations. Although the M and H population pairs were very close together, they were considered to be independent because the maximum foraging range of an A. gracilipes worker has been estimated to be 100 m (B. Hoffmann, unpublished data), and these pairs were separated by 400 and 450 m, respectively, so there is little possibility of workers mixing between populations. Moreover, in Arnhem Land, A. gracilipes queens have not been observed to partake in nuptial flights. Instead, they appear to mate inside their natal nest where they remain or walk a few metres to form new nests (Hoffmann 2015). Therefore, gene flow between geographically separated infestations is also unlikely.
Population surveys
We conducted population surveys to confirm the abundance categories (H, M & L) upon which we based our population selection. Surveys were undertaken in late July 2013 and 2014 following the methods of Cooling and Hoffmann (2015). See Online Resource 1 for details on the surveying process.
We quantified A. gracilipes abundance in all six populations using pitfall traps. Traps were placed in four plots in each population, spaced a minimum of 10 m apart and placed where A. gracilipes was seen to be present. In each plot, five pitfall traps (45 mm diameter) were arranged in a cross 1 m apart. Traps were 2/3 filled with a 1:1 mixture of ethylene glycol and water and left for 48 h. Anoplolepis gracilipes abundance in the traps was pooled for each plot and log transformed to meet the assumptions of normality. A general linear model (GLM) was used to compare A. gracilipes abundance among populations and years using SPSS v 22.0.0.0.
Queen collection
Queens were collected from each population in February 2013 and DNA and RNA were extracted for Illumina sequencing using Qiagen DNeasy Blood and Tissue, and RNeasy kit following the manufacturer’s protocol. One sample was extracted from each population, two per population type (L, M, H). Each sample consisted of five queens from different nests in the population that were pooled together prior to RNA and DNA extraction. These queens had been dissected as part of a previous experiment (Cooling 2016) and were preserved in RNAlater Stabilization Solution (AMBION, Inc., Austin, TX, USA) at −80 °C.
Viral survey
RNA quality was determined by the presence of an 18S rRNA peak using an Agilent 2100 Bioanalyzer. Barcoded TruSeq stranded total RNA libraries were sequenced by New Zealand Genomics Limited (NZGL) over four lanes of Illumina HiSeq 2500 [2 X100 bp paired-end (PE)]. The Illumina HiSeq sequencing returned just over 65 million reads (65 080 176), with an average of 10,846,696 ± 1,897,369 reads per sample (mean ± SD). Reads were of high quality as 94.6% had a Phred score of Q > 30. The RNA sequences were adapter and quality trimmed at a probability of 0.01 (Phred score of Q20) using the program FastQC 0.11.3 (Andrews 2015). To reduce the chance of erroneous sequences overinflating diversity estimates, all reads smaller than 50 bps were discarded, as were singletons (reads without a pair) with the program SolexaQA 3.0 (Cox et al. 2010). Transcriptome de novo assembly of normalized reads was performed with Trinity v 2.0.6 (Grabherr et al. 2013). The assembly resulted in 837,259 contigs with a mean length of 960 ± 24 bp (mean ± SD). These contigs were compared against the National Center for Biotechnology Information (NCBI) nt database downloaded from GenBank in November 2015. We used the BLASTn algorithm with a minimum e value of 0.0001. Searches were run on the Victoria University Science Faculty High Performance Computer with a BLAST v 2.2.30 installation (Zhang et al. 2000). Taxonomic assignment of the BLAST output was performed in MEGAN v. 4.7 (Huson et al. 2011) using the default settings. This filtering process resulted in taxonomy assignment to 425,022 contigs. Assigned viral sequences with a bit score greater than 100 were then extracted and searched with BLASTx against the nr database with the conservative search parameters of a minimum e value of 0.0001, Best Hit algorithm overhang value of 0.25 and Best Hit algorithm score edge value of 0.05 (Camacho et al. 2008). Taxonomy was assigned in MEGAN using the same conservative threshold as above.
Phylogenetic analyses of the three putative viral sequences assigned by MEGAN (TR44839, TR80102, TR17983) were conducted in MEGA v. 6.06 (Tamura et al. 2013). The contigs were aligned with their closest Genbank matches with ClustalW (Larkin et al. 2007). Maximum-likelihood trees with 1000 bootstrap replicates were generated using the Tamura 3-parameter model (the best-fitting model) for TR44839 (T92; lnL −1126.878) and TR17983 (T92; lnL −1398.077), and the Tamura 3-parameter model with a gamma distribution [T92 + G(0.95); lnL −1409.281] for TR80102 (Tamura 2002).
Virus confirmation assay
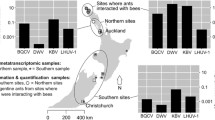
One-step RT-PCR assays were used to confirm the presence of viral sequences in queens and workers in two populations (M2, L2) that had matches to viral sequences. Additional queens collected in 2015 and workers collected in February 2013 were placed in RNAlater and frozen at −80 °C. The digestive systems of queens from 2015 were removed prior to extraction to rule out environmental contamination (Valles 2012). RNA was extracted using a GENEzol–chloroform–isopropanol protocol (see Online Resource 1 for the detailed protocol). Worker samples from each population contained approximately 40 individuals pooled from at least three nests. Queen samples each consisted of five individuals. We also tested for the following viruses commonly found in other ant and Hymenoptera species (Chantawannakul et al. 2006; Martin et al. 2012; Sébastien et al. 2015) using one-step RT-PCRs: Deformed wing virus (DWV), Acute bee paralysis virus (ABPV), Black queen cell virus (BQCV), Israeli acute paralysis virus (IAPV), Kashmir bee virus (KBV) and Linepithema humile virus-1 (LHUV-1). The contig TR44839 was also assessed using primers designed in the program Amplifix 1.7.0. (see Online Resource 1 for the viral primers used). Primer design was not possible for contigs TR80102 and TR17983.
Bacterial survey
DNA quantity and quality were checked using a Nanodrop and 1% agarose gel (Robertson 2003). PCR amplification of the V3–V4 hypervariable region of the 16S rRNA gene was achieved using Nextera adapter primers provided by NZGL (see Online Resource 1 for the primers used and the PCR process).
Primers and adapters were trimmed and paired-end reads joined and error corrected using PANDAseq, v 2.8 (Masella et al. 2012). Of the total reads, 94.5% were assembled. The sequence length after primer removal and joining was 418 ± 4 bp (mean ± SD), with an average of 43,668 ± 41,723 reads (mean ± SD) per sample. The remaining 262,009 sequences were further analysed using the program Quantitative Insights Into Microbial Ecology (QIIME, v 1.9.0; Caporaso et al. 2012). De novo chimeric sequences were removed (919 sequences, 0.35% of total sequences) and reads were clustered into operational taxonomic units (OTUs) at 97% sequence similarity with the de novo method with QIIME’s implementation of USEARCH, v 6.1.544 (Edgar 2010). A representative sequence was then picked for each OTU using the Greengenes 13.8 rep set, and taxonomy was assigned using Greengenes 13.8 taxonomy (McDonald et al. 2012). Sequences were aligned against the Greengenes database using PYNAST, v 1.2.2, and filtered using the Greengenes lanemask. OTUs representing less than 0.005% of the total number of sequences were discarded (Bokulich et al. 2013). Before comparative analyses, sequence libraries from individual populations were rarified in QIIME to 2767 reads: the size of the smallest library (population M1) (Fig. 2).
Variation in bacterial genus diversity among samples of different population types was explored using an Analysis of similarities (ANOSIM, with 9999 permutations) with a resemblance matrix derived using the Bray–Curtis similarity index (Clarke 1993). Data were square-root transformed prior to analysis to decrease the influence of very abundant species. The contribution of each bacterial genus to differences among population types was assessed using a similarity percentage analysis (SIMPER). ANOSIM and SIMPER were run in Primer-E v 6.1.13 (Clarke and Gorley 2006). QIIME log-likelihood ratio tests, using the Bonferroni procedure to correct for multiple comparisons, identified which bacterial genera differed between population types (Rice 1989).
Results
Population surveys
Anoplolepis gracilipes abundance varied significantly among the three population types (GLM, F = 87.851, p < 0.001) (Fig. 3; see Online Resource 2 for the surveyed population maps), with 96% of individuals in pitfall traps being from the two high-abundance populations (Fig. 3). Abundance within pairs of population types did not significantly differ from each other (see Online Resource 2). Population L1 did not differ significantly in A. gracilipes abundance from populations M1 or M2 (Tukey test, p > 0.05). Nevertheless, as populations M1 and M2 both had approximately 1300% more presence points than population L1 (i.e. a larger population; see Online Resource 2 for population survey results), we consider our classifications valid. The abundance of A. gracilipes within population types did not differ significantly between years (F = 0.052, p = 0.426).
Mean number of A. gracilipes workers ± SD per plot (n = 4/population) in each of the two low-abundance, medium-abundance and high-abundance populations. Anoplolepis gracilipes abundances of each population are shown for two sampling years (2013 and 2014). The x-axis has been split to accommodate the measure of A. gracilipes workers collected from high-abundance populations, whose abundances were two orders of magnitude higher than in other populations
Viral survey
Our MEGAN analysis assigned 95% of RNA reads as eukaryote, mostly matching ants (Fig. 4a). Only 0.004% of the 425,022 total reads had viral sequence similarity. Three candidate viral sequences were found from two populations, one from the medium-abundance population M2 (TR44839) and two from the low-abundance population L2 (TR17983, TR80102). These sequences matched to RNA polymerase genes, making them suitable for phylogenetic analysis and taxonomic assignment. The MEGA phylogenetic analysis grouped all three putative viral sequences with the Dicistroviridae family of viruses (Fig. 4b–d).
a Summary of BLASTn results showing taxonomic assignment for all samples combined based on a total of 425 022 reads. Maximum-likelihood phylogenetic trees (1000 bootstrap replicates) showing the relationship of sequence, b TR44839 from the medium-abundance population M2 (220 bp), c TR80102 from the low-abundance population L2 (240 bp) and d TR17983 (282 bp) from the low-abundance population L2. The scale shows changes per nucleotide
The contig TR44839 was 220 bp with ten hits and matched most closely with helicase of Israeli acute paralysis virus (see Online Resource 2 for the closest matches). TR17983 was 240 bp and TR80102 282 bp, with nine hits total and matched most closely with RNA polymerase of BQCV and KBV (see Online Resource 2 for more search results). NGS methods may not always find high numbers of sequences, even for taxa that are prevalent in ant populations (e.g. Sébastien et al. 2015), particularly if the virus is in low titre. Therefore, we decided to investigate these possible viral sequences further despite their low abundance.
RT-PCR confirmed the presence of sequence TR44839 (using custom-designed primers; see Online Resource 1) in queens from 2013 and 2015 and workers from 2013 in populations M2 and L2. However, although BQCV was found in pooled samples from L1 and M2, using primers designed by Chantawannakul et al. (2006), the viruses DWV, ABPV, IAPV, KBV and LHUV-1 were not (using published primers; see Online Resource 1).
Bacterial survey
The bacterial communities from five A. gracilipes queens from each of six populations were characterized using Illumina amplicon sequencing (261,039 chimera-checked, quality controlled sequences; n = 7,707–103,350 per library). At the genus level, 170 unique OTUs (97% sequence similarity) were found in the queens (45–119 OTUs per sample; Fig. 5). Of these, 58 OTUs were assigned to a known genus, and 14 of these genera made up more than 1% of the total sequences in at least one sample. The composition of bacterial genera did not differ between the three population types (ANOSIM, R = −0.056, p = 0.600).
SIMPER analysis showed that 90% of dissimilarity between high- and low-abundance populations was attributed to 109 bacterial genera, and 50% of the dissimilarity was attributed to 23 genera (Online Resource 2). Similar numbers were found for medium- and low-abundance populations, with 90% of the dissimilarity accounted for by 91 bacterial genera, and 50% of the dissimilarity by 27 bacterial genera. Thirteen bacterial OTUs differed significantly among population types (Table 1). The pathogen Candidatus Rhabdochlamydia (Candidatus refers to the fact this taxon has not been successfully maintained in culture; hereafter Rhabdochlamydia) was only present in high-abundance populations. The entomopathogen Serratia marcescens and the reproductive parasite Candidatus Cardinium (hereafter Cardinium) were more commonly found in a medium-abundance population. The order Rhizobales, which contains putative nutritional mutualists of ants, was significantly less common in the low-abundance populations (Table 1). Arsenophonus and Wolbachia, which were previously found in A. gracilipes (Sébastien et al. 2012), were not detected, possibly because the previous study investigated the microbial community in A. gracilipes workers, and not queens.
Discussion
Pathogens clearly affect a wide range of social insects and can dramatically change their population dynamics (Briano 2005; Arif et al. 2011; Cameron et al. 2011). Alternatively, mutualists may provide protection against some of these pathogens, as well as providing other benefits (Cox-Foster et al. 2007; Feldhaar 2011; Vasquez et al. 2012). Given the diversity, abundance and influence of pathogens and mutualists on well-studied social insects such as honey bees, bumble bees and other ant species, we similarly expected to see a range of pathogenic and mutualistic microbial taxa in A. gracilipes. Our aims were to investigate (1) pathogen and mutualist presence in A. gracilipes from Arnhem Land and (2) examine for evidence that declining and expanding populations differ in their bacterial and viral communities. We found that A. gracilipes hosted a multitude of microbes, including a possible unidentified dicistrovirus and Black queen cell virus, several potential bacterial mutualists (Acinetobacter, Enterococcus, Fructobacillus, Gluconobacter, Lactobacillus), pathogens (Rhabdochlamydia, Serratia marcescens) and a potential reproductive parasite (Cardinium).
This is the first study to record the presence of viral sequences in A. gracilipes. Although we do not know if this potential virus, or viruses, plays, or play, a role in the observed population declines of A. gracilipes, viral sequences were found in one low- and one medium-abundance population, which is suggestive of pathogen-induced population decline. More research is needed to characterize the potential novel virus and establish the prevalence of putative infection.
The three viral sequences aligned to partial genes were involved in replication (helicase, RNA polymerase). Phylogenetic trees grouped these sequences with the Dicistroviridae viral family and most closely to KBV, IAPV and BQCV. With the exception of BQCV, PCR analysis with published primers for KBV, IAPV and other common honey bee pathogens failed to detect these viruses in A. gracilipes. This result suggests that these sequences represent a new dicistrovirus or perhaps novel strains of a known species. Dicistroviruses have been implicated in colony collapse disorder in A. mellifera (Cox-Foster et al. 2007; de Miranda et al. 2010), as well as colony decline in S. invicta (Valles 2012; Porter et al. 2013). Several dicistroviruses affecting S. invicta are being investigated as candidates for biocontrol in that species (Valles 2012). Viruses often have high host specificity and high pathogenicity, making them ideal biocontrol agents (Arif et al. 2011; Porter et al. 2013).
As we did not test for active replication, we do not know if these viruses are parasitizing A. gracilipes, or were simply ingested by the ants from the environment. The latter seems unlikely as queens from 2015 had the gut dissected prior to sequencing. It is also unclear what role the viruses play within the ants. It is possible that these viruses are benign, as many insect viruses are asymptomatic (Liu et al. 2011; de Miranda et al. 2010; Valles 2012). In rare cases, viruses may even be beneficial to insects, such as the densovirus Helicoverpa armigera densovirus-1, which appears to enhance host resistance to a baculovirus, and the bacterium Bacillus thuringiensis (Xu et al. 2014).
Metagenomic surveys can identify bacteria that make up the microbiota of the host. But such studies can only speculate on what effect these bacteria have on host fitness. However, we can identify bacterial candidates for further study based on their function in other similar hosts, such as ants and hymenoptera. Though several potentially pathogenic and mutualistic bacteria were found, no obvious patterns in their occurrence suggested that these bacteria played a role in the observed population declines of A. gracilipes. However, the absence of a pattern found here may, in part, be due to low sample size. A potential bacterial pathogen, Rhabdochlamydia, was found in one of the high-abundance populations. Other members of the order Chlamydiales are often pathogenic (Corsaro et al. 2007). Rhabdochlamydia crassificans is known to be pathogenic in cockroaches, where it can cause swelling of the abdomen and infect the ovaries and fat bodies (Corsaro et al. 2007). No tissue swelling was apparent in the infected A. gracilipes queens. To the best of our knowledge, this bacterial genus has never before been isolated in ants, so its effects in ants are unknown.
We detected the lactic acid bacteria (LAB) genera Lactobacillus, Enterococcus and Fructobacillus in all samples. Enterococcus and Fructobacillus were both significantly more prevalent in high-abundance populations, whereas Lactobacillus was more prevalent in low- and medium-abundance populations. These bacteria are mutualistic in A. mellifera, with Enterococcus and Lactobacillus protecting their hosts from microbial pathogens (Vasquez et al. 2012). Fructobacillus may produce by-products that appear to promote the growth of core members of the gut microbiome (Rokop et al. 2015). Thus, it appears that all population types had at least one type of LAB in high prevalence, making it unlikely that the absence of these mutualists is responsible for the differing abundances observed among A. gracilipes populations. However, an interactive effect between different LAB taxa is possible (Rokop et al. 2015), so it remains possible that they have an influence on ant population dynamics.
Sequences with identity to Serratia were found. This bacterial genus has been isolated from the guts of the ants Camponotus japonica and Formica cinerea, which possibly acquired the bacteria from feeding on honeydew excreted from aphids (He et al. 2011; Sirvio and Pamilo 2010). Anoplolepis gracilipes are also known to feed on honeydew (Holway et al. 2002; Lach and Hoffmann 2011) and could possibly have acquired this bacterial taxon that way. Serratia marcescens is also known to be pathogenic in a suite of insects, including A. mellifera (Lauzon et al. 2003 and references therein). We also detected Gluconobacter at high prevalence in all populations. Gluconobacter is another potential mutualist in Hymenoptera with high sugar diets (Crotti et al. 2010).
Cardinium is a reproductive parasite in arthropods (Hunter et al. 2003) and has also been found in at least one other species of ant, Formica cinera (Sirvio and Pamilo 2010). Cardinium seems to be uncommon in ants (Russell et al. 2012; Zchori-Fein and Perlman 2004), unlike other reproductive parasites such as Wolbachia, which have been found in one-third of ant species examined (Russell 2012) including A. gracilipes (Sébastien et al. 2012; Gruber 2012). Like Wolbachia, any fitness impact that Cardinium has on its ant hosts is yet to be demonstrated (Russell 2012; Sirvio and Pamilo 2010).
The detected microbe, Acinetobacter, has been reported in many insect groups, including ants and other Hymenoptera (Ishak et al. 2011a; Paulson et al. 2014; Singh et al. 2012), as well as A. gracilipes (Gruber 2012). The roles Acinetobacter plays in most of these groups are not known. One previous study found no evidence for its influence on fitness parameters in a female wasp (Srinatha et al. 2015).
We examined the microbiome of queens from a single point in time, but other potential pathogens or mutualists may be found by examining other castes besides queens, such as the workers or brood (Ishak et al. 2011b; Gruber 2012). Likewise, the presence of specific pathogens can vary throughout the year, likely due to differences in temperature and/or rainfall. For example, SINV-1 has been found to be much more prevalent in spring and summer than at other times of the year (Valles et al. 2010). It is suspected that this difference is due to the inability of SINV-1 to replicate at cooler temperatures. Arnhem Land has consistently high temperatures throughout the year, so low temperatures may not influence their microbial dynamics. However, Arnhem Land has a distinct wet season with most rain falling between December and July (Lach and Hoffmann 2011). Rainfall may also change pathogen prevalence. Briano (2005) found a significant correlation between infection of Solenopsis fire ants with the microsporidian Kneallhazia solenopsae and rainfall.
Our study has identified several bacterial candidates for further study as mutualists or pathogens as well as a potentially novel virus. Viruses and bacteria have been successfully utilized for biocontrol programmes. For example, a Gammabaculovirus was introduced from Europe to North America where it effectively controls the invasive spruce sawfly, Diprion hercyniae (Arif et al. 2011; Szewczyk et al. 2006). Likewise, the bacterium Bacillus thuringiensis is the most commonly utilized microbe worldwide for the control of agricultural pests (Cannon 1993). The viability of the virus, SINV-3, as a biocontrol agent of the invasive ant, S. invicta, is currently being investigated (Valles 2012; Valles and Porter 2015); it has been successfully released into S. invicta colonies in Florida and California (Valles and Oi 2014). Though it is too early to say if the potential viruses found in A. gracilipes play a role in the observed population declines, such viruses may be promising biocontrol candidates.
References
Andrews S (2015) FastQC v 0.11.3 Babraham Bioinformatics. http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc/
Arif B, Escasa S, Pavlik L (2011) Biology and genomics of viruses within the genus Gammabaculovirus. Viruses 3:2214–2222
Bartram A, Lynch M, Stearns J, Moreno-Hagelsieb G, Neufeld J (2011) Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired-end Illumina reads. Appl Environ Microb 77(11):3846–3852
Bokulich N, Subramanian S, Faith J, Gevers D, Gordon J, Knight R, Mills D, Caporaso G (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10(1):57–59
Briano J (2005) Long-term studies of the red imported fire ant, Solenopsis invicta, infected with the microsporidia Vairimorpha invictae and Thelohania solenopsae in Argentina. Environ Entomol 34(1):124–132
Camacho C, Madden T, Ma N, Tao T, Agarwala R, Morgulis A (2008) BLAST® command line applications user manual. National Center for Biotechnology Information, Bethesda
Cameron S, Lozier J, Strange J, Koch J, Cordes N, Solter L, Griswold T (2011) Patterns of widespread decline in North American bumble bees. PNAS 108(2):662–667
Cannon R (1993) Prospects and progress for Bacillus thuringiensis based pesticides. Pest Manag Sci 37:331–335
Caporaso J, Lauber C, Walters W, Berg-Lyons D, Huntley J, Fierer N, Owens S, Betley J, Fraser L, Bauer M, Gormley N, Gilbert J, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Chantawannakul P, Ward L, Boonham N, Brown M (2006) A scientific note on the detection of honeybee viruses using real-time PCR (TaqMan) in Varroa mites collected from a Thai honeybee (Apis mellifera) apiary. J Invertebr Pathol 91:69–73
Clarke K (1993) Non-parametric multivariate analyses of changes in community structure. Aust Ecol 18:117–143
Clarke K, Gorley R (2006) PRIMER v 6.1.11: User manual and tutorial. PRIMER-E, Plymouth
Cooling M (2016) Population dynamics and pathogens of the invasive yellow crazy ant (Anoplolepis gracilipes) in Arnhem Land, Australia. Dissertation, Victoria University of Wellington
Cooling M, Hoffmann B (2015) Here today, gone tomorrow: declines and local extinctions of invasive ant populations in the absence of intervention. Biol Invasions 17:3351–3357
Corsaro D, Thomas V, Goy G, Venditti D, Radek R, Greub G (2007) ‘Candidatus Rhabdochlamydia crassificans’, an intracellular bacterial pathogen of the cockroach Blatta orientalis (Insecta: Blattodea). Syst Appl Microbiol 30:221–228
Cox M, Peterson D, Biggs P (2010) SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinform 11:485
Cox-Foster D, Conlan S, Holmes E, Palacios G, Evans J, Moran N, Quan P, Briese T, Hornig M, Geiser D, Martinson V, vanEngelsdorp D, Kalkstein A, Drysdale A, Hui J, Zhai J, Cui L, Hutchison S, Simons J, Egholm M, Pettis J, Lipkin W (2007) A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318:283–287
Crotti E, Rizzi Chouaia B, Ricci I, Favia G, Alma A, Sacchi L, Bourtzis K, Mandrioli M, Cherif A, Bandi C, Daffonchio D (2010) Acetic acid bacteria, newly emerging symbionts of insects. Appl Environ Microb 76(21):6963–6970
de Miranda J, Cordoni G, Budge G (2010) The acute bee paralysis virus-Kashmir bee virus-Israeli acute paralysis virus complex. J Invertebr Pathol 103:S30–S47
Edgar R (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461
Evans J, Schwarz R (2011) Bees brought to their knees: microbes affecting honey bee health. Trends Microbiol 19(12):614–620
Feldhaar H (2011) Bacterial symbionts as mediators of ecologically important traits of insect hosts. Ecol Entomol 36:533–543
Grabherr M, Haas B, Yassour M, Levin J, Thompson D et al (2013) Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat Biotechnol 26(7):644–652
Gruber M (2012) Genetic factors associated with variation in abundance of the invasive yellow crazy ant (Anoplolepis gracilipes). Dissertation, Victoria University of Wellington
Gruber M, Burne A, Abbott K, Pierce R, Lester P (2012) Population decline but increased distribution of an invasive ant genotype on a Pacific atoll. Biol Invasions 15(3):599–612
Haines I, Haines J, Cherrett J (1994) The impact and control of the Crazy ant, Anoplolepis longipes (Jerd.), in the Seychelles. In: Williams D (ed) Exotic ants: biology, impact, and control of introduced species. Westview Press, Boulder, pp 206–218
He H, Chen Y, Zhang Y, Wei C (2011) Bacteria associated with gut lumen of Camponotus japonicus. Environ Entomol 40(6):1405–1409
Hoffmann B (2015) Integrating biology into invasive species management is a key principle for eradication success: the case of yellow crazy ant Anoplolepis gracilipes in northern Australia. Bull Entomol Res 105:141–151
Holway D, Lach L, Suarez A, Tsutsui N, Case T (2002) The causes and consequences of ant invasions. Annu Rev Ecol Evol Syst 33:181–233
Hunter M, Perlman S, Kelly S (2003) A bacterial symbiont in the Bacteroidetes induces cytoplasmic incompatibility in the parasitoid wasp Encarsia pergandiella. Proc R Soc B 270:2185–2190
Huson D, Mitra S, Ruscheweyh H, Weber N, Schuster S (2011) Integrative analysis of environmental sequences using MEGAN4. Genome Res 21(9):1552–1560
Ishak H, Plowes R, Sen R, Kellner K, Meyer E, Estrada D, Dowd S, Mueller U (2011a) Bacterial diversity in Solenopsis invicta and Solenopsis geminata ant colonies characterized by 16S amplicon 454 pyrosequencing. Microb Ecol 61:821–831
Ishak H, Miller J, Sen R, Dowd S, Meyer E, Mueller U (2011b) Microbiomes of ant castes implicate new microbial roles in the fungus-growing ant Trachymyrmex septentrionalis. Sci Rep 1:204
Johansson H, Dhaygude K, Lindstrom S, Helantera H, Sundstrom L, Trontti K (2013) A metatranscriptomic approach to the identification of microbiota associated with the ant Formica exsecta. PLoS One 8(11):e79777
Lach L, Hoffmann B (2011) Are invasive ants better plant-defense mutualists? A comparison of foliage patrolling and herbivory in sites with invasive yellow crazy ants and native weaver ants. Oikos 120:9–16
Larkin M, Blackshields G, Brown N, Chenna R, McGettigan PA et al (2007) ClustalW and ClustalX version 2. Bioinformatics 23:2947–2948
Lauzon C, Bussert T, Sjogren R, Prokopy R (2003) Serratia marcescens as a bacterial pathogen of Rhagoletis pomonella flies (Diptera: Tephritidae). Eur J Entomol 100:87–92
Lester P, Gruber M (2016) Booms, busts and population collapses in invasive ants. Biol Invasions. doi:10.1007/s10530-016-1214-2
Liu S, Vijayendran D, Bonning B (2011) Next generation sequencing technologies for insect virus discovery. Viruses 3:1849–1869
Majer J (1984) Recolonisation by ants in rehabilitated open-cut mines in Northern Australia. Reclam Reveg Res 2:279–298
Martin S, Highfield A, Brettell L, Villalobos E, Budge G, Powell M, Nikaido S, Schroeder D (2012) Global honey bee viral landscape altered by a parasitic mite. Science 336:1304–1306
Masella A, Bartram A, Truszkowski J, Brown D, Neufeld J (2012) PANDAseq: paired-end assembler for Illumina sequences. BMC Bioinform 13:31
McDonald D, Price M, Goodrich J, Nawrocki E, DeSantis T, Probst A, Andersen G, Knight R, Hugenholtz P (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6(3):610–618
Paulson A, von Aderkas P, Perlman S (2014) Bacterial associates of seed-parasitic wasps (Hymenoptera: Megastigmus). BMC Microbiol 14:224
Porter S, Valles S, Oi D (2013) Host specificity and colony impacts of the fire ant pathogen, Solenopsis invicta virus 3. J Invertebr Pathol 114:1–6
Rice W (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Robertson C (2003) Spectrophotometry’s next generation technology. American Biotechnology Laboratory
Rokop Z, Horton M, Newton I (2015) Interactions between co-occurring lactic acid bacteria in the honey bee hive. Appl Environ Entomol. doi:10.1128/AEM.01259-15AEM.01259-15
Rose E, Harris R, Glare T (1999) Possible pathogens of social wasps (Hymenoptera: Vespidae) and their potential as biological control agents. N Z J Zool 26:179–190
Russell J (2012) The ants (Hymenoptera: Formicidae) are unique and enigmatic hosts of prevalent Wolbachia (Alphaproteobacteria) symbionts. Myrmecol News 16:7–23
Russell J, Moreau C, Goldman-Huertas B, Fujiwara M, Lohman D, Pierce N (2009) Bacterial gut symbionts are tightly linked with the evolution of herbivory in ants. PNAS 106(50):21236–21241
Russell J, Funaro C, Giraldo Y, Goldman-Huertas B, Suh D, Kronauer D, Moreau C, Pierce N (2012) A veritable menagerie of heritable bacteria from ants, butterflies and beyond: broad molecular surveys and a systematic review. PLoS One 7(12):e51027
Schmid-Hempel P (1998) Parasites in social insects. Princeton University Press, Princeton
Schmid-Hempel P, Eckhardt M, Goulson D, Heinzmann D, Lange C, Plischuk S, Escudero L, Salathe R, Scriven J, Schmid-Hempel P (2013) The invasion of southern South America by imported bumblebees and associated parasites. J Anim Ecol 83:823–837
Sébastien A, Gruber M, Lester P (2012) Prevalence and genetic diversity of three bacterial endosymbionts (Wolbachia, Arsenophonus, and Rhizobiales) associated with the invasive yellow crazy ant (Anoplolepis gracilipes). Insect Soc 59:33–40
Sébastien A, Lester P, Hall R, Wang J, Moore N, Gruber M (2015) Invasive ants carry novel viruses in their new range and form reservoirs for a honeybee pathogen. Biol Lett 11:20150610
Singh S, Priya N, Kumar J, Rana V, Ellango R, Joshi A, Priyadarshini G, Asokan R, Rajagopal R (2012) Diversity and phylogenetic analysis of endosymbiotic bacteria from field caught Bemisia tabaci from different locations of North India based on 16S rDNA library screening. Infect Genet Evol 12:411–419
Sirvio A, Pamilo P (2010) Multiple endosymbionts in populations of the ant Formica cinerea. BMC Evol Biol 10:335
Srinatha H, Jalali S, Sriram S, Chakravarthy A (2015) Isolation of microbes associated with field-collected populations of the egg parasitoid, Trichogramma chilonis capable of enhancing biotic fitness. Biocontrol Sci Technol 25(7):789–802
Szewczyk B, Hoyos-Carvajal L, Paluszek M, Skrzecz I, Lobo de Souza M (2006) Baculoviruses—re-emerging biopesticides. Biotechnol Adv 24:143–160
Tamura K (2002) Evolutionary distance estimation under heterogenous substitution pattern among lineages. Mol Biol Evol 19:1727–1736
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Valles S (2012) Positive-strand RNA viruses infecting the red imported fire ant, Solenopsis invicta. Psyche 2012:1–14
Valles S, Oi D (2014) Successful transmission of Solenopsis invicta virus 3 to field colonies of Solenopsis invicta (Hymenoptera: Formicidae). Fla Entomol 97:1244–1246
Valles S, Porter S (2015) Dose response of red imported fire ant colonies to Solenopsis invicta virus 3. Arch Virol 160:2407–2413
Valles S, Varone L, Ramirez L, Briano J (2009) Multiplex detection of Solenopsis invicta virus-1, -2, and -3. J Virol Methods 162:276–279
Valles S, Oi D, Porter S (2010) Seasonal variation and the co-occurrence of four pathogens and a group of parasites among monogyne and polygyne fire ant colonies. Biol Control 54:342–348
Valles S, Oi D, Yu F, Tan X, Buss E (2012) Metatranscriptomics and pyrosequencing facilitate discovery of potential viral natural enemies of the invasive Caribbean crazy ant Nylanderia pubens. PLoS One 7(2):e31828
Vasquez A, Forsgren E, Fries I, Paxton R, Flaberg E, Szekely L, Olofsson T (2012) Symbionts as major modulators of insect health: lactic acid bacteria and honeybees. PLoS One 7(3):e33188
Xu P, Liu Y, Graham R, Wilson K, Wu K (2014) Densovirus is a mutualistic symbiont of a global crop pest (Helicoverpa armigera) and protects against a baculovirus and Bt biopesticide. PLoS Pathog 10(10):e1004490
Young G, Bellis G, Brown G (2001) The crazy ant, Anoplolepis gracilipes (Smith) (Hymenoptera: Formicidae) in east Arnhem Land, Australia. Aust Entomol 28(3):97–104
Zchori-Fein E, Perlman S (2004) Distribution of the bacterial symbiont Cardinium in arthropods. Mol Ecol 13:2009–2016
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A greedy algorithm for aligning DNA sequences. J Comput Biol 7(1–2):203–214
Acknowledgements
We thank the many people that provided technical assistance, especially Daryl Lacey, Paul Augustin, Leanne Dzendolet, Tony Schultz and the many people involved with Conservation Volunteers Australia. Our thanks are due to Dhimurru Aboriginal Corporation and the traditional landowners of northeast Arnhem Land for access to the region and the staff of Rio Tinto for access to the mining leases and accommodation. This work was funded and supported by the Victoria University of Wellington Research Trust, CSIRO Australia, Rio Tinto Alcan, and the Dhimurru Aboriginal Corporation. M. Cooling was supported by a Victoria University of Wellington Doctoral Scholarship.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cooling, M., Gruber, M.A.M., Hoffmann, B.D. et al. A metatranscriptomic survey of the invasive yellow crazy ant, Anoplolepis gracilipes, identifies several potential viral and bacterial pathogens and mutualists. Insect. Soc. 64, 197–207 (2017). https://doi.org/10.1007/s00040-016-0531-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00040-016-0531-x