Abstract
Mountain plant species with wide elevational ranges are expected to be exposed to different selection pressures at low and high elevation. Estimating the extent of genetic population differentiation contributes to our understanding of selective forces shaping phenotypic variation of plants in response to changing climate conditions. Using a common garden experiment, we measured narrow-sense heritability (h 2) and quantitative trait differentiation (Q ST) in several growth-related, reproductive and phenological traits among low and high elevation populations of the semi-dry grassland species Ranunculus bulbosus from the Swiss Alps. Q ST values were compared to neutral genetic differentiation (F ST) based on AFLP markers and indicated divergent selection in most traits among all populations as well as among low and high elevation populations separately. Furthermore, pairwise Q ST and F ST estimates were not correlated suggesting that neutral marker differentiation is not a valuable proxy for quantitative trait differentiation. Neutral molecular differentiation increased with geographical distance whereas quantitative genetic differentiation did not follow an isolation by distance pattern. Plant traits did mostly not differ among low and high populations and pairwise Q ST–F ST comparisons indicated no divergent selection between the two elevations. Differing levels of trait differentiation in low and high elevation populations, however, indicated that different selective forces acted on plants at contrasting elevations suggesting that selection regimes might alter under climate warming.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In mountain habitats strong environmental variation over relatively short geographical distances (Körner 2003) is associated with highly divergent selective pressures on plant species and the emergence of locally adapted traits (Linhart and Grant 1996; Kawecki and Ebert 2004). Both abiotic factors, such as temperature, UV radiation, precipitation, and substrate, as well as biotic factors, such as competition, vary across elevations and potentially act as selective agents leading to plant differentiation (Choler et al. 2001; Till-Bottraud and Gaudeul 2002; Körner 2003; Blume et al. 2010). Besides the important role of natural selection in shaping phenotypic variation, neutral genetic processes, such as genetic drift, gene flow and random mutation, are also likely to influence population differentiation (Wright 1951; Slatkin 1977; Lande 1992). On the one hand, gene flow through pollen or seed dispersal is facilitated by the close proximity of populations along elevational gradients and has the potential to counteract the effects of divergent selection in populations, and thus to prevent local adaptation (reviewed in Slatkin 1987; Lenormand 2002; Raesaenen and Hendry 2008). Genetic drift, on the other hand, can induce random genetic variation through random changes in allele frequencies, especially in small populations (Spitze 1993; Lynch and Milligan 1994).
The relative importance of selection and non-selective neutral processes for among-population differentiation in quantitative traits can be evaluated by comparing quantitative genetic differentiation (Q ST; Spitze 1993) with neutral marker differentiation (F ST; Wright 1951). Higher relative divergence in the quantitative traits than in neutral markers (i.e. Q ST > F ST) indicates divergent selection favouring different genotypes in different populations whereas the opposite (i.e. Q ST < F ST) indicates stabilizing selection restricting divergence caused by neutral processes because similar plant trait values are selected for. If the two measures do not significantly differ (i.e. Q ST = F ST), neutral processes (drift, migration and mutation) alone may be sufficient to explain patterns of differentiation, although the relative effects of drift and natural selection on population differentiation cannot be separated (Merilä and Crnokrak 2001; McKay and Latta 2002; O’Hara and Merilä 2005; Whitlock 2008). Studies comparing Q ST and F ST allow eliminating neutral genetic processes as an explanation for divergence and have already provided valuable insights into plant trait responses to environmental heterogeneity (reviewed in McKay and Latta 2002; Leinonen et al. 2008; De Kort et al. 2013). For a more comprehensive understanding of the nature of selection, however, additional measures are needed, such as correlations of divergence with environmental differences (Whitlock 2008) or spatial patterns of differentiation, i.e. isolation by distance (IBD; Hutchison and Templeton 1999). Non-adaptive isolation by distance patterns may arise from decreasing homogenizing effects of gene flow with increasing geographical distance and lead to phenotypic divergence among geographically distant populations. The relative contributions of selection and neutral processes should then also depend on the spatial correlation between the selective environment and the geographical distance between populations.
Mountain ecosystems are considered vulnerable to the impacts of land use change (Fischer et al. 2008) and a rapidly changing climate (Begert et al. 2005; IPCC 2013). Owing to steep environmental gradients, they are particularly suited to investigate population differentiation and diversity (Till-Bottraud and Gaudeul 2002). Transplant experiments along elevational gradients have revealed local adaptation in fitness-related traits (Byars et al. 2007; Gonzalo-Turpin and Hazard 2009) and in leaf and reproductive phenological traits (Scheepens et al. 2010; Alberto et al. 2011). While several species have shown neutral genetic differentiation across elevational gradients (reviewed in Ohsawa and Ide 2008), only little differentiation was found between low and high elevation populations of three semi-dry nutrient-poor grassland species, probably due to high gene flow (Hahn et al. 2012; Matter et al. 2013). Yet, genetic differentiation was slightly greater among high than among low elevation populations (Hahn et al. 2012) as shown also for other species (Young et al. 2002; Herrera and Bazaga 2008; Byars et al. 2009) suggesting that gene flow underlies different processes at high and low elevation. Likewise, differentiation in quantitative traits and the magnitude of heritability might change along elevational gradients. Understanding the interplay of natural selection and neutral genetic processes as well as their variation at low and high elevations sheds light on the driving forces behind intraspecific differentiation and will gain in importance in the context of global change.
In this study, we assessed how natural selection and neutral processes influence genetic differentiation in low and high elevation populations of Ranunculus bulbosus. In a common garden experiment we grew seed families of R. bulbosus from 14 populations originating at 1,200 m and 1,800 m a.s.l. Vegetative and reproductive traits as well as flowering phenological traits were measured. Subsequently, narrow-sense heritability (h 2) and quantitative trait differentiation (Q ST) among all populations, as well as among low and high elevation populations separately, were determined. Neutral genetic differentiation (F ST) was calculated based on amplified fragment length polymorphism (AFLP) marker data of a previous project, which analysed genetic variation within and among a larger number of populations of R. bulbosus (Hahn et al. 2012) whereas our molecular analysis was based on a subset identical with the seed sampling populations.
Comparisons of overall Q ST and F ST estimates allowed testing the null hypothesis of neutrality in each trait. Furthermore, by comparing pairwise Q ijST and F ijST estimates of all low with all high elevation populations, we tested whether there was divergent selection between low and high elevation populations. Mantel correlations of pairwise Q ijST and F ijST estimates were used to test whether selection and drift are acting in the same direction (e.g. Leinonen et al. 2008). Finally, Mantel correlations between geographical distance and neutral molecular marker as well as quantitative trait differentiation were used to investigate isolation by distance patterns (Hutchison and Templeton 1999).
Materials and methods
Study species and plant material
The perennial herb Ranunculus bulbosus L. is a characteristic plant of semi-dry nutrient-poor calcareous grasslands ranging from 400 to 2,000 m a.s.l (colline to subalpine) in the European Alps with a distribution center in the montane belt (Aeschimann et al. 2004). Ranunculus bulbosus is an early-flowering insect-pollinated geophytic herb (Coles 1973; Sarukhan and Harper 1973; Steinbach and Gottsberger 1994).
In summer 2008, seeds were sampled from seven regions across the Swiss Alps, each containing one low elevation population at 1,200 m a.s.l. and one high elevation population at 1,800 m a.s.l. resulting in a total of 14 populations (Fig. 1; Table S1). The horizontal distance between the two sampling locations of a region was 1–18 km (median: 4.5 km; Table 1). Population sizes and plant densities at low and high elevation were similar (Hahn et al. 2012). Seeds of nine maternal plants (seed families) of each population were collected at a distance of at least four metres among individuals. They were air-dried and stored in paper bags at 4 °C. In spring 2009, seeds were germinated in a greenhouse at the ETH Zurich field station Lindau/Eschikon. Plants were raised in randomly arranged individual pots (800 cm3), filled with a 3:2 mixture of nutrient-poor commercial soil and sand.
Leaf material for the molecular analyses was collected from the same populations but from different mother plants than the seeds for the common garden experiment (Table S1). Circa 30 plants of each population were sampled with distances among individuals of at least four metres. Sampled leaf material was immediately dried with silica gel.
Quantitative genetic data
In March 2010, a common garden experiment was set up to determine quantitative genetic differentiation (Q ST) among populations. In this experiment, plants were cultivated under controlled environmental conditions using a half-sib design (Evanno et al. 2006), and thus the variation of phenotypes reflected their genetic differences (Briggs and Walters 1997; Petit et al. 2001). Three to four half-sibs of each of the nine R. bulbosus seed families per population (440 individuals in total) were arranged randomly in the greenhouse and rotated weekly.
At the start of the experiment, initial plant size was assessed as number of leaves, and length and width of the longest leaf. The product of them was used as covariate. From April to June 2010, flowering phenology of each plant was monitored weekly to record the date of the first open flower, the last open flower and the first mature fruit. Flowering duration was calculated as number of days between the first and last open flower of a plant. In July 2010, flowers were counted and length of the longest flower stalk was measured. In October 2010, length of the longest leaf was measured and specific leaf area (SLA) determined. The fresh leaf blades were scanned (HP All-InOne colour Scanner, Hewlett-Packard GmbH, Dübendorf, Switzerland) and their area was determined using LAMINA version 1.0.2 (Bylesjoe et al. 2008). The scanned leaf blades were weighed after oven drying at 60 °C for 48 h. SLA was calculated as the ratio of leaf area to dry weight of the leaf blade (Cornelissen et al. 2003). Moreover, leaf and bulb biomass were harvested, oven-dried and weighed.
Molecular genetic data
Population differentiation based on neutral genetic processes was determined using putatively neutral amplified fragment length polymorphism (AFLP) markers (Vos et al. 1995). For further details of genotyping, see Hahn et al. (2012).
Only polymorphic loci and loci with an allele frequency smaller than 1 – (3/N), where N is the total number of sampled individuals, were included in the analysis (Lynch and Milligan 1994). The data set for all populations comprised 125 loci, the separate sets for low elevation populations 121 loci and high elevation populations 122 loci. Because AFLP markers might also be part of or linked to coding gene regions, they might be under selection (Stinchcombe and Hoekstra 2008). Such markers need to be excluded because Q ST – F ST comparisons assume that F ST reflects neutral processes. A Bayesian outlier analysis in BayeScan 2.1 (Foll and Gaggiotti 2008), with 5,000 iterations and a thinning interval of 10, revealed that no loci were under divergent selection. The same result was previously also found for a slightly larger dataset (Hahn et al. 2012). All loci were therefore kept for subsequent analyses.
Statistical analyses
All statistical analyses were performed in R 2.15.0 (R Development Core Team 2012) with a full data set including all 14 populations as well as two data subsets including low and high elevation populations separately. To estimate between and within-population variances of quantitative traits, we ran linear mixed-effects models using the function lmer and the restricted maximum likelihood estimation (REML) in the R package lme4 (Bates et al. 2012). To meet the assumptions of normality of residuals and homoscedasticity, variables were transformed appropriately. The models included ‘population’ and ‘seed family’ nested within ‘population’ as random factors. Because data showed strong correlations with initial plant size, we used this variable as fixed covariate (see also Chun et al. 2011). Moreover, based on the full data set, we assessed genetic differentiation between low and high elevation plants using the same model with the addition of ‘altitude of origin’ as fixed factor.
Estimation of heritability, Q ST and F ST
Narrow-sense heritabilities (h 2) were calculated according to Petit et al. (2001) as
where V A is the additive genetic variance, which is the average genetic variance within populations. It was estimated by V A ≈ 4 V FAM where V FAM is the variance among seed families (Petit et al. 2001).
Quantitative genetic differentiation (Q ST) for each trait was calculated as
where V POP is the among-population variance. Accounting for the half-sib design, V GEN was estimated by V GEN = V A ≈ 4 V FAM. The 95 % confidence intervals of h 2 and Q ST estimates were calculated using a jack-knife procedure over seed families.
As a measure of neutral molecular differentiation, F ST was calculated (Wright 1951). Single locus F ST values were determined using BayeScan 2.1 (Foll and Gaggiotti 2008) and a bootstrap approach over loci with 5,000 iterations was applied to estimate the 95 % CI of the mean F ST across loci.
Overall Q ST – F ST comparisons
To test whether quantitative trait differentiation was significantly different from neutral genetic differentiation, an approach similar to the one proposed by Whitlock and Guillaume (2009) was applied. Analogously to an equivalence test, the difference of each resampled Q ST (based on the distribution of the jack-knife procedure over seed families) and resampled single marker F ST was calculated. After repeating this 126 times (equivalent to the total number of seed families), the sampling distribution of Q ST – F ST was generated. Q ST is significantly different from its expected value under neutral divergence among populations (F ST) if the 95 % confidence interval of the Q ST – F ST estimate does not contain zero.
Pairwise comparisons and Mantel tests
Population-pairwise Q ijST values were calculated using Eq. 2, but with the among-population variance V POP calculated as the variance of the estimates of the population means of the two populations, i.e.
where c is the contrast between the two populations in a mixed-effects model containing ‘population’ as a fixed effect (Palo et al. 2003; personal communication B. O’Hara). Population-pairwise F ijST values were calculated with Arlequin 3.5 (Excoffier et al. 2005).
First, population-pairwise Q ijST values were compared to pairwise F ijST in all low vs. high elevation population pairs as well as in all low vs. low and high vs. high pairs. Second, correlations between geographical distance, genetic (F ST) and phenotypic (Q ST) components were analysed using Mantel tests (Mantel 1967; Antoniazza et al. 2010). All Mantel tests were computed based on Legendre and Legendre (2012) with the R package vegan (Oksanen et al. 2013) and 1,000 permutations to assess the significance of the Mantel correlation coefficients (r). For Mantel tests with geographical distances, F ijST /(1 − F ijST ) respectively Q ijST /(1 − Q ijST ) were used while geographical distances were log-transformed (Rousset 1997).
Results
Plant growth and phenology
Based on the results of linear mixed-effects models, none of the investigated quantitative traits showed genetic differentiation between altitudes of origin (P > 0.05; Table 2), except for bulb biomass, which was greater in low than high elevation plants (P = 0.01). The variances due to the random effects population and seed family accounted for 10–72 % and 4–20 %, respectively, in the different traits.
Heritability, Q ST and F ST
The observed narrow-sense heritabilities (h 2) ranged from 0.291 to 0.675 in all populations (Table 3). Heritability of low and high elevation plants was similar in leaf length, bulb biomass, flowering start and flowering duration, but differed in the other six traits: SLA, number of flowers and fruiting start had larger h 2 whereas number of leaves, leaf biomass and flower stalk length had smaller h 2 in low as compared to high elevation populations.
The observed Q ST values ranged from 0.103 for leaf length to 0.474 for flowering start in all populations (Fig. 2; Table S2). All traits showed significant among-population differentiation (Q ST > 0) except for SLA in low elevation plants (Q ST = 0.003; 95 % confidence interval (CI): [0.000, 0.007]). Q ST values were larger in five (leaf biomass, bulb biomass, flower stalk length, flowering start and fruiting start) and smaller in three traits (SLA, number of flowers and flowering duration) in low as compared to high elevation populations. Q ST values of number of leaves and leaf length did not differ between altitudes of origin.
From the AFLP data, we obtained an F ST value, averaged over loci, of 0.085 (CI: [0.080, 0.091]) in all populations. The F ST of low elevation populations (F ST = 0.093; CI: [0.090, 0.097]) was similar to the F ST of high elevation populations (F ST = 0.095; CI: [0.092, 0.100]).
Overall Q ST–F ST comparisons
Considering the full data set with all populations, overall Q ST exceeded F ST in all measured traits with estimates ranging from 0.019 to 0.389 (average: 0.189; Fig. 3; Table S3). Also in the two subsets, Q ST exceeded F ST in all traits except for both subsets in leaf length and bulb biomass as well as SLA of low elevation populations. Q ST–F ST estimates in low elevation populations ranged from −0.090 to 0.447 with an average of 0.184 and in high elevation populations from −0.078 to 0.392 with an average of 0.183. In most traits, except for number of leaves and leaf length, Q ST–F ST estimates differed between low and high elevation populations indicated by non-overlapping CI’s (Fig. 3; Table S3).
Pairwise Q ijST and F ijST comparisons and Mantel tests
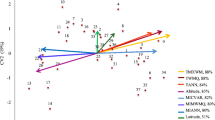
Pairwise F ijST estimates ranged from 0.030 to 0.205 (Table 1). Considering pairwise Q ijST and F ijST estimates of all low with all high elevation populations, as well as all low vs. low and high vs. high pairs, Q ijST did not differ from F ijST in any of the measured traits (Table S4). Pairwise Q ijST values were not correlated with pairwise F ijST values in any of the investigated traits (P > 0.23; Table 4A). Pairwise Q ijST were not correlated with geographical distance in any of the investigated traits (P > 0.21; Table 4B). In contrast, pairwise F ijST were positively correlated with geographical distance in all populations (Mantel’s r = 0.56, P < 0.01; Tables 2 and 4C), as well as in the subsets of low (Mantel’s r = 0.49, P < 0.05) and high elevation populations (Mantel’s r = 0.60, P < 0.05).
Discussion
The main objective of this study was to evaluate the relative importance of natural selection vs. genetic drift in determining population differentiation in low and high populations in a semi-dry grassland species. Random genetic drift is expected to affect all loci equally. Thus, we used AFLP markers and confirmed their selective neutrality in our data set, which was in line with the findings of Hahn et al. (2012). Since the overall Q ST–F ST comparisons may not adequately reflect variation in evolutionary forces acting at the inter-population level (Gutiérrez 2008), we combined this approach with population-pairwise Q ijST –F ijST comparisons.
Evidence for divergent selection
Considering low and high elevation populations together, all traits revealed significant heritability (Table 3) indicating evolutionary potential (Silvertown and Charlesworth 2001). Q ST values exceeded F ST values in all measured traits (Fig. 3), which indicates divergent selection and implies that different phenotypes are favoured in different populations (Merilä and Crnokrak 2001). This result is in line with a meta-analysis including 50 plant and animal species, which revealed that 70 % of the Q ST values exceeded their associated F ST values (Leinonen et al. 2008). The average Q ST–F ST difference among all populations in our study of 0.19 (Fig. 3; Table S3) is about 60 % higher than the average of 0.12 reported in the review of Leinonen et al. (2008) but substantially less than the difference of 0.58 in Rana temporaria (Palo et al. 2003) or 0.74 in Arabis fecunda (McKay et al. 2001). The authors of the latter study attributed the large estimate to geographic isolation and small effective population sizes of their rare endemic study species whereas R. bulbosus is a rather common species and distances among the sampled populations in the Swiss Alps are much shorter. These dissimilarities in abundance and sampling design might explain the smaller average Q ST–F ST differences in our study. While overall Q ST–F ST differences in half of the traits of R. bulbosus were larger than 0.21, they were very small in leaf length, bulb biomass and SLA. The latter two traits are expected to have smaller Q ST–F ST values because their generally relatively complex genetic background constrains strong evolutionary changes (De Kort et al. 2013) or it might indicate that these traits evolve neutrally. Highest divergent selection was found in reproductive phenology (flowering start and fruiting start). Phenological traits often vary across environmental gradients, such as advanced phenology at low as compared to high elevations, and hence are expected to be under divergent selection (Linhart and Grant 1996). Moreover, these traits are often associated with few large-effect genes (De Kort et al. 2013).
In line with the findings of Hahn et al. (2012), mean F ST values were generally low in all three data sets and showed a weak but significant IBD pattern (Table 4C). Moreover, a study on pollen flow in R. bulbosus along an elevation gradient in the Swiss Alps suggests high levels of contemporary gene flow, at least across small scales of <1 km (Matter et al. 2013). These findings suggest that gene flow constrained neutral genetic differentiation between more closely located populations. Under high levels of gene flow selection pressures or genetic drift must be very strong to leave a genetic imprint (Slatkin 1987) resulting in high Q ST values (De Kort et al. 2013). In our study, average Q ST of 0.27 was within the range of other insect-pollinated herbs such as Ranunculus reptans, Silene diclinis or Scabiosa columbaria (reviewed in Leinonen et al. 2008) but far below those of the annuals Arabidopsis thaliana (Q ST = 0.88; Kuittinen et al. 1997) and Arabis fecunda (Q ST = 0.94; McKay et al. 2001).
While neutral genetic markers followed an IBD pattern, quantitative genetic differentiation was not correlated with geographical distance in any of the investigated traits (Table 4B) indicating that geographical distance is not the driving selection agent in our study system. In contrast, quantitative genetic differentiation in amphibians increased with geographical distance among populations in Scandinavia (Palo et al. 2003; Hangartner et al. 2012). In these studies, geographical distance was also correlated with environmental clines (e.g. latitude or acidity) whereas in our paired sampling approach with evenly spread low and high elevation populations elevation was not correlated with geographical distance. Therefore, other factors which do not vary clinally with geographical distance seem to be responsible for the observed differentiation among populations, e.g. differences in land use (Stöcklin et al. 2009; Völler et al. 2013) or competition (Bischoff et al. 2006).
A positive correlation of Q ST and F ST would indicate that the degree of genetic differentiation in neutral markers reflects the degree of genetic differentiation in quantitative traits. Such correlations were found in a meta-analysis across different species (Leinonen et al. 2008). However, relatively few studies investigated the intraspecific correlation of pairwise Q ijST with F ijST . Some authors found positive correlations between pairwise Q ijST and F ijST (Morgan et al. 2001; Steinger et al. 2002) whereas others reported correlations that were only weak and inconsistent among traits (Palo et al. 2003; Hangartner et al. 2012). Likewise our results showed no Q ijST –F ijST correlation (Table 4A) indicating that selection, drift and gene flow did not influence molecular and quantitative differentiation in the same way. This limited predictability of quantitative traits from molecular markers might have important implications on conservation biological issues since management decisions are increasingly based purely on molecular analyses (e.g. Reed and Frankham 2001). Management plans should therefore also consider variation in quantitative traits, which might better reflect variation important for fitness (Lynch 1996).
Selection differences between low and high elevation populations
In addition to the divergent selection among all populations, we observed that the magnitude of selection differed between low and high elevation populations in eight out of ten traits (Fig. 3; Table S3). In number of flowers and flowering duration, divergent selection was stronger in high as compared to low elevation populations, which could be due to adaptation to the more heterogeneous local conditions at high elevations (Körner 2003). For SLA, high elevation populations were also under divergent selection whereas low elevation populations showed stabilizing selection (i.e. a negative Q ST–F ST estimate). Furthermore, the stronger differentiation in leaf biomass, flower stalk length, flowering start and fruiting start in low as compared to high elevation populations might be shaped by greater heterogeneity in competition or land use among low elevation habitats (Choler et al. 2001; Rudmann-Maurer et al. 2008). Differing patterns of divergent selection in low and high elevation populations might influence plant responses to climate change. As a consequence of shifting temperature regimes, current levels of quantitative differentiation at 1,200 m a.s.l. are expected to be observed at 1,800 m a.s.l. under predicted climate warming.
In contrast to the observed patterns of divergent selection within low and high elevation populations, comparisons of pairwise Q ijST –F ijST values in low vs. high elevation populations indicated no divergent selection between low and high elevation populations (Table S4). This is in line with the lack of genetic differentiation between altitudes of origin in the majority of mean trait values (Table 2). Therefore, elevation is likely not the main selective agent driving the observed quantitative differentiation.
An alternative approach to test for divergent selection as the main driver of phenotypic differentiation is the reciprocal transplantation of plants across a species range (e.g. Joshi et al. 2001). Individuals performing better under home-site conditions provide evidence that natural selection resulted in local adaptation, which has been considered a widespread phenomenon across climatic gradients (Santamaria et al. 2003; Becker et al. 2006; Macel et al. 2007) and also along elevational gradients (Byars et al. 2007; Gonzalo-Turpin and Hazard 2009). In contrast, but in line with the findings of this study, a transplant experiment with R. bulbosus between low and high elevation sites revealed no evidence for local adaptation although several traits showed quantitative genetic differentiation possibly due to the co-occurrence of environmental factors, other than elevation (Frei et al. 2014).
Accuracy of Q ST–F ST comparisons
Although Q ST–F ST comparisons provide a valuable tool to separate neutral processes from natural selection, they are not without criticism. To mention a few points among others, first, the indication of divergent selection is conservative, because several non-additive components of variance, such as maternal and dominance effects, may lower the Q ST estimate relative to F ST (Lopez-Fanjul et al. 2003; Goudet and Buchi 2006). While full-sib approaches contain twice the environmental component of variance common to family members as well as part of the dominance and epistatic components of variance, the half-sib design used here reduces this bias (Lynch and Walsh 1998; Merilä and Crnokrak 2001; Lopez-Fanjul et al. 2003). Furthermore, biased estimates of within-population components of variance through maternal effects might be of minor importance in our study because these effects generally diminish with plant age (Roach and Wulff 1987) and our plants were about one year old at the start of the experiment. Second, environmental effects, that mask underlying genetic divergence, might lead to under- or overestimation of quantitative trait differentiation (Conover et al. 2009). In our common garden experiment, we can exclude such direct environmental effects typical in studies on individuals collected from the wild (Pujol et al. 2008). Third, the precision of Q ST estimates increases with the number of populations investigated (O’Hara and Merilä 2005). Indeed, the confidence intervals of our Q ST estimates across all populations were often smaller than the ones calculated per elevation (Fig. 2; Table S2). Since Q ST estimates are rather high in most of the studied traits, we are confident that potential inaccuracies in the estimates of Q ST would not influence the results qualitatively.
Conclusions
By comparing quantitative trait to neutral molecular marker differentiation, we demonstrated that relatively strong divergent selection acted on several morphological and phenological traits among low and high elevation populations of the common semi-dry grassland species R. bulbosus from the Swiss Alps. Although pairwise Q ijST –F ijST comparisons indicated no divergent selection between the two elevations, varying degrees of trait differentiation within each elevation indicated differences in selective forces between low and high elevation habitats. Differing selection pressures at contrasting elevations, and thereby under different temperature regimes, imply that climate change might alter selection processes. If so, the considerable levels of heritability might facilitate respective plant responses and thereby foster population persistence in the longer term.
References
Aeschimann D, Lauber K, Moser DM, Theurillat JP (2004) Flora Alpina, 1st edn. Haupt, Berne
Alberto F, Bouffier L, Louvet JM, Lamy JB, Delzon S, Kremer A (2011) Adaptive responses for seed and leaf phenology in natural populations of sessile oak along an altitudinal gradient. J Evol Biol 24(7):1442–1454. doi:10.1111/j.1420-9101.2011.02277.x
Antoniazza S, Burri R, Fumagalli L, Goudet J, Roulin A (2010) Local adaptation maintains clinal variation in melanin-based coloration of european barn owls (Tyto alba). Evolution 64(7):1944–1954. doi:10.1111/j.1558-5646.2010.00969.x
Bates D, Maechler M, Bolker B (2012) lme4: linear mixed-effects models using S4 classes. R package. 0.999999-0
Becker U, Colling G, Dostal P, Jakobsson A, Matthies D (2006) Local adaptation in the monocarpic perennial Carlina vulgaris at different spatial scales across Europe. Oecologia 150(3):506–518. doi:10.1007/s00442-006-0534-9
Begert M, Schlegel T, Kirchhofer W (2005) Homogeneous temperature and precipitation series of Switzerland from 1864 to 2000. Int J Climatol 25(1):65–80. doi:10.1002/joc.1118
Bischoff A, Cremieux L, Smilauerova M, Lawson CS, Mortimer SR, Dolezal J, Lanta V, Edwards AR, Brook AJ, Macel M, Leps J, Steinger T, Muller-Scharer H (2006) Detecting local adaptation in widespread grassland species—the importance of scale and local plant community. J Ecol 94(6):1130–1142. doi:10.1111/j.1365-2745.2006.01174.x
Blume H-P, Brümmer GW, Horn R, Kandeler E, Kögel-Knabner I, Kretzschmar R, Star K, Wilke B-M (2010) Scheffer/Schachtschabel: Lehrbuch der Bodenkunde. Springer, Berlin
Briggs D, Walters S (1997) Plant variation and evolution, 3rd edn. Cambridge University Press, Cambridge
Byars SG, Papst W, Hoffmann AA (2007) Local adaptation and cogradient selection in the alpine plant, Poa hiemata, along a narrow altitudinal gradient. Evolution 61(12):2925–2941. doi:10.1111/j.1558-5646.2007.00248.x
Byars SG, Parsons Y, Hoffmann AA (2009) Effect of altitude on the genetic structure of an Alpine grass, Poa hiemata. Ann Bot 103(6):885–899. doi:10.1093/aob/mcp018
Bylesjoe M, Segura V, Soolanayakanahally RY, Rae AM, Trygg J, Gustafsson P, Jansson S, Street NR (2008) LAMINA: a tool for rapid quantification of leaf size and shape parameters. BMC Plant Biol 8:82–91. doi:10.1186/1471-2229-8-82
Choler P, Michalet R, Callaway RM (2001) Facilitation and competition on gradients in alpine plant communities. Ecology 82(12):3295–3308. doi:10.1890/0012-9658(2001)082[3295:facogi]2.0.co;2
Chun YJ, Le Corre V, Bretagnolle F (2011) Adaptive divergence for a fitness-related trait among invasive Ambrosia artemisiifolia populations in France. Mol Ecol 20(7):1378–1388. doi:10.1111/j.1365-294X.2011.05013.x
Coles SM (1973) Ranunculus bulbosus L. in Europe. Watsonia 9:207–228
Conover DO, Duffy TA, Hice LA (2009) The covariance between genetic and environmental influences across ecological gradients reassessing the evolutionary significance of countergradient and cogradient variation. In: Schlichting CD, Mousseau TA (eds) Year in evolutionary biology 2009, vol 1168. Annals of the New York Academy of Sciences, pp 100–129. doi:10.1111/j.1749-6632.2009.04575.x
Cornelissen JHC, Lavorel S, Garnier E, Díaz S, Buchmann N, Gurvich DE, Reich PB, ter Steege H, Morgan HD, van der Heijden MGA, Pausas JG, Poorter H (2003) A handbook of protocols for standardised and easy measurement of plant functional traits worldwide. Aust J Bot 51:335–380
De Kort H, Vandepitte K, Honnay O (2013) A meta-analysis of the effects of plant traits and geographical scale on the magnitude of adaptive differentiation as measured by the difference between Q ST and F ST. Evol Ecol 27(6):1081–1097. doi:10.1007/s10682-012-9624-9
Evanno G, Castella E, Goudet J (2006) Evolutionary aspects of population structure for molecular and quantitative traits in the freshwater snail Radix balthica. J Evol Biol 19(4):1071–1082. doi:10.1111/j.1420-9101.2006.01098.x
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinf 1:47–50
Fischer M, Rudmann-Maurer K, Weyand A, Stöcklin J (2008) Agricultural land use and biodiversity in the Alps. How cultural tradition and socioeconomically motivated changes are shaping grassland biodiversity in the Swiss Alps. Mt Res Dev 28(2):148–155
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180(2):977–993. doi:10.1534/genetics.108.092221
Frei ER, Ghazoul J, Matter P, Heggli M, Pluess AR (2014) Plant population differentiation and climate change: responses of grassland species along an elevational gradient. Glob Change Biol 20(2):441–455. doi:10.1111/gcb.12403
Gonzalo-Turpin H, Hazard L (2009) Local adaptation occurs along altitudinal gradient despite the existence of gene flow in the alpine plant species Festuca eskia. J Ecol 97(4):742–751. doi:10.1111/j.1365-2745.2009.01509.x
Goudet J, Buchi L (2006) The effects of dominance, regular inbreeding and sampling design on Q ST, an estimator of population differentiation for quantitative traits. Genetics 172(2):1337–1347. doi:10.1534/genetics.105.050583
Gutiérrez L (2008) Genetic diversity in cultivated and wild Hordeum specis. PhD Thesis, Iowa State University, Iowa, USA
Hahn T, Kettle CJ, Ghazoul J, Frei ER, Matter P, Pluess AR (2012) Patterns of genetic variation across altitude in three plant species of semi-dry grasslands. PLoS ONE 7(8):e41608. doi:10.1371/journal.pone.0041608
Hangartner S, Laurila A, Rasanen K (2012) Adaptive divergence in moor frog (Rana arvalis) populations along an acidification gradient: inferences from Q ST–F ST correlations. Evolution 66(3):867–881. doi:10.1111/j.1558-5646.2011.01472.x
Herrera CM, Bazaga P (2008) Adding a third dimension to the edge of a species’ range: altitude and genetic structuring in mountainous landscapes. Heredity 100(3):275–285. doi:10.1038/sj.hdy.6801072
Hutchison DW, Templeton AR (1999) Correlation of pairwise genetic and geographic distance measures: inferring the relative influences of gene flow and drift on the distribution of genetic variability. Evolution 53(6):1898–1914. doi:10.2307/2640449
IPCC (2013) Climate change 2013: the physical science basis. In: Stocker TF, Qin D, Plattner G-K, Tignor M, Allen SK et al (eds) Contribution of Working Group I to the fifth assessment report of the intergovernmental panel on climate change. Cambridge University Press, Cambridge
Joshi J, Schmid B, Caldeira M, Dimitrakopoulos P, Good J, Harris R, Hector A, Huss-Danell K, Jumpponen A, Minns A, Mulder C, Pereira J, Prinz A, Scherer-Lorenzen M, Siamantziouras A-SD, Terry A, Troumbis A, Lawton J (2001) Local adaptation enhances performance of common plant species. Ecol Lett 4:536–544
Kawecki TJ, Ebert D (2004) Conceptual issues in local adaptation. Ecol Lett 7(12):1225–1241. doi:10.1111/j.1461-0248.2004.00684.x
Körner C (2003) Alpine plant life: functional plant ecology of high mountain ecosystems, 2nd edn. Springer, Berlin, Germany
Kuittinen H, Mattila A, Savolainen O (1997) Genetic variation at marker loci and in quantitative traits in natural populations of Arabidopsis thaliana. Heredity 79:144–152. doi:10.1038/hdy.1997.137
Lande R (1992) Neutral theory of quantitative genetic variance in an island model with local extinction and colonization. Evolution 46(2):381–389. doi:10.2307/2409859
Legendre P, Legendre L (2012) Numerical Ecology. Elsevier, Amsterdam
Leinonen T, O`Hara RB, Cano JM, Merilä J (2008) Comparative studies of quantitative trait and neutral marker divergence: a meta-analysis. J Evol Biol 21:1–17
Lenormand T (2002) Gene flow and the limits to natural selection. Trends Ecol Evol 17(4):183–189. doi:10.1016/s0169-5347(02)02497-7
Linhart YB, Grant MC (1996) Evolutionary significance of local genetic differentiation in plants. Annu Rev Ecol Syst 27:237–277. doi:10.1146/annurev.ecolsys.27.1.237
Lopez-Fanjul C, Fernandez A, Toro MA (2003) The effect of neutral nonadditive gene action on the quantitative index of population divergence. Genetics 164(4):1627–1633
Lynch M (1996) A quantitative genetic perspective on conservation issues. In: Avise J, Hamrick J (eds) Conservation genetics: case histories from nature. Chapman & Hall, New York, pp 471–501
Lynch M, Milligan BG (1994) Analysis of population genetic-structure with RAPD markers. Mol Ecol 3(2):91–99. doi:10.1111/j.1365-294X.1994.tb00109.x
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Associates, Sunderland
Macel M, Lawson CS, Mortimer SR, Smilauerova M, Bischoff A, Cremieux L, Dolezal J, Edwards AR, Lanta V, Bezemer TM, van der Putten WH, Igual JM, Rodriguez-Barrueco C, Mueller-Schaerer H, Steinger T (2007) Climate vs. soil factors in local adaptation of two common plant species. Ecology 88(2):424–433. doi:10.1890/0012-9658(2007)88[424:cvsfil]2.0.co;2
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Matter P, Kettle CJ, Ghazoul J, Pluess AR (2013) Extensive contemporary pollen-mediated gene flow in two herb species, Ranunculus bulbosus and Trifolium montanum, along an altitudinal gradient in a meadow landscape. Ann Bot 111(4):611–621. doi:10.1093/aob/mct021
McKay JK, Latta RG (2002) Adaptive population divergence: markers, QTL and traits. Trends Ecol Evol 17(6):285–291. doi:10.1016/s0169-5347(02)02478-3
McKay JK, Bishop JG, Lin JZ, Richards JH, Sala A, Mitchell-Olds T (2001) Local adaptation across a climatic gradient despite small effective population size in the rare sapphire rockcress. Proc R Soc B Biol Sci 268(1477):1715–1721
Merilä J, Crnokrak P (2001) Comparison of genetic differentiation at marker loci and quantitative traits. J Evol Biol 14(6):892–903. doi:10.1046/j.1420-9101.2001.00348.x
Morgan KK, Hicks J, Spitze K, Latta L, Pfrender ME, Weaver CS, Ottone M, Lynch M (2001) Patterns of genetic architecture for life-history traits and molecular markers in a subdivided species. Evolution 55(9):1753–1761
O’Hara RB, Merilä J (2005) Bias and precision in Q ST estimates: problems and some solutions. Genetics 171(3):1331–1339. doi:10.1534/genetics.105.044545
Ohsawa T, Ide Y (2008) Global patterns of genetic variation in plant species along vertical and horizontal gradients on mountains. Glob Ecol Biogeogr 17(2):152–163. doi:10.1111/j.1466-8238.2007.00357.x
Oksanen J, Guillaume F, Kindt R, Legendre P, Minchin P, O`Hara RB, Simpson GL, Solymos P, Henry M, Stevens H, Wagner H (2013) vegan: community ecology package. 2.0-10
Palo JU, O’Hara RB, Laugen AT, Laurila A, Primmer CR, Merila J (2003) Latitudinal divergence of common frog (Rana temporaria) life history traits by natural selection: evidence from a comparison of molecular and quantitative genetic data. Mol Ecol 12(7):1963–1978. doi:10.1046/j.1365-294X.2003.01865.x
Petit C, Freville H, Mignot A, Colas B, Riba M, Imbert E, Hurtrez-Bousses S, Virevaire M, Olivieri I (2001) Gene flow and local adaptation in two endemic plant species. Biol Conserv 100(1):21–34. doi:10.1016/s0006-3207(00)00204-4
Pujol B, Wilson AJ, Ross RIC, Pannell JR (2008) Are Q ST – F ST comparisons for natural populations meaningful? Mol Ecol 17(22):4782–4785. doi:10.1111/j.1365-294X.2008.03958.x
Raesaenen K, Hendry AP (2008) Disentangling interactions between adaptive divergence and gene flow when ecology drives diversification. Ecol Lett 11(6):624–636. doi:10.1111/j.1461-0248.2008.01176.x
Reed DH, Frankham R (2001) How closely correlated are molecular and quantitative measures of genetic variation? A meta-analysis. Evolution 55(6):1095–1103
Roach DA, Wulff RD (1987) Maternal effects in plants. Annu Rev Ecol Syst 18:209–235. doi:10.1146/annurev.ecolsys.18.1.209
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145(4):1219–1228
Rudmann-Maurer K, Weyand A, Fischer M, Stocklin J (2008) The role of landuse and natural determinants for grassland vegetation composition in the Swiss Alps. Basic Appl Ecol 9(5):494–503. doi:10.1016/j.baae.2007.08.005
Santamaria L, Figuerola J, Pilon JJ, Mjelde M, Green AJ, De Boer T, King RA, Gornall RJ (2003) Plant performance across latitude: the role of plasticity and local adaptation in an aquatic plant. Ecology 84(9):2454–2461. doi:10.1890/02-0431
Sarukhan J, Harper JL (1973) Studies on plant demography: Ranunculus repens L., R. bulbosus L. and R. acris L.: I. population flux and survivorship. J Ecol 61(3):675–716. doi:10.2307/2258643
Scheepens JF, Frei ES, Stocklin J (2010) Genotypic and environmental variation in specific leaf area in a widespread Alpine plant after transplantation to different altitudes. Oecologia 164(1):141–150. doi:10.1007/s00442-010-1650-0
Silvertown J, Charlesworth D (2001) Introduction to plant population biology, 4th edn. Blackwell Science, Oxford, United Kingdom
Slatkin M (1977) Gene flow and genetic drift in a species subject to frequent local extinctions. Theor Popul Biol 12(3):253–262. doi:10.1016/0040-5809(77)90045-4
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236:787–792
Spitze K (1993) Population structure in Daphnia obtusa: quantitative genetic and allozymic variation. Genetics 135:367–374
Steinbach K, Gottsberger G (1994) Phenology and pollination biology of 5 Ranunculus species in Giessen, Central Germany. Phyton 34(2):203–218
Steinger T, Haldimann P, Leiss KA, Muller-Scharer H (2002) Does natural selection promote population divergence? A comparative analysis of population structure using amplified fragment length polymorphism markers and quantitative traits. Mol Ecol 11(12):2583–2590. doi:10.1046/j.1365-294X.2002.01653.x
Stinchcombe JR, Hoekstra HE (2008) Combining population genomics and quantitative genetics: finding the genes underlying ecologically important traits. Heredity 100(2):158–170. doi:10.1038/sj.hdy.6800937
Stöcklin J, Kuss P, Pluess AR (2009) Genetic diversity, phenotypic variation and local adaptation in the alpine landscape: case studies with alpine plant species. Bot Helv 119(2):125–133. doi:10.1007/s00035-009-0065-1
R Development Core Team (2012) R: a language and environment for statistical computing. 2.15.0. R Foundation for Statistical Computing, Vienna, Austria
Till-Bottraud I, Gaudeul M (2002) Intraspecific genetic diversity in alpine plants. In: Körner C, Spehn EM (eds) Mountain Biodiversity: a global assessment. Parthenon Publishing, New York, pp 23–34
Völler E, Auge H, Bossdorf O, Prati D (2013) Land use causes genetic differentiation of life-history traits in Bromus hordeaceus. Glob Change Biol 19(3):892–899
Vos P, Hogers R, Bleeker M, Reijans M, Vandelee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23(21):4407–4414. doi:10.1093/nar/23.21.4407
Whitlock MC (2008) Evolutionary inference from Q ST. Mol Ecol 17(8):1885–1896. doi:10.1111/j.1365-294X.2008.03712.x
Whitlock MC, Guillaume F (2009) Testing for spatially divergent selection: comparing Q ST to F ST. Genetics 183(3):1055–1063. doi:10.1534/genetics.108.099812
Wright S (1951) The genetical structure of populations. Ann Eugen 15:323–354
Young AG, Hill JH, Murray BG, Peakall R (2002) Breeding system, genetic diversity and clonal structure in the sub-alpine forb Rutidosis leiolepis F. Muell. (Asteraceae). Biol Conserv 106(1):71–78. doi:10.1016/s0006-3207(01)00230-0
Acknowledgments
We thank Sarah Burg, Constanze Conradin, Reinhard Frei, Ernest Hennig, Denise Kurth, Tamara Lebrecht, Jörg Leuenberger, Marianne Leuzinger, Monika Macsai, Philippe Matter, Annemarie Nazarek, and Marco Urech who helped collecting seeds, setting up the common garden, measuring and harvesting plants. We also thank all municipalities and farmers for the permission to work on their land. We thank Young Jin Chun and Bob O’Hara for statistical advice as well as Eva S. Frei, Martin Heggli and Irène Till-Bottraud for stimulating discussions and critical comments on the manuscript. We also thank the editor J. Stöcklin as well as J.F. Scheepens and an anonymous reviewer for their helpful comments on the manuscript. The AFLP fragment length readings were done in the Genetic Diversity Centre of ETH Zurich. The study was funded by the Swiss National Science Foundation (project no. 3100AO-116277) to J.G. and A.R.P., the Competence Center Environment and Sustainability (CCES) BioChange (http://www.cces.ethz.ch/projects/clench/BioChange) to J.G. and the Basler Stiftung für biologische Forschung to E.R.F.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the special issue Vegetation in cold environments under climate change.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Frei, E.R., Hahn, T., Ghazoul, J. et al. Divergent selection in low and high elevation populations of a perennial herb in the Swiss Alps. Alp Botany 124, 131–142 (2014). https://doi.org/10.1007/s00035-014-0131-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00035-014-0131-1