Abstract
Non-coding RNAs are important regulators of differentiation during embryogenesis as well as key players in the fine-tuning of transcription and furthermore, they control the post-transcriptional regulation of mRNAs under physiological conditions. Deregulated expression of non-coding RNAs is often identified as one major contribution in a number of pathological conditions. Non-coding RNAs are a heterogenous group of RNAs and they represent the majority of nuclear transcripts in eukaryotes. An evolutionary highly conserved sub-group of non-coding RNAs is represented by vault RNAs, named since firstly discovered as component of the largest known ribonucleoprotein complexes called “vault”. Although they have been initially described 30 years ago, vault RNAs are largely unknown and their molecular role is still under investigation. In this review we will summarize the known functions of vault RNAs and their involvement in cellular mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-coding RNAs represent the vast majority of transcriptional product of the human genome [1, 2]. The family of non-coding RNAs is composed of 19 different classes; among them transfer RNAs (tRNAs), tRNA-derived RNA fragments (tRFs), ribosomal RNAs (rRNAs), small nucleolar RNAs (snoRNAs), endogenous small interfering RNAs (endo-siRNAs), sno-derived RNAs (sdRNAs), transcription initiation RNAs (tiRNAs), miRNA-offset-RNAs (moRNAs), circular RNAs (circRNAs), vault RNAs, microRNAs (miRNAs), small interfering RNAs (siRNAs), small nuclear RNAs (snRNAs), extracellular RNAs (exRNAs), piwi-interacting RNAs (piRNAs), small Cajal body RNAs (scaRNAs), transcribed-ultraconserved regions (t-UCRs), long intergenic non-coding RNAs (lincRNAs), and long non-coding RNAs (lncRNAs) [3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25]. The role and function of tRNAs, rRNAs, microRNAs and lncRNAs, in particular, have been well examined both under physiological and pathological conditions [26]. In general, non-coding RNAs control all levels of genes’ regulation in eukaryotes, including the control of chromosome dynamics, splicing, RNA editing, translational inhibition and mRNA degradation [26]. Even transcription itself may be regulated by non-coding RNAs as outlined in several reports [27,28,29]. This is achieved on one hand, by control of chromosome dynamics and modifications and on the other hand, by regulation of RNA polymerase II activity. Therefore, non-coding RNAs are involved in regulation of accessibility of DNA sequences for the transcription machinery, as well as in modulation of the transcription rate of RNA polymerase II [30,31,32,33,34]. Furthermore, splicing of pre-mRNA transcripts, post-transcriptional regulation of expression rate as well as translation of mRNAs in cytoplasm and regulation of mRNA half-life are under control of non-coding RNAs [26, 27, 35]. In addition, some non-coding RNAs are known to be involved in intercellular communication and cell regulation [36, 37]; whereas, others are part of the antiviral defence by stimulating immune response and activating RNA interference pathway [38, 39].
In contrast, the molecular functions of vault RNAs are still not completely clear even after more than 30 years since their discovery [40, 41]. With a length between 88 and 140 nucleotides vault RNAs are longer than miRNAs, but they are still included as members of the short non-coding RNA group [41, 42].
In humans, four vault RNAs are encoded on chromosome 5q31 in two different loci. The VTRNA-1 locus (located between zinc-finger matrin-type 2 gene and proto-cadherin cluster) contains the genetic information for three vault RNAs (vault RNA1-1, vault RNA1-2 and vault RNA1-3) and VTRNA-2 locus (located between the genes coding for transforming growth factor beta 1 and SMAD family member 5) codes for vault RNA2-1 also known as pre-miR-886 [43,44,45]. All vault RNA genes are under control of a polymerase III type 2 promoter and they contain a box A and box B motif normally found in tRNA genes [41, 42]. Nevertheless, the promoters of the two vault RNA loci are not identical; therefore, expression patterns of the vault RNA genes are different [42]. Furthermore, epigenetic modifications such as promoter methylation are important regulators for vault RNAs expression especially for the VTRNA-2 gene [46, 47]. The distant regulatory elements of the VTRNA-1 promoter are characterized by differential CpG accessibility and this might be a hint for a cell-type-specific expression of the three vault RNAs under control of this promoter [48]. The internal promoter sequences box A and box B present in VTRNA-1 and VTRNA-2 enable binding of transcription factors TFIIIC and TFIIIB which facilitate polymerase III binding to the transcription starting site [49]. Vault RNAs transcription is also under control of cAMP response (CRE)- and tetradecanoyl-phorbol acetate response (TRE)-like elements [41, 42]. These elements represent binding sites for the transcription factors CREB and AP-1, respectively, which adapt key cellular processes such as differentiation, proliferation and survival to nutrient, growth factor and stress signaling [50, 51]. This could explain the observed differential vault RNA transcription rate upon viral infection, starvation and cancer [45, 52,53,54]. Furthermore, the short half-life time observed of around 1 h makes vault RNAs suitable as signaling molecules that quickly respond to stimuli [55, 56].
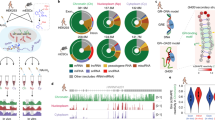
Vault RNAs were first identified as a component of vault particles [40] but most of the vault RNAs (around 95%) are not associated with these particles and therefore, vault RNAs are most probably also involved in other cellular processes and interactions [57, 58] (Fig. 1).
Vault RNAs are involved in different cellular processes and interactions. The vast majority (95%) of vault RNAs are not associated with the vault ribonucleoprotein complex; they are involved in regulation of important cellular pathways like cell death (intrinsic and extrinsic apoptosis; autophagy), proliferation, cellular differentiation, mRNA regulation and cell–cell communication. Besides this, vault RNAs influence the immune reaction and especially in cancer, they have an important role in rendering cells resistance to drug treatment
Vault particles
Vault particles are the largest known ribonucleoprotein complexes in eukaryotes [59, 60]. Structure and protein composition of vaults are highly conserved and present in high number (10,000–100,000/cell) in different species [43, 61,62,63,64]. Therefore, it is very likely that vault particles might play fundamental roles in eukaryotic cells [42]. Some reports connect the vault particles complex to a plethora of mechanisms that include: nuclear-pore complex formation, nuclear–cytoplasmic transport, cellular signaling, innate immune response, apoptosis resistance, DNA damage repair and development of multidrug resistance in cancer cells [43, 65,66,67,68,69,70,71,72] (Fig. 1).
The main component of vault particles is the major vault protein that is sufficient itself for the structural conformation characteristic of the vault particles [73,74,75,76,77]. The major vault protein has no homology to any other protein known, but it is highly conserved among different species (around 90% identity between mammalians and around 60% with lower organisms) [75, 78]. Interestingly, the major vault protein contains two Ca2+ binding sites at the N-terminal end which are necessary for correct folding and particle assembly but also to interact with other proteins like PTEN, thus connecting the vault particles to cellular signaling pathways [79]. Beside the major vault protein, two other proteins are present in the vaults particles; the poly-(adenosine-diphosphate ribose) polymerase—a member of the PARP family—and the telomerase-associated protein 1 (TEP1) [80,81,82]. In these vault particles, the vault RNAs are associated with the caps [43, 57, 75, 83]. The vast majority (around 90%) of vault particles in unstressed cells are located in the cytoplasm but vault particles are also found to be associated with the nuclear membrane. The distribution of vault particles varies in response to external stimuli and rapidly react towards extracellular changes with translocation to different subcellular compartments. Furthermore, under pathological conditions like cancer, a higher amount of vault particles are associated with the nuclear membrane and up to 5% of them are found within the nucleus [68, 84,85,86,87,88]. Based on this observation and the barrel-like structure of the vault particles, the hypothesis exist that vault particles have an important role in mediating shuttle processes between cytoplasm and nucleus, including nuclear import of tumor-suppressors like PTEN, nuclear hormone receptors as well as drug export. It is speculated that some of the cargos transported in vault particles are bound to the vault RNAs present in these complexes [66, 67, 89,90,91]. But up to now, the role of vault particles as transporter is still under discussion and further investigation is urgently needed because most of these studies used either immunoprecipitation of signaling complexes or yeast two hybrid systems, and it cannot be excluded that the found interaction with vault particles and vault RNAs occurred accidentally and is without any biological sense. A verification of vault particles as transporter in humans under physiological and/or pathological conditions (e.g., tumor) is still missing.
Studies on vault particles using knock-out mice
The role of major vault protein and vault particles has been addressed in relevant mice knock-out models [92,93,94]. In TEP1 knock-out mice vault particles were still present, but inside these vault particles, no vault RNAs have been found. Therefore, it was concluded that TEP1 is absolutely required for a stable association of vault RNAs with the vault complex [93]. TEP1 knock-out mice as well as major vault protein knock-out mice are viable, healthy and display no obvious abnormalities [93, 94]. The major vault protein knock-out mice express no vaults particles as expected and have been used in different studies to examine the role of vault particles [78, 84, 94, 95]. Surprisingly, embryonic stem cells and bone marrow cells derived from major vault protein knock-out mice showed no change in sensitivity to drugs when compared to wild-type mice cells. In addition, the activities of the multidrug resistance-related transporters P-glycoprotein, multidrug resistance-associated protein and breast cancer resistance protein were not altered in vault-deficient cells ruling out the possibility that these proteins compensate for the loss of vaults. Also, the response towards doxorubicin treatment was the same in major vault protein knock-out and wild-type mice in in vivo experiments [94]. These observations lead to the conclusion that at least in mice, vaults are not directly involved in drug resistance [78, 84]. In another study, the major vault protein knock-out mice were used to address the role of vaults in regard to dendritic cells. Development and function of dendritic cells, derived from mononuclear bone marrow cells, appeared normal in knock-out mice. In-vivo immunization assays showed that neither T-cell-mediated immune response nor T-cell-dependent humoral response were affected by major vault protein knock-out, indicating intact antigen-presenting and migration capacities of dendritic cells. Obviously, in mice vault particles are not required for primary dendritic cell functions [95]. This observation is in contrast with findings in humans where major vault protein and vault particles are up-regulated during the development of human dendritic cells. Moreover, major vault protein-specific antibodies, presumably interfering with the function of major vault protein or vaults, resulted in reduced expression levels of dendritic cell markers, co-stimulatory molecules and decreased capacity to induce T-cell proliferative and interferon-ɣ-releasing responses [96]. Recently, the major vault protein was identified as a suppressor for NF-κB signaling in macrophages [97]. Global as well as myeloid-specific major vault protein gene knock-out intensified high-fat diet-induced obesity, insulin resistance, hepatic steatosis and atherosclerosis in mice via NF-κB signaling pathway. Furthermore, increased macrophage infiltration and inflammatory responses in the microenvironments have been observed [97]. Another study used peripheral blood mononuclear cells (PBMCs) from major vault protein knock-out mice and evaluated an essential role of major vault protein for the induction of early antiviral cytokines (like IL-6 and IL-8) in the context of double-stranded RNA- or virus-induced pro-inflammatory response [98]. In the following sections, we will focus on the role of vault particles and vault RNAs in humans.
Vault RNAs, vault particles and drug resistance
One of the roles of the vault particles is the contribution to mediate drug resistance mechanisms by transporting the drugs from their intracellular targets to the extracellular compartment and also in drug sequestration [78]. In an elegant experiment, expression of major vault proteins was prevented by a siRNA approach in human bladder cancer cells under doxycycline treatment. This resulted in inhibition of cytosolic doxorubicin sequestration in perinuclear lysosomes and enhanced accumulation of the drug in the nucleus as well as increased cytotoxicity [99]. Based on the fact that nuclear PTEN is involved in the maintenance of chromosomal stability [100], its nuclear transportation by vaults particles could also play a role in drug resistance mechanisms by counteracting drug-induced DNA damage [101].
In most cell lines, vault RNA1-1 has the highest expression level of all vault RNA transcripts [102]. In multidrug-resistant cells, the level of vault RNA1-1 is not altered but expression rate of vault RNA1-3 is raised and an increased association of vault RNA1-3 with vaults particles has been observed [43, 102]. However, the molecular details behind this observation are still not clear. In general, vault RNAs bound to the vault particles have the capacity to interact with drugs via specific binding sites [103]. For instance, in cancer patients who developed resistance to chemotherapy, the number of vault particles is increased, in agreement with their observed role, in in vitro models [78, 103,104,105]. Another relevant example is given by mitoxantrone resistance in osteosarcoma, glioblastoma and leukemia where drug failure is based on direct binding of the drug to vtRNA1-1 and vtRNA1-2 [103, 104].
Besides sequestering drugs, vault RNAs are processed into several small RNAs (Fig. 1). Among them small-vault RNAs, account in a second way for multidrug resistance in cancer patients by down-regulating CYP3A4, the key enzyme in drug metabolism [106]. Interestingly, the introduction of 5-methyl-cytosine by the RNA methyltransferase NSUN2-dependent leads to the cleavage of vault RNAs in a Dicer-dependent mechanism; thus, the resulting small-vault RNAs regulate their target genes in a miRNA-like fashion [106,107,108].
Furthermore, vault RNAs can induce drug resistance in an indirect way by influencing cell proliferation and preventing cell death as described in the following sections.
Vault RNAs and proliferation
Drug resistance can also arise by the increase in cell proliferation rate [109, 110]. Vault RNAs have been found to influence cell proliferation in different ways and in a cell-type-specific manner without the participation of vault particles (Fig. 1).
In breast cancer, vault RNA1-1 interacts directly with the RNA/DNA-binding protein polypyrimidine tract binding splicing factor (PSF) [105]. PSF is an important regulatory nuclear protein that acts as a component of spliceosomes via the RNA-binding domain and furthermore regulates transcription of genes via the DNA-binding domain; e.g., PSF controls the transcription of P450-linked side-chain cleaving enzyme (CYP11A1) and regulates this by the steroid pathway; in addition PSF inhibits transcription of proto-oncogene G antigen 6 (GAGE6) [111,112,113]. Following the binding of vault RNA1-1 to PSF RNA-binding domain, the transcriptional repression of GAGE6 via the DNA-binding domain is released and transcription of the proto-oncogene proceeds [114]. Induced expression of GAGE6 results in increased cell proliferation and causes drug resistance [105]. Vault RNA2-1 interacts with and blocks the pro-apoptotic interferon-inducible protein kinase R (PKR). PKR is a central protein for cellular response to different stress signals such as pathogens, starvation, cytokines and irradiation. PKR activates different central pathways like JNK, NF-ϰB, PP2A, p38 and inhibits the eukaryotic translation initiation factor eIF2α by phosphorylation [115]. In normal cells, this inhibits further cellular mRNA translation based on AUG initiation codons and in parallel activates the tumor-suppressor PP2A which blocks cell-cycle, as well as proliferation and leads ultimately to cell death [55]. In different cancer cells, active PKR fails to induce phosphorylation of eIF2α and PP2A, so that apoptosis is not triggered but PKR promotes still the pro-survival NF-ϰB pathway [116,117,118]. Therefore, the reduced expression levels of vault RNA2-1 found in cancer cell lines and cancer patients specimens result in activation of PKR and subsequent increased cell proliferation as well as drug resistance [119]. Consequently, vault RNA2-1 seems to act as tumor suppressor in contrast to oncogenic effects of vault RNA1-1 [46, 47, 120,121,122,123].
Vault RNAs and apoptosis
Vault RNA1-1 is involved in inhibiting the intrinsic as well as extrinsic apoptosis pathway in several cancer cell lines as demonstrated in in vitro experiments [54, 124]. To address the role of vault RNAs in apoptotic mechanisms, cells have been treated with an autophagy inhibitor and cell death induced by serum starvation. Cells with knock-out vault RNA1-1 gene were more susceptible to programmed cell death; whereas, re-expression of vault RNA1-1 restored apoptosis resistance of the cells. The mechanism underlying the blocking of apoptosis seems to be related to a short stretch within the central domain of vault RNA1-1 and cannot be exerted by other vault RNA members. Furthermore, it was demonstrated that regulation of programmed cell death is independent of vault particles and relay only on vault RNA1-1 [54]. The protective effects of vault RNA1-1 against programmed cell death have been observed after triggering the intrinsic (via staurosporine, etoposide) as well as extrinsic (via Fas ligand) apoptosis pathway [54]. Increased vault RNA1-1 expression activates the pro-survival PI3K-/AKT- and ERK1/2 MAPK-signaling pathways and by this counteract cell death [124]. In Epstein–Barr virus (EBV)-infected B-cells, the latent membrane protein 1 (LMP1) of EBV up-regulates the NF-ϰB pathway that results in increased expression of vault RNA1-1. In this context, vault RNA1-1 inhibits the extrinsic and intrinsic apoptotic pathways and enables cell proliferation by further activation of NF-ϰB pathway and up-regulation of the expression of Bcl-xL [54].
Vault RNAs and autophagy
Autophagy is besides apoptosis another catabolic pathway essential in homeostasis of cells [125]. Both mechanisms are interconnected by several molecular nodes and a close cross-talk exists [54]. In direct proximity of the VTRNA-1 locus is the proto-cadherin cluster that encodes for the proto-cadherin family, which is involved in autophagy [126]. Therefore, it seems indicative that also vault RNAs might be involved in autophagy [52] (Fig. 1). Autophagic process is necessary for cleaning out unnecessary or dysfunctional components in cells and recycle nutrients and energy. All cargos that cannot be degraded by the ubiquitin–proteasome system are cleaved via autophagy in the lysosomes [127,128,129]. In contrast to apoptosis that results in cell death [130], autophagy in cancer can facilitate tumor cell survival in stress conditions (e.g., under hypoxia or starving conditions) by providing energy and nutrients [131]. An established marker for the autophagic state of a cell is the intracellular levels of p62 [132]. The selective autophagy receptor p62 [133, 134] is of pivotal importance to the autophagic process by recognizing cargos for the autophagic process, triggering autophagosome formation and exerting a regulatory role in autophagy [127, 135,136,137,138]. Vault RNA1-1 binds directly to p62 preventing its oligomerization, a prerequisite for autophagy. This results in inhibition of p62-dependent autophagy and aggregate clearance [52, 139]. Another role of p62 is the cross-talk between autophagy and apoptosis [140, 141] and increased levels of monomeric p62, upon autophagy inhibition via vault RNA1-1, could modulate the balance between the two catabolic pathways. In addition, p62 is involved in the regulation of inflammatory pathways, especially the autophagic defence against invading bacteria and viruses [142]. Most probably, viruses target p62 by up-regulation vault RNAs to decrease the autophagic processes in parallel with inhibition of interferon responses as outlined further below [53].
Vault RNAs, cellular differentiation and development
It is well established that the levels of non-coding RNAs, including vault RNAs, are highly regulated during development and cellular differentiation since they are essential to these processes [143]. One example is based on the above-mentioned NSUN2-dependent 5-methyl-cytosine modification of vault RNA1.1 and vault RNA1.3 [107, 144] which was recently reported to influence cell differentiation [107, 108]. The serine/arginine-rich splicing factor 2 (SRSF2) binds to the non-methylated form of vault RNA1-1 with higher affinity and counteracts the processing by NSUN2 [108]. Therefore, the expression level of SRSF2 and NSUN2 and their binding to vault RNA1-1 orchestrates the production of small-vault RNAs. The lack of NSUN2-mediated methylation of vault RNA1-1 results in reduced amount of small-vault RNAs and results in changes in epidermal differentiation program of keratinocytes [107, 108]. It is well established also that lack of NSUN2-dependent 5-methyl-cytosine modification in other non-coding RNAs modifies the physiologic situation too; e.g., aberrant 5-methyl-cytosine modification of tRNAs impairs the translation machinery and causes neuro-developmental deficits [145, 146].
The regulated expression of a small-vault RNA derived from vault RNA2-1 (called small-vault RNA2-1a) has been shown to modulate early developmental processes in the central nervous system and has an important role in human brain development as well as aging. The small-vault RNA2-1a has the highest expression level early in post-natal developmental stages and the amount decreases after 1 year with low levels being detected at the oldest ages examined [147].
Vault RNA-derived small RNAs
Another peculiar characteristic of vault RNAs is that they can be processed into several small RNAs and the cleavage process of vault RNAs is mediated by RNA methyltransferase NSUN2. The introduction of 5-methyl-cytosine by NSUN2 is a prerequisite for DICER-dependent cleavage process of vault RNAs and the resulting small-vault RNAs regulate their target genes in a miRNA-like fashion [106,107,108] as the aforementioned down-regulation of CYP3A4 by small RNAs resulting in altered dug metabolism [106] as well as the role of small-vault RNAs for epidermal differentiation program of keratinocytes [107, 108]. In both cases, the small-vault RNAs are processed from vault RNA1-1. Another example for the role of small-vault-dependent RNAs was recently reported in prostate cancer. Vault RNA2-1 produces two small RNAs (snc886-3p and snc886-5p) that are found to be reduced in tumor tissues compared to the surrounding normal tissues. Based on PAR-CLIP (photoactivatable ribonucleoside-enhanced crosslinking and immunoprecipitation) and knock-out experiments of microRNA biogenesis enzymes, it was demonstrated that vault RNA2-1 cleavage is based on DICER but independent of DROSHA and the resulting small-vault RNAs are associated with argonaute proteins [148] in a similar process of miRNAs biogenesis [149]. As functional proof of action, over-expression of snc886-3p in relevant in vitro and in vivo systems, resulted in down-regulation of mRNAs containing complementary sequences to the seed sequence of the small-vault RNA in their 3′-UTRs. This led to reduced cell cycle progression, increased apoptosis [148, 150] and this seems in agreement with the view of vault RNA2-1 as tumor suppressor [46, 47, 120,121,122,123]. In Parkinson disease, a small-vault RNA derived from vault RNA2-1 is up-regulated in early stages of the disease and this small-vault RNA is most probably involved in the process of brain development as outlined in detail above [151].
Furthermore, small RNAs derived from vault RNAs and associated with the argonaute complex have been identified also in breast, prostate, lung and lymphoid tissue [106, 148]. These findings support the hypothesis of a cleavage of vault RNAs into small RNAs which influence mRNA stability and/or regulate translation like miRNAs. However, the main role of small-vault RNAs need further investigation and it will be of valuable interest if these small RNAs can regulate transcription in a tissue and cell-type-specific fashion as miRNAs [149].
Furthermore, small-vault RNAs are secreted by cells and they are present in high numbers in exosomes (Fig. 1). Therefore, small-vault RNAs are most probably also involved in cell–cell signaling [106, 107, 152].
Vault RNAs, viral infection and immune system
Viral infections induce vault RNA expression [45, 153] and this was observed in in vitro models for different virus families including γ-herpesviridae (Herpes simplex virus 1), paramyxovirus (Sendai and Epstein–Barr virus), Kaposi's sarcoma-associated herpes and influenza-A virus [44, 45, 53, 54]. Most of these viruses are known to reduce the autophagic capacity of their host cells that is a consequence of high expression levels of vaults RNAs as mentioned above [53, 154]. In addition, transcriptional induction of vault RNAs upon infection, has been associated with expression of latent membrane protein 1 for EBV and non-structural protein NS1 of influenza virus, respectively, with the aim to prevent cells from apoptosis and suppress PKR-mediated innate immunity [53, 54]. Therefore, high expression levels of vault RNAs result in an increased viral load. Viruses are known to hijack cells and their cellular replication machinery to maximize viral replication while inhibiting cellular defence mechanisms [155]. Up-regulation of vault RNA levels seems to be a very efficient way to escape targeted viral degradation via autophagy and subsequent MHC class II antigen presentation [156] and in parallel force the cell to enter a pro-proliferative state that counteracts cellular suicide programs as well as support rapid virus replication [157]. Therefore, it is not surprising that vault RNAs are hijacked and used by viruses. This underlines the important and central role of vault RNAs in regulating cellular processes (Fig. 1).
Another effect of viral infection is the reduction of cellular DUSP11 expression. DUSP11-mediated de-phosphorylation of the 5′-end of vault RNAs initiates the degradation of these RNAs [158, 159]. Therefore, an infection-dependent reduction of DUSP11 levels results in accumulation of vault RNAs that in turn trigger an innate immune response via retinoic acid-inducible gene-1 (RIG-1) receptors [160]. By this, at least one of the anti-viral defence mechanisms against RNA virus is activated [161].
Vault RNAs as diagnostic and prognostic markers
In Parkinson disease, down-regulation of miR-7, miR-34b/c and miR-133b [162,163,164] as well as up-regulation of a small-vault RNA derived from vault RNA2-1 is common in brain areas that are affected by this disease [147]. Increased expression of vault RNA2-1 occurs early in the course of disease and could perhaps be used as a diagnostic marker.
Hyper-methylation of vault RNA2-1 gene is correlated with poor prognosis and overall survival in several cancers; e.g., gastric, oesophageal, lung, prostate, acute myeloid leukemia and myelodysplastic syndrome. Therefore, vault RNA2-1 could act as tumor suppressor [46, 47, 120,121,122,123, 150] and the expression level of vault RNA2-1 could be used as a prognostic marker.
In addition, the expression level of the major vault protein has been correlated with therapy resistance, prognosis and overall survival in several blood cancers (acute myeloid leukemia, acute lymphoblastic leukemia, adult T-cellss leukemia and multiple myeloma) [165,166,167,168,169,170,171,172,173,174,175,176]. In solid tumors, expression level of the major vault protein is a good prediction factor for response to chemotherapy in bladder cancer [177], melanoma [178] and for determining the aggressive phenotype of testicular germ-cell tumors [179] and glioblastoma [180].
Conclusion and perspectives
The old simplistic view that non-coding RNAs only play functional roles in protein synthesis as integral components (rRNA) or reaction substrates (tRNA) of the ribosome has dramatically evolved during the last 2 decades with emerging concepts linking different classes of non-coding RNAs to physiology and disease. The non-coding RNA group of vault RNAs, which is composed of only four members in human, exert an important role within the cell. Although until recently not all functions and processes have been unveiled in detail, it is already clear that vault RNAs add another level of regulation to the network of non-coding and coding RNAs. As outlined in this review, vault RNAs are involved in transferring extracellular stimuli into signals inside the cell; they regulate central signaling pathways and cell–cell communication. Furthermore, vault RNAs play a substantial role in immunity response, influencing proliferation, apoptosis and autophagy as well as being involved in drug resistance mechanisms (Fig. 1). All these functions are under vault RNAs regulation either via direct interaction with proteins or via post-transcriptional regulation of mRNAs. In particular, in the context of cancer, vault RNAs appear to have a critical role and a better understanding of their biology in this disease could offer a new prospect for cancer treatment and prevention of drug resistance.
References
Geisler S, Coller J (2013) RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol 14:699–712. https://doi.org/10.1038/nrm3679
Morris KV, Mattick JS (2014) The rise of regulatory RNA. Nat Rev Genet 15:423–437. https://doi.org/10.1038/nrg3722
Hahne JC, Valeri N (2018) Non-coding RNAs and resistance to anticancer drugs in gastrointestinal tumors. Front Oncol 8:226. https://doi.org/10.3389/fonc.2018.00226
Eddy SR (2001) Non-coding RNA genes and the modern RNA world. Nat Rev Genet 2:919–929. https://doi.org/10.1038/35103511
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531. https://doi.org/10.1038/nrg1379
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227. https://doi.org/10.1038/nature07672
Langenberger D, Bermudez-Santana C, Hertel J, Hoffmann S, Khaitovich P, Stadler PF (2009) Evidence for human microRNA-offset RNAs in small RNA sequencing data. Bioinformatics 25:2298–2301. https://doi.org/10.1093/bioinformatics/btp419
Taft RJ, Glazov EA, Cloonan N, Simons C, Stephen S, Faulkner GJ, Lassmann T, Forrest AR, Grimmond SM, Schroder K, Irvine K, Arakawa T, Nakamura M, Kubosaki A, Hayashida K, Kawazu C, Murata M, Nishiyori H, Fukuda S, Kawai J, Daub CO, Hume DA, Suzuki H, Orlando V, Carninci P, Hayashizaki Y, Mattick JS (2009) Tiny RNAs associated with transcription start sites in animals. Nat Genet 41:572–578. https://doi.org/10.1038/ng.312
Taft RJ, Glazov EA, Lassmann T, Hayashizaki Y, Carninci P, Mattick JS (2009) Small RNAs derived from snoRNAs. RNA 15:1233–1240. https://doi.org/10.1261/rna.1528909
Wilusz JE, Sunwoo H, Spector DL (2009) Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 23:1494–1504. https://doi.org/10.1101/gad.1800909
Ling H, Fabbri M, Calin GA (2013) MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov 12:847–865. https://doi.org/10.1038/nrd4140
Claycomb JM (2014) Ancient endo-siRNA pathways reveal new tricks. Curr Biol 24:R703–R715. https://doi.org/10.1016/j.cub.2014.06.009
Guo JU, Agarwal V, Guo H, Bartel DP (2014) Expanded identification and characterization of mammalian circular RNAs. Genome Biol 15:409. https://doi.org/10.1186/s13059-014-0409-z
An Y, Furber KL, Ji S (2017) Pseudogenes regulate parental gene expression via ceRNA network. J Cell Mol Med 21:185–192. https://doi.org/10.1111/jcmm.12952
Azlan A, Dzaki N, Azzam G (2016) Argonaute: the executor of small RNA function. J Genet Genom 43:481–494. https://doi.org/10.1016/j.jgg.2016.06.002
Beermann J, Piccoli MT, Viereck J, Thum T (2016) Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev 96:1297–1325. https://doi.org/10.1152/physrev.00041.2015
de Almeida RA, Fraczek MG, Parker S, Delneri D, O’Keefe RT (2016) Non-coding RNAs and disease: the classical ncRNAs make a comeback. Biochem Soc Trans 44:1073–1078. https://doi.org/10.1042/BST20160089
Evans JR, Feng FY, Chinnaiyan AM (2016) The bright side of dark matter: lncRNAs in cancer. J Clin Investig 126:2775–2782. https://doi.org/10.1172/JCI84421
Geiger J, Dalgaard LT (2017) Interplay of mitochondrial metabolism and microRNAs. Cell Mol Life Sci 74:631–646. https://doi.org/10.1007/s00018-016-2342-7
Granados-Riveron JT, Aquino-Jarquin G (2016) The complexity of the translation ability of circRNAs. Biochim Biophys Acta 1859:1245–1251. https://doi.org/10.1016/j.bbagrm.2016.07.009
Khurana E, Fu Y, Chakravarty D, Demichelis F, Rubin MA, Gerstein M (2016) Role of non-coding sequence variants in cancer. Nat Rev Genet 17:93–108. https://doi.org/10.1038/nrg.2015.17
Qi P, Zhou XY, Du X (2016) Circulating long non-coding RNAs in cancer: current status and future perspectives. Mol Cancer 15:39. https://doi.org/10.1186/s12943-016-0524-4
Quinn JJ, Chang HY (2016) Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet 17:47–62. https://doi.org/10.1038/nrg.2015.10
Taal BG, Audisio RA, Bleiberg H, Blijham GH, Neijt JP, Veenhof CH, Duez N, Sahmoud T (1993) Phase II trial of mitomycin C (MMC) in advanced gallbladder and biliary tree carcinoma. An EORTC Gastrointestinal Tract Cancer Cooperative Group Study. Ann Oncol 4:607–609. https://doi.org/10.1093/oxfordjournals.annonc.a058597
Yu M, Lu B, Zhang J, Ding J, Liu P, Lu Y (2020) tRNA-derived RNA fragments in cancer: current status and future perspectives. J Hematol Oncol 13:121. https://doi.org/10.1186/s13045-020-00955-6
Mattick JS, Makunin IV (2006) Non-coding RNA. Hum Mol Genet 15(Spec No 1):R17–R29. https://doi.org/10.1093/hmg/ddl046
Kaikkonen MU, Lam MT, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90:430–440. https://doi.org/10.1093/cvr/cvr097
Mattick JS (2003) Challenging the dogma: the hidden layer of non-protein-coding RNAs in complex organisms. BioEssays 25:930–939. https://doi.org/10.1002/bies.10332
Corey DR (2005) Regulating mammalian transcription with RNA. Trends Biochem Sci 30:655–658. https://doi.org/10.1016/j.tibs.2005.09.007
Yang Z, Zhu Q, Luo K, Zhou Q (2001) The 7SK small nuclear RNA inhibits the CDK9/cyclin T1 kinase to control transcription. Nature 414:317–322. https://doi.org/10.1038/35104575
Michels AA, Fraldi A, Li Q, Adamson TE, Bonnet F, Nguyen VT, Sedore SC, Price JP, Price DH, Lania L, Bensaude O (2004) Binding of the 7SK snRNA turns the HEXIM1 protein into a P-TEFb (CDK9/cyclin T) inhibitor. EMBO J 23:2608–2619. https://doi.org/10.1038/sj.emboj.7600275
Yik JH, Chen R, Pezda AC, Zhou Q (2005) Compensatory contributions of HEXIM1 and HEXIM2 in maintaining the balance of active and inactive positive transcription elongation factor b complexes for control of transcription. J Biol Chem 280:16368–16376. https://doi.org/10.1074/jbc.M500912200
Li Q, Price JP, Byers SA, Cheng D, Peng J, Price DH (2005) Analysis of the large inactive P-TEFb complex indicates that it contains one 7SK molecule, a dimer of HEXIM1 or HEXIM2, and two P-TEFb molecules containing Cdk9 phosphorylated at threonine 186. J Biol Chem 280:28819–28826. https://doi.org/10.1074/jbc.M502712200
Haaland RE, Herrmann CH, Rice AP (2005) siRNA depletion of 7SK snRNA induces apoptosis but does not affect expression of the HIV-1 LTR or P-TEFb-dependent cellular genes. J Cell Physiol 205:463–470. https://doi.org/10.1002/jcp.20528
Yoon JH, Abdelmohsen K, Kim J, Yang X, Martindale JL, Tominaga-Yamanaka K, White EJ, Orjalo AV, Rinn JL, Kreft SG, Wilson GM, Gorospe M (2013) Scaffold function of long non-coding RNA HOTAIR in protein ubiquitination. Nat Commun 4:2939. https://doi.org/10.1038/ncomms3939
Ramon YCS, Segura MF, Hummer S (2019) Interplay between ncRNAs and cellular communication: a proposal for understanding cell-specific signaling pathways. Front Genet 10:281. https://doi.org/10.3389/fgene.2019.00281
Klinge CM (2018) Non-coding RNAs in breast cancer: intracellular and intercellular communication. Noncoding RNA. https://doi.org/10.3390/ncrna4040040
Qiu L, Wang T, Tang Q, Li G, Wu P, Chen K (2018) Long non-coding RNAs: regulators of viral infection and the interferon antiviral response. Front Microbiol 9:1621. https://doi.org/10.3389/fmicb.2018.01621
Wang P (2018) The opening of Pandora’s Box: an emerging role of long noncoding RNA in viral infections. Front Immunol 9:3138. https://doi.org/10.3389/fimmu.2018.03138
Kedersha NL, Rome LH (1986) Isolation and characterization of a novel ribonucleoprotein particle: large structures contain a single species of small RNA. J Cell Biol 103:699–709. https://doi.org/10.1083/jcb.103.3.699
Kickhoefer VA, Searles RP, Kedersha NL, Garber ME, Johnson DL, Rome LH (1993) Vault ribonucleoprotein particles from rat and bullfrog contain a related small RNA that is transcribed by RNA polymerase III. J Biol Chem 268:7868–7873
Stadler PF, Chen JJ, Hackermuller J, Hoffmann S, Horn F, Khaitovich P, Kretzschmar AK, Mosig A, Prohaska SJ, Qi X, Schutt K, Ullmann K (2009) Evolution of vault RNAs. Mol Biol Evol 26:1975–1991. https://doi.org/10.1093/molbev/msp112
Kickhoefer VA, Rajavel KS, Scheffer GL, Dalton WS, Scheper RJ, Rome LH (1998) Vaults are up-regulated in multidrug-resistant cancer cell lines. J Biol Chem 273:8971–8974. https://doi.org/10.1074/jbc.273.15.8971
Nandy C, Mrazek J, Stoiber H, Grasser FA, Huttenhofer A, Polacek N (2009) Epstein–barr virus-induced expression of a novel human vault RNA. J Mol Biol 388:776–784. https://doi.org/10.1016/j.jmb.2009.03.031
Mrazek J, Kreutmayer SB, Grasser FA, Polacek N, Huttenhofer A (2007) Subtractive hybridization identifies novel differentially expressed ncRNA species in EBV-infected human B cells. Nucleic Acids Res 35:e73. https://doi.org/10.1093/nar/gkm244
Helbo AS, Treppendahl M, Aslan D, Dimopoulos K, Nandrup-Bus C, Holm MS, Andersen MK, Liang G, Kristensen LS, Gronbaek K (2015) Hypermethylation of the VTRNA1-3 Promoter is associated with poor outcome in lower risk myelodysplastic syndrome patients. Genes (Basel) 6:977–990. https://doi.org/10.3390/genes6040977
Treppendahl MB, Qiu X, Sogaard A, Yang X, Nandrup-Bus C, Hother C, Andersen MK, Kjeldsen L, Mollgard L, Hellstrom-Lindberg E, Jendholm J, Porse BT, Jones PA, Liang G, Gronbaek K (2012) Allelic methylation levels of the noncoding VTRNA2-1 located on chromosome 5q31.1 predict outcome in AML. Blood 119:206–216. https://doi.org/10.1182/blood-2011-06-362541
Helbo AS, Lay FD, Jones PA, Liang G, Gronbaek K (2017) Nucleosome positioning and NDR structure at RNA polymerase III promoters. Sci Rep 7:41947. https://doi.org/10.1038/srep41947
Schramm L, Hernandez N (2002) Recruitment of RNA polymerase III to its target promoters. Genes Dev 16:2593–2620. https://doi.org/10.1101/gad.1018902
Angel P, Imagawa M, Chiu R, Stein B, Imbra RJ, Rahmsdorf HJ, Jonat C, Herrlich P, Karin M (1987) Phorbol ester-inducible genes contain a common cis element recognized by a TPA-modulated trans-acting factor. Cell 49:729–739. https://doi.org/10.1016/0092-8674(87)90611-8
Shaywitz AJ, Greenberg ME (1999) CREB: a stimulus-induced transcription factor activated by a diverse array of extracellular signals. Annu Rev Biochem 68:821–861. https://doi.org/10.1146/annurev.biochem.68.1.821
Horos R, Buscher M, Kleinendorst R, Alleaume AM, Tarafder AK, Schwarzl T, Dziuba D, Tischer C, Zielonka EM, Adak A, Castello A, Huber W, Sachse C, Hentze MW (2019) The small non-coding vault RNA1-1 acts as a riboregulator of autophagy. Cell 176(1054–1067):e12. https://doi.org/10.1016/j.cell.2019.01.030
Li F, Chen Y, Zhang Z, Ouyang J, Wang Y, Yan R, Huang S, Gao GF, Guo G, Chen JL (2015) Robust expression of vault RNAs induced by influenza A virus plays a critical role in suppression of PKR-mediated innate immunity. Nucleic Acids Res 43:10321–10337. https://doi.org/10.1093/nar/gkv1078
Amort M, Nachbauer B, Tuzlak S, Kieser A, Schepers A, Villunger A, Polacek N (2015) Expression of the vault RNA protects cells from undergoing apoptosis. Nat Commun 6:7030. https://doi.org/10.1038/ncomms8030
Lee K, Kunkeaw N, Jeon SH, Lee I, Johnson BH, Kang GY, Bang JY, Park HS, Leelayuwat C, Lee YS (2011) Precursor miR-886, a novel noncoding RNA repressed in cancer, associates with PKR and modulates its activity. RNA 17:1076–1089. https://doi.org/10.1261/rna.2701111
Park JL, Lee YS, Song MJ, Hong SH, Ahn JH, Seo EH, Shin SP, Lee SJ, Johnson BH, Stampfer MR, Kim HP, Kim SY, Lee YS (2017) Epigenetic regulation of RNA polymerase III transcription in early breast tumorigenesis. Oncogene 36:6793–6804. https://doi.org/10.1038/onc.2017.285
Kickhoefer VA, Poderycki MJ, Chan EK, Rome LH (2002) The La RNA-binding protein interacts with the vault RNA and is a vault-associated protein. J Biol Chem 277:41282–41286. https://doi.org/10.1074/jbc.M206980200
Berger W, Steiner E, Grusch M, Elbling L, Micksche M (2009) Vaults and the major vault protein: novel roles in signal pathway regulation and immunity. Cell Mol Life Sci 66:43–61. https://doi.org/10.1007/s00018-008-8364-z
Kong LB, Siva AC, Rome LH, Stewart PL (1999) Structure of the vault, a ubiquitous cellular component. Structure 7:371–379. https://doi.org/10.1016/s0969-2126(99)80050-1
Tanaka H, Kato K, Yamashita E, Sumizawa T, Zhou Y, Yao M, Iwasaki K, Yoshimura M, Tsukihara T (2009) The structure of rat liver vault at 3.5 angstrom resolution. Science 323:384–388. https://doi.org/10.1126/science.1164975
Kickhoefer VA, Rome LH (1994) The sequence of a cDNA encoding the major vault protein from Rattus norvegicus. Gene 151:257–260. https://doi.org/10.1016/0378-1119(94)90667-x
Scheffer GL, Wijngaard PL, Flens MJ, Izquierdo MA, Slovak ML, Pinedo HM, Meijer CJ, Clevers HC, Scheper RJ (1995) The drug resistance-related protein LRP is the human major vault protein. Nat Med 1:578–582. https://doi.org/10.1038/nm0695-578
Vasu SK, Kedersha NL, Rome LH (1993) cDNA cloning and disruption of the major vault protein alpha gene (mvpA) in Dictyostelium discoideum. J Biol Chem 268:15356–15360
Vasu SK, Rome LH (1995) Dictyostelium vaults: disruption of the major proteins reveals growth and morphological defects and uncovers a new associated protein. J Biol Chem 270:16588–16594. https://doi.org/10.1074/jbc.270.28.16588
Chugani DC, Rome LH, Kedersha NL (1993) Evidence that vault ribonucleoprotein particles localize to the nuclear pore complex. J Cell Sci 106(Pt 1):23–29
Kitazono M, Sumizawa T, Takebayashi Y, Chen ZS, Furukawa T, Nagayama S, Tani A, Takao S, Aikou T, Akiyama S (1999) Multidrug resistance and the lung resistance-related protein in human colon carcinoma SW-620 cells. J Natl Cancer Inst 91:1647–1653. https://doi.org/10.1093/jnci/91.19.1647
Chung JH, Ginn-Pease ME, Eng C (2005) Phosphatase and tensin homologue deleted on chromosome 10 (PTEN) has nuclear localization signal-like sequences for nuclear import mediated by major vault protein. Cancer Res 65:4108–4116. https://doi.org/10.1158/0008-5472.CAN-05-0124
Kim E, Lee S, Mian MF, Yun SU, Song M, Yi KS, Ryu SH, Suh PG (2006) Crosstalk between Src and major vault protein in epidermal growth factor-dependent cell signalling. FEBS J 273:793–804. https://doi.org/10.1111/j.1742-4658.2006.05112.x
Shimamoto Y, Sumizawa T, Haraguchi M, Gotanda T, Jueng HC, Furukawa T, Sakata R, Akiyama S (2006) Direct activation of the human major vault protein gene by DNA-damaging agents. Oncol Rep 15:645–652
Kowalski MP, Dubouix-Bourandy A, Bajmoczi M, Golan DE, Zaidi T, Coutinho-Sledge YS, Gygi MP, Gygi SP, Wiemer EA, Pier GB (2007) Host resistance to lung infection mediated by major vault protein in epithelial cells. Science 317:130–132. https://doi.org/10.1126/science.1142311
Ryu SJ, An HJ, Oh YS, Choi HR, Ha MK, Park SC (2008) On the role of major vault protein in the resistance of senescent human diploid fibroblasts to apoptosis. Cell Death Differ 15:1673–1680. https://doi.org/10.1038/cdd.2008.96
Vollmar F, Hacker C, Zahedi RP, Sickmann A, Ewald A, Scheer U, Dabauvalle MC (2009) Assembly of nuclear pore complexes mediated by major vault protein. J Cell Sci 122:780–786. https://doi.org/10.1242/jcs.039529
Stephen AG, Raval-Fernandes S, Huynh T, Torres M, Kickhoefer VA, Rome LH (2001) Assembly of vault-like particles in insect cells expressing only the major vault protein. J Biol Chem 276:23217–23220. https://doi.org/10.1074/jbc.C100226200
Poderycki MJ, Kickhoefer VA, Kaddis CS, Raval-Fernandes S, Johansson E, Zink JI, Loo JA, Rome LH (2006) The vault exterior shell is a dynamic structure that allows incorporation of vault-associated proteins into its interior. Biochemistry 45:12184–12193. https://doi.org/10.1021/bi0610552
Kedersha NL, Heuser JE, Chugani DC, Rome LH (1991) Vaults. III. Vault ribonucleoprotein particles open into flower-like structures with octagonal symmetry. J Cell Biol 112:225–235. https://doi.org/10.1083/jcb.112.2.225
Rome L, Kedersha N, Chugani D (1991) Unlocking vaults: organelles in search of a function. Trends Cell Biol 1:47–50. https://doi.org/10.1016/0962-8924(91)90088-q
Mrazek J, Toso D, Ryazantsev S, Zhang X, Zhou ZH, Fernandez BC, Kickhoefer VA, Rome LH (2014) Polyribosomes are molecular 3D nanoprinters that orchestrate the assembly of vault particles. ACS Nano 8:11552–11559. https://doi.org/10.1021/nn504778h
Mossink MH, van Zon A, Scheper RJ, Sonneveld P, Wiemer EA (2003) Vaults: a ribonucleoprotein particle involved in drug resistance? Oncogene 22:7458–7467. https://doi.org/10.1038/sj.onc.1206947
Yu Z, Fotouhi-Ardakani N, Wu L, Maoui M, Wang S, Banville D, Shen SH (2002) PTEN associates with the vault particles in HeLa cells. J Biol Chem 277:40247–40252. https://doi.org/10.1074/jbc.M207608200
Ame JC, Spenlehauer C, de Murcia G (2004) The PARP superfamily. BioEssays 26:882–893. https://doi.org/10.1002/bies.20085
Mikyas Y, Makabi M, Raval-Fernandes S, Harrington L, Kickhoefer VA, Rome LH, Stewart PL (2004) Cryoelectron microscopy imaging of recombinant and tissue derived vaults: localization of the MVP N termini and VPARP. J Mol Biol 344:91–105. https://doi.org/10.1016/j.jmb.2004.09.021
Harrington L, McPhail T, Mar V, Zhou W, Oulton R, Bass MB, Arruda I, Robinson MO (1997) A mammalian telomerase-associated protein. Science 275:973–977. https://doi.org/10.1126/science.275.5302.973
Kong LB, Siva AC, Kickhoefer VA, Rome LH, Stewart PL (2000) RNA location and modeling of a WD40 repeat domain within the vault. RNA 6:890–900. https://doi.org/10.1017/s1355838200000157
van Zon A, Mossink MH, Scheper RJ, Sonneveld P, Wiemer EA (2003) The vault complex. Cell Mol Life Sci 60:1828–1837. https://doi.org/10.1007/s00018-003-3030-y
Slesina M, Inman EM, Moore AE, Goldhaber JI, Rome LH, Volknandt W (2006) Movement of vault particles visualized by GFP-tagged major vault protein. Cell Tissue Res 324:403–410. https://doi.org/10.1007/s00441-006-0158-8
Slesina M, Inman EM, Rome LH, Volknandt W (2005) Nuclear localization of the major vault protein in U373 cells. Cell Tissue Res 321:97–104. https://doi.org/10.1007/s00441-005-1086-8
van Zon A, Mossink MH, Houtsmuller AB, Schoester M, Scheffer GL, Scheper RJ, Sonneveld P, Wiemer EA (2006) Vault mobility depends in part on microtubules and vaults can be recruited to the nuclear envelope. Exp Cell Res 312:245–255. https://doi.org/10.1016/j.yexcr.2005.10.016
van Zon A, Mossink MH, Schoester M, Houtsmuller AB, Scheffer GL, Scheper RJ, Sonneveld P, Wiemer EA (2003) The formation of vault-tubes: a dynamic interaction between vaults and vault PARP. J Cell Sci 116:4391–4400. https://doi.org/10.1242/jcs.00749
Chung JH, Eng C (2005) Nuclear-cytoplasmic partitioning of phosphatase and tensin homologue deleted on chromosome 10 (PTEN) differentially regulates the cell cycle and apoptosis. Cancer Res 65:8096–8100. https://doi.org/10.1158/0008-5472.CAN-05-1888
Kitazono M, Okumura H, Ikeda R, Sumizawa T, Furukawa T, Nagayama S, Seto K, Aikou T, Akiyama S (2001) Reversal of LRP-associated drug resistance in colon carcinoma SW-620 cells. Int J Cancer 91:126–131. https://doi.org/10.1002/1097-0215(20010101)91:1%3c126::aid-ijc1018%3e3.0.co;2-8
Abbondanza C, Rossi V, Roscigno A, Gallo L, Belsito A, Piluso G, Medici N, Nigro V, Molinari AM, Moncharmont B, Puca GA (1998) Interaction of vault particles with estrogen receptor in the MCF-7 breast cancer cell. J Cell Biol 141:1301–1310. https://doi.org/10.1083/jcb.141.6.1301
Liu Y, Snow BE, Hande MP, Baerlocher G, Kickhoefer VA, Yeung D, Wakeham A, Itie A, Siderovski DP, Lansdorp PM, Robinson MO, Harrington L (2000) Telomerase-associated protein TEP1 is not essential for telomerase activity or telomere length maintenance in vivo. Mol Cell Biol 20:8178–8184. https://doi.org/10.1128/mcb.20.21.8178-8184.2000
Kickhoefer VA, Liu Y, Kong LB, Snow BE, Stewart PL, Harrington L, Rome LH (2001) The Telomerase/vault-associated protein TEP1 is required for vault RNA stability and its association with the vault particle. J Cell Biol 152:157–164. https://doi.org/10.1083/jcb.152.1.157
Mossink MH, van Zon A, Franzel-Luiten E, Schoester M, Kickhoefer VA, Scheffer GL, Scheper RJ, Sonneveld P, Wiemer EA (2002) Disruption of the murine major vault protein (MVP/LRP) gene does not induce hypersensitivity to cytostatics. Cancer Res 62:7298–7304
Mossink MH, de Groot J, van Zon A, Franzel-Luiten E, Schoester M, Scheffer GL, Sonneveld P, Scheper RJ, Wiemer EA (2003) Unimpaired dendritic cell functions in MVP/LRP knockout mice. Immunology 110:58–65. https://doi.org/10.1046/j.1365-2567.2003.01708.x
Schroeijers AB, Reurs AW, Scheffer GL, Stam AG, de Jong MC, Rustemeyer T, Wiemer EA, de Gruijl TD, Scheper RJ (2002) Up-regulation of drug resistance-related vaults during dendritic cell development. J Immunol 168:1572–1578. https://doi.org/10.4049/jimmunol.168.4.1572
Ben J, Jiang B, Wang D, Liu Q, Zhang Y, Qi Y, Tong X, Chen L, Liu X, Zhang Y, Zhu X, Li X, Zhang H, Bai H, Yang Q, Ma J, Wiemer EAC, Xu Y, Chen Q (2019) Major vault protein suppresses obesity and atherosclerosis through inhibiting IKK-NF-kappaB signaling mediated inflammation. Nat Commun 10:1801. https://doi.org/10.1038/s41467-019-09588-x
Peng N, Liu S, Xia Z, Ren S, Feng J, Jing M, Gao X, Wiemer EA, Zhu Y (2016) Inducible major vault protein plays a pivotal role in double-stranded RNA- or virus-induced proinflammatory response. J Immunol 196:2753–2766. https://doi.org/10.4049/jimmunol.1501481
Herlevsen M, Oxford G, Owens CR, Conaway M, Theodorescu D (2007) Depletion of major vault protein increases doxorubicin sensitivity and nuclear accumulation and disrupts its sequestration in lysosomes. Mol Cancer Ther 6:1804–1813. https://doi.org/10.1158/1535-7163.MCT-06-0372
Shen WH, Balajee AS, Wang J, Wu H, Eng C, Pandolfi PP, Yin Y (2007) Essential role for nuclear PTEN in maintaining chromosomal integrity. Cell 128:157–170. https://doi.org/10.1016/j.cell.2006.11.042
Bassi C, Ho J, Srikumar T, Dowling RJ, Gorrini C, Miller SJ, Mak TW, Neel BG, Raught B, Stambolic V (2013) Nuclear PTEN controls DNA repair and sensitivity to genotoxic stress. Science 341:395–399. https://doi.org/10.1126/science.1236188
van Zon A, Mossink MH, Schoester M, Scheffer GL, Scheper RJ, Sonneveld P, Wiemer EA (2001) Multiple human vault RNAs. Expression and association with the vault complex. J Biol Chem 276:37715–37721. https://doi.org/10.1074/jbc.M106055200
Gopinath SC, Wadhwa R, Kumar PK (2010) Expression of noncoding vault RNA in human malignant cells and its importance in mitoxantrone resistance. Mol Cancer Res 8:1536–1546. https://doi.org/10.1158/1541-7786.MCR-10-0242
Gopinath SC, Matsugami A, Katahira M, Kumar PK (2005) Human vault-associated non-coding RNAs bind to mitoxantrone, a chemotherapeutic compound. Nucleic Acids Res 33:4874–4881. https://doi.org/10.1093/nar/gki809
Chen J, OuYang H, An X, Liu S (2018) Vault RNAs partially induces drug resistance of human tumor cells MCF-7 by binding to the RNA/DNA-binding protein PSF and inducing oncogene GAGE6. PLoS ONE 13:e0191325. https://doi.org/10.1371/journal.pone.0191325
Persson H, Kvist A, Vallon-Christersson J, Medstrand P, Borg A, Rovira C (2009) The non-coding RNA of the multidrug resistance-linked vault particle encodes multiple regulatory small RNAs. Nat Cell Biol 11:1268–1271. https://doi.org/10.1038/ncb1972
Hussain S, Sajini AA, Blanco S, Dietmann S, Lombard P, Sugimoto Y, Paramor M, Gleeson JG, Odom DT, Ule J, Frye M (2013) NSun2-mediated cytosine-5 methylation of vault noncoding RNA determines its processing into regulatory small RNAs. Cell Rep 4:255–261. https://doi.org/10.1016/j.celrep.2013.06.029
Sajini AA, Choudhury NR, Wagner RE, Bornelov S, Selmi T, Spanos C, Dietmann S, Rappsilber J, Michlewski G, Frye M (2019) Loss of 5-methylcytosine alters the biogenesis of vault-derived small RNAs to coordinate epidermal differentiation. Nat Commun 10:2550. https://doi.org/10.1038/s41467-019-10020-7
Zhao X, Yang L, Hu J (2011) Down-regulation of miR-27a might inhibit proliferation and drug resistance of gastric cancer cells. J Exp Clin Cancer Res 30:55. https://doi.org/10.1186/1756-9966-30-55
Nussinov R, Tsai CJ, Jang H (2017) A new view of pathway-driven drug resistance in tumor proliferation. Trends Pharmacol Sci 38:427–437. https://doi.org/10.1016/j.tips.2017.02.001
Urban RJ, Bodenburg Y, Kurosky A, Wood TG, Gasic S (2000) Polypyrimidine tract-binding protein-associated splicing factor is a negative regulator of transcriptional activity of the porcine p450scc insulin-like growth factor response element. Mol Endocrinol 14:774–782. https://doi.org/10.1210/mend.14.6.0485
Urban RJ, Bodenburg YH, Wood TG (2002) NH2 terminus of PTB-associated splicing factor binds to the porcine P450scc IGF-I response element. Am J Physiol Endocrinol Metab 283:E423–E427. https://doi.org/10.1152/ajpendo.00057.2002
Patton JG, Porro EB, Galceran J, Tempst P, Nadal-Ginard B (1993) Cloning and characterization of PSF, a novel pre-mRNA splicing factor. Genes Dev 7:393–406. https://doi.org/10.1101/gad.7.3.393
Li L, Feng T, Lian Y, Zhang G, Garen A, Song X (2009) Role of human noncoding RNAs in the control of tumorigenesis. Proc Natl Acad Sci USA 106:12956–12961. https://doi.org/10.1073/pnas.0906005106
Garcia MA, Gil J, Ventoso I, Guerra S, Domingo E, Rivas C, Esteban M (2006) Impact of protein kinase PKR in cell biology: from antiviral to antiproliferative action. Microbiol Mol Biol Rev 70:1032–1060. https://doi.org/10.1128/MMBR.00027-06
Prasad S, Ravindran J, Aggarwal BB (2010) NF-kappaB and cancer: how intimate is this relationship. Mol Cell Biochem 336:25–37. https://doi.org/10.1007/s11010-009-0267-2
Kim SH, Gunnery S, Choe JK, Mathews MB (2002) Neoplastic progression in melanoma and colon cancer is associated with increased expression and activity of the interferon-inducible protein kinase, PKR. Oncogene 21:8741–8748. https://doi.org/10.1038/sj.onc.1205987
Terada T, Maeta H, Endo K, Ohta T (2000) Protein expression of double-stranded RNA-activated protein kinase in thyroid carcinomas: correlations with histologic types, pathologic parameters, and Ki-67 labeling. Hum Pathol 31:817–821. https://doi.org/10.1053/hupa.2000.8443
Kunkeaw N, Jeon SH, Lee K, Johnson BH, Tanasanvimon S, Javle M, Pairojkul C, Chamgramol Y, Wongfieng W, Gong B, Leelayuwat C, Lee YS (2013) Cell death/proliferation roles for nc886, a non-coding RNA, in the protein kinase R pathway in cholangiocarcinoma. Oncogene 32:3722–3731. https://doi.org/10.1038/onc.2012.382
Cao J, Song Y, Bi N, Shen J, Liu W, Fan J, Sun G, Tong T, He J, Shi Y, Zhang X, Lu N, He Y, Zhang H, Ma K, Luo X, Lv L, Deng H, Cheng J, Zhu J, Wang L, Zhan Q (2013) DNA methylation-mediated repression of miR-886–3p predicts poor outcome of human small cell lung cancer. Cancer Res 73:3326–3335. https://doi.org/10.1158/0008-5472.CAN-12-3055
Lee HS, Lee K, Jang HJ, Lee GK, Park JL, Kim SY, Kim SB, Johnson BH, Zo JI, Lee JS, Lee YS (2014) Epigenetic silencing of the non-coding RNA nc886 provokes oncogenes during human esophageal tumorigenesis. Oncotarget 5:3472–3481. https://doi.org/10.18632/oncotarget.1927
Lee KS, Park JL, Lee K, Richardson LE, Johnson BH, Lee HS, Lee JS, Kim SB, Kwon OH, Song KS, Kim YS, Ashktorab H, Smoot DT, Jeon SH, Kim SY, Lee YS (2014) nc886, a non-coding RNA of anti-proliferative role, is suppressed by CpG DNA methylation in human gastric cancer. Oncotarget 5:3944–3955. https://doi.org/10.18632/oncotarget.2047
Lee YS (2015) A novel type of non-coding RNA, nc886, implicated in tumor sensing and suppression. Genom Inform 13:26–30. https://doi.org/10.5808/GI.2015.13.2.26
Bracher L, Ferro I, Pulido-Quetglas C, Ruepp MD, Johnson R, Polacek N (2020) Human vtRNA1-1 levels modulate signaling pathways and regulate apoptosis in human cancer cells. Biomolecules. https://doi.org/10.3390/biom10040614
Acevo-Rodriguez PS, Maldonado G, Castro-Obregon S, Hernandez G (2020) Autophagy Regulation by the translation machinery and its implications in cancer. Front Oncol 10:322. https://doi.org/10.3389/fonc.2020.00322
Hanson HH, Kang S, Fernandez-Monreal M, Oung T, Yildirim M, Lee R, Suyama K, Hazan RB, Phillips GR (2010) LC3-dependent intracellular membrane tubules induced by gamma-protocadherins A3 and B2: a role for intraluminal interactions. J Biol Chem 285:20982–20992. https://doi.org/10.1074/jbc.M109.092031
Cha-Molstad H, Yu JE, Feng Z, Lee SH, Kim JG, Yang P, Han B, Sung KW, Yoo YD, Hwang J, McGuire T, Shim SM, Song HD, Ganipisetti S, Wang N, Jang JM, Lee MJ, Kim SJ, Lee KH, Hong JT, Ciechanover A, Mook-Jung I, Kim KP, Xie XQ, Kwon YT, Kim BY (2017) p62/SQSTM1/sequestosome-1 is an N-recognin of the N-end rule pathway which modulates autophagosome biogenesis. Nat Commun 8:102. https://doi.org/10.1038/s41467-017-00085-7
Demishtein A, Fraiberg M, Berko D, Tirosh B, Elazar Z, Navon A (2017) SQSTM1/p62-mediated autophagy compensates for loss of proteasome polyubiquitin recruiting capacity. Autophagy 13:1697–1708. https://doi.org/10.1080/15548627.2017.1356549
Dikic I (2017) Proteasomal and autophagic degradation systems. Annu Rev Biochem 86:193–224. https://doi.org/10.1146/annurev-biochem-061516-044908
Zhao J, Lu Y, Shen HM (2012) Targeting p53 as a therapeutic strategy in sensitizing TRAIL-induced apoptosis in cancer cells. Cancer Lett 314:8–23. https://doi.org/10.1016/j.canlet.2011.09.040
Degenhardt K, Mathew R, Beaudoin B, Bray K, Anderson D, Chen G, Mukherjee C, Shi Y, Gelinas C, Fan Y, Nelson DA, Jin S, White E (2006) Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 10:51–64. https://doi.org/10.1016/j.ccr.2006.06.001
Klionsky DJ, Abdelmohsen K, Abe A et al (2016) Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12:1–222. https://doi.org/10.1080/15548627.2015.1100356
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A, Stenmark H, Johansen T (2005) p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 171:603–614. https://doi.org/10.1083/jcb.200507002
Pankiv S, Clausen TH, Lamark T, Brech A, Bruun JA, Outzen H, Overvatn A, Bjorkoy G, Johansen T (2007) p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem 282:24131–24145. https://doi.org/10.1074/jbc.M702824200
Sahani MH, Itakura E, Mizushima N (2014) Expression of the autophagy substrate SQSTM1/p62 is restored during prolonged starvation depending on transcriptional upregulation and autophagy-derived amino acids. Autophagy 10:431–441. https://doi.org/10.4161/auto.27344
Turco E, Witt M, Abert C, Bock-Bierbaum T, Su MY, Trapannone R, Sztacho M, Danieli A, Shi X, Zaffagnini G, Gamper A, Schuschnig M, Fracchiolla D, Bernklau D, Romanov J, Hartl M, Hurley JH, Daumke O, Martens S (2019) FIP200 claw domain binding to p62 promotes autophagosome formation at ubiquitin condensates. Mol Cell 74(330–346):e11. https://doi.org/10.1016/j.molcel.2019.01.035
Johansen T, Lamark T (2011) Selective autophagy mediated by autophagic adapter proteins. Autophagy 7:279–296. https://doi.org/10.4161/auto.7.3.14487
Rogov V, Dotsch V, Johansen T, Kirkin V (2014) Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol Cell 53:167–178. https://doi.org/10.1016/j.molcel.2013.12.014
Itakura E, Mizushima N (2011) p62 Targeting to the autophagosome formation site requires self-oligomerization but not LC3 binding. J Cell Biol 192:17–27. https://doi.org/10.1083/jcb.201009067
Moscat J, Diaz-Meco MT (2009) p62 at the crossroads of autophagy, apoptosis, and cancer. Cell 137:1001–1004. https://doi.org/10.1016/j.cell.2009.05.023
Moscat J, Karin M, Diaz-Meco MT (2016) p62 in cancer: signaling adaptor beyond autophagy. Cell 167:606–609. https://doi.org/10.1016/j.cell.2016.09.030
Deretic V, Saitoh T, Akira S (2013) Autophagy in infection, inflammation and immunity. Nat Rev Immunol 13:722–737. https://doi.org/10.1038/nri3532
Skreka K, Schafferer S, Nat IR, Zywicki M, Salti A, Apostolova G, Griehl M, Rederstorff M, Dechant G, Huttenhofer A (2012) Identification of differentially expressed non-coding RNAs in embryonic stem cell neural differentiation. Nucleic Acids Res 40:6001–6015. https://doi.org/10.1093/nar/gks311
Squires JE, Patel HR, Nousch M, Sibbritt T, Humphreys DT, Parker BJ, Suter CM, Preiss T (2012) Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res 40:5023–5033. https://doi.org/10.1093/nar/gks144
Frye M, Blanco S (2016) Post-transcriptional modifications in development and stem cells. Development 143:3871–3881. https://doi.org/10.1242/dev.136556
Blanco S, Bandiera R, Popis M, Hussain S, Lombard P, Aleksic J, Sajini A, Tanna H, Cortes-Garrido R, Gkatza N, Dietmann S, Frye M (2016) Stem cell function and stress response are controlled by protein synthesis. Nature 534:335–340. https://doi.org/10.1038/nature18282
Minones-Moyano E, Friedlander MR, Pallares J, Kagerbauer B, Porta S, Escaramis G, Ferrer I, Estivill X, Marti E (2013) Upregulation of a small vault RNA (svtRNA2-1a) is an early event in Parkinson disease and induces neuronal dysfunction. RNA Biol 10:1093–1106. https://doi.org/10.4161/rna.24813
Fort RS, Garat B, Sotelo-Silveira JR, Duhagon MA (2020) vtRNA2-1/nc886 produces a small rna that contributes to its tumor suppression action through the microRNA pathway in prostate cancer. Noncoding RNA. https://doi.org/10.3390/ncrna6010007
O’Brien J, Hayder H, Zayed Y, Peng C (2018) Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (Lausanne) 9:402. https://doi.org/10.3389/fendo.2018.00402
Fort RS, Matho C, Geraldo MV, Ottati MC, Yamashita AS, Saito KC, Leite KRM, Mendez M, Maedo N, Mendez L, Garat B, Kimura ET, Sotelo-Silveira JR, Duhagon MA (2018) Nc886 is epigenetically repressed in prostate cancer and acts as a tumor suppressor through the inhibition of cell growth. BMC Cancer 18:127. https://doi.org/10.1186/s12885-018-4049-7
Somel M, Guo S, Fu N, Yan Z, Hu HY, Xu Y, Yuan Y, Ning Z, Hu Y, Menzel C, Hu H, Lachmann M, Zeng R, Chen W, Khaitovich P (2010) MicroRNA, mRNA, and protein expression link development and aging in human and macaque brain. Genome Res 20:1207–1218. https://doi.org/10.1101/gr.106849.110
van Balkom BW, Eisele AS, Pegtel DM, Bervoets S, Verhaar MC (2015) Quantitative and qualitative analysis of small RNAs in human endothelial cells and exosomes provides insights into localized RNA processing, degradation and sorting. J Extracell Vesicles 4:26760. https://doi.org/10.3402/jev.v4.26760
Motsch N, Pfuhl T, Mrazek J, Barth S, Grasser FA (2007) Epstein-Barr virus-encoded latent membrane protein 1 (LMP1) induces the expression of the cellular microRNA miR-146a. RNA Biol 4:131–137. https://doi.org/10.4161/rna.4.3.5206
Jackson WT (2015) Viruses and the autophagy pathway. Virology 479–480:450–456. https://doi.org/10.1016/j.virol.2015.03.042
Weitzman MD, Fradet-Turcotte A (2018) Virus DNA replication and the host DNA damage response. Annu Rev Virol 5:141–164. https://doi.org/10.1146/annurev-virology-092917-043534
Oh JE, Lee HK (2013) Autophagy as an innate immune modulator. Immune Netw 13:1–9. https://doi.org/10.4110/in.2013.13.1.1
Eisenreich W, Rudel T, Heesemann J, Goebel W (2019) How viral and intracellular bacterial pathogens reprogram the metabolism of host cells to allow their intracellular replication. Front Cell Infect Microbiol 9:42. https://doi.org/10.3389/fcimb.2019.00042
Burke JM, Kincaid RP, Nottingham RM, Lambowitz AM, Sullivan CS (2016) DUSP11 activity on triphosphorylated transcripts promotes Argonaute association with noncanonical viral microRNAs and regulates steady-state levels of cellular noncoding RNAs. Genes Dev 30:2076–2092. https://doi.org/10.1101/gad.282616.116
Burke JM, Sullivan CS (2017) DUSP11—an RNA phosphatase that regulates host and viral non-coding RNAs in mammalian cells. RNA Biol 14:1457–1465. https://doi.org/10.1080/15476286.2017.1306169
Zhao Y, Ye X, Dunker W, Song Y, Karijolich J (2018) RIG-I like receptor sensing of host RNAs facilitates the cell-intrinsic immune response to KSHV infection. Nat Commun 9:4841. https://doi.org/10.1038/s41467-018-07314-7
Ablasser A, Bauernfeind F, Hartmann G, Latz E, Fitzgerald KA, Hornung V (2009) RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III-transcribed RNA intermediate. Nat Immunol 10:1065–1072. https://doi.org/10.1038/ni.1779
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E, Hannon G, Abeliovich A (2007) A MicroRNA feedback circuit in midbrain dopamine neurons. Science 317:1220–1224. https://doi.org/10.1126/science.1140481
Junn E, Lee KW, Jeong BS, Chan TW, Im JY, Mouradian MM (2009) Repression of alpha-synuclein expression and toxicity by microRNA-7. Proc Natl Acad Sci USA 106:13052–13057. https://doi.org/10.1073/pnas.0906277106
Minones-Moyano E, Porta S, Escaramis G, Rabionet R, Iraola S, Kagerbauer B, Espinosa-Parrilla Y, Ferrer I, Estivill X, Marti E (2011) MicroRNA profiling of Parkinson’s disease brains identifies early downregulation of miR-34b/c which modulate mitochondrial function. Hum Mol Genet 20:3067–3078. https://doi.org/10.1093/hmg/ddr210
Filipits M, Pohl G, Stranzl T, Suchomel RW, Scheper RJ, Jager U, Geissler K, Lechner K, Pirker R (1998) Expression of the lung resistance protein predicts poor outcome in de novo acute myeloid leukemia. Blood 91:1508–1513
Filipits M, Stranzl T, Pohl G, Heinzl H, Jager U, Geissler K, Fonatsch C, Haas OA, Lechner K, Pirker R (2000) Drug resistance factors in acute myeloid leukemia: a comparative analysis. Leukemia 14:68–76. https://doi.org/10.1038/sj.leu.2401634
Pirker R, Pohl G, Stranzl T, Suchomel RW, Scheper RJ, Jager U, Geissler K, Lechner K, Filipits M (1999) The lung resistance protein (LRP) predicts poor outcome in acute myeloid leukemia. Adv Exp Med Biol 457:133–139. https://doi.org/10.1007/978-1-4615-4811-9_15
Xu D, Arestrom I, Virtala R, Pisa P, Peterson C, Gruber A (1999) High levels of lung resistance related protein mRNA in leukaemic cells from patients with acute myelogenous leukaemia are associated with inferior response to chemotherapy and prior treatment with mitoxantrone. Br J Haematol 106:627–633. https://doi.org/10.1046/j.1365-2141.1999.01611.x
Oh EJ, Kahng J, Kim Y, Kim M, Lim J, Kang CS, Min WS, Cho B, Lee A, Lee KY, Kim WI, Shim SI, Han K (2003) Expression of functional markers in acute lymphoblastic leukemia. Leuk Res 27:903–908. https://doi.org/10.1016/s0145-2126(03)00026-2
Valera ET, Scrideli CA, Queiroz RG, Mori BM, Tone LG (2004) Multiple drug resistance protein (MDR-1), multidrug resistance-related protein (MRP) and lung resistance protein (LRP) gene expression in childhood acute lymphoblastic leukemia. Sao Paulo Med J 122:166–171. https://doi.org/10.1590/s1516-31802004000400007
Volm M, Mattern J, Koomagi R (1997) Expression of lung resistance-related protein (LRP) in non-small cell lung carcinomas of smokers and non-smokers and its predictive value for doxorubicin resistance. Anticancer Drugs 8:931–936. https://doi.org/10.1097/00001813-199711000-00003
Ohno N, Tani A, Uozumi K, Hanada S, Furukawa T, Akiba S, Sumizawa T, Utsunomiya A, Arima T, Akiyama S (2001) Expression of functional lung resistance-related protein predicts poor outcome in adult T-cell leukemia. Blood 98:1160–1165. https://doi.org/10.1182/blood.v98.4.1160
Sakaki Y, Terashi K, Yamaguchi A, Kawamata N, Tokito Y, Mori H, Umehara M, Yoshiyama T, Ohtsubo H, Arimura K, Arima N, Tei C (2002) Human T-cell lymphotropic virus type I Tax activates lung resistance-related protein expression in leukemic clones established from an adult T-cell leukemia patient. Exp Hematol 30:340–345. https://doi.org/10.1016/s0301-472x(02)00775-0
Filipits M, Drach J, Pohl G, Schuster J, Stranzl T, Ackermann J, Konigsberg R, Kaufmann H, Gisslinger H, Huber H, Ludwig H, Pirker R (1999) Expression of the lung resistance protein predicts poor outcome in patients with multiple myeloma. Clin Cancer Res 5:2426–2430
Rimsza LM, Campbell K, Dalton WS, Salmon S, Willcox G, Grogan TM (1999) The major vault protein (MVP), a new multidrug resistance associated protein, is frequently expressed in multiple myeloma. Leuk Lymphoma 34:315–324. https://doi.org/10.3109/10428199909050956
Schwarzenbach H (2002) Expression of MDR1/P-glycoprotein, the multidrug resistance protein MRP, and the lung-resistance protein LRP in multiple myeloma. Med Oncol 19:87–104. https://doi.org/10.1385/MO:19:2:87
Diestra JE, Condom E, Del Muro XG, Scheffer GL, Perez J, Zurita AJ, Munoz-Segui J, Vigues F, Scheper RJ, Capella G, Germa-Lluch JR, Izquierdo MA (2003) Expression of multidrug resistance proteins P-glycoprotein, multidrug resistance protein 1, breast cancer resistance protein and lung resistance related protein in locally advanced bladder cancer treated with neoadjuvant chemotherapy: biological and clinical implications. J Urol 170:1383–1387. https://doi.org/10.1097/01.ju.0000074710.96154.c9
Schadendorf D, Makki A, Stahr C, van Dyck A, Wanner R, Scheffer GL, Flens MJ, Scheper R, Henz BM (1995) Membrane transport proteins associated with drug resistance expressed in human melanoma. Am J Pathol 147:1545–1552
Mandoky L, Geczi L, Doleschall Z, Bodrogi I, Csuka O, Kasler M, Bak M (2004) Expression and prognostic value of the lung resistance-related protein (LRP) in germ cell testicular tumors. Anticancer Res 24:1097–1104
Tews DS, Nissen A, Kulgen C, Gaumann AK (2000) Drug resistance-associated factors in primary and secondary glioblastomas and their precursor tumors. J Neurooncol 50:227–237. https://doi.org/10.1023/a:1006491405010
Funding
This research received no external funding.
Author information
Authors and Affiliations
Contributions
Conceptualization, JCH, AL and NV; writing, original draft preparation, review and editing, JCH, AL and NV. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
N.V. received speaker honorarium from the companies Bayer, Eli-Lilly, Pfizer and Merck. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Code availability
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hahne, J.C., Lampis, A. & Valeri, N. Vault RNAs: hidden gems in RNA and protein regulation. Cell. Mol. Life Sci. 78, 1487–1499 (2021). https://doi.org/10.1007/s00018-020-03675-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03675-9