Abstract
Circular RNAs (circRNAs) are single-stranded and covalently closed back-splicing products of pre-mRNAs. They can be derived from exons, introns, or exons with intron retained between exons of transcripts, as well as antisense transcripts. CircRNAs have been reported to function as microRNA sponges, regulate gene transcription mediated by RNA polymerase II, and modulate the splicing or stability of mRNA. However, emerging studies demonstrate that they affect the behavior of proteins via direct interactions with them. Here, we summarize that by binding directly with proteins; circRNAs can facilitate their nuclear or cytoplasmic localizations, regulate their functions or stability, promote or inhibit the interactions between them, or influence the interactions between them and DNA. Furthermore, these circRNA-binding proteins contain transcription factors, RNA processing proteins, proteases, and some other RNA-binding proteins. As a consequence, circRNAs are involved in the regulation of multiple physiological or pathological processes, including tumorigenesis, atherosclerosis, wound repair, cardiac senescence, myocardial ischemia/reperfusion injury, and so forth. Nonetheless, it is worthwhile to further explore more types of proteins that interact with circRNAs, which would be helpful in revealing other unknown biological functions of circRNAs that guide the variation in behavior of cellular proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
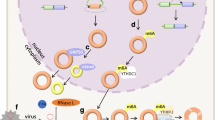
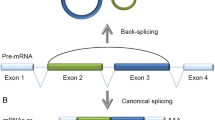
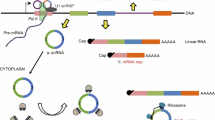
Circular RNAs (circRNAs), which are products generated from back-splicing of pre-mRNA transcripts, are single-stranded and covalently closed where the 3′ and 5′ RNA ends are joined together. Although circRNAs were discovered more than two decades ago, they were formerly regarded as byproducts of aberrant splicing, and only a few circRNAs have been found and characterized [1,2,3,4]. Benefiting from the advancement of high-throughput sequencing technology and computational algorithms, circular RNAs were recently identified as a large class of RNAs in animals [5, 6]. CircRNAs can be circularized natural antisense transcripts (e.g., antisense to the cerebellar degeneration-related protein 1 transcript (CDR1as)) [7] or products from back-spliced exons [e.g., sex-determining region Y (Sry)] that have been defined as exonic circRNAs [4, 8]. However, a class of circular intronic RNAs that are derived from intronic sequences were identified and characterized in human cells and termed as ciRNAs [9]. Furthermore, another kind of circRNAs, which are composed of circularized exons with intron retained between exons, were named Exon–Intron circular RNAs (EIciRNAs) [10]. Recently, a class of fusion circRNAs were found to be generated from transcribed exons of distinct genes during aberrant chromosomal translocations in cancer cells [11]. Possibly due to back-splicing being far less efficient than canonical splicing in cells, circRNAs are prevalent on average only about 1% of their linear cognate RNAs [12, 13]. CircRNAs have been reported to play key roles in various physiological or pathological processes, such as myogenesis [14], brain function [15], epithelial–mesenchymal transition [16], ischemic myocardial injury [17], vascular diseases [18], tumorigenesis and tumor progression [19,20,21,22], and so forth.
Although many circRNAs are considered to be non-coding RNAs [23,24,25], emerging studies have demonstrated that a number of endogenous circRNAs possess translation potential [14, 26,27,28,29,30]. A well-defined function of circRNAs is binding with microRNAs (miRNAs) and sequestering them away from their target mRNAs as competitive endogenous RNAs. In 2013, a human cytoplasmic circRNA CDR1as was found to harbor 63 conserved binding sites for miR-7, preventing their function as a sponge, and leading to expression release of miR-7-inhibited mRNA [5]. Additionally, another circRNA, Sry, serves as a sponge of miR-138 [6]. Lately, an increasing number of studies confirmed that circRNAs act as binding platforms for the miRNA–AGO complex and function as a class of post-transcriptional regulators [31,32,33]. However, circRNAs have also been reported to be involved in the transcription regulation of their parent coding genes, a process that is mediated by RNA polymerase II. Ci-ankrd52, as an example of ciRNA, largely accumulates to the transcription sites of its parental gene ankrd52, associates with elongation Pol II machinery, and acts as a positive regulator of Pol II-mediated ankrd52 transcription [9]. Besides this, EIciRNAs also promote Pol II-mediated parental gene transcription by interacting with U1 snRNA [10], a small nuclear RNA that promotes early transcriptional events by enhancing preinitiation complex assembly [34, 35]. Furthermore, circRNA can regulate the splicing of its cognate mRNA [36] or bind with mRNA and enhance its stability [37].
Except for the association with RNAs, increasing numbers of studies have shown that circRNAs function via direct interaction with cellular proteins. Although the biological implications and functions of circRNA–protein interactions have been reviewed earlier elsewhere [38], we focus here on the behavior of cellular proteins affected by direct binding of circRNAs, including subcellular localization of protein, protein–protein interaction, protein function or stability, and protein–DNA interaction. In addition, the working models of how circRNAs regulate the behavior and function of proteins, as well as the prediction and validation methods of circRNA–protein interactions, are also introduced in this article. Moreover, we categorize the proteins directly bound by circRNAs, together with the corresponding biological or physiological processes they are involved in.
CircRNAs regulate the subcellular localization of proteins
CircRNAs promote the nuclear localization of proteins
Emerging studies have demonstrated that through direct interaction, circRNAs can modulate the nuclear localization of proteins as well as their corresponding biological functions [39,40,41,42] (Fig. 1a). Circ-Amotl1, a circular RNA originating from exon 3 of the angiomotin like 1 (AMOTL1) gene, is expressed at high levels in breast carcinoma and breast cancer cell lines [39]. It not only enhances cell proliferation and colony formation, but also inhibits cell apoptosis. Moreover, tumor growth assay by nude mouse xenograft models showed that circ-Amotl1 promotes tumorigenesis. Mechanistically, circ-Amotl1 interacts with c-Myc protein, a well-known oncogenic transcription factor [43], and facilitates its nuclear translocalization. Consequently, expression of c-Myc-regulated target genes is promoted, resulting in enhanced cancer growth.
CircRNAs regulate the subcellular localization of proteins. a Circ-Amotl1 can promote the nuclear localization of c-Myc, STAT3, PDK1, and AKT1 by direct interaction with them. Through its different regions, circ-Dnmt1 can bind to p53 and Auf1, and facilitate their nuclear localization. b Circ-Foxo3 is predominantly localized in the cytoplasm and interacts with ID1, E2F1, HIF1α, and FAK, leading to their cytoplasm retention. Through the interaction with HuR protein, circAGO2 facilitates its translocation from the nucleus to the cytoplasm. Please see more detail in the text. STAT3 signal transducer and activator of transcription 3, PDK1 phosphoinositide-dependent kinase 1, AKT1 AKT serine/threonine kinase 1, ID1 inhibitor of DNA binding 1, E2F1 E2F transcription factor 1, HIF1α hypoxia inducible factor 1 subunit alpha, FAK Focal adhesion kinase, HuR human antigen R
By RNA immunoprecipitation and pull-down assays, it has been demonstrated that specific physical association also exists between circ-Amotl1 and transcription factor STAT3 [40]. As a consequence, circ-Amotl1 increases the nuclear translocation of STAT3. Subsequently, STAT3 binds to the Dnmt3a promoter and activates its transcription. As a DNA methyltransferase, Dnmt3a can methylate the promoter of miR-17 and suppress the expression of miR-17-5p. Furthermore, the inhibitory effect of miR-17-5p on its target genes Stat3, Dnmt3, and extracellular matrix gene fibronectin are eliminated. By promoting the expression of these genes and enhancing cell proliferation, survival, adhesion, and migration, circ-Amotl1 accelerates wound repair, which was demonstrated in a mouse wound healing model [40].
Additionally, circ-Amotl1 is highly expressed in neonatal human cardiac tissue, and its ectopic expression decreased doxorubicin-induced cardiomyopathy and cardiomyocytes death in mice [41]. AKT1, one kind of serine/threonine kinase, has been reported to be involved in the transduction of the anti-apoptotic signaling cascade and inhibit cell death in cardiomyocytes [44]. Nonetheless, phosphoinositide-dependent kinase 1 (PDK1) is an upstream kinase that can phosphorylate AKT1 in cardiomyocytes [45]. Circ-Amotl1 can physically interact with both PDK1 and AKT1, facilitate their nuclear translocation, and promote phosphorylation of AKT1. Subsequently, more phosphorylated AKT1 are localized in the nucleus, where they directly phosphorylate downstream intracellular proteins that facilitate cell proliferation and survival [41]. Therefore, circ-Amotl1 may reinforce the cardio-protective role mediated by AKT1.
Another circRNA circ-Dnmt1, which is derived from back-splicing between exon 7 and exon 6 of Dnmt1, is highly expressed in breast cancer cell lines and breast carcinoma tissues [42]. Circ-Dnmt1 can bind to both p53 and Auf1, although through different regions, and promote their nuclear translocation. Consequently, on one hand, nuclear p53 facilitates autophagy [46, 47]. On the other hand, Auf1 nuclear translocation results in a decrease of its cytosol accumulation, which causes the release of its suppression to Dnmt1 mRNA’s stability in the cytoplasm [48]. Subsequently, upregulated Dnmt1 inhibits p53 transcription in the nucleus by binding to its regulatory region [49]. Therefore, circ-Dnmt1 plays a central role in increasing cell autophagy and enhancing cell proliferation, survival, and tumor growth, possibly by regulating the localization and expression of tumor suppressor p53.
CircRNAs promote the cytoplasmic localization of proteins
In addition to nuclear localization, cytoplasm localization of proteins is also regulated by circRNAs [50, 51] (Fig. 1b). Circ-Foxo3, a highly expressed circular RNA in the heart tissues of elderly patients and mice, is derived from exon 2 of Foxo3 [50]. Integrated in vitro and in vivo assays have revealed the role of circ-Foxo3 in senescence. Ectopic expression of circ-Foxo3 not only promoted the cellular senescence of mouse embryonic fibroblasts, but also exacerbated doxorubicin-induced cardiomyopathy in mice. Besides, circ-Foxo3 is predominantly localized in the cytoplasm, where it interacts with the anti-senescence proteins ID1 and E2F1 and anti-stress proteins FAK and HIF1α, leading to cytoplasm retention of these proteins. Consequently, their function as transcription factors (ID1, E2F1, and HIF1α) in the nucleus and anti-stress protein (FAK) in the mitochondria was repressed, which facilitates cellular senescence. Therefore, circ-Foxo3 enhances cardiac senescence by regulating the subcellular localization of these protein factors.
Another circRNA, circAGO2 derived from the partial first intron of AGO2, enhances the cytoplasm localization of HuR protein [51]. CircAGO2 is upregulated in several cancer types, such as gastric cancer, colorectal cancer, prostate cancer, and neuroblastoma, and exhibits an oncogenic role in tumorigenesis in vitro and in vivo. As an RNA-binding protein, HuR can stabilize mRNAs by specifically binding to the AU-rich elements located in their 3′ untranslated regions (UTRs), a major target region in miRNA-mediated post-transcriptional regulation [52]. CircAGO2 can interact with HuR and facilitate its translocation from the nucleus to the cytoplasm, thereby increasing its binding to the 3′-UTR regions of target mRNAs. As a consequence, the binding of AGO2 protein to the mRNAs is restrained and the gene silencing effect mediated by AGO2–miRNA complexes is suppressed. Therefore, through HuR-mediated inhibition of the function of AGO2–miRNA complexes and release of the expression of their downstream target genes, circAGO2 promotes cancer progression [51].
CircRNAs regulate the function or stability of proteins
Besides subcellular localization, the function of proteins can also be regulated by circRNAs via their direct interaction. An autophagy-related circular RNA (ACR), also named mmu_circRNA_006636, was implicated in the regulation of cardiomyocyte autophagy and myocardial infarction [53]. It suppressed autophagy and cell death not only in cultured cardiomyocytes but also in the hearts of mice upon ischemia/reperfusion injury. Mechanistic studies demonstrated that ACR physically binds to DNA methyltransferase Dnmt3B and represses Dnmt3B-mediated methylation of the Pink1 promoter, leading to increased Pink1 expression (Fig. 2a). Furthermore, Pink1 phosphorylates FAM65B at serine 46, which then restrains autophagy and cell death in cardiomyocytes. Therefore, ACR exerts its resistance effect on autophagy via the Pink1/FAM65B axis and reduces myocardial ischemia/reperfusion injury [53].
CircRNAs regulate the function or stability of proteins. a ACR physically binds to Dnmt3B and suppresses its function as a DNA methyltransferase. b CircMbl attenuates the cellular function of MBNL1 as a sponge of MBNL1 proteins. c CircPABPN1 functions as a sponge of HuR proteins and sequesters them away from the PABPN1 mRNA, which results in attenuated translation of PABPN1. d CircNOL10 interacts with SCML1 and increases its stability by inhibiting its ubiquitination. Please see more detail in the text. “perpendicular sign” indicates an inhibition effect. ACR autophagy-related circular RNA, MBNL1 Muscleblind like splicing regulator 1, SCML1 Scm polycomb group protein like 1
Muscleblind like splicing regulator 1 (MBNL1) is another protein whose function is affected by direct binding of circRNA [54]. MBNL1 not only modulates the alternative splicing of pre-mRNAs as a splicing factor, but also regulates circRNA biogenesis [54]. It strongly and specifically binds to the highly conserved binding sites in the flanking introns of the second exon of MBNL1 in flies and humans. As a consequence, it promotes circularization of the exon and production of circular RNA circMbl. Nevertheless, circRNA biogenesis and pre-mRNA splicing compete with each other, probably because both circularization of exons and linear splicing depend on canonical splicing machinery [54, 55]. Additionally, a direct and strong interaction exists between circMbl and MBNL1, which serves to explain a possible feedback regulation of MBNL1 by circMbl. Once cellular MBNL1 protein is excessive, it enhances the biogenesis of circMbl, which results in the reduction of MBNL1 mRNA for the competition. Moreover, circMbl attenuates the cellular function of MBNL1 by binding and sponging out the excess MBNL1 protein (Fig. 2b).
In addition to cytoplasm localization [51], one recent study reported that circular RNA also influences the function of HuR directly [56], which has been demonstrated to regulate the stability and translation of target mRNAs [57, 58]. Through RNA immunoprecipitation (RIP) followed by microarray analysis, a mass of HuR-binding circular RNAs were identified in HeLa cells [56]. Hsa_circ_0031288, which is also named circPABPN1 because it originates from the exon 6 of PABPN1, is one of the most enriched circRNAs by HuR. RIP and polysome distribution analysis revealed that HuR can bind to the PABPN1 mRNA and promote its translation, respectively. However, circPABPN1, maybe functioning as a sponge of HuR (Fig. 2c), sequesters HuR away from the PABPN1 mRNA, therefore leading to attenuation of PABPN1 translation and protein expression [56].
Furthermore, the stability of proteins can also be modulated by circRNA. CircNOL10, a circRNA formed via circularization of exons 6–12 of NOL10 pre-mRNA, has low expression in lung cancer tissues and inhibits lung cancer progression in vitro and in vivo [59]. Mass spectrometer detection after an RNA pull-down assay of circNOL10-binding proteins revealed that circNOL10 interacts with the transcription factor Scm polycomb group protein like 1 (SCML1) (Fig. 2d). Moreover, circNOL10 enhances the expression of SCML1 in lung cancer cells by inhibiting its ubiquitination. Subsequently, several members of the HN polypeptide family, which are regulated by SCML1 at the transcriptional level, affect mitochondrial function and multiple tumor signaling pathways, eventually contributing to decreased cell cycle progression and cell proliferation, and increased apoptosis. Consequently, circNOL10 restrains lung cancer development via the SCML1/HN polypeptide family axis.
CircRNAs affect the interaction between proteins
CircRNAs promote the interaction between proteins
Recently, it was discovered that the interaction between proteins can be reinforced by circRNAs [60, 61]. One study demonstrated that circ-Foxo3 is conserved in human and mouse and suppresses cell cycle progression and cell proliferation by strengthening the interaction between p21 and CDK2 [60]. p21, a known cyclin-dependent kinase inhibitor, can interact with CDK2 and restrain the formation of the CDK2–cyclin E complex, therefore blocking the transition of cells from the G1 to the S phase [62]. However, circ-Foxo3 can bind to both p21 and CDK2 with high affinity, reinforce the interaction between them (Fig. 3a), and enhance the inhibitory effect of p21 on CDK2 activity. As a downstream consequence, the formation of CDK2–cyclin A complexes are also reduced, resulting in failure of the cells to pass through the S phase. Therefore, circ-Foxo3 controls cell cycle entry of cells by forming ternary complexes with p21 and CDK2 [60].
CircRNAs affect the interaction between proteins. a Circ-Foxo3 can interact with both p21 and CDK2, strengthen their interaction, and heighten the inhibitory effect of p21 on CDK2 activity. b Circ-Foxo3 can bind to both MDM2 and p53, reinforce their interaction, and facilitate MDM2-mediated poly-ubiquitination of p53. c CircANRIL competitively binds to PES1 and prevents the assembly of the PeBoW protein complex that is consistsed of PES1, BOP1, and WDR12. Please see more detail in the text. “down arrow” indicates a promotion effect, but “perpendicular sign” indicates an inhibition effect. CDK2 cyclin dependent kinase 2, MDM2 MDM2 proto-oncogene, PES1 Pescadillo ribosomal biogenesis factor 1, BOP1 BOP1 ribosomal biogenesis factor, WDR12 WD repeat domain 12
Another study revealed that circ-Foxo3 is down-regulated in specimens of patients with breast carcinoma, and its ectopic expression in breast cancer cells inhibited xenograft tumor growth and prolonged mouse lifespan [61]. MDM2, a nuclear-localized E3 ubiquitin ligase, was identified as a protein that interacts with circ-Foxo3 and regulates its intracellular effect. MDM2 has been reported to promote the protein degradation of p53 [63] and Foxo3 [64] via the ubiquitin–proteasome pathway. Circ-Foxo3 can interact with both MDM2 and p53, leading to enhanced interaction between them [61] (Fig. 3b). Subsequently, MDM2 mediates the poly-ubiquitination of p53 and promotes its degradation. However, circ-Foxo3 has only a low binding affinity to Foxo3, another target protein of MDM2, therefore failing to promote the binding of MDM2 to it. As a consequence of the circ-Foxo3–MDM2–p53 interaction, Foxo3 is relieved from the poly-ubiquitination and degradation modulated by MDM2. Upregulated Foxo3 then activates the expression of its downstream target PUMA, which contributes to increased cell apoptosis [65, 66].
CircRNAs suppress the interaction between proteins
However, one recent study reported that circRNA suppresses the interaction of proteins by competitive binding [67]. CircANRIL is derived from an ANRIL transcript on chromosome 9p21 and contains exon 5, 6, and 7, where exon 7 is covalently joined to exon 5. It was found to bind Pescadillo ribosomal biogenesis factor 1 (PES1) and affect ribosome biogenesis [67]. A causal correlation of circANRIL expression with 9p21 haplotypes in peripheral blood mononuclear cells from coronary artery disease patients indicates an atheroprotection role of circANRIL. PES1, which constitutes the nucleolar PeBoW complex with Bop1 and WDR12 [68], was identified as a protein with the strongest binding to circANRIL. As a result of circANRIL’s competitive binding to the C-terminal domain of PES1 (Fig. 3c), the assembly and function of the PeBoW protein complex in pre-rRNA processing and the maturation of the 60S ribosomal subunit are arrested. Subsequently, nucleolar stress and p53 accumulation are induced, leading to increased apoptosis and decreased proliferation that restrain the development of atherosclerosis [69, 70]. Therefore, by controlling the pathway associated with atherogenesis, circANRIL contributes to atheroprotection [67].
CircRNAs influence the interaction between protein and DNA
In addition to the interaction between proteins, interaction between protein and DNA was also found to be promoted [71] or inhibited [72] by circRNA. FLI1 exonic circular RNA (FECR1) is a circRNA generated by the circularization of exons 4, 3, and 2 of FLI1, which functions potentially as an oncogenic driver in multiple types of cancer [71]. In accordance with the high abundance of FLI1 in advanced and metastatic breast cancer tissues, FECR1 is also expressed in breast cancer tissues and promotes the invasion of breast cancer cells. The investigations of molecular mechanisms revealed that on one hand, FECR1 binds to the promoter of DNMT1, a kind of methyltransferase responsible for maintaining the methylation of DNA, and downregulates its transcription. On the other hand, FECR1 interacts with the GC-rich region of the FLI1 promoter and further recruits demethylase TET1 to it, resulting in the demethylation of DNA [73] (Fig. 4a). Together, FECR1 serves to induce widespread DNA demethylation of the FLI1 promoter and epigenetically activates FLI1’s expression. In turn, the oncoprotein FLI1 contributes to metastasis in breast cancer [71].
CircRNAs influence the interaction between protein and DNA. a FECR1 binds to the FLI1 promoter and further recruits the demethylase TET1 to it, which mediates the DNA demethylation and activates the transcription of FLI1. b Cia-cGAS competitively binds to cGAS and restrains the interaction between genomic DNA and cGAS that is pivotal to the enzymatic activity of cGAS. Please see more detail in the text. “down arrow” indicates a promotion effect, but “perpendicular sign” indicates an inhibition effect. FECR1 FLI1 exonic circular RNA, TET1 tet methylcytosine dioxygenase 1, cGAS cyclic GMP-AMP synthase, cia-cGAS an circular RNA antagonist for cGAS
Nevertheless, another study reported that circRNA suppresses the interaction between protein and DNA through competitive binding [72]. Cyclic GMP–AMP synthase (cGAS) is a cytosolic DNA sensor, whose binding with DNA allows it to catalyze the synthesis of a second messenger cGAMP and stimulate downstream STING signaling [74]. The cGAMP/STING axis then induces the production of type I interferons (IFNs), which mediate the exhaustion of quiescent hematopoietic stem cells (HSCs) [75, 76]. A circular RNA antagonist for cGAS named cia-cGAS is composed of exons 4, 5, and 6 of mouse D430042O09Rik gene transcripts, with exon 6 linked to exon 4 [72]. Cia-cGAS is highly expressed in the nucleus of long-term hematopoietic stem cells (LT-HSCs) and contributes to the maintenance of quiescent HSCs. Its deficiency in mice results in the increase of type I IFNs in LT-HSCs and significant reduction of dormant LT-HSCs in bone marrow. The functional mechanism studies revealed that cia-cGAS harbors a stronger affinity to cGAS than that of its self-genomic DNA (Fig. 4b). Through the competitive binding to cGAS with genomic DNA, cia-cGAS blocks the enzymatic activity of cGAS in the nucleus of LT-HSCs and restrains cGAS-catalyzed production of type I IFNs. As a consequence, cia-cGAS defends dormant HSCs from cGAS/Type I IFNs-modulated exhaustion [72].
Prediction, screening, and validation of proteins directly bound by circRNAs
Physical association between protein and circular RNA can be predicted on the basis of bioinformatics analysis, to which Burton Yang’s group has devoted a lot of work [39, 41, 42, 61]. The first step in their prediction was to determine the secondary structure of circRNA with lowest free energy using biological software such as Mfold. The best putative secondary structure defined with dot-bracket notation was then analyzed for tertiary structure prediction by software such as RNAComposer. Subsequently, NPDock was utilized to perform in silico molecular docking analysis of circRNA and protein, which is based on the tertiary structure of the circRNA and the 3D structure of the protein. Additionally, the secondary structure of circRNAs can also be predicted using the RNAstructure web server [59, 77] or the Vienna RNA package [67, 72, 78], and the prediction of protein–circRNA binding has also been performed with the catRAPID [59, 67, 79] or the RNA–Protein Interaction Prediction (RPISeq) software [67, 80].
The experimental method of cross-linking and immunoprecipitation followed by RNA sequencing (CLIP-Seq) is commonly used to identify cellular circRNAs that bind to one known protein [56]. However, researchers also searched for potential binding proteins of one known circular RNA by means of a proteomic screen. Holdt et al. constructed an expression vector of circANRIL with BoxB RNA hairpin sequences inserted into the coding sequence of exon 6, without impairing the circularization of exon 5 and 7 [67, 81]. Due to the high affinity of bacteriophage l transcriptional anti-terminator protein N (λN-peptide) to the BoxB RNA hairpin, λN-peptide coupled to d-desthiobiotin can mediate the pull-down of circANRIL–protein complexes in cellular lysates by streptavidin beads [82]. The captured proteins bound to circANRIL were then identified by mass spectrometric analyses [67]. Additionally, to identify the proteins interacting with one specific circRNA, Chen et al. performed an RNA pull-down assay by incubating the biotin-labeled circRNA probe with cell lysates and enriching the circRNA–protein complexes with streptavidin beads, and then detected the proteins by mass spectrometry assays [51].
One regular method to validate the interaction between circRNA and protein is the RNA immunoprecipitation (RIP) assay, in which antibodies against protein are utilized to immunoprecipitate the protein factor associated with the circRNA, followed by quantitative PCR analysis for the circRNA [39,40,41,42, 50, 53, 54, 56, 60, 61, 67, 71, 72]. Another regular method is the RNA pull-down assay, in which a complementary and biotinylated probe to circRNA is used, and the circRNA in cell lysates is pulled down by streptavidin beads; the protein bound to the circRNA is then co-precipitated and detected by western blot assay [39,40,41,42, 50, 51, 53, 56, 59,60,61, 67]. Expression of circRNA within cells or tissues can also be detected by fluorescent in situ hybridization (FISH) with a mixture of dye-labeled DNA oligo probes paired with circRNA [39, 50, 51, 59, 61, 72]. Moreover, a combination of FISH to circRNA and immunofluorescence staining with an antibody against protein makes it feasible to observe the colocalization of circRNA and protein with confocal laser scanning microscopy [39, 50, 51, 61, 72]. Additionally, the blocking oligo, which is complementary to the binding sites of protein on circRNA and aims to impair protein–circRNA binding, can be delivered into cells for checking the downstream variation of cellular function, further confirming the protein–circRNA interaction [40,41,42]. Besides, the electrophoretic mobility shift assay (EMSA) after the incubation of circRNA and protein was considered as another technique to prove the direct circRNA–protein interaction [51, 59, 72].
Conclusions and perspectives
As a large class of long non-coding RNAs in mammalians, circRNAs are known for their function as miRNA sponges. However, it is worth noting that unlike CDR1as and Sry, which have 67 and 16 putative binding sites for miR-7 [5] and miR-138 [6], respectively, a single predicted miRNA binding site within a circRNA sequence is not adequate to justify its sponge effect on the miRNA. Furthermore, the relative expression levels of circRNA and miRNA in the cells should be taken into account when the sponge function of the circRNA is claimed. Moreover, experimental verification, such as luciferase reporter assays for the test of circRNA’s suppression to miRNA’s knockdown potential, as well as demonstration of direct binding between circRNA and miRNA [6], would be essential for the evaluation of circRNA’s sponge efficiency to miRNA. Additionally, circRNAs have been revealed to regulate the stability of mRNA via direct interaction with it [37], or to affect transcription by binding with DNA promoters of genes [10].
However, in this article, we have comprehensively summarized four working models on how circRNAs modulate the action of proteins by direct interaction. One is to promote the nuclear localization [39,40,41,42] or cytoplasmic localization of proteins [50, 51] (Fig. 1); the second is to regulate the function [53, 54, 56] or stability [59] of proteins (Fig. 2); the third is to facilitate [60, 61] or restrain [67] the interaction between proteins (Fig. 3); and the fourth is to affect the interaction between protein and DNA [71, 72] (Fig. 4). The circRNAs and the proteins they interact with, together with the biological processes they are implicated in, are summarized in Table 1. The proteins bound by circRNAs can be classified as transcription factors [39, 40, 42, 50, 59], RNA processing proteins [54, 67], proteases [41, 53, 60, 61, 71, 72], and RNA-binding proteins [51, 56]. Accordingly, circRNAs are implicated in the regulation of multiple physiologic and pathologic processes, including tumorigenesis, atherosclerosis, wound repair, cardiac senescence, myocardial ischemia/reperfusion injury, and so forth (Table 1).
In view of distinct circRNA–protein complexes that exist in mammalian cells [24], interactions between circRNAs and protein factors, as well as the caused variation of cellular biological functions, would be far beyond what we described in this article. For instance, the regulation of circRNAs to proteins′ action in other organelles except for the cell nucleus and cytosol, including the endoplasmic reticulum, Golgi apparatus, and mitochondria, has not been reported. In addition, it would be worthwhile to investigate whether circRNAs can affect protein modification unreported, such as protein acetylation or phosphorylation, except for the protein localization, activity or stability reviewed in this article. Besides protein–protein or protein–DNA interactions, studies focused on whether circRNAs affect the interactions between proteins and mRNAs or other non-coding RNAs should also be given importance. Furthermore, the identification of other types of unexplored proteins interacting with circRNAs, such as the chromatin regulatory proteins that control chromatin architecture, will further unveil more unknown functions of circRNAs. Finally, the improvement in methods or techniques for identifying unexplored proteins directly interacting with circRNAs would propel the discovery of circRNAs′ guidance to protein behaviors.
References
Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD, Kinzler KW, Vogelstein B (1991) Scrambled exons. Cell 64:607–613
Cocquerelle C, Daubersies P, Majerus MA, Kerckaert JP, Bailleul B (1992) Splicing with inverted order of exons occurs proximal to large introns. EMBO J 11:1095–1098
Cocquerelle C, Mascrez B, Hetuin D, Bailleul B (1993) Mis-splicing yields circular RNA molecules. FASEB J 7:155–160
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, Goodfellow P, Lovell-Badge R (1993) Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73:1019–1030
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388
Hansen TB, Wiklund ED, Bramsen JB, Villadsen SB, Statham AL, Clark SJ, Kjems J (2011) miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J 30:4414–4422
Pasman Z, Been MD, Garcia-Blanco MA (1996) Exon circularization in mammalian nuclear extracts. RNA 2:603–610
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, Zhu S, Yang L, Chen LL (2013) Circular intronic long noncoding RNAs. Mol Cell 51:792–806
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L, Zhu P, Chang Z, Wu Q, Zhao Y, Jia Y, Xu P, Liu H, Shan G (2015) Exon–intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22:256–264
Guarnerio J, Bezzi M, Jeong JC, Paffenholz SV, Berry K, Naldini MM, Lo-Coco F, Tay Y, Beck AH, Pandolfi PP (2016) Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell 165:289–302
Zhang Y, Xue W, Li X, Zhang J, Chen S, Zhang JL, Yang L, Chen LL (2016) The biogenesis of nascent circular RNAs. Cell Rep 15:611–624
Wilusz JE (2018) A 360 degrees view of circular RNAs: from biogenesis to functions. Wiley Interdiscip Rev RNA 9:e1478
Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade M, Laneve P, Rajewsky N, Bozzoni I (2017) Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol Cell 66(22–37):e29
Piwecka M, Glazar P, Hernandez-Miranda LR, Memczak S, Wolf SA, Rybak-Wolf A, Filipchyk A, Klironomos F, Cerda Jara CA, Fenske P, Trimbuch T, Zywitza V, Plass M, Schreyer L, Ayoub S, Kocks C, Kuhn R, Rosenmund C, Birchmeier C, Rajewsky N (2017) Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357:eaam8526
Zhang X, Yan Y, Lin W, Li A, Zhang H, Lei X, Dai Z, Li X, Li H, Chen W, Chen F, Ma J, Xie Q (2019) Circular RNA Vav3 sponges gga-miR-375 to promote epithelial–mesenchymal transition. RNA Biol 16:118–132
Li M, Ding W, Tariq MA, Chang W, Zhang X, Xu W, Hou L, Wang Y, Wang J (2018) A circular transcript of ncx1 gene mediates ischemic myocardial injury by targeting miR-133a-3p. Theranostics 8:5855–5869
Hall IF, Climent M, Quintavalle M, Farina FM, Schorn T, Zani S, Carullo P, Kunderfranco P, Civilini E, Condorelli G, Elia L (2019) Circ_Lrp6, a circular RNA enriched in vascular smooth muscle cells, acts as a sponge regulating miRNA-145 function. Circ Res 124:498–510
Qiu M, Xia W, Chen R, Wang S, Xu Y, Ma Z, Xu W, Zhang E, Wang J, Fang T, Hu J, Dong G, Yin R, Wang J, Xu L (2018) The circular RNA circPRKCI promotes tumor growth in lung adenocarcinoma. Can Res 78:2839–2851
Li Z, Yanfang W, Li J, Jiang P, Peng T, Chen K, Zhao X, Zhang Y, Zhen P, Zhu J, Li X (2018) Tumor-released exosomal circular RNA PDE8A promotes invasive growth via the miR-338/MACC1/MET pathway in pancreatic cancer. Cancer Lett 432:237–250
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, Wu YM, Dhanasekaran SM, Engelke CG, Cao X, Robinson DR, Nesvizhskii AI, Chinnaiyan AM (2019) The landscape of circular RNA in cancer. Cell 176(869–881):e813
Chen S, Huang V, Xu X, Livingstone J, Soares F, Jeon J, Zeng Y, Hua JT, Petricca J, Guo H, Wang M, Yousif F, Zhang Y, Donmez N, Ahmed M, Volik S, Lapuk A, Chua MLK, Heisler LE, Foucal A, Fox NS, Fraser M, Bhandari V, Shiah YJ, Guan J, Li J, Orain M, Picard V, Hovington H, Bergeron A, Lacombe L, Fradet Y, Tetu B, Liu S, Feng F, Wu X, Shao YW, Komor MA, Sahinalp C, Collins C, Hoogstrate Y, de Jong M, Fijneman RJA, Fei T, Jenster G, van der Kwast T, Bristow RG, Boutros PC, He HH (2019) Widespread and functional RNA circularization in localized prostate cancer. Cell 176(831–843):e822
Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA, Roslan S, Schreiber AW, Gregory PA, Goodall GJ (2015) The RNA binding protein quaking regulates formation of circRNAs. Cell 160:1125–1134
Schneider T, Hung LH, Schreiner S, Starke S, Eckhof H, Rossbach O, Reich S, Medenbach J, Bindereif A (2016) CircRNA–protein complexes: iMP3 protein component defines subfamily of circRNPs. Sci Rep 6:31313
Zhang J, Zhang X, Li C, Yue L, Ding N, Riordan T, Yang L, Li Y, Jen C, Lin S, Zhou D, Chen F (2019) Circular RNA profiling provides insights into their subcellular distribution and molecular characteristics in HepG2 cells. RNA Biol 16:220–232
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, Jin Y, Yang Y, Chen LL, Wang Y, Wong CC, Xiao X, Wang Z (2017) Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res 27:626–641
Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, Shenzis S, Samson M, Dittmar G, Landthaler M, Chekulaeva M, Rajewsky N, Kadener S (2017) Translation of CircRNAs. Mol Cell 66(9–21):e27
Wesselhoeft RA, Kowalski PS, Parker-Hale FC, Huang Y, Bisaria N, Anderson DG (2019) RNA circularization diminishes immunogenicity and can extend translation duration in vivo. Mol Cell 74:508–520
Zhao J, Lee EE, Kim J, Yang R, Chamseddin B, Ni C, Gusho E, Xie Y, Chiang CM, Buszczak M, Zhan X, Laimins L, Wang RC (2019) Transforming activity of an oncoprotein-encoding circular RNA from human papillomavirus. Nat Commun 10:2300
van Heesch S, Witte F, Schneider-Lunitz V, Schulz JF, Adami E, Faber AB, Kirchner M, Maatz H, Blachut S, Sandmann CL, Kanda M, Worth CL, Schafer S, Calviello L, Merriott R, Patone G, Hummel O, Wyler E, Obermayer B, Mucke MB, Lindberg EL, Trnka F, Memczak S, Schilling M, Felkin LE, Barton PJR, Quaife NM, Vanezis K, Diecke S, Mukai M, Mah N, Oh SJ, Kurtz A, Schramm C, Schwinge D, Sebode M, Harakalova M, Asselbergs FW, Vink A, de Weger RA, Viswanathan S, Widjaja AA, Gartner-Rommel A, Milting H, Dos Remedios C, Knosalla C, Mertins P, Landthaler M, Vingron M, Linke WA, Seidman JG, Seidman CE, Rajewsky N, Ohler U, Cook SA, Hubner N (2019) The translational landscape of the human heart. Cell 178:242–260
Hutvagner G, Simard MJ (2008) Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Biol 9:22–32
Yu CY, Li TC, Wu YY, Yeh CH, Chiang W, Chuang CY, Kuo HC (2017) The circular RNA circBIRC6 participates in the molecular circuitry controlling human pluripotency. Nat Commun 8:1149
Yang R, Xing L, Zheng X, Sun Y, Wang X, Chen J (2019) The circRNA circAGFG1 acts as a sponge of miR-195-5p to promote triple-negative breast cancer progression through regulating CCNE1 expression. Mol Cancer 18:4
Kwek KY, Murphy S, Furger A, Thomas B, O’Gorman W, Kimura H, Proudfoot NJ, Akoulitchev A (2002) U1 snRNA associates with TFIIH and regulates transcriptional initiation. Nat Struct Biol 9:800–805
Damgaard CK, Kahns S, Lykke-Andersen S, Nielsen AL, Jensen TH, Kjems J (2008) A 5′ splice site enhances the recruitment of basal transcription initiation factors in vivo. Mol Cell 29:271–278
Conn VM, Hugouvieux V, Nayak A, Conos SA, Capovilla G, Cildir G, Jourdain A, Tergaonkar V, Schmid M, Zubieta C, Conn SJ (2017) A circRNA from SEPALLATA3 regulates splicing of its cognate mRNA through R-loop formation. Nat Plants 3:17053
Zhu P, Zhu X, Wu J, He L, Lu T, Wang Y, Liu B, Ye B, Sun L, Fan D, Wang J, Yang L, Qin X, Du Y, Li C, He L, Ren W, Wu X, Tian Y, Fan Z (2019) IL-13 secreted by ILC2 s promotes the self-renewal of intestinal stem cells through circular RNA circPan3. Nat Immunol 20:183–194
Du WW, Zhang C, Yang W, Yong T, Awan FM, Yang BB (2017) Identifying and characterizing circRNA–protein interaction. Theranostics 7:4183–4191
Yang Q, Du WW, Wu N, Yang W, Awan FM, Fang L, Ma J, Li X, Zeng Y, Yang Z, Dong J, Khorshidi A, Yang BB (2017) A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ 24:1609–1620
Yang ZG, Awan FM, Du WW, Zeng Y, Lyu J, Wu Gupta S, Yang W, Yang BB (2017) The circular RNA interacts with STAT3, increasing its nuclear translocation and wound repair by modulating Dnmt3a and miR-17 function. Mol Ther 25:2062–2074
Zeng Y, Du WW, Wu Y, Yang Z, Awan FM, Li X, Yang W, Zhang C, Yang Q, Yee A, Chen Y, Yang F, Sun H, Huang R, Yee AJ, Li RK, Wu Z, Backx PH, Yang BB (2017) A circular RNA binds to and activates AKT phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics 7:3842–3855
Du WW, Yang W, Li X, Awan FM, Yang Z, Fang L, Lyu J, Li F, Peng C, Krylov SN, Xie Y, Zhang Y, He C, Wu N, Zhang C, Sdiri M, Dong J, Ma J, Gao C, Hibberd S, Yang BB (2018) A circular RNA circ-DNMT1 enhances breast cancer progression by activating autophagy. Oncogene 37:5829–5842
Lin CY, Loven J, Rahl PB, Paranal RM, Burge CB, Bradner JE, Lee TI, Young RA (2012) Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151:56–67
Baldanzi G, Filigheddu N, Cutrupi S, Catapano F, Bonissoni S, Fubini A, Malan D, Baj G, Granata R, Broglio F, Papotti M, Surico N, Bussolino F, Isgaard J, Deghenghi R, Sinigaglia F, Prat M, Muccioli G, Ghigo E, Graziani A (2002) Ghrelin and des-acyl ghrelin inhibit cell death in cardiomyocytes and endothelial cells through ERK1/2 and PI 3-kinase/AKT. J Cell Biol 159:1029–1037
Di R, Wu X, Chang Z, Zhao X, Feng Q, Lu S, Luan Q, Hemmings BA, Li X, Yang Z (2012) S6 K inhibition renders cardiac protection against myocardial infarction through PDK1 phosphorylation of Akt. Biochem J 441:199–207
White E (2016) Autophagy and p53. Cold Spring Harb Perspect Med 6:a026120
Tasdemir E, Chiara Maiuri M, Morselli E, Criollo A, D’Amelio M, Djavaheri-Mergny M, Cecconi F, Tavernarakis N, Kroemer G (2008) A dual role of p53 in the control of autophagy. Autophagy 4:810–814
Torrisani J, Unterberger A, Tendulkar SR, Shikimi K, Szyf M (2007) AUF1 cell cycle variations define genomic DNA methylation by regulation of DNMT1 mRNA stability. Mol Cell Biol 27:395–410
Georgia S, Kanji M, Bhushan A (2013) DNMT1 represses p53 to maintain progenitor cell survival during pancreatic organogenesis. Genes Dev 27:372–377
Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, Li X, Yang BB (2017) Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J 38:1402–1412
Chen Y, Yang F, Fang E, Xiao W, Mei H, Li H, Li D, Song H, Wang J, Hong M, Wang X, Huang K, Zheng L, Tong Q (2019) Circular RNA circAGO2 drives cancer progression through facilitating HuR-repressed functions of AGO2-miRNA complexes. Cell Death Differ 26:1346–1364
Fan XC, Steitz JA (1998) Overexpression of HuR, a nuclear-cytoplasmic shuttling protein, increases the in vivo stability of ARE-containing mRNAs. EMBO J 17:3448–3460
Zhou LY, Zhai M, Huang Y, Xu S, An T, Wang YH, Zhang RC, Liu CY, Dong YH, Wang M, Qian LL, Ponnusamy M, Zhang YH, Zhang J, Wang K (2019) The circular RNA ACR attenuates myocardial ischemia/reperfusion injury by suppressing autophagy via modulation of the Pink1/FAM65B pathway. Cell Death Differ 26:1299–1315
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S (2014) circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56:55–66
Starke S, Jost I, Rossbach O, Schneider T, Schreiner S, Hung LH, Bindereif A (2015) Exon circularization requires canonical splice signals. Cell Rep 10:103–111
Abdelmohsen K, Panda AC, Munk R, Grammatikakis I, Dudekula DB, De S, Kim J, Noh JH, Kim KM, Martindale JL, Gorospe M (2017) Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol 14:361–369
Barnhart MD, Moon SL, Emch AW, Wilusz CJ, Wilusz J (2013) Changes in cellular mRNA stability, splicing, and polyadenylation through HuR protein sequestration by a cytoplasmic RNA virus. Cell Rep 5:909–917
Liu L, Zhuang R, Xiao L, Chung HK, Luo J, Turner DJ, Rao JN, Gorospe M, Wang JY (2017) HuR enhances early restitution of the intestinal epithelium by increasing Cdc42 translation. Mol Cell Biol 37:e00574-16
Nan A, Chen L, Zhang N, Jia Y, Li X, Zhou H, Ling Y, Wang Z, Yang C, Liu S, Jiang Y (2019) Circular RNA circNOL10 inhibits lung cancer development by promoting SCLM1-mediated transcriptional regulation of the humanin polypeptide family. Adv Sci 6:1800654
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB (2016) Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res 44:2846–2858
Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, Yang BB (2017) Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ 24:357–370
Harper JW, Adami GR, Wei N, Keyomarsi K, Elledge SJ (1993) The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 75:805–816
Haupt Y, Maya R, Kazaz A, Oren M (1997) Mdm2 promotes the rapid degradation of p53. Nature 387:296–299
Yang JY, Zong CS, Xia W, Yamaguchi H, Ding Q, Xie X, Lang JY, Lai CC, Chang CJ, Huang WC, Huang H, Kuo HP, Lee DF, Li LY, Lien HC, Cheng X, Chang KJ, Hsiao CD, Tsai FJ, Tsai CH, Sahin AA, Muller WJ, Mills GB, Yu D, Hortobagyi GN, Hung MC (2008) ERK promotes tumorigenesis by inhibiting FOXO3a via MDM2-mediated degradation. Nat Cell Biol 10:138–148
Ekoff M, Kaufmann T, Engstrom M, Motoyama N, Villunger A, Jonsson JI, Strasser A, Nilsson G (2007) The BH3-only protein Puma plays an essential role in cytokine deprivation induced apoptosis of mast cells. Blood 110:3209–3217
Sun L, Huang Y, Liu Y, Zhao Y, He X, Zhang L, Wang F, Zhang Y (2018) Ipatasertib, a novel Akt inhibitor, induces transcription factor FoxO3a and NF-kappaB directly regulates PUMA-dependent apoptosis. Cell Death Dis 9:911
Holdt LM, Stahringer A, Sass K, Pichler G, Kulak NA, Wilfert W, Kohlmaier A, Herbst A, Northoff BH, Nicolaou A, Gabel G, Beutner F, Scholz M, Thiery J, Musunuru K, Krohn K, Mann M, Teupser D (2016) Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat Commun 7:12429
Grimm T, Holzel M, Rohrmoser M, Harasim T, Malamoussi A, Gruber-Eber A, Kremmer E, Eick D (2006) Dominant-negative Pes1 mutants inhibit ribosomal RNA processing and cell proliferation via incorporation into the PeBoW-complex. Nucleic Acids Res 34:3030–3043
Guevara NV, Kim HS, Antonova EI, Chan L (1999) The absence of p53 accelerates atherosclerosis by increasing cell proliferation in vivo. Nat Med 5:335–339
Wu G, Cai J, Han Y, Chen J, Huang ZP, Chen C, Cai Y, Huang H, Yang Y, Liu Y, Xu Z, He D, Zhang X, Hu X, Pinello L, Zhong D, He F, Yuan GC, Wang DZ, Zeng C (2014) LincRNA-p21 regulates neointima formation, vascular smooth muscle cell proliferation, apoptosis, and atherosclerosis by enhancing p53 activity. Circulation 130:1452–1465
Chen N, Zhao G, Yan X, Lv Z, Yin H, Zhang S, Song W, Li X, Li L, Du Z, Jia L, Zhou L, Li W, Hoffman AR, Hu JF, Cui J (2018) A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol 19:218
Xia P, Wang S, Ye B, Du Y, Li C, Xiong Z, Qu Y, Fan Z (2018) A circular RNA protects dormant hematopoietic stem cells from DNA sensor cGAS-mediated exhaustion. Immunity 48(688–701):e687
Good CR, Panjarian S, Kelly AD, Madzo J, Patel B, Jelinek J, Issa JJ (2018) TET1-mediated hypomethylation activates oncogenic signaling in triple-negative breast cancer. Can Res 78:4126–4137
Galluzzi L, Vanpouille-Box C, Bakhoum SF, Demaria S (2018) SnapShot: CGAS-STING signaling. Cell 173(276–276):e271
Essers MA, Offner S, Blanco-Bose WE, Waibler Z, Kalinke U, Duchosal MA, Trumpp A (2009) IFNalpha activates dormant haematopoietic stem cells in vivo. Nature 458:904–908
Sato T, Onai N, Yoshihara H, Arai F, Suda T, Ohteki T (2009) Interferon regulatory factor-2 protects quiescent hematopoietic stem cells from type I interferon-dependent exhaustion. Nat Med 15:696–700
Bellaousov S, Reuter JS, Seetin MG, Mathews DH (2013) RNAstructure: web servers for RNA secondary structure prediction and analysis. Nucleic Acids Res 41:W471–W474
Gruber AR, Lorenz R, Bernhart SH, Neubock R, Hofacker IL (2008) The Vienna RNA websuite. Nucleic Acids Res 36:W70–74
Bellucci M, Agostini F, Masin M, Tartaglia GG (2011) Predicting protein associations with long noncoding RNAs. Nat Methods 8:444–445
Muppirala UK, Honavar VG, Dobbs D (2011) Predicting RNA–protein interactions using only sequence information. BMC Bioinf 12:489
De Gregorio E, Preiss T, Hentze MW (1999) Translation driven by an eIF4G core domain in vivo. EMBO J 18:4865–4874
Hirsch JD, Eslamizar L, Filanoski BJ, Malekzadeh N, Haugland RP, Beechem JM, Haugland RP (2002) Easily reversible desthiobiotin binding to streptavidin, avidin, and other biotin-binding proteins: uses for protein labeling, detection, and isolation. Anal Biochem 308:343–357
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (81772552, 81572714) and the Fundamental Research Funds for the Central Universities of China (531118010140).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Luo, J., Liu, H., Luan, S. et al. Guidance of circular RNAs to proteins’ behavior as binding partners. Cell. Mol. Life Sci. 76, 4233–4243 (2019). https://doi.org/10.1007/s00018-019-03216-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-019-03216-z