Abstract
Annexin A10 is the latest identified member of the annexin family of Ca2+- and phospholipid-binding proteins. In previous studies, downregulation of annexin A10 was correlated with dedifferentiation, invasion, and tumor progression, pointing to a possible tumor suppressor role. However, the biochemical characteristics and functions of annexin A10 remain unknown. We show that annexin A10 displays biochemical characteristics atypical for an annexin, indicating a Ca2+- and membrane-binding-independent function. Annexin A10 co-localizes with the mRNA-binding proteins SFPQ and PSPC1 at paraspeckles, an only recently discovered nuclear body, and decreases paraspeckle numbers when overexpressed in HeLa cells. In addition, annexin A10 relocates to dark perinucleolar caps upon transcriptional inhibition of RNA polymerase II. We mapped the cap-binding function of annexin A10 to the proximal part of the core domain, which is missing in the short isoform of annexin A10, and show its independence from the remaining functional type II Ca2+-binding site. In contrast to this, paraspeckle recruitment required additional core regions and was negatively affected by the mutation of the last type II Ca2+-binding site. Additionally, we show that overexpression of annexin A10 in HeLa cells increases their sensitivity to apoptosis and reduces colony formation. The identification of unique nuclear and biochemical characteristics of annexin A10 points towards its membrane-independent role in paraspeckle-associated mRNA regulation or processing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Annexins are a highly conserved superfamily of proteins expressed in tissues of organisms ranging from protists to higher plants and vertebrates. Ca2+-dependent, reversible binding to negatively charged phospholipids are key properties of the superfamily and require annexin-specific type II Ca2+-binding sites. The structural organization of annexins generally differentiates two distinct domains: a highly conserved C-terminal core region that is formed by annexin repeats and harbors the Ca2+-binding sites and the N-terminal region that is unique to each member. The N-terminal annexin domains display variable length and amino acid compositions, often contain sites for posttranslational modification, and provide platforms for annexin-binding ligands. Annexins are involved in a large number of cellular processes that are primarily associated with membrane–cytoskeleton scaffolding, vesicle trafficking, and Ca2+-mediated signaling events, but also play an inhibitory role in inflammation and coagulation [14].
The cDNA of vertebrate annexin A10 (anxA10) was identified in 1999 [35], but detailed analyses of both biochemical properties and intracellular functions of this novel annexin are still lacking. Non-pathological expression of anxA10 is restricted to a small subset of tissues, mainly the epithelia of the gastrointestinal tract [30, 31]. Homozygous deletion and down-regulation of anxA10 expression correlate with poor survival in gastric and bladder cancer, thus implicating it in tumor suppression [26, 27, 30, 31, 33, 36, 43]. Interestingly, sequence analysis of anxA10 revealed that the protein is so far the only annexin family member to harbor only one Ca2+-binding site [35]. This distinct feature and the potential function of anxA10 in tumor suppression prompted us to investigate the biochemical characteristics of this atypical annexin.
Here, we demonstrate that, due to low Ca2+-binding capacities, membrane association of anxA10 is less likely to explain intracellular in vivo functions of the protein. Instead, analysis of subcellular localization indicates that anxA10 localizes to nuclear substructures and affects paraspeckle numbers. The localization to subnuclear bodies implicates that anxA10 might participate in transcriptional regulation and mRNA processing. Thus, anxA10 broadens the functional spectrum so far associated with annexin proteins.
Materials and methods
Expression constructs and antibodies
IMAGENES clone IRAUp969B12196D was used to amplify the full-length human anxA10 cDNA. The PCR fragment was cloned, via EcoRI sites immediately upstream of the initial ATG and immediately downstream of the stop codon, into the bacterial expression vector pET-23a(+) (Novagen) modified for expression without the T7 tag [15]. For the expression of anxA10 fused C-terminally to GFP (anxA10GFP), the cDNA of anxA10 was amplified via the same forward primer and a reverse primer with a SalI site replacing the stop codon and cloned into the pEGFP-N3 expression vector (Clontech). Mutant anxA10 expression constructs were generated by site-directed mutagenesis. The anxA1GFP and anxA2GFP constructs were previously published [49, 64]. To generate an annexin A2-mCherry expression construct, mCherry was swapped with the GFP-tag. The bacterial expression construct for anxA8 was described previously [15]. All constructs were verified by sequencing.
Goat polyclonal anti-anxA10 antibodies were obtained from Abcam, rabbit polyclonal anti-anxA10 antibodies were kindly provided by R. Morgan. Rabbit polyclonal anti-caveolin antibodies were obtained from Becton–Dickinson, mouse monoclonal anti-PSF (SFPQ), rabbit polyclonal anti-GAPDH, and rabbit polyclonal anti-PSPC1 antibodies from Santa Cruz, and rabbit monoclonal anti-fibrillarin antibodies from Cell Signaling Technologies. Rabbit polyclonal anti-GFP serum was purchased from Invitrogen. DRAQ5 (Biostatus) was used to visualize DNA.
Expression and purification of human annexins
Human anxA8 was purified as previously described [15]. Full-length anxA10 was expressed in E. coli BL21(DE3)pLysS (Stratagene) transformed with pET-23a(+)mod./anxA10. Bacteria were grown in LB-medium containing ampicillin at 37 °C to an OD600 of 0.6. Protein expression was induced by 1 mM isopropyl-β-D-thiogalactose for 3 h. Bacteria were lysed in buffer containing 50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1 mM DTT, 1 mM EGTA, and 0.5 mM phenylmethylsulfonyl fluoride (PMSF) by 3 freeze/thaw cycles and subsequent sonication. Cleared lysate was added to brain extract (Folch fraction I of bovine brain; Sigma) liposomes with a final concentration of 2 mg/ml in the presence of 25 μM E64 and 3 mM CaCl2. Incubation for 1 h at room temperature was followed by pelleting of liposomes containing bound anxA10 at 100,000g for 30 min at 4 °C. AnxA10 was released into the supernatant via resuspension of the liposome pellet in lysis buffer containing 50 mM EGTA and recentrifugation. The protein was dialyzed against buffer containing 50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1 mM DTT, and 0.5 mM PMSF.
Phospholipid binding assay and PIP Strips
Binding of anxA10 to brain extract liposomes was analyzed as described previously [15, 58]. Briefly, 30 μg anxA10 were incubated in binding buffer containing 50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1 mM DTT, and 0.5 mM PMSF in the presence of different Ca2+ concentrations (1 mM EGTA; 0.5, 1, 2 or 5 mM CaCl2) for 20 min at room temperature and pre-cleared at 100,000g for 30 min at 4 °C. Supernatants were incubated for 1 h at room temperature with brain extract liposomes at a final concentration of 2 mg/ml. Liposomes were then pelleted at 100,000g for 30 min at 4 °C and washed in binding buffer. Liposome-associated anxA10 was detected by SDS-PAGE and Coomassie staining. Quantification of the signal intensities was carried out by densitometric analysis. Data were collected from four independent experiments and the statistical significance of differences in the liposome-bound A10 levels was evaluated by repeated measures one-way ANOVA followed by Dunnett’s multiple comparison test. P < 0.05 indicated a significant difference.
Phosphoinositide phosphate (PIP) Strips (Echelon Biosciences) were blocked with 1 % nonfat-dry milk in PBS for 1 h at room temperature. Purified anxA10 was added to the PIP Strips at a final concentration of 2 μg/ml and incubated overnight at 4 °C in blocking buffer containing 50 or 500 μM Ca2+. PIP Strips were subsequently washed with PBS containing 0.1 % (v/v) Tween 20. The respective Ca2+ concentrations were maintained during all subsequent incubation steps. Bound anxA10 was detected by conventional, indirect immunoblotting using anti-anxA10 antibody.
Actin co-sedimentation assay
Analysis of actin binding was performed as described previously [15, 23]. Globular actin purified from skeletal muscle (Cytoskeleton) was diluted to 1 mg/ml in General Actin Buffer (5 mM Tris–HCl, pH 8.0, 0.2 mM CaCl2, 0.2 mM ATP). Polymerization to F-actin was induced by addition of a 1/50 volume of polymerization solution (2.5 M KCl, 100 mM MgCl2, 50 mM ATP) followed by incubation for 1 h at room temperature. Then, 50 μg of purified annexins were incubated in a final volume of 200 μl reaction buffer (5 mM Tris–HCl, pH 8.0, 0.2 mM CaCl2, 1 mM MgCl2, 0.33 mM ATP, 0.5 mM DTT) in the presence of 2 mM CaCl2 or 2 mM EGTA for 15 min at room temperature and then pre-cleared by centrifugation at 350,000g for 15 min at 4 °C. Supernatants were incubated with 25 μg F-actin for 1 h at room temperature. F-actin was spun down at 350,000g for 30 min at 4 °C. Pellets were washed in 200 μl reaction buffer supplemented with 2 mM CaCl2 or 2 mM EGTA and re-pelleted. Equal protein amounts were analyzed by SDS-PAGE and Coomassie staining. The levels of F-actin associated annexins were quantified densitometrically. Data were collected from three independent experiments and the statistical significance of differences was evaluated by Student’s t test. P < 0.05 indicated a significant difference.
Cell culture and transfection
HeLa and MDCK II (Sigma-Aldrich) cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) High Glucose (PAA) supplemented with 10 % fetal calf serum, glutamine, and antibiotics at 7 % CO2 and 37 °C. Cells grown to 50–80 % confluency were transfected with Lipofectamine 2000 (Invitrogen) as recommended by the manufacturer. For selection of stably transfected clones, transfected cells were cultured in the presence of 800 μg/ml G418 for 4 weeks. Expression of the constructs was verified by fluorescence microscopy and immunoblotting.
Cell fractionations
All steps were performed at 4 °C. Confluent cells from a 10-cm dish were first washed then scraped into cold PBS and pelleted (3 min, 800g, 4 °C). Cell pellets were resuspended in 500 μl cold lysis buffer (25 mM MES pH 6.5, 150 mM NaCl, supplemented with fresh protease inhibitor cocktail (Roche) and the indicated concentrations of CaCl2.) Cells were passed 10 times through a 23-gauge needle and incubated on ice for 20 min. Nuclei were removed by centrifugation (5 min, 800g, 4 °C) followed by a second centrifugation of the supernatant to remove cytoskeletal and mitochondrial components (10 min, 10,000g, 4 °C). Cytosolic and membrane fractions were separated by a 1-h centrifugation step at 100,000g and 4 °C. The cytosolic (supernatant) fraction was removed and denatured in SDS sample buffer. The membrane pellet was resuspended in 400 μl lysis buffer with supplements. After a second centrifugation (10 min, 10,000g, 4 °C), the supernatant was discarded and the membrane pellet resuspended and denatured in 2 × SDS sample buffer. For immunoblot analysis, one-tenth of the cytosolic and membrane fractions from anxA10GFP expressing MDCK cells were separated via electrophoresis. For the detection of endogenous anxA10 in the untransfected MDCK cells, it was necessary to load complete membrane fractions on the gel. Membrane and cytosol fractions were separated by SDS-PAGE, transferred onto a PVDF membrane and probed by immunoblotting for endogenous anxA10 and GFP as well as the respective membrane and cytosol marker proteins caveolin and GAPDH. Quantification of membrane-associated anxA10 signal intensities normalized to GAPDH or caveolin, respectively, was carried out by densitometric analysis. Statistical significance of results obtained from either five (anxA10GFP) or three (endogenous anxA10) independent experiments was evaluated using ANOVA followed by Tukey’s multiple comparison test.
Colony-formation assay
HeLa cells stably expressing either anxA10GFP or GFP were seeded in 6-well plates (103 cells/well). Cells were grown in DMEM + 1 % fetal calf serum with or without 10 ng/ml EGF, fixed, and stained 2 weeks after plating with DIFF-QUIK Staining Set (Siemens, Germany). Colonies containing more than 40 cells were quantified using ImageJ software. Data were collected from four independent experiments and the statistical significance was evaluated by Student′s t test. P < 0.05 indicated a significant difference.
Detection of doxorubicin-induced apoptosis
Apoptosis assays were performed with HeLa cell lines stably transfected with GFP or anxA10GFP. Three differently expressing subclones were combined for experimental working cultures. 1.2 × 106 cells were seeded per 6-well plate 1 day prior to stimulation. Stimulation was performed with 5 μM doxorubicin-hydrochloride (suitable for fluorescence; Sigma-Aldrich) for 24 h while unstimulated samples were used as controls. Cells were pelleted (3 min, 800g, 4 °C), resuspended in 8 M urea (50–80 μl) supplemented with Complete Protease Inhibitor (Roche), and sonicated 3–5 times on ice with a tip sonicator (amplitude 50, duty cycle 50). For the analysis of apoptosis, identical amounts of protein were separated by SDS-PAGE on a 12 % gel and analyzed by immunoblotting with anti-poly ADP ribose polymerase (PARP) and anti-GAPDH as a loading control. Protein detection and quantification were performed with appropriate IRDye 800 and IRDye 680 CW-conjugated secondary antibodies and the LI-COR Odyssey infrared fluorescence imaging system and software (LI-COR). PARP band intensities were normalized to GAPDH levels and PARP fragmentation was expressed as the percentage of cleaved PARP over total PARP. The statistical significance was evaluated by repeated measures one-way ANOVA followed by either Tukey’s or Dunnett’s multiple comparison test. P < 0.05 indicated a significant difference.
Immunofluorescences
Cap-formation was induced through incubation with 5 μg/ml actinomycin D (ActD; Calbiochem) or 30 μg/ml α-amanitin (Sigma) [55], or with 5 μM doxorubicin (Doxo; Sigma). After incubation, cells were fixed with 4 % paraformaldehyde (PFA) + 0.1 % Triton X-100 (TX-100) in PBS, followed by 10 min at RT with 4 % PFA in PBS. For endogenous anxA10 and paraspeckle-staining with anti-SFPQ antibodies, cells were fixed for 3 min with 4 % PFA + 0.1 % TX-100 in PBS. Cells were subsequently blocked with 2 % BSA in PBS for 15 min and incubated with primary antibodies at 4 °C overnight. Secondary antibody as well as DRAQ5 staining were performed at RT for 45 min. Confocal microscopy was carried out using a LSM 510 META or a LSM 780 confocal microscope (Zeiss) equipped with a Plan-Apochromat ×63/1.4 oil immersion objective.
For quantification of nuclear and cytosolic fluorescence signals, images were recorded with identical laser and detector settings. Regions of interest (ROIs) of identical size were placed in the nucleus and cytosol of transfected cells and their integrated fluorescence intensities were determined using the MetaMorph software (Universal Imaging). For each cell, the ratio of nuclear to cytosolic intensity was calculated, with ratios >1 indicating nuclear enrichment. Data were collected from more than 300 cells (at least 50 cells/experiment/annexin) in three independent transfection experiments. Statistical significance of the results was evaluated by paired Student′s t test (two-tailed P value).
To correlate the signal strengths of SFPQ and endogenous anxA10 in subnuclear foci, the line profile analysis tool of the Zeiss Zen software was used creating intensity value profiles of the respective signals along a line which was drawn through several subnuclear foci.
To quantitate the average number of paraspeckles per cell, the nucleus was defined by DNA staining and nuclear PSPC1 was visualized by immunofluorescence. Confocal z stacks of nuclei were acquired and maximum projections of color-separated images were converted to grayscale images. After thresholding, remaining signals within the nuclear area were binarized and the “analyze particles” plugin of the ImageJ software was used to digitally quantify paraspeckle numbers of nuclei. For each subclone, randomized images of 90 nuclei obtained from three independent experiments were scored. Statistical analysis of the differences was performed utilizing ANOVA followed by Tukey’s multiple comparison test.
Live cell imaging of Ca2+-induced membrane binding
Cells co-expressing anxA10GFP and annexin A2mCherry were imaged in medium buffered with 20 mM HEPES in a heated chamber. To increase intracellular Ca2+-levels, 1 μM thapsigargin was added to the medium and images were recorded by time-lapse microscopy every 30 s over a duration of 5 min. Another image was recorded before addition of 5 μM ionomycin.
Results
Biochemical characterization of anxA10
To elucidate its biochemical properties, we recombinantly expressed human anxA10 protein in E. coli, as this had been shown previously to be a suitable method to prepare annexins while retaining their typical properties [25, 39]. To ensure that the purified protein was indeed full-length anxA10, we subjected the molecule to analysis by electrospray mass spectrometry. An experimentally determined molecular mass of 37,277 ± 1 Da was in full accordance with the predicted mass of 37,278 Da for human anxA10 [35], indicating the presence of full-length protein (Suppl. Fig. 1).
To characterize the Ca2+-dependency of phospholipid binding, we performed small-scale liposome pelleting assays with purified anxA10 in the presence of increasing Ca2+ concentrations, using brain extract liposomes that contained approximately 50 % phosphatidylserine (Fig. 1a). As expected, no interaction of anxA10 with the liposomes was observed in the presence of the Ca2+-chelator EGTA. Binding of anxA10 to liposomes was barely detectable at 0.5 mM free Ca2+ and increased proportional with rising Ca2+ concentrations. Quantitative analysis confirmed that a significant increase in Ca2+-dependent liposome association was only observable at Ca2+ concentrations higher than 0.5 mM. However, even in the presence of 5 mM Ca2+, anxA10 association with liposomes still remained incomplete (Fig. 1a, lower panel). AnxA10 that had been bound at 5 mM Ca2+ could be re-extracted with a subsequent EGTA wash, indicating that the weak association observed in this assay was reversible and only dependent on Ca2+. In addition to their interaction with negatively charged phospholipids, some annexins bind preferentially to phosphoinositides, and this binding is thought to play a role in membrane subdomain organization [47]. Therefore, we performed an in vitro PIP Strip analysis with the purified protein. In agreement with the liposome pelleting assays, no interaction was observed in the presence of low (50 μM) Ca2+ concentrations (Fig. 1b, left panel). Increasing the Ca2+ concentration to 500 μM resulted in binding to phosphatidylserine but also revealed a weak but specific interaction with immobilized PI(3)P and PI(4)P (Fig. 1b, right panel).
Biochemical analysis of human anxA10. a Ca2+-dependent phospholipid binding of recombinantly expressed human anxA10 (rh anxA10). Purified anxA10 was incubated with brain extract liposomes in the presence of either 1 mM EGTA or increasing amounts of Ca2+. Bound anxA10 at 5 mM Ca2+ was additionally re-extracted with 50 mM EGTA. Liposomes were spun down and equal protein amounts in the pellets (P) or the supernatant (SN) were analyzed by SDS-PAGE and Coomassie staining. Levels of liposome-associated anxA10 were densitometrically quantified. Mean values ± SEM from four independent experiments are shown (lower panel). **statistically significant difference (P < 0.01) as revealed by ANOVA followed by Dunnett’s post test. b AnxA10 binds specifically to PI(3)P and PI(4)P. Membranes with immobilized phospholipids (PIP strips) were incubated overnight with 2 μg/ml anxA10 in the presence of either 50 or 500 μM Ca2+. Bound anxA10 was detected by immunoblotting with an anxA10-specific antibody. In the presence of 500 μM CaCl2, anxA10 showed a specific adsorption to immobilized phosphatidylinositol-3-phosphate (PI(3)P), phosphatidylinositol-4-phosphate (PI(4)P), and phosphatidylserine (PS). c AnxA10 does not bind to F-actin. Purified anxA8 or anxA10 was mixed with polymerized F-actin (input) in the presence of either 2 mM EGTA or 2 mM Ca2+. Actin filaments together with bound proteins were recovered by high speed centrifugation. Equal protein amounts of the pellets were analyzed by SDS-PAGE and Coomassie staining. Ca2+-dependent F-actin binding of anxA8 served as a control for the co-sedimentation assay. Levels of F-actin-associated annexins were densitometrically quantified. Mean values ± SEM from three independent experiments are shown. **statistically significant difference (P < 0.01) as revealed by ANOVA followed by Student’s t-test
Several members of the annexin family bind to and, in the case of annexin A2, also bundle F-actin in a Ca2+-dependent manner [19, 20]. Because the C-terminal domain of anxA10 shares sequence similarities with the postulated annexin A2 F-actin binding motif, which was mapped to the very C-terminus [10], we investigated whether F-actin binding properties could also be observed for anxA10. To address this, we performed an in vitro actin co-sedimentation assay in the presence of 2 mM Ca2+, used because F-actin binding might only be seen in the presence of high Ca2+ concentrations, as has been shown for annexin A5 [59, 60], with anxA8 as a positive control of an F-actin binding annexin. AnxA8 showed almost no association with F-actin in the presence of EGTA, but was clearly detectable in the F-actin pellet at 2 mM Ca2+, and densitometric analysis confirmed that the increase in F-actin-associated anxA8 was statistically significant (Fig. 1c). In contrast, we detected only negligible co-pelleting of anxA10 with F-actin in the absence or presence of Ca2+.
The results of our biochemical characterization point to the intriguing possibility that anxA10 might play a Ca2+- and consequentially membrane-binding-independent role in cellular functions. To investigate the cellular functions of anxA10, we chose a human epithelial cervical cancer cell line (HeLa) and the epithelial canine kidney cell line MDCK. While HeLa cells displayed no detectable anxA10 expression at the protein level, we were able to detect a protein signal of identical size as the bacterially expressed anxA10 (Fig. 2a) in MDCK cells. This signal was specifically downregulated by siRNA designed to target canine anxA10 (suppl. Fig. 2).
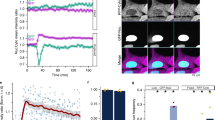
Membrane distribution of anxA10. a Different expression levels of anxA10 in MDCK and HeLa cells. Bacterially expressed anxA10 (0.2 μg purified protein) or 150 μg protein of total cell lysates were loaded and analyzed by immunoblotting. An immunoreactive band of similar size to the recombinant protein (rh anxA10) was detected in MDCK but not in HeLa cells. GAPDH was used as a loading control. AnxA10GFP shows identical membrane binding as endogenous anxA10. MDCK cells were separated into cytosolic (Cyt) and membrane (Mem) fractions in the presence of increasing Ca2+ concentrations. Complete membrane fractions were loaded to achieve a sufficient anxA10 signal (b). For analysis of anxA10GFP, one-tenth of the cytosolic and membrane fractions from anxA10GFP-expressing MDCK cells (c) were loaded. Fractions were analyzed by immunoblotting against anxA10 and GFP. GAPDH was used as a marker protein for cytosolic fractions, caveolin for membrane fractions (upper panels). Lower panels show the relative distribution of endogenous anxA10 or ectopically expressed anxA10GFP in the respective fractions. Results are mean values ± SEM from either five (anxA10GFP) or three (endogenous anxA10) independent experiments. ANOVA followed by Tukey’s post test revealed no statistical difference of anxA10 levels in the membrane fractions. d Live cell imaging of Ca2+-sensitive membrane binding of annexins. HeLa cells co-expressing anxA10GFP and anxA2mCherry were first stimulated with 1 μM thapsigargin to increase intracellular Ca2+-levels. Subsequently, 5 μM ionomycin was added to allow influx of extracellular Ca2+. Time series of fluorescent images were recorded every 30 s and selected frames are shown. Note that anxA2mCherry was found at the plasma membrane already 5 min after addition of thapsigargin, whereas anxA10GFP remained cytosolic even after cell permeabilization with the ionophore. Bar 20 μM. The movie can be viewed in suppl. Mov1
To overcome restrictions due to the limited amount of endogenous anxA10, we created cell lines that stably overexpressed a C-terminally GFP-tagged anxA10 construct (anxA10GFP) for subsequent functional assays. To avoid clonal effects, cell populations were created from a mix of three subclones with varying anxA10GFP expression levels. To confirm that the Ca2+-dependent membrane-binding properties of the tagged protein resembled those of endogenous anxA10, we separated both MDCK cells and MDCK cells stably overexpressing anxA10GFP into cytosolic and membrane fractions in the presence of increasing Ca2+ concentrations. Soluble GAPDH and the membrane protein caveolin served as marker proteins to ensure the purity of the fractions. As shown in Fig. 2b, the endogenous anxA10 was recovered almost completely in the soluble fraction and did not significantly translocate to the membrane fractions even at high Ca2+ concentrations. When cells expressing the GFP-tagged protein were analyzed, a similar distribution pattern was seen, with the anxA10GFP remaining mostly in the cytosolic fraction even in the presence of 2 mM Ca2+. No significant increase in the membrane fractions was seen with rising Ca2+ concentrations (Fig. 2c). To confirm this in vivo, we co-expressed anxA10GFP and anxA2mCherry in HeLa cells and followed their behavior in response to thapsigargin, a drug that inhibits Ca2+ re-uptake into intracellular Ca2+ stores and thus moderately increases cytosolic Ca2+ levels [57], and a subsequent ionomycin stimulus using live cell fluorescence microscopy. A series of confocal images was recorded every 30 s; frames were taken before stimulation, during 5 min after addition of 1 μM thapsigargin, and during 5 min after the subsequent addition of 5 μM ionomycin (Fig. 2d, complete movie in suppl. Mov 1). The expression levels of both proteins varied considerably among individual cells, but localization patterns were identical across cells. In non-stimulated cells, anxA10GFP was found predominantly in the nucleus and, to a lesser extent, in the cytosol, whereas anxA2mCherry was excluded from the nucleus and showed a diffuse, predominantly cytosolic localization with partial plasma membrane binding. The striking difference in the nuclear localization was most likely due to the nuclear export sequence found in the N-terminal part of anxA2 [8]. After treating the cells with thapsigargin, anxA2mCherry readily translocated to the plasma membrane, whereas anxA10 remained unchanged. Upon subsequent stimulation with ionomycin, which permeabilizes the membrane and induces a strong Ca2+ influx from the extracellular space, anxA2mCherry fluorescence at the plasma membrane increased further and additional association with intracellular vesicular structures was observed. Even under these conditions, anxA10GFP did not translocate to membranes, reflecting the low Ca2+ sensitivity already seen in the in vitro liposome membrane binding as well as membrane fractionation experiments.
AnxA10 overexpression in human epithelial cancer cells reduced clonogenic ability and increased sensitivity to doxorubicin-induced apoptosis
AnxA10 expression is downregulated in a number of human epithelial carcinomas as well as tumor-derived cell lines [26, 27, 30, 33, 36, 43, 61]. We therefore tested if the described effects of anxA10 on cancer cell proliferation and survival could be observed in anxA10GFP-overexpressing HeLa cells, which do not express detectable levels of endogenous anxA10. For this, we performed a long-term clonogenic assay, which reflects the ability of an individual cell to grow into a colony. Large differences in the percentage of cells that gave rise to colonies after seeding were seen for HeLa cells stably expressing either anxA10GFP or GFP (Fig. 3a) in the presence or absence of EGF. As shown in Fig. 3b, under both conditions reduced numbers of colonies were observed in anxA10-overexpressing cells and statistical analysis revealed that expression of anxA10GFP was indeed associated with a significant decrease in clonogenicity.
AnxA10 expression reduces clonogenic growth and resistance to apoptosis in HeLa cells. a Western blot showing stable expression of anxA10GFP and GFP in HeLa cells. b Clonogenic survival is reduced in HeLa cells stably transfected with anxA10GFP. HeLa cells stably expressing GFP (left) or anxA10GFP (right) were grown without (w/o, upper) or in the presence of 10 ng/ml (lower) epidermal growth factor (EGF). Representative images of four independent experiments are shown. Numbers of colonies consisting of >40 cells were determined and the statistical significance was evaluated by Student′s t test with P < 0.05 indicating a significant difference. c Immunoblots of AnxA10GFP-expressing HeLa cells left untreated (unstim.) or stimulated for 24 h with 5 μM doxorubicin (Doxo). PARP fragmentation was analyzed by immunoblotting with GAPDH as a loading control. Levels of the 89-kDa cleaved PARP were calculated as a percentage of the total PARP (full-length plus cleaved). The graph shows mean (±SEM) levels of cleaved PARP from six independent experiments. Statistical analysis was performed using ANOVA followed by Tukey’s multiple comparison test. *P < 0.05
To further confirm a role of anxA10 in tumor formation, we analyzed the sensitivity of stably anxA10GFP-expressing HeLa cells to treatment with doxorubicin, a drug widely used in cancer chemotherapy. AnxA10GFP-overexpressing cells were stimulated for 24 h with 5 μM doxorubicin and compared with stimulated GFP-expressing control cells. To detect apoptosis, lysates were probed for cleavage products of PARP, an enzyme involved in DNA damage repair and the maintenance of genomic stability. PARP cleavage, a process mediated by activation of caspase, is commonly used to detect apoptotic cell death [41]. Whereas no difference in the amount of PARP cleavage was detected in unstimulated cells expressing anxA10GFP vs. those with just GFP expression, significantly higher amounts of the large PARP (89-kDa) fragment were detected in anxA10GFP-expressing cells compared to control cells after 24 h of doxorubicin treatment (Fig. 3c). This indicates that anxA10 is able to increase cytotoxin sensitivity in epithelial cancer cells.
Nuclear anxA10GFP is recruited to dark nucleolar caps upon transcriptional inhibition of RNA polymerase II and localizes to paraspeckles
To shed some light on the mechanism by which anxA10 achieves its anti-tumorigenic properties, we used confocal fluorescence microscopy and screened transiently transfected cells for enrichment of anxA10GFP at cellular substructures. Previous studies detected anxA10 in the nuclei of gastric epithelial cells [26, 32]. As mentioned above, anxA10GFP also showed a prominent nuclear localization, but some signal was also found throughout the cytosol. To confirm a specific enrichment, we quantified the fluorescence signals of anxA10 in the cytoplasm and nucleoplasm of HeLa cells and compared them with those of anxA1GFP, an annexin of similar size that is also able to enter the nucleus, but shows no enrichment under resting conditions [48]. While anxA1GFP was equally distributed in both cytosol and nucleoplasm (1.0 ± 0.048 nuclear/cytosolic), anxA10GFP was higher in the nucleoplasm and, upon quantification, showed a 70 % enrichment compared to the cytosol (1.7 ± 0.029; Fig. 4a).
Enrichment of anxA10GFP in the nucleus and at subnuclear foci. a Confocal immunofluorescence images of PFA-fixed HeLa cells showing nuclear enrichment of anxA10GFP in contrast to anxA1GFP. Bar 20 μm. Signal intensities of identically sized regions from the nucleus and cytosol within each cell were measured and expressed as the ratio of nuclear versus cytosolic fluorescence. The bar graph shows the mean ± SEM of three independent transfection experiments. Statistical analysis using a paired t test revealed a 70 % enrichment of anxA10GFP in the nucleus of transiently transfected HeLa cells and equal distribution of anxA1GFP between cytosol and nucleus. ***P < 0.001. b Confocal images of nuclear anxA10GFP localization upon doxorubicin (Doxo) treatment. HeLa cells transiently expressing anxA10GFP were incubated for 4 h with 5 μM Doxo. AnxA10GFP (green) translocates to distinct nuclear foci surrounding nucleoli. Doxo intercalates DNA and fluorescence emission can be used to visualize DNA (red). Bar 5 μm. c Translocation of anxA10GFP upon inhibition of transcription with actinomycin D (ActD). MDCK cells transiently transfected for 24 h with anxA10GFP (green) were incubated for 2 h with 5 μg/ml ActD. DNA was visualized with DRAQ5 (red). Inset high magnification of a single nucleolus surrounded by condensed chromatin and anxA10GFP-positive foci. Bar 5 μm
In light of our results showing increased doxorubicin-induced apoptosis in anxA10-overexpressing cells, we were interested to know if incubation with doxorubicin would also lead to changes in subcellular anxA10 distribution. We found that, after 4 h of doxorubicin stimulation, anxA10GFP-expressing HeLa cells showed a striking translocation of the GFP-tagged protein to half-moon-shaped nuclear foci. These foci seemingly localized around nucleoli but were distinct from condensed chromosomal DNA, which was evident through DNA staining with fluorescently detectable doxorubicin (Fig. 4b). Similar foci have been described to form around nucleoli following transcriptional inhibition caused by apoptosis, mitosis, or RNA Pol II-inhibiting drugs [50]. These so-called ‘nucleolar or perinucleolar caps’ contain proteins and RNA of segregated nucleolar substructures, but also accumulate a distinct subset of nucleoplasmic proteins, including components of nuclear bodies, such as paraspeckles and PML bodies. Perinucleolar caps can be divided into dark and light subtypes, which can also be distinguished by their specific protein contents [55]. To determine whether the nuclear anxA10 foci represent nucleolar caps, we stimulated anxA10GFP-expressing MDCK cells for 2 h with 5 μg/ml actinomycin D (ActD), a drug that induces transcriptional inhibition and cap-formation through DNA intercalation and the inhibition of topoisomerase II. ActD treatment of transiently transfected cells resulted in the efficient recruitment of anxA10GFP to nuclear foci (Fig. 4c). As seen previously with doxorubicin, the visualization of DNA using the intercalating stain DRAQ5 clearly showed that anxA10 translocation to cap structures was not just dependent on DNA-accumulation around nucleoli, as areas of increased DRAQ5 fluorescence did not overlap with the anxA10GFP signal.
To determine whether the ActD-induced anxA10-foci indeed represented perinucleolar caps, we counterstained with antibodies against SFPQ (Splicing factor, proline- and glutamine-rich) and fibrillarin, two known marker-proteins of the light and dark subtypes of nucleolar caps, respectively [55]. After ActD stimulation, the fibrillarin-positive light nucleolar caps (LNCs) were clearly distinct from anxA10GFP and SFPQ-positive caps (Fig. 5a) as were coilin-positive caps and PML-bodies (data not shown). In contrast, anxA10GFP showed a striking co-localization with SFPQ, revealing its compartmentalization in dark nucleolar caps (DNCs). To exclude artifactual translocation into these caps due to anxA10 overexpression or GFP-fusion, we confirmed that the endogenous anxA10 was also enriched in SFPQ-positive DNCs upon ActD incubation (Fig. 5b).
Nucleoplasmic anxA10 is recruited to dark nucleolar caps (DNCs) upon transcriptional inhibition. a Translocation of anxA10GFP. Under unstimulated conditions (upper panel, unstim.), anxA10GFP and polypyrimidine tract binding protein-associated splicing factor (SFPQ, blue) distribute throughout the nucleoplasm, whereas fibrillarin (red) is found in nucleoli. Transcriptional inhibition induced with 5 μg/ml actinomycin D (ActD) for 2 h (lower panel) resulted in translocation of fibrillarin to light nucleolar caps (LNCs), whereas SFPQ was enriched at dark nucleolar caps (DNCs). Note that anxA10GFP co-localizes strongly with SFPQ in DNCs. Inserts depict high magnification images of protein distribution at a single nucleolus. b Enrichment of endogenous anxA10 at DNCs. Endogenous anxA10 in nuclei of MDCK cells treated for 6 h with 5 μg/ml ActD was visualized by immunofluorescence staining (green), and showed extensive co-localization with SFPQ (red). Bar 5 μm, line delineates outline of the nucleus
Previous studies show that SFPQ-recruitment to DNCs is caused by RNA-polymerase II (Pol II) inhibition [55]. To conclusively answer if anxA10, like SFPQ, is recruited to DNCs as a result of Pol II-inhibition, we stimulated cells for 6 h with α-amanitin, a Pol II-specific inhibitor [28] and co-immunofluorescently stained the cells against GFP and either anti-SFPQ or anti-fibrillarin. AnxA10GFP was observed in SFPQ-positive caps and remained clearly separate from fibrillarin-positive LNCs (Fig. 6), confirming that anxA10 recruitment to DNCs is indeed caused by Pol II inhibition. To see whether the RNA Pol II-induced translocation into caps was a feature shared with other annexins, we further tested anxA1- and anxA2GFP, two members of the annexin family with reported nuclear and RNA-associated functions [21, 22, 29, 34], for cap recruitment. No translocation of either anxA1- or anxA2-GFP to nucleolar caps was detected in response to ActD-induced transcriptional inhibition (suppl. Fig. 3).
Translocation of anxA10GFP to DNCs is dependent on RNA Pol II. Immunofluorescent images showing MDCK cells transiently expressing anxA10GFP that were treated with α-amanitin (6 h, 30 μg/ml) and then co-stained for fibrillarin (upper panel) or SFPQ (lower panel). AnxA10GFP is not found in fibrillarin-positive light nucleolar caps but in SFPQ-positive DNCs upon RNA Pol II inhibition. Bar 5 μm
The co-localization of anxA10GFP with DNCs prompted us to examine a correlation with other proteins that are enriched at this compartment. Interestingly, DNCs are strongly enriched with proteins normally found in paraspeckles, a recently discovered nuclear body [12, 55]. We immunostained anxA10GFP-expressing MDCK cells for SFPQ and found that resting cells displayed a co-localization of the GFP-tagged construct with SFPQ-positive foci that were clustered in the nucleoplasm (Fig. 7a, upper panel) and might represent paraspeckles. To confirm this, we also immunostained the anxA10GFP-expressing MDCK cells for paraspeckle component 1 (PSPC1). PSPC1, together with SFPQ and p54NRB, is one of three drosophila behavior/human splicing (DBHS) proteins that are tightly linked to paraspeckle structure and function [13]. In accordance with the previously observed SFPQ co-staining, we detected anxA10GFP in association with PSPC1-positive paraspeckles (Fig. 7a, lower panel). We were also able to visualize endogenous anxA10 in SFPQ-positive foci by immunostaining, thus ruling out an artificial recruitment of the exogenously expressed anxA10GFP (Fig. 7b, upper panel). Profile analysis of a line running through the nuclear region confirmed peak intensity correlation of the SFPQ and anxA10 signals (Fig. 7b, lower panel). Paraspeckle proteins are so far only defined through their co-localization with the canonical DBHS family proteins, anxA10 therefore seems to represent a novel component of this compartment.
AnxA10 localizes to paraspeckles in the nuclei of resting cells. a MDCK cells transiently expressing anxA10GFP were co-stained for the paraspeckle components SFPQ (upper panel, red signal) and PSPC1 (lower panel, red signal). Insets show that nucleoplasmic anxA10GFP foci (green signal) in resting cells co-localize with SFPQ- and PSPC1-positive paraspeckles. b Endogenous anxA10 in nuclei of MDCK cells (upper panel, green signal) was co-stained with SFPQ (upper panel, red signal). The graph shows respective signal intensities along the indicated line through the nuclear region of the cell, covering an 18-μm distance (lower panel). Bar 5 μm; line delineates outline of the nucleus
We next assessed whether upregulating the anxA10 expression level would have an impact on the number of paraspeckles. Comparison of the number of PSPC1-positive nucleolar foci in individual HeLa subclones stably overexpressing GFP or anxA10GFP revealed that, on average, significantly fewer paraspeckles were found in nuclei of cells overexpressing anxA10GFP (Fig. 8).
Overexpression of anxA10GFP reduces paraspeckle numbers in HeLa cells. Quantification of the average number of paraspeckles in individual subclones stably expressing GFP or anxA10GFP. PSPC1-positive foci per nucleus were digitally counted and the differences in the average numbers were statistically analyzed using ANOVA followed by Tukey’s multiple comparison test. *P < 0.05, **P < 0.01, ***P < 0.001
Domain-requirements for anxA10 recruitment to DNCs and paraspeckles
To address which protein region of anxA10 mediates localization to the nuclear sub-compartments, we created GFP-tagged mutants of anxA10 and analyzed them by confocal fluorescence microscopy for DNC co-localization (Fig. 9). We first introduced a mutation into the second annexin-repeat (E146A, anxA10GFP-CM), which is sufficient to abolish the Ca2+-affinity of type II sites [24], but found no effect on DNC-binding of this construct. To test whether the unique N-terminal domain is involved in cap recruitment, we utilized a deletion mutant lacking amino acids 1–16, which only retains the anxA10 core domain (anxA10-core). Co-localization screening again showed no impairment of anxA10-core recruitment to DNCs, after ActD-induced transcriptional inhibition.
AnxA10 recruitment to DNCs is mediated by the proximal part of the core domain. Confocal fluorescent images of MDCK cells transiently expressing various anxA10GF constructs (CM, core, short, and proxcore; see text). Localization to DNCs was determined by counterstaining with antibodies against SFPQ (red) after 2 h of ActD treatment. Note that only the short anxA10 isoform, lacking the N-terminal domain as well as the first and half of the second annexin-repeats, is not enriched at DNCs. Bar 5 μm
The core domain of annexins is a highly conserved structure that is composed of four annexin repeats, forming a tightly packed disc. We were averse to creating anxA10 deletion mutants of this domain because deletions are likely to result in changes to the core conformation and thus possible disruption of interaction sites. We therefore tested the cap-binding abilities of a short anxA10 isoform (anxA10-short), which lacks the N-terminal domain as well as the first and half of the second annexin repeats [44]. Intriguingly, our results showed that the GFP-tagged short isoform of anxA10 was observed in the nucleus, but was not able to translocate to DNCs in response to ActD treatment. Thus, the part of the core sequence not found in the short isoform carries the binding site responsible for cap recruitment. To further test this hypothesis, we created a GFP-tagged construct that only expressed the proximal anxA10 core fragment missing in the short isoform (aa F17-Q134, anxA10-proxcore); this construct was found enriched at ActD-induced DNCs (Fig. 9).
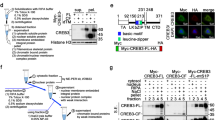
Cap-binding of PSPC1 and SFPQ, two members of the paraspeckle-binding DBHS-family, is mediated by the conserved coiled-coil interaction domains of these proteins; however, this domain was necessary but not sufficient for the recruitment of PSPC1 to paraspeckles, which required the presence of an additional RNA-binding motif [11, 42, 55]. To investigate if a similar paraspeckle binding pattern was observed for anxA10, we tested the above-described anxA10 mutants, including the short anxA10-isoform, for co-localization with paraspeckles using PSPC1 as a marker protein. In these experiments, we used HeLa cells, which are a well-established system for paraspeckle studies and show a distinct paraspeckle-staining pattern [13]. Full-length anxA10GFP (Fig. 10, wt) showed the paraspeckle enrichment already observed in MDCK cells. Identical to our cap-binding experiments, we observed that deletion of the N-terminal domain did not affect paraspeckle-binding (Fig. 10, core). Interestingly though, neither anxA10-short (Fig. 10, short) nor the cap-binding-competent proximal core construct, encompassing the first one and a half annexin repeats (Fig. 10, proxcore), showed paraspeckle recruitment. While the type II-site-mutated CM construct could be detected at paraspeckles, its enrichment appeared less pronounced (Fig. 10, CM). Figure 11a summarizes the domain requirements of anxA10 for DNC- and paraspeckle recruitment in HeLa and MDCK cells, respectively, and indicates that DNC translocation is dependent on a motif present in the proximal core domain, whereas paraspeckle binding requires an additional binding sequence. In order to investigate a correlation of anxA10 paraspeckle localization and enhanced doxorubicin-induced apoptosis, we measured the PARP cleavage upon 24 h doxorubicin treatment in HeLa cells stably expressing the anxA10 mutants. Interestingly, differences in PARP cleavage efficiency were observed. However, irrespective of the subnuclear localization, all mutants tested elicited significantly higher levels of cleaved PARP compared to GFP-expressing control cells. (Fig. 11b). Thus, there seems to be no direct correlation between the presence of anxA10 in paraspeckles and the tumor suppressor activity.
Localization of the various anxA10 mutants in paraspeckles. Confocal immunofluorescence (GFP, green) images of HeLa cells transiently expressing the various anxA10GFP constructs (wt, CM, core, short, and proxcore). Recruitment of anxA10-mutants and the short anxA10 isoform to paraspeckles was determined by co-staining with antibodies against PSPC1 (red). Note that anxA10-CM shows reduced accumulation, and neither anxA10-short nor the anxA10-proxcore is found in paraspeckles. Bar 5 μm
AnxA10 subnuclear localization and tumor suppressor activity are not directly linked. a Schematic representation of the various anxA10 mutants and short I soform. aa amino acids; filled circles represent functional type II Ca2+-binding sites, whereas open circles indicate non-functional type II Ca2+-binding sites. Annexin repeats in the core domain are indicated with roman numerals. C-terminally attached GFP is not shown. The table indicates the presence (+) or absence (−) in paraspeckles and DNCs. b HeLa cells stably expressing GFP or the GFP-tagged anxA10 mutants were treated for 24 h with 5 μM doxorubicin (Doxo). The graph shows mean (±SEM) levels of cleaved PARP as a percentage of the total PARP from three independent experiments. To statistically verify the observed increase in doxorubicin-induced apoptosis between GFP and the respective anxA10GFP construct expressing cells, ANOVA followed by Dunnetts’s multiple comparison test was performed. *P < 0.05, **P < 0.01
Discussion
Members of the annexin protein family are involved in cell differentiation, tumorigenesis, apoptosis, infection, and immunity [14]. With this wide range of functions, it is not surprising that changes in annexin expression coincide with changes in the metastatic behavior of cells. AnxA10 is the latest identified member of the vertebrate family of annexin [35]. While mostly uncharacterized in its biochemical properties and intracellular functions, homozygous deletion and downregulation of anxA10 expression is correlated with poor survival in gastric and bladder cancer patients [26, 32, 36]. In accordance with these findings, anxA10 reintroduction into gastric cancer cells resulted in reduced proliferation, while downregulation in anxA10-expressing epithelial cells facilitated migration and proliferation. AnxA10 has therefore been proposed to fulfill a tumor-suppressor function [26, 27, 32, 36].
Our biochemical characterization of anxA10 indicates that the type II Ca2+-binding site mutations in repeats 1, 3, and 4 drastically affect the Ca2+-mediated membrane-binding properties of anxA10. In contrast to annexins harboring more than one active type II Ca2+-binding site, which typically bind to acidic phospholipids at Ca2+ concentrations in the low micromolar range, anxA10 binding to liposomes containing negatively charged phospholipids only occurred at mM Ca2+ concentrations. Ca2+ concentrations needed for membrane-binding of anxA10 are unlikely to be reached in the cytosol under physiological conditions [4], thus intracellular monomeric anxA10 is unlikely to associate with intracellular membranes in a Ca2+-regulated manner. In this regard, anxA10 is the second member of the annexin family that is not regulated by intracellular Ca2+, a characteristic so far only shown for anxA9, in which all type II sites have been rendered inactive [16]. However, it cannot be ruled out that these annexins might still show Ca2+-independent interaction with cholesterol-rich membrane subdomains, as has been described for anxA2 [18].
The results obtained in the live cell imaging of anxA10GFP argue against any prominent translocation to either membranous compartments or the cytoskeleton, even under conditions such as apoptosis or membrane repair when high Ca2+ levels are reached. It is therefore tempting to speculate that annexins bearing primarily inactive type II Ca2+-binding sites are involved in cellular processes other than membrane organization and trafficking.
The large decrease in clonogenic survival seen upon expression of anxA10 in HeLa cells, which lack endogenous anxA10, clearly shows that anxA10 expression is linked to restricted cellular growth and points to a role of anxA10 as a tumor suppressor, in line with previous findings in cancer cell lines as well as studies correlating anxA10 downregulation with poor prognosis for gastric cancer patients [26, 27, 32, 36]. A putative role in antitumor activities is further strengthened by our observations that HeLa cells ectopically expressing anxA10 are more sensitive to doxorubicin-induced apoptosis.
Despite the fact that nuclear localization has been reported for several vertebrate annexins [14], functions of the annexin family in the nuclear context have been scarcely investigated. While annexins A1, A10, and A11 partially localize to the nucleoplasm [9, 26, 48], the recruitment of other annexins is often transient and related to cell cycle [1, 29] and differentiation events [51] as well as to cellular stress responses [17, 52]. In addition, nuclear enrichment of anxA1 was proposed as a diagnostic marker for several types of carcinomas [29, 63].
The present immunofluorescence studies show that GFP-labeled anxA10 is enriched in the nucleoplasm of resting epithelial cells and, in response to doxorubicin treatment, concentrates in dark nuclear caps. The composition of DNCs shows a striking enrichment of proteins normally associated with an only recently discovered type of nuclear body, the paraspeckle. Paraspeckles are nuclear sub-compartments that localize to the interchromatin space and are formed around two large non-protein coding mRNAs that provide the scaffold to which the structural core proteins SFPQ, p54NRB, and PSPC1, members of the DBHS protein family, are bound [13, 38]. Double immunofluorescent experiments in transcriptionally active cells showed that anxA10GFP co-localizes with the paraspeckle proteins SFPQ and PSPC1 in punctate nucleoplasmic foci. SFPQ co-localization was also observed for the endogenous anxA10. As novel proteins associating with paraspeckles have so far been characterized via their co-localization with one of these intrinsic components [13], we therefore propose anxA10 to be a novel paraspeckle component.
GFP-tagged anxA10 mutants as well as the short isoform of anxA10 first described by Peng et al. [44] enabled us to localize the subnuclear binding properties of anxA10. We found that both DNC- and paraspeckle-binding are mediated by the anxA10 core-domain and independent of the N-terminus. The exclusion of anxA10-short from both nuclear subcompartments indicates that the first two annexin repeats of anxA10, which are missing in this shorter isoform, contain a binding site necessary for this interaction. Interestingly, a construct only expressing these missing repeats was able to localize to DNCs in response to transcriptional inhibition but was not found at paraspeckles. Thus, the DNC-binding sequence located in the first two annexin repeats may be necessary but not sufficient to mediate paraspeckle binding. Our results imply the existence of at least two independent domains necessary for paraspeckle recruitment of anxA10. Intriguingly, these findings are similar to those described for the paraspeckle protein PSPC1. While PSPC1 is effectively recruited to DNCs via its coiled-coil domain [11], recent studies on the structure of DBHS protein heterodimers using PSPC1 truncation mutants revealed that the extended C-terminal coiled-coil motif is also essential for paraspeckle targeting. These domains emanating from DBHS protein dimers are supposed to mediate the formation of longer oligomers, thus driving paraspeckle assembly [11, 42]. In addition, paraspeckle localization requires the presence of functional RNA-binding domains in the DBHS core [7, 55]. Furthermore, most of the novel paraspeckle proteins recently identified in an extensive screen also carry putative RNA binding domains [37]. We therefore speculate that the distal anxA10 core might contain a so far unidentified RNA binding site required for paraspeckle recruitment. The observation that the Ca2+-binding-deficient mutant of anxA10 displayed a less pronounced paraspeckle co-localization might imply a Ca2+-dependent conformational change that might facilitate paraspeckle binding but is not essential for DNC recruitment.
Paraspeckles play a role in the nuclear retention of A-to-I-edited mRNAs [2, 3], and are thought to represent a regulation mechanism for the control of protein expression in differentiation and stress response events [2, 3, 46, 62]. Considering the tumor-suppressor characteristics of anxA10 as well as the currently described functions of paraspeckle- and DNC-enriched proteins, our results imply that anxA10 may fulfill a radically different function than that of other annexins, as a tissue-specific nuclear factor in paraspeckle-regulated expression and differentiation processes.
Although several paraspeckle proteins are related to diseases such as cancer [5, 40, 45, 54, 56], as yet the functions of paraspeckles in the context of cell growth and differentiation remain enigmatic. The enrichment of hyperedited RNAs in paraspeckles has been discussed as a storage pool for transcripts that might function in the stress response [46]. Prolonged stress eventually leads to apoptotic cell death. However, a functional correlation of paraspeckle formation and apoptosis is not established and our mutational analysis of anxA10 suggests no direct correlation of subnuclear localization and pro-apoptotic functions.
The restriction of anxA10 expression to certain epithelial cells argues against a structural role in de novo paraspeckle formation. This is supported by our findings that anxA10 expression negatively affects paraspeckle numbers in otherwise anxA10-negative HeLa cells. Paraspeckles are assembled around the long noncoding RNA NEAT1 [6, 53]. Of the two NEAT isoforms generated by alternative 3′-end processing, NEAT1_2 but not NEAT1_1 appears to be essential for paraspeckle generation. NEAT1 isoform ratios are affected by the essential paraspeckle protein HNRNPK through inhibition of binding of the pre-messenger RNA Cleavage Factor I (CFIm) to NEAT1_1, thereby favoring NEAT1_2 accumulation [37]. It is tempting to speculate that anxA10 might function in balancing NEAT1 isoform ratios, e.g., by regulating the CFIm complex. However, we cannot rule out that anxA10 negatively regulates the assembly of paraspeckles through interaction with paraspeckle proteins essential in paraspeckle formation.
References
Barwise JL, Walker JH (1996) Subcellular localization of annexin V in human foreskin fibroblasts: nuclear localization depends on growth state. FEBS Lett 394:213–216
Chen LL, Carmichael GG (2009) Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear noncoding RNA. Mol Cell 35:467–478
Chen LL, Carmichael GG (2008) Gene regulation by SINES and inosines: biological consequences of A-to-I editing of Alu element inverted repeats. Cell Cycle 7:3294–3301
Clapham DE (2007) Calcium signaling. Cell 131:1047–1058
Clark J, Lu YJ, Sidhar SK, Parker C, Gill S, Smedley D, Hamoudi R, Linehan WM, Shipley J, Cooper CS (1997) Fusion of splicing factor genes PSF and NonO (p54nrb) to the TFE3 gene in papillary renal cell carcinoma. Oncogene 15:2233–2239
Clemson CM, Hutchinson JN, Sara SA, Ensminger AW, Fox AH, Chess A, Lawrence JB (2009) An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell 33:717–726
Dye BT, Patton JG (2001) An RNA recognition motif (RRM) is required for the localization of PTB-associated splicing factor (PSF) to subnuclear speckles. Exp Cell Res 263:131–144
Eberhard DA, Karns LR, VandenBerg SR, Creutz CE (2001) Control of the nuclear-cytoplasmic partitioning of annexin II by a nuclear export signal and by p11 binding. J Cell Sci 114:3155–3166
Farnaes L, Ditzel HJ (2003) Dissecting the cellular functions of annexin XI using recombinant human annexin XI-specific autoantibodies cloned by phage display. J Biol Chem 278:33120–33126
Filipenko NR, Waisman DM (2001) The C terminus of annexin II mediates binding to F-actin. J Biol Chem 276:5310–5315
Fox AH, Bond CS, Lamond AI (2005) P54nrb forms a heterodimer with PSP1 that localizes to paraspeckles in an RNA-dependent manner. Mol Biol Cell 16:5304–5315
Fox AH, Lam YW, Leung AK, Lyon CE, Andersen J, Mann M, Lamond AI (2002) Paraspeckles: a novel nuclear domain. Curr Biol 12:13–25
Fox AH, Lamond AI (2010) Paraspeckles. Cold Spring Harb Perspect Biol 2:a000687
Gerke V, Creutz CE, Moss SE (2005) Annexins: linking Ca2+ signalling to membrane dynamics. Nat Rev Mol Cell Biol 6:449–461
Goebeler V, Ruhe D, Gerke V, Rescher U (2006) Annexin A8 displays unique phospholipid and F-actin binding properties. FEBS Lett 580:2430–2434
Goebeler V, Ruhe D, Gerke V, Rescher U (2003) Atypical properties displayed by annexin A9, a novel member of the annexin family of Ca2+ and lipid binding proteins. FEBS Lett 546:359–364
Han EK, Tahir SK, Cherian SP, Collins N, Ng SC (2000) Modulation of paclitaxel resistance by annexin IV in human cancer cell lines. Br J Cancer 83:83–88
Harder T, Kellner R, Parton RG, Gruenberg J (1997) Specific release of membrane-bound annexin II and cortical cytoskeletal elements by sequestration of membrane cholesterol. Mol Biol Cell 8:533–545
Hayes MJ, Merrifield CJ, Shao D, Ayala-Sanmartin J, Schorey CD, Levine TP, Proust J, Curran J, Bailly M, Moss SE (2004) Annexin 2 binding to phosphatidylinositol 4,5-bisphosphate on endocytic vesicles is regulated by the stress response pathway. J Biol Chem 279:14157–14164
Hayes MJ, Rescher U, Gerke V, Moss SE (2004) Annexin-actin interactions. Traffic 5:571–576
Hirata A, Hirata F (2002) DNA chain unwinding and annealing reactions of lipocortin (annexin) I heterotetramer: regulation by Ca2+ and Mg2+. Biochem Biophys Res Commun 291:205–209
Hirata A, Hirata F (1999) Lipocortin (Annexin) I heterotetramer binds to purine RNA and pyrimidine DNA. Biochem Biophys Res Commun 265:200–204
Jones PG, Moore GJ, Waisman DM (1992) A nonapeptide to the putative F-actin binding site of annexin-II tetramer inhibits its calcium-dependent activation of actin filament bundling. J Biol Chem 267:13993–13997
Jost M, Thiel C, Weber K, Gerke V (1992) Mapping of three unique Ca2+-binding sites in human annexin II. Eur J Biochem 207:923–930
Jost M, Weber K, Gerke V (1994) Annexin II contains two types of Ca2+-binding sites. Biochem J 298(Pt 3):553–559
Kim J, Kim MA, Jee CD, Jung EJ, Kim WH (2009) Reduced expression and homozygous deletion of annexin A10 in gastric carcinoma. Int J Cancer 125:1842–1850
Kim JK, Kim PJ, Jung KH, Noh JH, Eun JW, Bae HJ, Xie HJ, Shan JM, Ping WY, Park WS, Lee JY, Nam SW (2010) Decreased expression of annexin A10 in gastric cancer and its overexpression in tumor cell growth suppression. Oncol Rep 24:607–612
Lindell TJ, Weinberg F, Morris PW, Roeder RG, Rutter WJ (1970) Specific inhibition of nuclear RNA polymerase II by alpha-amanitin. Science 170:447–449
Liu J, Rothermund CA, Ayala-Sanmartin J, Vishwanatha JK (2003) Nuclear annexin II negatively regulates growth of LNCaP cells and substitution of ser 11 and 25 to glu prevents nucleo-cytoplasmic shuttling of annexin II. BMC Biochem 4:10–25
Liu SH, Lin CY, Peng SY, Jeng YM, Pan HW, Lai PL, Liu CL, Hsu HC (2002) Down-regulation of annexin A10 in hepatocellular carcinoma is associated with vascular invasion, early recurrence, and poor prognosis in synergy with p53 mutation. Am J Pathol 160:1831–1837
Lopez-Pedrera C, Villalba JM, Siendones E, Barbarroja N, Gomez-Diaz C, Rodriguez-Ariza A, Buendia P, Torres A, Velasco F (2006) Proteomic analysis of acute myeloid leukemia: identification of potential early biomarkers and therapeutic targets. Proteomics 6(Suppl 1):S293–S299
Lu SH, Chen YL, Shun CT, Lai JN, Peng SY, Lai PL, Hsu HC (2011) Expression and prognostic significance of gastric-specific annexin A10 in diffuse- and intestinal-type gastric carcinoma. J Gastroenterol Hepatol 26:90–97
Meng LX, Li Q, Xue YJ, Guo RD, Zhang YQ, Song XY (2007) Identification of gastric cancer-related genes by multiple high throughput analysis and data mining. Zhonghua Wei Chang Wai Ke Za Zhi 10:169–172
Mickleburgh I, Burtle B, Hollas H, Campbell G, Chrzanowska-Lightowlers Z, Vedeler A, Hesketh J (2005) Annexin A2 binds to the localization signal in the 3′ untranslated region of c-myc mRNA. FEBS J 272:413–421
Morgan RO, Jenkins NA, Gilbert DJ, Copeland NG, Balsara BR, Testa JR, Fernandez MP (1999) Novel human and mouse annexin A10 are linked to the genome duplications during early chordate evolution. Genomics 60:40–49
Munksgaard PP, Mansilla F, Brems Eskildsen AS, Fristrup N, Birkenkamp-Demtroder K, Ulhoi BP, Borre M, Agerbaek M, Hermann GG, Orntoft TF, Dyrskjot L (2011) Low ANXA10 expression is associated with disease aggressiveness in bladder cancer. Br J Cancer 105:1379–1387
Naganuma T, Nakagawa S, Tanigawa A, Sasaki YF, Goshima N, Hirose T (2012) Alternative 3′-end processing of long noncoding RNA initiates construction of nuclear paraspeckles. EMBO J 31:4020–4034
Nakagawa S, Hirose T (2012) Paraspeckle nuclear bodies-useful uselessness? Cell Mol Life Sci 69:3027–3036
Nelson MR, Creutz CE (1995) Comparison of the expression of native and mutant bovine annexin IV in Escherichia coli using four different expression systems. Protein Expr Purif 6:132–140
Notari M, Neviani P, Santhanam R, Blaser BW, Chang JS, Galietta A, Willis AE, Roy DC, Caligiuri MA, Marcucci G, Perrotti D (2006) A MAPK/HNRPK pathway controls BCR/ABL oncogenic potential by regulating MYC mRNA translation. Blood 107:2507–2516
Oliver FJ, de la Rubia G, Rolli V, Ruiz–Ruiz MC, de Murcia G, Murcia JM (1998) Importance of poly(ADP-ribose) polymerase and its cleavage in apoptosis. Lesson from an uncleavable mutant. J Biol Chem 273:33533–33539
Passon DM, Lee M, Rackham O, Stanley WA, Sadowska A, Filipovska A, Fox AH, Bond CS (2012) Structure of the heterodimer of human NONO and paraspeckle protein component 1 and analysis of its role in subnuclear body formation. Proc Natl Acad Sci USA 109:4846–4850
Patsos G, Germann A, Gebert J, Dihlmann S (2010) Restoration of absent in melanoma 2 (AIM2) induces G2/M cell cycle arrest and promotes invasion of colorectal cancer cells. Int J Cancer 126:1838–1849
Peng SY, Ou YH, Chen WJ, Li HY, Liu SH, Pan HW, Lai PL, Jeng YM, Chen DC, Hsu HC (2005) Aberrant expressions of annexin A10 short isoform, osteopontin and alpha-fetoprotein at chromosome 4q cooperatively contribute to progression and poor prognosis of hepatocellular carcinoma. Int J Oncol 26:1053–1061
Pordzik S, Petrovici K, Schmid C, Kroell T, Schweiger C, Kohne CH, Schmetzer H (2011) Expression and prognostic value of FAS receptor/FAS ligand and TrailR1/TrailR2 in acute myeloid leukemia. Hematology 16:341–350
Prasanth KV, Prasanth SG, Xuan Z, Hearn S, Freier SM, Bennett CF, Zhang MQ, Spector DL (2005) Regulating gene expression through RNA nuclear retention. Cell 123:249–263
Raynal P, Pollard HB (1994) Annexins: the problem of assessing the biological role for a gene family of multifunctional calcium- and phospholipid-binding proteins. Biochim Biophys Acta 1197:63–93
Raynal P, van Bergen en Henegouwen PM, Hullin F, Ragab-Thomas JM, Fauvel J, Verkleij A and Chap H (1992) Morphological and biochemical evidence for partial nuclear localization of annexin 1 in endothelial cells. Biochem Biophys Res Commun 186:432-439
Rescher U, Zobiack N, Gerke V (2000) Intact Ca2+-binding sites are required for targeting of annexin 1 to endosomal membranes in living HeLa cells. J Cell Sci 113:3931–3938
Reynolds RC, Montgomery PO, Hughes B (1964) Nucleolar “caps” produced by actinomycin D. Cancer Res 24:1269–1277
Rick M, Ramos Garrido SI, Herr C, Thal DR, Noegel AA, Clemen CS (2005) Nuclear localization of Annexin A7 during murine brain development. BMC Neurosci 6:25–38
Sacre SM, Moss SE (2002) Intracellular localization of endothelial cell annexins is differentially regulated by oxidative stress. Exp Cell Res 274:254–263
Sasaki YT, Ideue T, Sano M, Mituyama T, Hirose T (2009) MENepsilon/beta noncoding RNAs are essential for structural integrity of nuclear paraspeckles. Proc Natl Acad Sci USA 106:2525–2530
Schiffner S, Zimara N, Schmid R, Bosserhoff AK (2011) P54nrb is a new regulator of progression of malignant melanoma. Carcinogenesis 32:1176–1182
Shav-Tal Y, Blechman J, Darzacq X, Montagna C, Dye BT, Patton JG, Singer RH, Zipori D (2005) Dynamic sorting of nuclear components into distinct nucleolar caps during transcriptional inhibition. Mol Biol Cell 16:2395–2413
Sui Y, Yang Z, Xiong S, Zhang L, Blanchard KL, Peiper SC, Dynan WS, Tuan D, Ko L (2007) Gene amplification and associated loss of 5′ regulatory sequences of CoAA in human cancers. Oncogene 26:822–835
Thastrup O, Cullen PJ, Drobak BK, Hanley MR, Dawson AP (1990) Thapsigargin, a tumor promoter, discharges intracellular Ca2+ stores by specific inhibition of the endoplasmic reticulum Ca2+-ATPase. Proc Natl Acad Sci USA 87:2466–2470
Thiel C, Weber K, Gerke V (1991) Characterization of a Ca2+-binding site in human annexin II by site-directed mutagenesis. J Biol Chem 266:14732–14739
Tzima E, Trotter PJ, Orchard MA, Walker JH (2000) Annexin V relocates to the platelet cytoskeleton upon activation and binds to a specific isoform of actin. Eur J Biochem 267:4720–4730
Tzima E, Trotter PJ, Orchard MA, Walker JH (1999) Annexin V binds to the actin-based cytoskeleton at the plasma membrane of activated platelets. Exp Cell Res 251:185–193
Xu Y, Man X, Lv Z, Li D, Sun Z, Chen H, Wang Z, Luo Y, Xu H (2012) Loss of heterozygosity at chromosomes 1p35-pter, 4q, and 18q and protein expression differences between adenocarcinomas of the distal stomach and gastric cardia. Hum Pathol 43:2308–2317
Zhang Z, Carmichael GG (2001) The fate of dsRNA in the nucleus: a p54(nrb)-containing complex mediates the nuclear retention of promiscuously A-to-I edited RNAs. Cell 106:465–475
Zhu F, Xu C, Jiang Z, Jin M, Wang L, Zeng S, Teng L, Cao J (2010) Nuclear localization of annexin A1 correlates with advanced disease and peritoneal dissemination in patients with gastric carcinoma. Anat Rec (Hoboken) 293:1310–1314
Zobiack N, Gerke V, Rescher U (2001) Complex formation and submembranous localization of annexin 2 and S100A10 in live HepG2 cells. FEBS Lett 500:137–140
Acknowledgments
We thank Annette Janning for technical assistance. Rabbit polyclonal anti-annexin A10 antibody was generously provided by Reginald Morgan (Department of Biochemistry and Molecular Biology, University of Oviedo, Spain). This work was supported by grants from the Interdisciplinary Clinical Research Centre of the University of Münster (IZKF, RE2/017/10) and the German Research Foundation (DFG, RE2611/2-1, SFB 629/A1) to V.G. and U.R.
Conflict of interest
The authors declare no competing financial interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Quiskamp, N., Poeter, M., Raabe, C.A. et al. The tumor suppressor annexin A10 is a novel component of nuclear paraspeckles. Cell. Mol. Life Sci. 71, 311–329 (2014). https://doi.org/10.1007/s00018-013-1375-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-013-1375-4