Abstract
Sprouty2 is an important inhibitor of cell proliferation and signal transduction. In this study, we found a bimodal expression of Sprouty2 protein during cell cycle progression after exit from quiescence, whereas elevated Sprouty4 expression in the G1 phase stayed high throughout the rest of the cell cycle. Induction of the mitogen-activated protein kinase via activated Ras was crucial for increased Sprouty2 expression at the G0/G1 transition. Following the first peak, accelerated proteasomal protein degradation caused a transient attenuation of Sprouty2 abundance during late G1. Since the decline in its expression was abolished by dominant negative c-Cbl and the timely restricted interaction between Sprouty2 and c-Cbl disappeared at the second peak of Sprouty2 expression, we conclude that the second phase in the cell cycle-specific expression profile of Sprouty2 is solely dependent on ubiquitination by c-Cbl. Our results suggest that Sprouty2 abundance is the result of strictly coordinated activities of Ras and c-Cbl.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cell proliferation is a highly regulated process initiated as a coordinated response to a combination of signals generated by the presence of growth factors. Amplification and transduction of these signals are requirements to enable the receiving cell to exit from quiescence (G0) and progress through an organized sequence of distinct phases, called the cell cycle.
To avoid inappropriate signalling, several negative feedback loops which attenuate and terminate the cellular stimulation induced by growth signals are implemented. One of these negative feedback loops includes the members of the Sprouty (Spry) protein family. Spry was discovered as an antagonist of receptor tyrosine kinase (RTK)-mediated signalling in Drosophila [1]. Subsequent studies demonstrated that Spry proteins are involved in regulatory pathways counteracting growth factor-induced processes in several organisms including Xenopus [2], chicken [3, 4], zebrafish [5], mouse [6] and humans [7]. Also mammalian Spry proteins function mainly as inhibitors of RTK-mediated processes, and their expression has been shown to repress lung branching morphogenesis [7, 8], angiogenesis [9, 10], cell growth [11–14] and migration [10, 12–14].
In normal cells many control mechanisms are involved in regulating the expression of the Spry proteins. Growth factor-mediated increases in Spry mRNA levels [9, 11, 12, 15, 16] and the observation that Spry proteins are expressed mainly at sites of excessive FGF signalling during organogenesis [3, 4, 8, 17, 18] led to the postulation of a negative feedback loop. Additionally, several mechanisms prevent inappropriate Spry expression by influencing its protein stability. The E3 ubiquitin ligases c-Cbl, SIAH2 and NEDD4 were recently shown to interact with Spry2 and in consequence Spry2 is ubiquitinated and degraded by the proteasome [19–22].
In this study we investigated the expression of Spry2 and Spry4 after initiating quiescent cells to progress through the cell cycle. The data presented reveal an unexpected biphasic expression profile of Spry2 during cell cycle progression. The results also demonstrate a major role of Ras-activated MAPK induction and c-Cbl-mediated protein degradation in controlling the temporal fluctuations of Spry2 expression.
Materials and methods
Plasmid constructions and generation of recombinant adenoviruses
All Ras-expressing plasmids used were based on constructs kindly provided by Dr. Rojas (Instituto de Salud Carlos III, Madrid, Spain). The coding sequences of human H-, K- and N-Raswt and K-RasG12V were subcloned into the pADlox vector. Using PCR site-directed mutagenesis H-RasG12V, N-RasQ61R, K-RasS17N, H-RasS17N, N-RasS17N, K-RasS35T and K-RasY40C were constructed. Using vectors kindly provided by Dr. Langdon (University of Western Australia, Crawley, Australia), c-Cblwt and c-CblΔ70Z proteins tagged with haemagglutinin (HA-c-Cblwt and HA-c-CblΔ70Z) were cloned into pADlox via BamH1 sites. Using appropriately designed primers, YFP and CFP were amplified from pEYFP-N1 and pECFP-C1 vectors (Clontech) as templates and matching restriction sites were introduced into pADlox, before full-length HA-cCblwt and hSpry2wt were fused in frame to the 3′ end of YFP or CFP vectors to obtain YFP-HA-c-Cblwt and CFP-hSpry2. All constructs were verified by restriction digestion and sequencing.
Recombinant viruses were produced as described previously [23] and used at a multiplicity of infection (MOI) of 50 unless indicated otherwise.
Cell culture and cell treatments
Normal embryonic lung fibroblasts (WI-38 at passage 16) were purchased from ATCC (Rockville, MD) and maintained in DMEM supplemented with 10% FCS (Fisher Scientific). For cell cycle analysis, WI-38 cells were serum-deprived for 3 days and released from quiescence by adding medium containing 20% serum.
Of 15 NSCLC cell lines analysed, 10 were established at our institute from surgical specimens, as follows: one histologically confirmed adenocarcinoma (VL-1), seven squamous cell carcinomas (VL-3 and VL-5 to VL-10) and two large-cell carcinomas (VL-2 and VL-4). The cell lines were used at passage numbers between 15 and 30. Six of the ten cell lines (VL-1 to VL-4, VL-9 and VL-10) were derived from primary tumours and four (VL5 to VL8) from lymph node metastases. Additionally, five adenocarcinoma cell lines (A-427, A-549, Calu-3, Calu-6 and SK-LU-1) were purchased from ATCC (Rockville, MD). Of the 15 cell lines, 6 harboured a mutated K-Ras (VL-2, VL-4, A-427, A-549, Calu6, SK-LU-1). The mutation analyses have been described in detail by Sutterluty et al. [13].
FACS analysis was performed as described previously [23]. The proteasomal inhibitors LLnL and MG132 (Sigma, St Louis, MO) were added to a final concentration of 100 μM. All growth factors used were applied at a final concentration of 10 ng/ml. FGFs and IGF-1 were purchased from Strathmann Biotec (Hamburg, Germany), EGF from Sigma, and PDGF-AA from Chemicon International (Temecula, CA).
MAPK was inhibited by incubating the cells with the MEK inhibitor U0126 at a final concentration of 20 μM for 10 h. For measurement of protein half-lives, protein translation was inhibited by the addition of 10 μg/ml cycloheximide (CHX) for the indicated times.
Phosphatase treatment
Cell lysates were incubated for 30 min at 37°C with 1 U of shrimp alkaline phosphatase (Roche). Control incubations were performed in the absence of phosphatase.
Antibodies, western blotting, immunoprecipitation
The antisera against hSpry2 and hSpry4 were produced and purified as described previously [13]. hSpry4 was raised against NH2-terminal 220 amino acids of human Spry4 and the specificity of the antibodies was verified (data not shown). Antibodies against the following proteins were used: pERK1/2, p110PI3K (Cell Signaling, Danvers, MA), total ERK1/2 (Acris Antibodies, Hiddenhausen, Germany), β-actin (Novus Biologicals, Littleton, CO), AU5 (Bethyl Laboratories, Montgomery, TX), and cyclin A (H-432), cyclin B1 (GNS1), cyclin D1 (M-20), HA epitope (12CA5) (Santa Cruz Biotechnology, Santa Cruz, CA).
Immunoblotting was performed as described previously [13]. Signals were quantified using ImageQuant software (Molecular Dynamics, Sunnyvale, CA). The immunoprecipitation procedure was as previously described [23].
RNA extraction and northern blotting
RNA preparation and northern blotting procedures followed published methods [24]. As probes we used the coding sequence of hSpry2, hSpry4, ribosomal protein S6 (rPS6) and GAPDH.
Fluorescence resonance energy transfer
At 48 h prior to analysis WI-38 cells were infected at a MOI of 5. Fluorescence resonance energy transfer (FRET) analysis was performed as described previously [25]. For quantitative analysis calculated FRET images were exported and analysed using ImageQuant software. Line scans were performed and the area under the curve was measured.
Results
Spry2 protein shows a biphasic expression through the cell cycle
As an initial experiment, we analysed Spry2 and Spry4 expression during cell cycle progression after serum-deprived WI-38 cells were initiated to commit to cell division. At selected time points cells were harvested and cell synchrony was monitored by flow cytometry and immunoblotting using antibodies against cyclins (Fig. 1a). Serum withdrawal resulted in an efficient block in G0 and more than 70% of the cells re-entered a new cell cycle in response to serum addition. At the 4 h time point increased cyclin D1 levels showed that cells had progressed through G1 phase. At 22 h after release, G1/S transition was indicated by a strong increase in cyclin A protein levels although the DNA content was not considerably augmented. After 28 h the majority of cells were in S phase. About 32 h after serum release characteristic accumulation of cells with doubled DNA content and a peak in cylin B1 protein levels showed that the released cells were in G2/M phase. Then the cells underwent mitosis and the cell population was again in G1 phase 42 h after the serum induction (Fig. 1a).
Spry2 and Spry4 expression after cell cycle induction of serum-deprived WI-38 cells. WI-38 cells were synchronized in the G0/G1 phase by culture in medium without serum. They were released from quiescence by adding medium containing 20% FCS. a At the indicated time points, cells were analysed by FACS. In parallel isolated total protein was separated by SDS-PAGE, and probed with antibodies recognizing cyclin D1, cyclin A and cyclin B1 proteins. b The same samples were probed with Spry2- and Spry4-specific antibodies. Equal loading was confirmed using β-actin as a control. Spry expression levels were determined by densitometry analysis and normalized to β-actin. Starved cells were set arbitrarily to 1. c In parallel, cytoplasmic RNA was extracted from part of the cells and northern blotting was performed using 32P-labelled cDNA probes specific for Spry2, Spry4 and rpS6. Since intensity signals derived from hybridization with rpS6 changed, methylene blue staining and hybridization with GAPDH were added as further loading controls. The expression level ratios of Spry2 and Spry4 were calculated and normalized to rpS6 and GAPDH, respectively. The mean values of both standardizations are indicated. Starved cells were set arbitrarily at 1 in each case. d Calculated Spry2 protein and mRNA expression levels during the cell cycle of WI-38. Each column represents the mean of at least three independent experiments including the standard deviation
The total abundance of Spry2 and Spry4 varied specifically as cells progressed through the cell cycle. While antibodies detected Spry2 and Spry4 proteins in logarithmically growing cells, the corresponding bands were not present in quiescent cells (Fig. 1b). Additionally, northern blot analysis revealed that Spry2 and Spry4 mRNA expression was almost undetectable in serum-deprived cells (Fig. 1c). In early G1 phase as cells emerged from quiescence, both Spry2 and Spry4 mRNA and protein levels were strongly induced (Fig. 1b, c). Subsequently, Spry4 protein levels increased during G1 phase and remained high throughout the rest of the cell cycle, while induction of Spry2 in early G1 phase was followed by a substantial reduction in the protein levels at the G1/S transition. During S and G2 phases, Spry2 protein levels were again elevated and then fell when cells entered the next G1 phase (Fig. 1b). As illustrated in Fig. 1d, these results reveal a previously unreported bimodal expression pattern of Spry2 protein during cell-cycle progression after exit from quiescence.
In contrast to the observed differences in Spry2 and Spry4 protein abundance through the cell cycle, Spry2 and Spry4 mRNA expression patterns resembled each other. After a distinct induction in early G1 phase, the mRNA levels fell slightly (1.6-fold) and showed only modest variations as cells progressed further through the cell cycle. These findings indicate that the regulatory mechanisms responsible for the characteristics of Spry2 and Spry4 protein expression profiles as cells progress from late G1 to S and G2 phase are different and are mostly posttranscriptional, while Spry2 and Spry4 at the G0/G1 phase are mainly induced at the mRNA level and are probably controlled by similar mechanisms.
Growth factor-mediated upregulation of Spry2 expression correlates with MAPK activation
To explore the molecular pathways responsible for the cell cycle-specific expression of Spry2 and Spry4 proteins, we first focused on the requirements for their induction at the G0/G1 boundary. According to several reports, Spry expression is induced in response to mitogenic signals. Thus, we investigated the potential of several growth factors—mainly of the FGF family—to increase Spry expression and to activate MAPK and/or PI3K cascades.
WI-38 cells were mitogen-deprived for 72 h and subsequently growth factors were added for 5 h. While Spry2 expression was induced in response to FCS, EGF, FGF2 and FGF9, Spry4 protein was clearly elevated only after stimulation with FCS (Fig. 2a). Consistent with the observed upregulations at the G0/G1 transition (compare Fig. 1b and Fig. 1c), mitogen-induced expression of Spry2 and Spry4 proteins was paralleled by increased mRNA levels (Fig. 2b), implying that Spry expression in response to growth factors is mainly increased at the mRNA level. Since the degree of Spry mRNA increase is about twofold less than the quantitative differences observed at the protein level (compare also Fig. 2c), we cannot exclude the possibility that additional control mechanisms at the protein level intensify activation of mRNA expression.
Growth factor-mediated induction of Spry protein and mRNA expression in human WI-38 fibroblasts. Normal human WI-38 fibroblasts were serum-deprived for 3 days before the indicated growth factors were added for 5 h (a, b), and 5 and 20 min (d), respectively. Protein lysates were prepared for immunoblotting (a, d) using the indicated antibodies. mRNA was analysed via northern blot (b) using 32P-labelled full-length Spry2, Spry4 and rPS6 as probes. The expression level ratios of Spry were calculated after densitometric analysis normalized to either β-actin (a) or rPS6 and GAPDH (b). Serum-deprived cells were set to 1. c Summary of three or four experiments. The means and SDs of calculated and normalized Spry2 expression levels are shown
In parallel to Spry expression studies, the influence of the external signals on activation of MAPK and PI3K pathways were tested by monitoring phosphorylation of ERK and ribosomal protein S6 (rPS6) 5 min and 20 min after growth factor addition (Fig. 2d). This analysis of RTK-mediated signalling pathways revealed that FCS, EGF, FGF2 and FGF9 were able not only to increase Spry2 abundance, but also to induce MAPK pathway strongly and immediately (after 5 min). The other tested mitogens FGF5, KGF, FGF10, FGF18, PDGF and IGF failed to induce Spry2 expression and immediate ERK phosphorylation. A weak activation of ERK signals after 20 min was observed in response to all tested factors. In contrast, activation of the PI3K pathway did not coincide with Spry2 expression. Immediate phosphorylation of rPS6 was observed in response to FCS, EGF and FGF2, and 20 min after mitogen supply additionally FGF9 and IGF induced PI3K-connected pathways. Thus we conclude that exclusive activation of PI3K fails to induce Spry protein expressions.
These results indicate that activation of MAPK and PI3K pathways is not sufficient to induce Spry4 expression, while mitogen-dependent upregulation of Spry2 abundance strongly coincides with MAPK activation.
Ras-mediated MAPK activation induces Spry2 expression
Transduction of growth factor signals usually involves activation of Ras proteins as a key event. In order to examine whether the mitogen-induced Spry abundance is a direct consequence of activated Ras signalling, we expressed the constitutively activated, oncogenic Ras forms H-RasG12V, K-RasG12V and N-RasQ61R in WI-38 cells (Fig. 3). Additionally, recombinant adenoviruses bearing dominant negative forms of the Ras proteins (H-RasS17N, K-RasS17N or N-RasS17N) were generated. As a control, cells were infected with recombinant adenoviruses expressing LacZ. The cells were either serum-deprived or cultivated in medium containing 10% serum. Adenoviruses were added to the cells at the time of serum removal or medium change.
The influence of K-Ras on expression of Spry2 and Spry4. a Serum-deprived and logarithmically growing WI-38 cells were infected using adenoviruses expressing different oncogenic Ras proteins. LacZ and the respective dominant-negative RasS17N adenoviruses were included in the experiment. Cells were harvested after 3 days and equal amounts of protein were separated via SDS-PAGE, transferred onto a nitrocellulose membrane and immunoblotting was performed using the indicated antibodies. A representative immunoblot of two independent experiments is shown. Protein levels were quantified by densitometric analyses and normalized to β-actin and serum-starved LacZ-infected cells. b Endogenous Spry protein expression was analysed in 15 NSCLC-derived tumour cell lines. All cells were serum-deprived for 2 days before harvesting. Equal amounts of protein were used for immunoblotting. After densitometric analysis the protein levels were normalized to β-actin. WI-38 cells were arbitrarily set to 1. The results are presented as a box and whiskers diagram and statistical analyses were performed using the Mann–Whitney U test. *P < 0.05. c Serum-deprived WI-38 cells were infected with adenoviruses expressing the indicated Ras mutations: upper row activation of Ras/MAPK was measured via immunoblotting using phospho-ERK-specific antibody; lower row binding of Ras to PI3K was determined via AU5 immunoprecipitation and subsequent immunoblotting using p110PI3K antibodies. d In parallel, expression of Spry2 and Spry4 was determined in serum-deprived WI-38 cells (left panel) and in logarithmically growing WI-38 cells (right panel) infected with the indicated adenoviruses via immunoblotting using specific antibodies. The Spry2 and Spry4 ratios were calculated via densitometric analysis and normalized to β-actin and LacZ-infected cells, respectively. e WI-38 cells were serum-deprived and infected with adenoviruses expressing lacZ or K-RasG12V. At 10 h prior to harvesting MEK inhibitor U0126 or the organic solvent DMSO were added to the cells. Immunoblotting was performed using the indicated antibodies (⊥ serum-deprived cells, log logarithmically growing cells)
Immunoblot analysis of mitogen-deprived cells expressing the dominant active forms of Ras revealed that Spry2 and Spry4 protein levels were elevated in response to activation of Ras-mediated signalling (Fig. 3a) but differed in the degree of induction. While the abundance of Spry2 in cells expressing either N-RasQ61R, K-RasG12V or H-RasG12V was about eightfold higher than in the LacZ-treated reference cells, Spry4 protein levels increased only two- to threefold in response to Ras activation. Dominant-negative H-RasS17N, K-RasS17N and N-RasS17N reduced the expression of Spry2 and Spry4 proteins especially in logarithmically growing cells (Fig. 3a). These findings demonstrate that all three activated Ras family members elevate Spry2 and to a lesser extent also Spry4 expression, and indicate a central role of Ras in the upregulation of Spry2 and Spry4 as cells exit quiescence.
Based on these results, we sought to determine if Spry protein expression correlated with constitutively active K-Ras mutations in cell lines. Therefore Spry2 and Spry4 expression levels were determined in 15 serum-depleted NSCLC-derived cell lines, 6 of which harboured a K-Ras mutation [13]. For normalization, serum-arrested normal human lung fibroblasts WI-38 were chosen. All of the cell lines expressed Spry proteins but levels differed considerably. As illustrated in Fig. 3b, the levels of both Spry2 and Spry4 were clearly and significantly upregulated in the cell lines harbouring activating K-Ras mutations. Consistent with the quantitative difference in activation of Spry2 and Spry4 in response to expression of activated Ras, the average Spry2 expression in cells harbouring an activating mutation in the K-Ras gene was ninefold higher than the levels in K-Ras wt cells, while Spry4 levels differed only about sixfold between these two groups of cell lines. These results emphasize the importance of Ras activation in regulation of Spry proteins.
To test if Ras-mediated activation of Spry2 expression is dependent on signalling via MAPK pathway, viruses expressing constitutively active K-Ras versions carrying mutations in the effector loop in such a way as to exclusively bind Raf (K-RasG12V,T35S) or PI3K (K-RasG12V,Y40C) were generated [26]. At 48 h after infection, the effects of their expression on the MAPK and PI3K pathways were analysed. In accordance with the literature, K-RasG12V activated both pathways, while K-RasG12V,T35S failed to bind p110PI3K and K-RasG12V,Y40C was unable to activate ERK (Fig. 3c). Additionally, viruses expressing dominant-negative K-RasS17N and LacZ were included as controls. In serum-deprived cells (Fig. 3d, left panel) and logarithmically growing cells (Fig. 3d, right panel) the K-Ras variants able to activate ERK phosphorylation clearly induced Spry2 expression, while activated K-Ras exclusively signalling via PI3K pathway had no influence on Spry2 expression. Spry4 was also induced by the K-Ras mutants known to activate the MAPK pathway. In addition K-RasG12V,Y40C caused a moderate elevation in Spry4 protein levels. To confirm the importance of MAPK activation for Ras-dependent induction of Spry2 expression, serum-deprived cells were infected with adenoviruses expressing K-RasG12V and treated with the MEK inhibitor U0126 in order to inhibit MAPK activation. In line with the data generated using K-Ras effector mutants, constitutively active K-Ras failed to induce Spry2 levels when MAPK activation was inhibited (Fig. 3e), demonstrating that MAPK activation is essential for Ras-mediated elevation of Spry2 expression.
Downregulation of Spry2 expression during G1 phase involves protein degradation
To investigate more fully the decline in Spry2 expression at the G1/S boundary, we first visualized its abundance during G1 phase progression at several time points. Spry2 protein levels increased as cells progressed through early G1 phase and peaked between 6 and 8 h after cell cycle induction of quiescent cells (Fig. 4a). Subsequently, the Spry2 levels declined until the cells entered S phase at about 24 h as indicated by induction of cyclin A expression. Again, Spry4 expression remained elevated after reaching its plateau at about 6 h. Since the mRNA expression profiles indicated that this decrease in Spry2 expression was mainly controlled at the posttranscriptional level (Fig. 1), we sought to determine if differences in protein stability were responsible for the decline in Spry2 protein levels observed after the peak at 8 h. Therefore serum-deprived cells were serum-induced for 4 and 10 h before protein synthesis was blocked by the addition of CHX. The protein half-life in early G1 phase was about 2 h, but the protein stability was reduced (half-life 1.3 h) later in G1 phase (Fig. 4b). These findings indicate that regulation of protein degradation is an important mechanism reducing Spry2 expression during later G1 phase.
Influence of proteasome-mediated degradation on Spry protein expression between G0 and S phase. WI-38 cells were serum-deprived for 3 days to mediate a cell cycle arrest in G0/G1. Cell cycle entry was induced by medium containing 20% FCS. a Total protein was isolated at the indicated time points after serum addition. Equal amounts of the protein lysates were separated by SDS-PAGE and blotted onto nitrocellulose membrane. Spry-specific antibodies were used to detect the levels of Spry2 and Spry4 proteins. Cyclin A and β-actin were used as controls. b At the time points 4 h (left panel) and 10 h (right panel) after serum induction CHX was added to the cells. Total protein was isolated at the indicated times and equal amounts of protein were analysed by immunoblotting. A representative blot is shown. Densitometric data for Spry2 protein levels were normalized to β-actin and half-lives were calculated from three independent experiments using ImageQuant and GraphPad Prism software. c At 6 h after cell cycle release from quiescence, 100 μM of the proteasome inhibitors LLnL and MG-132 (dissolved in DMSO) were added for 3, 6 or 9 h, and compared to DMSO-treated WI-38 cells. The cells were harvested at the indicated times, processed for immunoblotting and stained with Spry2- and Spry4-specific antibodies. Note the appearance of a Spry2 band migrating more slowly (arrow) after proteasome inhibitor treatment. The ratios of the abundances of Spry2 and Spry4 proteins were calculated by densitometric analysis and normalized to β-actin and quiescent, untreated cells, respectively. d Extracts from cells harvested 15 h after serum induction and 9 h after addition of proteasome inhibitors were either mock-treated or incubated with shrimp alkaline phosphatase. (D DMSO-treated cells, L LLnL-treated cells, M MG-132-treated cells)
The next experiment sought to clarify if the disappearance of Spry2 protein was due to proteasomal degradation (Fig. 4c). To this end serum-deprived cells were released for 6 h before the proteasome inhibitors LLnL and MG-132 were added for 3, 6 and 9 h. The control cells were mock-treated with the solvent DMSO. In line with the previous experiments, in the controls Spry2 protein levels fell as cells progressed through G1 phase (compare lanes 3, 6 and 9 of Fig. 4c), indicating that DMSO had no influence on Spry2 expression. Treatment with both proteasome inhibitors increased Spry2 protein levels as early as after 3 h. Additionally, the continued presence of LLnL and MG-132 resulted in the accumulation of a band migrating more slowly. The observation that after treatment with shrimp alkaline phosphatase the electrophoretic mobility of the Spry2 accumulated in the presence of proteasomal inhibitors shifted almost completely to the most rapidly migrating form (Fig. 4d) indicates that predominantly a phosphorylated form of Spry2 is degraded by the proteasome.
In contrast to their effect on Spry2, there was no influence of the proteasome inhibitors on Spry4 protein levels. These findings indicate that protein degradation via the proteasome is an important mechanism in the regulation of Spry2 expression during the cell cycle.
Fluctuations in Spry2 expression through G1/S phases are mediated by c-Cbl
Cell cycle-specific degradation of proteins is mostly mediated by regulated ubiquitination via E3 ligases [27]. One of the reported interactions of Spry2, possibly responsible for the observed decline in G1 phase, involves the E3 ligase c-Cbl. To test the influence of c-Cbl on Spry2 expression, we modulated c-Cbl activity by overexpressing a c-Cbl wt protein and a c-Cbl mutant (c-CblΔ70Z) described as dominant-negative with respect to degradation of EGFR [28]. Therefore adenoviruses expressing HA-c-Cblwt and HA-c-CblΔ70Z proteins were generated.
To test the influence of c-Cbl on the cell cycle-specific expression pattern of Spry2, WI-38 cells were arrested in G0 phase. At 24 h after serum removal the cells were infected with either a control virus expressing LacZ or the viruses expressing the c-Cbl proteins. After 48 h cells were initiated to exit quiescence and at different time points during the cell cycle harvested for immunoblot analysis. In parallel, FACS analysis was performed 0 and 24 h after serum addition and revealed that the expressed proteins had no pronounced effect on cell cycle progression (Fig. 5a). In addition, independently of introduced c-Cbl, cyclin A expression was induced 22 h after release of the arrested cells (Fig. 5b). Expression of c-Cblwt and c-CblΔ70Z was comparable at all phases of the cell cycle, but weak in quiescent cells (Fig. 5b).
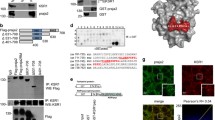
Influence of c-Cbl on Spry2 cell cycle-specific expression. Serum-deprived WI-38 cells were infected with the indicated viruses and induced by adding medium containing 20% serum. a Cells were trypsinized and prepared for FACS analysis at 0 h and after 24 h of induction. b In parallel the cells were harvested at the indicated time points representing the indicated phases of the cell cycle. Total protein was isolated, separated by SDS-PAGE and blotted onto nitrocellulose membrane and probed using the indicated antibodies. The ratios of Spry2 and Spry4 were calculated by densitometric analysis and normalized to β-actin. Arrested control cells were arbitrarily set to 1 (asterisk nonspecific band recognized by the 12CA5 antibody, arrows specific bands for the HA-cCbls). c The calculated Spry2 protein levels of WI-38 cells infected with viruses expressing lacZ, c-Cbl wt or dominant-negative cCblΔ70Z mutant are compared. Each column represents the means of at least three independent experiments and includes the standard deviation. *P < 0.05 (Mann–Whitney U test). d Cells infected with viruses coding for LacZ or the dominant-negative cCblΔ70Z were serum-induced for 22 h before CHX was added for the indicated times. Protein half-lives were calculated after densitometric analysis of two independent results (wt cCblwt, Δ dominant-negative cCblΔ70Z, Z LacZ control virus)
In the G0 and early G1 phases (after 6 h), Spry2 expression was influenced neither by increased expression of the c-Cblwt protein nor by the dominant-negative c-CblΔ70Z.
In arrested cells, the average Spry2 protein levels were low and increased about 15- to 20-fold when cells progressed through early G1 phase, although slight variability (see Fig. 5c) was detected. In contrast, in cells harvested at the G1/S transition (22 h after serum addition), the reduced Spry2 expression observed in control cells was significantly lower in the presence of additional c-Cblwt protein (p = 0.028). Expression of the dominant-negative mutant was able to override the usually observed decline of Spry2 in late G1 phase (Fig. 5b). As in cells in the G1 phase, the average expression of Spry2 was about 17-fold greater than in arrested cells, while lacZ cells expressed less than half the amount of Spry2 protein. In accordance with the previous observation that the diminished expression of Spry2 protein in G1 phase is mainly caused by increased protein degradation, measurements of Spry2 protein half-lives during the G1/S transition (22 h after serum addition) revealed that expression of dominant-negative c-Cbl caused stabilization of Spry2 (Fig. 5d). As cells progressed through S phase the strong effects of both c-Cbl proteins on Spry2 expression were substantially reduced. In cells infected with the control and the c-Cblwt virus, Spry2 levels were again elevated, while the levels in cells expressing the dominant-negative c-Cbl remained stably high.
Since Spry4 does not interact with c-Cbl [29, 30], variations in c-Cbl activity had no influence on its protein expression. Independent of c-Cbl expression, Spry4 abundance was increased in early G1 phase and remained elevated at later time points.
These findings demonstrate that G1 phase-specific degradation of Spry2 by c-Cbl is crucial for mediating the biphasic Spry2 expression pattern during the cell cycle.
Spry2 interaction with c-Cbl is limited to G1 phase
To investigate if the restricted influence of c-Cbl on the Spry2 levels is connected to limited availability of c-Cbl protein in certain cell cycle phases, we first analysed its expression throughout the cell cycle. As shown in Fig. 6a, c-Cbl expression showed only minimal fluctuations during the different cell cycle phases. Only in the S/G2 phase, when Cbl-mediated degradation had almost no influence on Spry2 levels, were the c-Cbl levels slightly elevated. We therefore sought to determine if the two proteins interact during the respective time periods during G1 phase using FRET analyses (Fig. 6b). To this end we synthesized adenoviruses expressing CFP-tagged Spry2 and a YFP-c-Cblwt fusion protein. Quiescent WI-38 cells were double-infected with both adenoviruses 24 h before induction of the cell cycle. To stabilize the Spry2-cCbl interaction, LLnL was added 2 h before the FRET analyses. At both G1 time points (10 and 22 h after serum addition) we calculated a prominent and significantly increased FRET signal in all cells coexpressing CFP-Spry2 and YFP-c-Cbl (Fig. 6b, c). The same combination failed to produce any detectable signal later in the cell cycle when cells progress through the G2 phase (measured 32 h after cell cycle induction; Fig. 6b, c). In serum-starved cells the percentage of cells expressing detectable amounts of the combination CFP-Spry2 and YFP-c-Cbl were clearly reduced (about 1% compared to about 15% at the three other time points), but these cells exhibited an evident FRET signal (Fig. 6b), although the intensity of the signal was noticeably weaker than that measured at the time points during G1 phase (Fig. 6b, c). Therefore we conclude that c-Cbl and Spry2 primarily interact during G1 phase of the cell cycle.
FRET analysis of Spry2 and c-Cbl interaction during specific cell cycle phases. WI-38 cells were serum-deprived for 3 days and synchronized in G0/G1 phase. a By addition of medium containing 20% serum, cells were released and analysed by immunoblotting at the indicated times. b At 48 h prior to FRET analyses, serum-starved cells were infected with adenoviruses expressing CFP-labelled Spry2 and YFP-labelled HA-c-Cblwt. Quiescent cells were induced using 20% serum for the indicated times. In all cases the proteasome inhibitor LLnL was added to a final concentration of 50 μM 2 h prior to the FRET analyses. Images were acquired with filters for CFP (cyan), YFP (yellow), and for FRET using identical microscope and camera settings. After determination of the correction factor for CFP (donor) and YFP (acceptor), the intensity of the FRET signal (green) was calculated [25]. Representative examples are shown. c Intensity profiles (lower panels) of the FRET signals along an arbitrary line across the cell (upper panels) presented as calculated grey level (y-axis) versus pixel distance (x-axis). For statistical analysis the areas under the curve of the line scans were quantified using ImageQuant. Each bar represents the mean ± SD of at least eight images from three independent experiments. *P < 0.05 (Mann–Whitney U test)
The timely overlap of the physical interaction between c-Cbl and Spry2 and stabilization of Spry2 by dominant-negative c-Cbl (see Fig. 5) strongly suggest that ubiquitination by c-Cbl mediates G1-specific degradation of Spry2.
Discussion
Initiation and passage of cells through the cell cycle is strictly regulated by a coordinated network of signal transduction and cell-cycle progression pathways. Failures at key decision checkpoints during G0/G1/S transitions lead to uncontrolled multiplication of cells, and thus can induce tumour formation [31]. Sprouty proteins are negative regulators of signal transduction as well as cell proliferation and are therefore considered to play a role as tumour suppressors in several cancers. Consequently, Spry proteins have been found deregulated in various tumours [13, 32–36].
In this study, we assessed the regulation of Spry2 and Spry4 expression during the cell cycle after exit from quiescence. Expression of a protein in a certain time window within the cell cycle usually means that its particular tasks are exclusively needed and desired at that particular time. In this regard, a distinguishable cell cycle expression profile of Spry2 and Spry4 is likely to reflect distinct functions during cell cycle progression. According to its expression, Spry4 functions immediately after serum induction throughout the cell cycle while Spry2 is only needed in early G1 phase and after cells have entered S phase. During late G1 phase Spry2 tasks are not needed for or even obstruct cell cycle progression. Correspondingly, enforced Spry2 expression interferes with cell proliferation of different cell lines including WI-38 cells [13] and reduced Spry2 expression in WI-38 cells released from quiescence accelerates progression through G1 phase, while expression of Spry4 fails to inhibit proliferation of logarithmically growing WI-38 (unpublished observations). Accordingly, other studies have demonstrated that Spry4 is not able to interfere with proliferation of pancreatic tumour cells [37] or prostate cancer cells [34]. Therefore we conclude that during late G1 phase Spry2 exerts a specific inhibitory function on cell proliferation. Consistent with this finding, a previous study has shown that Spry2 inhibits G1/S phase transition by decreasing phosphorylation of Akt via PTEN [38]. Since the abolishing of Spry2 degradation in late G1 phase by expression of dominant-negative Cbl does not affect G1/S transition (Fig. 5), it is likely that the mechanisms involving Spry2 during late G1 phase can be compensated for by another Cbl substrate.
Concerning the mechanisms responsible for regulation of Spry proteins, the results presented allow functional attribution of certain control mechanisms to distinct phases of the cell cycle. In agreement with the proposed negative feedback loop, elevated Spry2 and Spry4 expression immediately after serum induction is directly or indirectly dependent on Ras activation. Expression of both proteins was induced by any of the three oncogenic Ras family members. Correspondingly, NSCLC-derived cell lines carrying mutated K-Ras express significantly more Spry2 and Spry4 protein than cells harbouring wt K-Ras. In line with these findings, knock-in of an oncogenic RasG12D variant in murine fetal lungs led to an increase in Spry2 and Spry4 expression [39]. Furthermore, our findings demonstrate that although both Spry2 and Spry4 expression is upregulated by Ras, the mechanisms involved are different.
When quiescent cells were incubated with growth factors, the increase in Spry2 protein levels caused by the tested mitogens coincided with immediate activation of ERK phosphorylation. Accordingly Spry2 and Spry4 are among the upregulated mRNAs when activated Raf-1 is expressed in human fibroblasts [40]. Other studies have also shown that growth factor-induced Spry2 expression can be reduced by inhibitors of MEK-1 [15, 16]. Similar to the previous results showing that an inhibitor of the PI3K pathway fails to reduce Spry2 expression, IGF was not able to augment Spry2 protein levels but induced PI3K-mediated signalling. Our results obtained by ectopic expression of activated K-Ras mutated in the effector loop confirmed that Ras-induced Spry2 expression is solely mediated by Raf/ERK signals and further indicated that PI3K effector pathways have no impact on Spry2 expression.
In contrast, Spry4 expression was clearly induced solely in the presence of serum. Even growth factors which activated ERK and PI3K pathways failed to augment Spry4 protein levels. Therefore we conclude that different or additional signals are required for elevation of Spry4 protein levels. A more complex regulation of Spry4 is also indicated by published data showing that Wnt signalling [41, 42] is able to induce Spry4 expression.
Mitogen-mediated upregulation of Spry expressions was observed at the mRNA level, indicating that mechanisms at the mRNA level play an essential role in the regulation of Spry levels in the early cell cycle phases. Published promoter studies have shown that in WI-38 cells the 4-kb promoter element proximal to the Spry2 transcription start is not sensitive to growth factors [43]. Therefore it is likely that the increase in Spry2 mRNA expression at the G0/G1 boundary is dependent on additional regulatory elements elsewhere in the genome, in the intron or within the mRNA of Spry2.
Later in the cell cycle, regulated protein degradation by the proteasome is mainly responsible for the fluctuations in the profile of Spry2 expression levels. Spry4 levels remained high throughout G1/S/G2 phase and insensitive to inhibition of proteasomal degradation. Degradation via the proteasome is largely dependent on conjugation of ubiquitin to the lysines of the substrate proteins by so-called E3 ubiquitin ligases [27]. c-Cbl protein has a proven E3 ubiquitin ligase activity [44] and has been shown to interact with Spry2 but not with Spry4 [29, 30]. In accordance with several reports that degradation of Spry2 by c-Cbl is dependent on growth factor-mediated phosphorylation of tyrosine55 [20, 45], in serum-deprived cells, the interaction of c-Cbl with Spry2 was only weak and modulated c-Cbl activity had no influence on Spry2 expression. In contrast, the G1-specific decline in Spry2 protein was abrogated by the expression of dominant-negative c-CblΔ70Z. But when cells progressed through S phase, Spry2 levels were again insensitive to c-Cbl activity, indicating that only the transient G1-specific decline in Spry2 was mediated by cell cycle-specific ubiquitin ligase activity of c-Cbl. Future experiments will elucidate what mechanisms are responsible for phase-restricted degradation of Spry2 by c-Cbl.
Modifications to Spry2 may be involved in the temporal restriction of its degradation. Phosphorylation by MNK-1 has been reported to stabilize Spry2 and interfere with its degradation by c-Cbl [46]. Therefore it is possible that a conformational change in the complex between Spry2 and c-Cbl upon MNK-1 phosphorylation in late S phase is responsible for the second phase of Spry2 expression. Furthermore, the interaction between c-Cbl and Spry2 can be abrogated by the action of a phosphatase. PP2A and SHP2 are phosphatases known to interact with Spry2 [47, 48]. Although none of the two enzymes has a known S-phase-specific function, both molecules are potential candidates for regulation of the Spry2-cCbl interactions. PP2A is a serin/threonine phosphatase which, like c-Cbl, binds to a Spry2 motif including amino acids 50 to 60 [48]. Therefore it is conceivable that PP2A replaces c-Cbl and thereby stabilizes Spry2. In contrast, SHP2 is a tyrosine phosphatase which can remove the phosphate at tyrosine 55 of Spry2 [47] thereby weakening the interaction between c-Cbl and Spry2.
It is also possible that modifications to c-Cbl could regulate its interaction with Spry2. In the case of the EGF receptor, activation of Cdc42 enhances sequestration of Cbl proteins in a Cdc42-p85Cool-1 complex, which causes stabilization of the receptor [49]. A similar mechanism resulting in diminished levels of c-Cbl available for Spry2 interaction could also be responsible for stabilization of Spry2 when cells enter S phase.
Since the induction of mRNA expression in early G1 phase was less pronounced than the upregulation of the Spry proteins measured in parallel, we conclude that in other phases of the cell cycle protein degradation is also involved in adjusting the appropriate Spry expression level. Our findings indicate that c-Cbl activity has a negligible influence on Spry levels during G0 and early G1 phase, indicating that other control mechanisms, such as ubiquitination by Siah2 or Nedd4, could be more important for reducing Spry levels in these stages of the cell cycle.
In summary, our results demonstrate that the coordinated action of Ras- and c-Cbl-mediated pathways is responsible for a newly discovered bimodal expression of Spry2 during the cell cycle.
References
Hacohen N, Kramer S, Sutherland D, Hiromi Y, Krasnow MA (1998) Sprouty encodes a novel antagonist of FGF signaling that patterns apical branching of the Drosophila airways. Cell 92:253–263
Nutt SL, Dingwell KS, Holt CE, Amaya E (2001) Xenopus Sprouty2 inhibits FGF-mediated gastrulation movements but does not affect mesoderm induction and patterning. Genes Dev 15:1152–1166
Chambers D, Mason I (2000) Expression of sprouty2 during early development of the chick embryo is coincident with known sites of FGF signalling. Mech Dev 91:361–364
Minowada G, Jarvis LA, Chi CL, Neubuser A, Sun X, Hacohen N, Krasnow MA, Martin GR (1999) Vertebrate Sprouty genes are induced by FGF signaling and can cause chondrodysplasia when overexpressed. Development 126:4465–4475
Furthauer M, Lin W, Ang SL, Thisse B, Thisse C (2002) Sef is a feedback-induced antagonist of Ras/MAPK-mediated FGF signalling. Nat Cell Biol 4:170–174
de Maximy AA, Nakatake Y, Moncada S, Itoh N, Thiery JP, Bellusci S (1999) Cloning and expression pattern of a mouse homologue of drosophila sprouty in the mouse embryo. Mech Dev 81:213–216
Tefft JD, Lee M, Smith S, Leinwand M, Zhao J, Bringas P Jr, Crowe DL, Warburton D (1999) Conserved function of mSpry-2, a murine homolog of Drosophila sprouty, which negatively modulates respiratory organogenesis. Curr Biol 9:219–222
Mailleux AA, Tefft D, Ndiaye D, Itoh N, Thiery JP, Warburton D, Bellusci S (2001) Evidence that SPROUTY2 functions as an inhibitor of mouse embryonic lung growth and morphogenesis. Mech Dev 102:81–94
Impagnatiello MA, Weitzer S, Gannon G, Compagni A, Cotten M, Christofori G (2001) Mammalian sprouty-1 and -2 are membrane-anchored phosphoprotein inhibitors of growth factor signaling in endothelial cells. J Cell Biol 152:1087–1098
Lee SH, Schloss DJ, Jarvis L, Krasnow MA, Swain JL (2001) Inhibition of angiogenesis by a mouse sprouty protein. J Biol Chem 276:4128–4133
Gross I, Bassit B, Benezra M, Licht JD (2001) Mammalian sprouty proteins inhibit cell growth and differentiation by preventing ras activation. J Biol Chem 276:46460–46468
Lee CC, Putnam AJ, Miranti CK, Gustafson M, Wang LM, Vande Woude GF, Gao CF (2004) Overexpression of sprouty 2 inhibits HGF/SF-mediated cell growth, invasion, migration, and cytokinesis. Oncogene 23:5193–5202
Sutterluty H, Mayer CE, Setinek U, Attems J, Ovtcharov S, Mikula M, Mikulits W, Micksche M, Berger W (2007) Down-regulation of sprouty2 in non-small cell lung cancer contributes to tumor malignancy via extracellular signal-regulated kinase pathway-dependent and -independent mechanisms. Mol Cancer Res 5:509–520
Yigzaw Y, Cartin L, Pierre S, Scholich K, Patel TB (2001) The C terminus of sprouty is important for modulation of cellular migration and proliferation. J Biol Chem 276:22742–22747
Ozaki K, Kadomoto R, Asato K, Tanimura S, Itoh N, Kohno M (2001) ERK pathway positively regulates the expression of Sprouty genes. Biochem Biophys Res Commun 285:1084–1088
Sasaki A, Taketomi T, Wakioka T, Kato R, Yoshimura A (2001) Identification of a dominant negative mutant of Sprouty that potentiates fibroblast growth factor- but not epidermal growth factor-induced ERK activation. J Biol Chem 276:36804–36808
Chambers D, Medhurst AD, Walsh FS, Price J, Mason I (2000) Differential display of genes expressed at the midbrain–hindbrain junction identifies sprouty2: an FGF8-inducible member of a family of intracellular FGF antagonists. Mol Cell Neurosci 15:22–35
Mason JM, Morrison DJ, Basson MA, Licht JD (2006) Sprouty proteins: multifaceted negative-feedback regulators of receptor tyrosine kinase signaling. Trends Cell Biol 16:45–54
Nadeau RJ, Toher JL, Yang X, Kovalenko D, Friesel R (2007) Regulation of Sprouty2 stability by mammalian Seven-in-Absentia homolog 2. J Cell Biochem 100:151–160
Rubin C, Litvak V, Medvedovsky H, Zwang Y, Lev S, Yarden Y (2003) Sprouty fine-tunes EGF signaling through interlinked positive and negative feedback loops. Curr Biol 13:297–307
Wong ES, Lim J, Low BC, Chen Q, Guy GR (2001) Evidence for direct interaction between Sprouty and Cbl. J Biol Chem 276:5866–5875
Edwin F, Anderson K, Patel TB (2010) HECT domain-containing E3 ubiquitin ligase Nedd4 interacts with and ubiquitinates Sprouty2. J Biol Chem 285:255–264
Sutterluty H, Chatelain E, Marti A, Wirbelauer C, Senften M, Muller U, Krek W (1999) p45SKP2 promotes p27Kip1 degradation and induces S phase in quiescent cells. Nat Cell Biol 1:207–214
Sutterluety H, Bartl S, Doetzlhofer A, Khier H, Wintersberger E, Seiser C (1998) Growth-regulated antisense transcription of the mouse thymidine kinase gene. Nucleic Acids Res 26:4989–4995
Macheiner D, Gauglhofer C, Rodgarkia-Dara C, Grusch M, Brachner A, Bichler C, Kandioler D, Sutterluty H, Mikulits W, Schulte-Hermann R, Grasl-Kraupp B (2009) NORE1B is a putative tumor suppressor in hepatocarcinogenesis and may act via RASSF1A. Cancer Res 69:235–242
Rodriguez-Viciana P, Warne PH, Khwaja A, Marte BM, Pappin D, Das P, Waterfield MD, Ridley A, Downward J (1997) Role of phosphoinositide 3-OH kinase in cell transformation and control of the actin cytoskeleton by Ras. Cell 89:457–467
Reed SI (2003) Ratchets and clocks: the cell cycle, ubiquitylation and protein turnover. Nat Rev Mol Cell Biol 4:855–864
Thien CB, Walker F, Langdon WY (2001) RING finger mutations that abolish c-Cbl-directed polyubiquitination and downregulation of the EGF receptor are insufficient for cell transformation. Mol Cell 7:355–365
Haglund K, Schmidt MH, Wong ES, Guy GR, Dikic I (2005) Sprouty2 acts at the Cbl/CIN85 interface to inhibit epidermal growth factor receptor downregulation. EMBO Rep 6:635–641
Mason JM, Morrison DJ, Bassit B, Dimri M, Band H, Licht JD, Gross I (2004) Tyrosine phosphorylation of Sprouty proteins regulates their ability to inhibit growth factor signaling: a dual feedback loop. Mol Biol Cell 15:2176–2188
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100:57–70
Kwabi-Addo B, Wang J, Erdem H, Vaid A, Castro P, Ayala G, Ittmann M (2004) The expression of Sprouty1, an inhibitor of fibroblast growth factor signal transduction, is decreased in human prostate cancer. Cancer Res 64:4728–4735
McKie AB, Douglas DA, Olijslagers S, Graham J, Omar MM, Heer R, Gnanapragasam VJ, Robson CN, Leung HY (2005) Epigenetic inactivation of the human sprouty2 (hSPRY2) homologue in prostate cancer. Oncogene 24:2166–2174
Wang J, Thompson B, Ren C, Ittmann M, Kwabi-Addo B (2006) Sprouty4, a suppressor of tumor cell motility, is down regulated by DNA methylation in human prostate cancer. Prostate 66:613–624
Lo TL, Yusoff P, Fong CW, Guo K, McCaw BJ, Phillips WA, Yang H, Wong ES, Leong HF, Zeng Q, Putti TC, Guy GR (2004) The ras/mitogen-activated protein kinase pathway inhibitor and likely tumor suppressor proteins, sprouty 1 and sprouty 2 are deregulated in breast cancer. Cancer Res 64:6127–6136
Fong CW, Chua MS, McKie AB, Ling SH, Mason V, Li R, Yusoff P, Lo TL, Leung HY, So SK, Guy GR (2006) Sprouty 2, an inhibitor of mitogen-activated protein kinase signaling, is down-regulated in hepatocellular carcinoma. Cancer Res 66:2048–2058
Jaggi F, Cabrita MA, Perl AK, Christofori G (2008) Modulation of endocrine pancreas development but not beta-cell carcinogenesis by Sprouty4. Mol Cancer Res 6:468–482
Edwin F, Singh R, Endersby R, Baker SJ, Patel TB (2006) The tumor suppressor PTEN is necessary for human sprouty 2 mediated inhibition of cell proliferation. J Biol Chem 281:4816–4822
Shaw AT, Meissner A, Dowdle JA, Crowley D, Magendantz M, Ouyang C, Parisi T, Rajagopal J, Blank LJ, Bronson RT, Stone JR, Tuveson DA, Jaenisch R, Jacks T (2007) Sprouty-2 regulates oncogenic K-ras in lung development and tumorigenesis. Genes Dev 21:694–707
Courtois-Cox S, Genther Williams SM, Reczek EE, Johnson BW, McGillicuddy LT, Johannessen CM, Hollstein PE, MacCollin M, Cichowski K (2006) A negative feedback signaling network underlies oncogene-induced senescence. Cancer Cell 10:459–472
Taniguchi K, Ayada T, Ichiyama K, Kohno R, Yonemitsu Y, Minami Y, Kikuchi A, Maehara Y, Yoshimura A (2007) Sprouty2 and Sprouty4 are essential for embryonic morphogenesis and regulation of FGF signaling. Biochem Biophys Res Commun 352:896–902
Winn RA, Marek L, Han SY, Rodriguez K, Rodriguez N, Hammond M, Van Scoyk M, Acosta H, Mirus J, Barry N, Bren-Mattison Y, Van Raay TJ, Nemenoff RA, Heasley LE (2005) Restoration of Wnt-7a expression reverses non-small cell lung cancer cellular transformation through frizzled-9-mediated growth inhibition and promotion of cell differentiation. J Biol Chem 280:19625–19634
Ding W, Bellusci S, Shi W, Warburton D (2003) Functional analysis of the human Sprouty2 gene promoter. Gene 322:175–185
Thien CB, Langdon WY (2001) Cbl: many adaptations to regulate protein tyrosine kinases. Nat Rev Mol Cell Biol 2:294–307
Hall AB, Jura N, DaSilva J, Jang YJ, Gong D, Bar-Sagi D (2003) hSpry2 is targeted to the ubiquitin-dependent proteasome pathway by c-Cbl. Curr Biol 13:308–314
DaSilva J, Xu L, Kim HJ, Miller WT, Bar-Sagi D (2006) Regulation of sprouty stability by Mnk1-dependent phosphorylation. Mol Cell Biol 26:1898–1907
Hanafusa H, Torii S, Yasunaga T, Matsumoto K, Nishida E (2004) Shp2, an SH2-containing protein-tyrosine phosphatase, positively regulates receptor tyrosine kinase signaling by dephosphorylating and inactivating the inhibitor Sprouty. J Biol Chem 279:22992–22995
Lao DH, Yusoff P, Chandramouli S, Philp RJ, Fong CW, Jackson RA, Saw TY, Yu CY, Guy GR (2007) Direct binding of PP2A to Sprouty2 and phosphorylation changes are a prerequisite for ERK inhibition downstream of fibroblast growth factor receptor stimulation. J Biol Chem 282:9117–9126
Wu WJ, Tu S, Cerione RA (2003) Activated Cdc42 sequesters c-Cbl and prevents EGF receptor degradation. Cell 114:715–725
Acknowledgments
This study was supported by Herzfelder’sche Familienstiftung 07 and the Fonds der Stadt Wien für Innovative Interdisziplinäre Krebsforschung, project number K-7/07.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mayer, CE., Haigl, B., Jantscher, F. et al. Bimodal expression of Sprouty2 during the cell cycle is mediated by phase-specific Ras/MAPK and c-Cbl activities. Cell. Mol. Life Sci. 67, 3299–3311 (2010). https://doi.org/10.1007/s00018-010-0379-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0379-6