Abstract
Presynaptic differentiation takes place over three interrelated acts involving the biogenesis and trafficking of molecular complexes of active zone material, the “trapping” or stabilization of active zone sites, and the subsequent development of mature synapses. Although the identities of proteins involved with establishing presynaptic specializations have been increasingly delineated, the exact functional mechanisms by which the active zone is assembled remain poorly understood. Here, we discuss a theoretical model for how the trapping stage of presynaptic differentiation might occur in developing neurons. We suggest that subsets of active zone proteins containing polyglutamine domains undergo concentration-dependent prion-like conversions as they accumulate at the plasma membrane. This conversion might serve to aggregate the proteins into a singular structure, which is then able to recruit scaffolding agents necessary for regulated synaptic transmission. A brief informatics analysis in support of this ‘Q’ assembly hypothesis—across commonly used models of synaptogenesis—is presented.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Non-pathogenic prion proteins are able to assume two different functionally relevant conformations, one of which is transmissible to other proteins of the same type and causes self-aggregation. Nearly 6 years ago, Si et al. [1] reported that a brain-specific isoform of the Aplysia californica cytoplasmic polyadenylation element binding protein (CPEB) has prion-like features that are likely to contribute to how it normally maintains spatially restricted changes in long-term synaptic strength (i.e., a universally accepted mechanism for how animals learn and remember). They proposed a model wherein CPEB accumulation in response to synaptic activity triggers a semi-permanent, wholesale conversion of all of the local CPEB molecules to the ‘prion’ state. To our knowledge, no other brain-specific proteins with an ability to undergo prion modification as part of their normal function have been suggested since, although it is becoming clear that prion-like transitions in protein folding might serve as a post-translational modification in general biology [2].
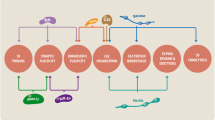
Aggregate forming domains in prion proteins very often exhibit amino acid (AA) sequence compositions that are biased towards glutamine (Q) or asparagine (N) residues. A search of the UniProt Protein Knowledgebase reveals that mammalian presynaptic active zone (AZ) and postsynaptic density (PSD) proteins are defined by a narrow Q content range of only ~5–6% (average ± SD, AZ, 5.8 ± 0.02%, n = 20; PSD, 4.9 ± 0.02%, n = 17). Against this backdrop, however, we find that there are three structurally important AZ proteins in humans (h) and mice (m) with putative prion-determinant regions: Bassoon, Piccolo, and MALS1. These specific ~60–100 AA regions are highly Q enriched (25–50%), are nestled in general areas of the protein where the majority of the protein’s total Q residues are concentrated, and appear to border coiled-coil and PDZ protein interaction motifs (Fig. 1, top 3 panel sets). Such regions are not evident in any other AZ or PSD proteins including RIM1α, CAST/ERC2, MUNC13-1, liprin-α1, CASK, Mint1, Shank1-3, Homer1-3, SAPAP1-4, the MAGUKS PSD-93, PSD-95, SAP97 or SAP102, BEGAIN, MAGI-2, or GKAP (Swiss-Prot and TrEMBL). Bassoon and Piccolo are known to be homo- and heterodimerizing binding partners [3–10], and regions of the proteins that harbor Q domains have been shown, inexplicably, to form dense aggregates upon over-expression in neurons and heterologous cells [6–8]. In the case of Bassoon—where this phenomenon has been best documented—painstaking analysis of various deletion constructs reveals that AAs ~2000–2600 (i.e., Bassoon’s first potential prionic area, Fig. 1) are crucial for clustering behavior [7, 8]. While full-length Bassoon and Bassoon (2,088–2,564) exhibit aggregation, Bassoon (∆2,088–2,564) displays only diffuse intracellular labeling [7]. Intriguingly, fragments of Bassoon containing the first prionic area form agglomerates that remain functional in so far as they are still able to recruit other AZ constituents [8]. Conversely, fragments not containing this area are unable to be anchored to the cytomatrix associated with the AZ [9] and render a significant fraction of synapses inactive [10].
The amino acid sequence for each AZ protein was evenly divided into ten segments. The percentage of all of the protein’s Q residues that could be found in each individual segment is plotted as a temperature map. By and large, local Q domains (specified on the top of each map) were found in “hot” spots of the proteins where overall amino acid composition was biased towards Q residues. For perspective, known interaction motifs for each AZ protein are also delineated in white. The amino acid length of each protein is shown on the bottom right. All regional sizes are approximate. h human, m mouse, dm drosophila, ce C. elegans
Regarding MALS1, little is known about its presynaptic function beyond participation in a tripartite complex with CASK and Mint1 via each member’s L27 domain [11]. It has not been determined, for instance, if Piccolo and MALS1 bind one another via their consensus type I PDZ domains; ligands for these modules have yet to be identified in the AZ proper. But, given the proteins’ proximity to one another and the large stretch of AAs dedicated to the MALS1 PDZ motif, this would not be an unreasonable expectation. PDZ-PDZ interactions have already been found to occur between PSD-95 and nNOS that mediate nNOS’s synaptic localization and generally have been hypothesized to organize macromolecular signaling complexes at synaptic membranes [12].
Drosophila melanogaster (dm) does not have orthologs of Bassoon or Piccolo, despite the fact that the majority of proteins that contribute to human presynaptic architecture are well conserved in insects [13]. Another coiled-coil protein—Bruchpilot—has been suggested to take on some of Bassoon’s functions at the fly AZ along with the fly ortholog of RIM1, dmRIM (comparatively, a much larger protein showing regions of homology with mammalian Piccolo [14, 15]). Bruchpilot, in some ways reminiscent of the role that Bassoon plays at photoreceptor ribbons [16], is absolutely essential for the structural integrity of the AZ as evidenced by the lack of T bars in brp mutants [17, 18]. dmRIM, in contrast to mammalian RIM1α, does not function as a Rab3 effector, hinting at a different biological role for the protein in invertebrates versus vertebrates [19]. Both Bruchpilot and dmRIM exhibit several putative prion-determinant regions that are unusually rich in Q or Q/N (33–52%). As was the case for Bassoon-Piccolo-MALS1, these specific domains are found in larger areas of the protein where the majority of the protein’s Q’s are clustered (Fig. 1, fourth set of panels). Q domains are not evident in the Drosophila MALS1 ortholog, Veli (TrEMBL). The N-terminal domain of Bruchpilot displays significant sequence homology with ERC [17]. Although not experimentally demonstrated, Bruchpilot and dmRIM are thought to be potential binding partners in flies as ERC and RIM1α are in mammals (D. Wagh, personal communication). Of note, Bruchpilot has also been recently shown to form ordered, clear vesicle dotted agglomerates upon over-expression. These free-floating AZs appear to be unconnected to plasma membrane [20].
Another invertebrate model organism C. elegans does not have direct homologs to Bassoon and Piccolo, but does have a RIM homolog named UNC-10. Like dmRIM, worm UNC-10 does not function as a Rab3 effector [19], again suggesting that the role of invertebrate RIM is fundamentally different from that of vertebrate RIM. SYD-2 (liprin-α), as demonstrated by electron microscopy, is centered within the very base of the worm AZ and is thought to be the main scaffolding organizer in worm presynaptic assembly [21, 22]. Its Drosophila counterpart, too, appears to surround the core of Bruchpilot-defined AZs [23]. Interestingly, UNC-10 and SYD-2 show a number of potential prion-determinant regions enriched for Q or Q/N (20–58%) that are located in larger areas with biased Q/N expression (Fig. 1, last set of panels). Similar to mammalian and insect Q domain containing proteins, UNC-10 and SYD-2 have been found to associate [24]. Q domains are not evident in the C. elegans MALS1 homolog, LIN-7 (TrEMBL, NCBI).
The presence of putative prion-like ‘Q’ or ‘Q/N’ domains in subsets of interacting AZ proteins in humans, mice, flies, and worms implies a conserved mechanism of presynaptic trapping, whereby intra- and inter-protein Q aggregates might lock and define the location of formal presynaptic macromolecular assemblies. In this model, the steady membrane accumulation of the aforementioned AZ molecules would eventually trigger a protein folding switch that—in a regulated manner—serves to establish the core of the AZ and to recruit a critical mass of peripheral scaffolding agents needed for neurotransmission. The signals that would initiate this synaptogenic event would vary, but possibly include Ca2+ considering its role in neurodevelopment and the presence of C2 domains in each AZ protein subset. The virtual insolubility during biochemical purification and lack of mobility in response to activity exhibited by Bassoon, Piccolo, Bruchpilot, etc. and their ubiquitous presence at central, retinal, and peripheral synapses, indicate that these proteins are universal structural anchors for synapses across neurobiology [3, 25] (please also see Chiang et al., Soc Neurosci Abstr #497.4, 2009). Q domains might serve as one motif by which they achieve their function, although we would caution that there is as of yet only circumstantial evidence that the synaptic proteins surveyed in the current work possess canonical prion activity or the capacity to form higher order oligomers in response to physiologically relevant stimuli. Moreover, it has not escaped our attention that should such a ‘Q’ aggregation mechanism prove true for presynaptic trapping/assembly, neurons would need to be equipped with, presumably, an elaborate set of molecular machinery that would: (1) regulate the timing, place, and extent of prion switching to avoid a self-destructive positive feedback loop, and (2) be able to deconstruct AZ masses when necessary. In a nod to these considerations, Buchner, Nieratschker, and colleagues have recently reported the existence of a kinase in Drosophila (i.e., SRPK79D) that prevents premature aggregation of Bruchpilot [20].
In closing, we would emphasize that the current contribution represents a hypothesis designed to trigger new experimental work in the synaptic biology field. In light of the data presented herein, along with noting the remarkable degree of conservation of Bruchpilot and RIM Q domains across Drosophila subspecies (e.g., D. simulans, D. sechellia, D. yakuba, D. erecta, D. ananassae, D. pseudoobscura, D. willistoni, D. mojavensis, D. virilis), and other insects that have been sequenced (i.e., Anopheles gambiae, Nasonia vitripennis) (NCBI), we believe that it is a hypothesis worth carefully exploring.
References
Si K, Lindquist S, Kandel ER (2003) A neuronal isoform of the aplysia CPEB has prion-like properties. Cell 115:879–891
Shorter J, Lindquist S (2005) Prions as adaptive conduits of memory and inheritance. Nat Rev Genet 6:435–450
tom Dieck S, Sanmarti-Vila L, Langnaese K, Richter K, Kindler S, Soyke A, Wex H, Smalla KH, Kämpf U, Fränzer JT, Stumm M, Garner CC, Gundelfinger ED (1998) Bassoon, a novel zincfinger CAG/glutamine-repeat protein selectively localized at the active zone of presynaptic nerve terminals. J Cell Biol 142:499–509
Zhai RG, Vardinon-Friedman H, Cases-Langhoff C, Becker B, Gundelfinger ED, Ziv NE, Garner CC (2001) Assembling the presynaptic active zone: a characterization of an active one precursor vesicle. Neuron 29:131–143
Fujimoto K, Shibasaki T, Yokoi N, Kashima Y, Matsumoto M, Sasaki T, Tajima N, Iwanaga T, Seino S (2002) Piccolo, a Ca2+ sensor in pancreatic beta-cells. Involvement of cAMP-GEFII.Rim2.Piccolo complex in cAMP-dependent exocytosis. J Biol Chem 277:50497–50502
Fenster SD, Kessels MM, Qualmann B, Chung WJ, Nash J, Gundelfinger ED, Garner CC (2003) Interactions between Piccolo and the actin/dynamin-binding protein Abp1 link vesicle endocytosis to presynaptic active zones. J Biol Chem 278:20268–20277
Dresbach T, Torres V, Wittenmayer N, Altrock WD, Zamorano P, Zuschratter W, Nawrotzki R, Ziv NE, Garner CC, Gundelfinger ED (2006) Assembly of active zone precursor vesicles: obligatory trafficking of presynaptic cytomatrix proteins Bassoon and Piccolo via a trans-Golgi compartment. J Biol Chem 281:6038–6047
Jose M, Nair DK, Altrock WD, Dresbach T, Gundelfinger ED, Zuschratter W (2008) Investigating interactions mediated by the presynaptic protein bassoon in living cells by Foerster’s resonance energy transfer and fluorescence lifetime imaging microscopy. Biophys J 94:1483–1496
Dresbach T, Hempelmann A, Spilker C, tom Dieck S, Altrock WD, Zuschratter W, Garner CC, Gundelfinger ED (2003) Functional regions of the presynaptic cytomatrix protein bassoon: significance for synaptic targeting and cytomatrix anchoring. Mol Cell Neurosci 23:279–291
Altrock WD, tom Dieck S, Sokolov M, Meyer AC, Sigler A, Brakebusch C, Fässler R, Richter K, Boeckers TM, Potschka H, Brandt C, Löscher W, Grimberg D, Dresbach T, Hempelmann A, Hassan H, Balschun D, Frey JU, Brandstätter JH, Garner CC, Rosenmund C, Gundelfinger ED (2003) Functional inactivation of a fraction of excitatory synapses in mice deficient for the active zone protein bassoon. Neuron 37:787–800
Butz S, Okamoto M, Südhof TC (1998) A tripartite protein complex with the potential to couple synaptic vesicle exocytosis to cell adhesion in brain. Cell 94:773–782
Brenman JE, Chao DS, Gee SH, McGee AW, Craven SE, Santillano DR, Wu Z, Huang F, Xia H, Peters MF, Froehner SC, Bredt DS (1996) Interaction of nitric oxide synthase with the postsynaptic density protein PSD-95 and alpha1-syntrophin mediated by PDZ domains. Cell 84:757–767
Yanay C, Morpurgo N, Linial M (2008) Evolution of insect proteomes: insights into synapse organization and synaptic vesicle life cycle. Genome Biol 9:R27
Wang Y, Südhof TC (2003) Genomic definition of RIM proteins: evolutionary amplification of a family of synaptic regulatory proteins. Genomics 81:126–137
Wang X, Kibschull M, Laue MM, Lichte B, Petrasch-Parwez E, Kilimann MW (1999) Aczonin, a 550-kD putative scaffolding protein of presynaptic active zones, shares homology regions with Rim and Bassoon and binds profilin. J Cell Biol 147:151–162
Dick O, tom Dieck S, Altrock WD, Ammermüller J, Weiler R, Garner CC, Gundelfinger ED, Brandstätter JH (2003) The presynaptic active zone protein bassoon is essential for photoreceptor ribbon synapse formation in the retina. Neuron 37:775–786
Wagh DA, Rasse TM, Asan E, Hofbauer A, Schwenkert I, Dürrbeck H, Buchner S, Dabauvalle MC, Schmidt M, Qin G, Wichmann C, Kittel R, Sigrist SJ, Buchner E (2006) Bruchpilot, a protein with homology to ELKS/CAST, is required for structural integrity and function of synaptic active zones in Drosophila. Neuron 49:833–844
Kittel RJ, Wichmann C, Rasse TM, Fouquet W, Schmidt M, Schmid A, Wagh DA, Pawlu C, Kellner RR, Willig KI, Hell SW, Buchner E, Heckmann M, Sigrist SJ (2006) Bruchpilot promotes active zone assembly, Ca2+ channel clustering, and vesicle release. Science 312:1051–1054
Fukuda M (2004) Alternative splicing in the first alpha-helical region of the Rab-binding domain of Rim regulates Rab3A binding activity: is Rim a Rab3 effector protein during evolution? Genes Cells 9:831–842
Nieratschker V, Schubert A, Jauch M, Bock N, Bucher D, Dippacher S, Krohne G, Asan E, Buchner S, Buchner E (2009) Bruchpilot in ribbon-like axonal agglomerates, behavioral defects, and early death in SRPK79D kinase mutants of Drosophila. PLoS Genet 5:e1000700
Zhen M, Jin Y (1999) The liprin protein SYD-2 regulates the differentiation of presynaptic termini in C. elegans. Nature 401:371–375
Patel MR, Lehrman EK, Poon VY, Crump JG, Zhen M, Bargmann CI, Shen K (2006) Hierarchical assembly of presynaptic components in defined C. elegans synapses. Nat Neurosci 9:1488–1498
Fouquet W, Owald D, Wichmann C, Mertel S, Depner H, Dyba M, Hallermann S, Kittel RJ, Eimer S, Sigrist SJ (2009) Maturation of active zone assembly by Drosophila Bruchpilot. J Cell Biol 186:129–145
Yeh E, Kawano T, Weimer RM, Bessereau JL, Zhen M (2005) Identification of genes involved in synaptogenesis using a fluorescent active zone marker in Caenorhabditis elegans. J Neurosci 25:3833–3841
Tao-Cheng JH (2006) Activity-related redistribution of presynaptic proteins at the active zone. Neuroscience 141:1217–1224
Acknowledgments
This work was supported by La Universidad de Antofagasta (Dirección de Investigacion, #1314, #1315) and El Fondo Nacional de Desarrollo Científico y Tecnológico (FONDECYT #1070462).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fernandez, F., Torres, V. & Zamorano, P. An evolutionarily conserved mechanism for presynaptic trapping. Cell. Mol. Life Sci. 67, 1751–1754 (2010). https://doi.org/10.1007/s00018-010-0343-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0343-5