Abstract
Polydnavirus-encoded IκB-like proteins are similar to insect and mammalian IκB, and an immunosuppressive function in the host cells has been inferred to these proteins. Here we show that the expression of one of these IκB-like viral genes, the TnBVank1, in the Drosophila germline affects the localization of gurken, bicoid, and oskar mRNAs whose gene products are relevant for proper embryonic patterning. The altered localization of these mRNAs is suggestive of general defects in the intracellular, microtubule-based, trafficking routes. Analysis of microtubule motor proteins components such as the dynein heavy chain and the kinesin heavy chain revealed defects in the polarized microtubule network. Interestingly, the TnBVANK1 viral protein is uniformly distributed over the entire oocyte cortex, and appears to be anchored to the microtubule ends. Our data open up a very interesting issue on novel function(s) played by the ank gene family by interfering with cytoskeleton organization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During oviposition, parasitic wasps (parasitoids) inject factors that disrupt the host immune reaction and endocrine balance in order to create a suitable environment for the development of their progeny [1]. These host-regulation factors include venom and ovarian secretions, which, in many parasitoid species of lepidopteran larvae, contain viruses of the family Polydnaviridae (PDV) [2]. These viruses are associated with a large number of species of ichneumonid and braconid wasps and are classified in the genera Ichnovirus and Bracovirus, based on morphological and genomic differences [3]. The PDVs are integrated as proviruses and replicate only in the ovary to form free viral particles, which contain viral genes encoding virulence factors, while those genes controlling the viral machinery producing the nucleocapsids are not encapsidated [4, 5]. The free PDV particles delivered in the host body at the oviposition, along with the parasitoid egg and venom, infect a number of tissues and express virulence factors that trigger host immune suppression and developmental regulation [1, 2]. To date, the sequences of eight PDV genomes have been completed, and several partial data sets are available for other PDV genomes [6]. Convergent evolution has shaped the genomic features shared by different PDV genera, such as a low density of coding sequences that show homology with eukaryotic genes and often organized in gene families [7].
One of these gene families, present both in bracoviruses and ichnoviruses, encodes viral ankyrin (vankyrin) proteins, showing significant sequence similarity (approx. 50%) with members of the IκB protein family, inhibitors of NF-κB signaling pathways in insects and vertebrates [8]. Because of the lack of the N- and C-terminal regulatory domains controlling their signal-induced and basal degradation, these vankyrin proteins appear to irreversibly bind and inhibit host NF-κB transcription factors [9–14]. Activation of the NF-κB transcription factors has been reported as being influenced by changes in the microtubule cytoskeleton network [15]. A molecular link between cytoskeletal dynamics and NF-κB-dependent gene regulation is also suggested by the finding that dynein/dynactin complex plays a role in the nuclear accumulation of neuronal NF-κB [16]. Furthermore, Shrum et al. [17] have recently reported that nuclear translocation and activation of NF-κB-dependent gene expression depends on endogenous dynein, in a variety of cell types and in response to diverse activating stimuli, suggesting that dynein-dependent transport of NF-κB may be a conserved mechanism in the NF-κB activation pathway.
In light of these findings, and considering that the cytoskeleton dynamics plays a key role in the cellular immune response and is very often targeted by parasitoid-, VLP (virus like-particles)- and PDV-encoded virulence factors [2, 18, 19], we wanted to explore if vankyrin proteins might interfere with cytoskeleton stability and dynamics.
To address this issue, we decided to express the Toxoneuron nigriceps bracovirus ANK1 protein (TnBVANK1) in Drosophila melanogaster. We generated Drosophila transgenic lines carrying the TnBVank1 gene and analyzed the effects produced by TnBVANK1 expression on oocyte development. Drosophila oogenesis is a well-characterized model system for studying basic questions in developmental and cell biology. The Drosophila ovary consists of repeated units, the ovarioles, each containing a chronologically ordered string of egg chambers developing into mature eggs during oogenesis. Each egg chamber, composed by one oocyte and 15 nurse cells (germline derived cells) surrounded by a monolayer of somatic follicle cells, develops through 14 stages [20]. A stage 1 egg chamber arises from a specialized region, located at the tip of each ovariole and called germarium. During oogenesis, through the connecting ring canals, the nurse cells transfer to the oocyte mRNAs, proteins, organelles, and lipidic droplets. Maternal cues required for embryonic development are specifically placed in the oocyte by microtubule-dependent transport [21, 22]. During oogenesis the microtubule cytoskeleton undergoes stage-specific remodeling [21]. In the early stages of egg-chamber development, up to stage 6, the minus-ends of the germline microtubules are organized in the oocyte by a microtubule organizing center (MTOC) positioned at the posterior pole and their plus-ends extend through the ring canals into the nurse cells. After stage 6, the germline microtubules rearrange and new microtubules nucleate from both the anterior and lateral cortex and extend in all directions [23, 24]. More recently, in vivo analysis of the plus end directed osk mRNA movements, further indicates that in the oocyte at mid-oogenesis the microtubule cytoskeleton is only slightly polarized, with about a 20% excess of microtubules with their plus ends pointing posteriorly [25].
In this study we report that the expression of the TnBVank1 gene in the Drosophila germline cells disrupts the localization of gurken, bicoid and oskar mRNAs, whose correct positioning within the oocyte depends on microtubules cytoskeleton. Our results indicate that this viral protein interferes with proper microtubule and microtubule-motor protein functions.
Materials and methods
Fly strains
Stocks were raised on standard cornmeal/yeast/agar medium at 25°C, and crosses were made at 18°C. The nanos Gal4 stock was obtained from the Bloomington Stock Center (stock #4937) and has the genotype w 1118;P{GAL4:VP16-nos.UTR}MVD1. The following stocks were also used: yw 67c23 /yw 67c23; KHC-lacZ insertion line KZ503 [26]; and Nod-lacZ insertion line NZ143.2 [27], kindly provided by Daniel St. Johnston.
Production of transgenic Drosophila
The TnBVank1 cDNA [11] was inserted into UASp transformation vector specifically designed to allow GAL4 inducible expression in somatic and germline cells [28, 29], which was kindly provided by Pernille Rørth. Transgenic flies were generated as previously described using a yw 67c23 strain as a recipient stock [30]. Several transgenic lines were isolated and the appropriate crosses performed to obtain homozygous lines carrying two copies of the transgene.
Gal4-driven expression in germline cells
GAL4-driven expression was induced by crossing females of the genotype w 1118 ; +/+ P{GAL4:VP16-nos.UTR}MVD1 to males TnBVank1/Y; TnBVank1/TnBVank1; +/+, and to males yw 67c23 as control. The crosses were performed at 18°C and freshly eclosed females of the genotype TnBVank1/w 1118; TnBVank1/+; P{GAL4:VP16-nos.UTR}MVD1/+ and w 1118 /yw 67c23; +/+; P{GAL4:VP16-nos.UTR}MVD1/+ as a control were collected, crossed with yw 67c23 males and the crosses were transferred daily to fresh yeasted food and kept at 29°C for 8 days before dissection.
Immunofluorescence microscopy
Ovaries were dissected in phosphate buffer saline (PBS 1×) pH 7.5, fixed for 20 min at room temperature in 4% paraformaldehyde in PBS1× pH 7.5 freshly prepared and the ovaries were then kept at room temperature to ensure that the microtubules did not depolymerize. After three 5-min washes in PBS + 0.1% Triton X-100 (PBT), egg chambers were dissected with needles and were permeabilized overnight at 4°C in PBS + 1% Triton X-100. The next day the egg chambers were washed three times for 5 min each in PBT and one time 15 min in PBT + 3% BSA. Egg chambers were then incubated for 4 h at room temperature on a rotating wheel with primary antibody diluted in PBT + 3% BSA. After incubation with primary antibody, the egg chambers were washed three times for 15 min each in PBT and one time 15 min in PBT + 3% BSA, and then were incubated 2 h at room temperature on a rotating wheel with secondary antibody diluted in PBT + 3% BSA. After several washes in PBT, egg chambers were mounted in Fluoromount G (Electron Microscopy Sciences) and subsequently were analyzed with conventional epifluorescence on a Nikon Eclipse 90i microscope and with a TCS SL Leica confocal system.
Primary antibodies were used at the following concentrations: mouse monoclonal anti-Gurken, 1:50 (Developmental Studies Hybridoma Bank); mouse monoclonal P1H4 anti-dynein heavy chain, 1:500 (kindly provided by Tom Hays); mouse monoclonal anti-β-gal, 1:25 (Developmental Studies Hybridoma Bank); rabbit polyclonal anti-oskar, 1:200 (kindly provided by Trudi Schüpbach); mouse monoclonal anti-β-tubulin, 1:25 (Developmental Studies Hybridoma Bank). The rabbit polyclonal antibody anti-TnBVANK1 was raised against two TnBVANK1 peptides, one located at the N-terminal domain (LLGERNELGNNFFHE) and the other at the C-terminal domain (NDKKMMEILKKNGAK). This antibody was used 1:500 for immunohistochemistry and 1:5,000 for Western blot. Secondary antibodies used were FITC-conjugated anti-mouse (1:250, Invitrogen); Cy3-conjugated anti-rabbit (1:1,000, Sigma); Alexa Fluor 546-conjugated anti-mouse (1:200, Molecular Probes). Nuclear staining was carried out after immunodetection by incubating for 2 h the egg chambers with To-Pro-3 (Molecular Probes) at 1μg/ml in PBT 1× pH 7.5 and, after several washes with PBT, the egg chambers were mounted as indicated above. TRITC-Phalloidin staining was carried out after immunodetection by incubating the egg chambers for 20 min at room temperature with TRITC-Phalloidin (40 μg/ml in PBS, Sigma) and after several washes with PBT, the egg chambers were mounted as indicated above.
In situ hybridization
Whole-mount RNA in situ hybridization procedure was a modification of the protocol of Tautz and Pfeifle [31], using 65°C as the hybridization temperature. Digoxigenin-labeled (Roche) antisense RNA probes were synthesized using as a template the full-length of the gurken cDNA (provided by Trudi Schüpbach), the bicoid DNA region (−40/+230), and the full-length of the oskar cDNA (kindly provided by Anne Ephrussi). Digoxygenin-labeled sense RNA probe was also used as control (data not shown). The egg chambers were mounted in Fluoromount-G and analyzed with a Nikon Eclipse 90i microscope.
Collection of eggs
Eggs laid by transgenic females obtained from the crosses described above were collected on standard apple juice agar plate and the hatching larvae were monitored for 3 days after oviposition. Lethality was measured by the ratio between the number of unhatched eggs and the total number of eggs laid in each plate.
Protein extracts and Western-blot analysis
The ovaries from ten TnBVank1/w 1118; TnBVank1/+; P{GAL4:VP16-nos.UTR}MVD1/+ females and from ten control w 1118 /yw 67c23; +/+; P{GAL4:VP16-nos.UTR}MVD1/+ females were collected and quickly frozen in liquid nitrogen and stored at −80°C. The ovaries were transferred in 75 μl cold 2× Laemmli sample buffer [32], homogenized by sonication and boiled for 5 min. After centrifugation for 10 min at 14,000 rpm at 4°C, the supernatant material was collected. Fifteen microliters of this soluble material was used and mixed with an equal volume of 1× Laemmli sample buffer previously boiled for 5 min. The soluble proteins were then loaded on 12% polyacrylamide gel. Protein transfer to membranes and Western blotting were performed as previously described [33]. The TnBVANK1 protein was detected by using the rabbit anti-TnBVANK1 antibody diluted 1:5,000. The primary antibody was detected by using goat anti-rabbit secondary antibody, conjugate with horseradish peroxidase (Santa Cruz Biotechnology) at the dilution of 1:7,000 and ECL detection kit (GE Healthcare).
Drug treatment
Ovaries were dissected in PBS 1× pH 7.5 and transferred into 24-well dishes with 1 ml per well of Schneider’s complete media supplemented with 15% Fetal Bovine Serum and 0.6× penicillin/streptomycin. Drugs were added at the following concentration: paclitaxel 3.2 μM (Sigma), nocodazole 2 μM (Sigma). A total of 17.7 μl of DMSO per well was used as a control since stock solutions of the three drugs were prepared in DMSO and kept frozen until use. The egg chambers were then incubated for 5 h at 25°C with drugs. After incubation, the egg chambers were fixed for 20 min at room temperature in 4% paraformaldehyde in PBS 1× pH 7.5 freshly prepared and then processed following the standard immunostaining procedures. This protocol is an adaptation of Prasad et al. [34].
Results
Expression of TnBVank1 induces a maternal effect lethal phenotype
The TnBVank1 gene of T. nigriceps codes for a protein showing an average 30% identity with Cactus, the Drosophila homologue of mammalian IκB, and with other IκB proteins from several species [11]. TnBVANK1 is 155 aa long, is made of three ankyrin repeats and does not contain the N-terminal IKK target motif that mediates the signal-induced degradation of Cactus or the C-terminal PEST domain, which is involved in rapid protein turnover and is present in all Cactus/IκB proteins [11]. To test the effect of the viral TnBVANK1 protein on the Drosophila oocyte development, we used the GAL4/UAS binary system that allows conditional and tissue-specific expression of genes of interest in Drosophila [28]. We produced Drosophila transgenic lines carrying TnBVank1 under the control of the UASp sequences [29], and the expression of the transgene was induced in germline cells using the nanos-GAL4:VP16 Drosophila stock [35]. To detect the TnBVANK1 protein, a polyclonal antibody (anti-TnBVANK1) was raised against two synthetic peptides of TnBVANK1 (see Materials and methods). The presence of TnBVANK1 in the ovaries of TnBVank1, nanos-GAL4:VP16 transgenic flies was assessed through regular Western blot on whole cell lysate. A single band in the expected size range of 17 kDa, corresponding to the predicted TnBVANK1 protein, was detected (Fig. 1a). No signal was instead revealed in ovarian extracts from females carrying only the nanosGal4:VP16 transcriptional activator gene, hereafter denoted as wild-type females and used as control. We also analyzed the TnBVANK1 protein distribution during oogenesis by immunostaining. As shown in Fig. 1b, the TnBVANK1 protein was detected in the germline cells throughout oogenesis, starting from the germarium and up to late egg chamber developmental stages.
Expression of TnBVank1 transgene during Drosophila melanogaster oogenesis. a Western blot of whole cell lysate using anti-TnBVANK1 antibody shows a single band in the size range of 17 kDa, corresponding to the predicted TnBVANK1 protein. No signal was instead revealed from control ovaries (we refer at these as wild-type) carrying the nanos-Gal4:VP16 transcriptional activator gene, without the heterologous gene TnBVank1. b Expression of TnBVANK1 protein (in green) during different stages of oogenesis in egg chambers extracted from females expressing TnBVank1. Immmunostaining was performed using anti-TnBVANK1 antibody. The expression of TnBVANK1 protein is detected in the germline cells from the germarium until late stages egg chambers. Anterior is to the left. St stage
We found that the germline expression of TnBVank1 interferes with embryonic development. The expression of one copy of the TnBVank1 gene induced a maternal effect semi-lethal phenotype. Lethality of 56.3 and 42.0%, was observed with two different transgenic lines, where the transgene was inserted on the X or on the II chromosome, respectively. This lethality was neither observed in embryos from control females carrying the heterologous gene without the Gal4 transcriptional activator gene, nor in the strain carrying only the Gal4 gene. This indicates that the lethality is indeed caused by the expression of the transgene. We also tested the effect of gene dosage by producing transgenic females that carry the Gal4 gene and two copies of TnBVank1. In this genetic condition we observed 99.2% lethality, which indicates that the TnBVank1 gene dosage is critical. Thus, all the experiments hereafter described in the paper were carried out by co-expressing the two copies of TnBVank1 transgene, one on the X and the other on the II chromosome.
Interestingly, the expression of two copies of the TnBvank1 gene causes ventralization of the eggshell (Fig. 2). The Drosophila wild-type eggshell has a clear dorsoventral polarity, marked by a pair of dorsal appendages at a dorsoanterior position (Fig. 2a). As a consequence of the TnBVANK1 activity, the eggs showed defects in dorsal appendages. In some eggs the dorsal appendages were fused at the base (Fig. 2b), which is a characteristic of weakly ventralized eggs. More severely ventralized eggs were found, distinguished by complete fusion of the dorsal appendages resulting in a single appendage on the dorsal midline (Fig. 2c). Furthermore, the majority of the eggs (65%, n = 100) showed the most extreme ventralized phenotype with the loss of any appendages material (Fig. 2d).
Ventralized eggshell phenotypes produced by germline expression of two copies of the TnBVank1 transgene. a Wild-type egg showing two dorsal appendages anteriorly. b–d Eggs produced by transgenic females carrying two copies of the TnBVank1 gene and the nanos-Gal4:VP16 transactivator. Eggs are short, rounded, and have defects in dorsal appendages. b The dorsal appendages are closed together and fused at the base. c The appendages are completely fused. d The egg with the most extreme ventralized phenotype does not show any appendages. Anterior is up in all the panels
Expression of TnBVank1 affects gurken mRNA localization and protein accumulation
The ventralized phenotype caused by TnBVank1 expression indicates an altered Gurken-Egfr signaling [36–38]. During oogenesis, this signaling pathway regulates the fate of dorsal follicle cells, and leads to the definition of two separate cell populations that will guide the production of the two dorsal appendages [39, 40]. Proper gurken (grk) mRNA localization during Drosophila oogenesis is necessary for the activation of Egfr signaling. grk mRNA is transported from the nurse cells into the oocyte, where it is properly localized by the minus-end-directed dynein motor protein along the polarized microtubule network [21, 24]. Therefore, we examined the effect of TnBVank1 expression on the grk mRNA localization. In early oogenesis of wild-type egg chambers, grk mRNA is localized to the posterior pole of the oocyte (Fig. 3a) where Grk instructs follicle cells to adopt a posterior cell fate [41, 42]. During mid-oogenesis, grk mRNA is localized to a dorsoanterior domain in the cytoplasm directly overlying the oocyte nucleus (stages 9 and 10 Fig. 3c, e) [36]. In ovaries expressing TnBVank1, the proper localization of grk mRNA is affected. In early egg chambers this mRNA was not completely localized in the oocyte, and in all the egg chambers analyzed the grk mRNA signal was still detected in the nurse cells (Fig. 3b). In mid-oogenesis, at stage 9 grk mRNA was found mostly as a diffuse signal throughout the oocyte cytoplasm, rather than concentrated above the oocyte nucleus, in 84% (n = 100) of the egg chambers (Fig. 3d). Furthermore, in 86% (n = 100) of stage 10B egg chambers grk mRNA was not detected, suggesting that in the absence of a correct localization grk mRNA may be dispersed in the whole oocyte cytoplasm (Fig. 3f). It has been demonstrated that grk mRNA is specifically translated, while mislocalized grk mRNA is transcriptionally repressed [43–45]. In wild-type egg chambers at mid-oogenesis, Grk protein distribution mirrors that of the mRNA, which is translated adjacent to the oocyte nucleus (Fig. 3g), at the dorsoanterior corner of the oocyte (Fig. 3g, i) [37]. In the ovaries expressing TnBVank1, Grk was restricted to the oocyte at earlier stages (Fig. 3h). However, the levels of Grk were variably reduced throughout oogenesis, and in 87% (n = 100) of the stage 10B egg chambers Grk protein was undetectable (Fig. 3j). The reduced level of Grk explains the ventralized eggshell phenotype produced by the expression of TnBVank1. Furthermore, our findings suggest that the TnBVANK1 protein may interfere with the microtubules-dependent transport of grk mRNA.
Defects in gurken mRNA localization and Gurken protein expression during oogenesis caused by expression of TnBVank1. In situ hybridization showing grk mRNA localization in wild-type ovaries (a, c, e) and in ovaries expressing TnBVank1 (b, d, f). a At early stages, grk mRNA is localized in the oocyte. At stage 6 it is localized within the oocyte in an anterior-cortical ring and then from early stage 9 (c) until stage 10B (e) to a dorsoanterior domain. b In early egg chambers the transport of grk mRNA into the oocyte is partly affected, and the grk mRNA signal is still detected in the nurse cells. At stage 9 grk mRNA (d) is found as a diffuse signal throughout the oocyte cytoplasm. f At stage 10B grk mRNA is not detected. In situ hybridizations were performed in parallel with wild-type controls, and all samples were stained for equal lengths of time. Imunolocalization of Grk protein as assayed by using anti-Grk antibody in wild-type ovaries (g, i) and in ovaries expressing TnBVank1(h, j). g From the germarium until stage 8 Grk is localized in an anterior cortical ring around the oocyte nucleus. From early stage 9 Grk is localized to the dorsoanterior corner of the oocyte. i In stage 10B egg chamber Grk is localized to the dorsoanterior corner of the oocyte, directly above the oocyte nucleus. h From germarium until early stage 8 Grk is restricted to the oocyte but its levels are reduced. j Stage 10B egg chamber in which Grk protein is absent. In all panels, the egg chambers are oriented with the anterior region toward the left. In panels e, f, i, j, the dorsal side is at the top. Scale bars 20 μm. St stage
Localization of bicoid and oskar mRNAs are also affected by TnBVank1 expression
The spatial information essential for the development of the future embryo is encoded also by other determinants such as bicoid and oskar that respectively define the anterior and posterior regions of the embryo [46]. These determinants are directed to the correct subcellular sites by a microtubule-dependent localization of their mRNAs in the oocyte [36, 47, 48]. Therefore, we investigated whether also bicoid (bcd) and oskar (osk) mRNA positioning was affected by the TnBVANK1 protein. bcd mRNA is produced during early stages of oogenesis in the nurse cells, assembled into large particles and transported to the oocyte. At stage 9 bcd mRNA is mostly localized at the anterior corners of the oocyte (Fig. 4a). During stage 10B it is redistributed into a disc-like pattern in the anterior cortex of the oocyte and intense labeling is also observed around the nurse cell nuclei (Fig. 4c). In 85% (n = 100) of stage 9 egg chambers expressing TnBVank1, bcd mRNA was not localized to the anterior corners of the oocyte but appeared diffuse throughout the cytoplasm (Fig. 4b). Then, at stage 10B, in 86% (n = 100) of the egg chambers bcd mRNA was not found at the anterior margin of the oocyte and remained concentrated within the germline cells (Fig. 4d).
Defects in bicoid and oskar mRNA localization during oogenesis caused by expression of TnBVank1. In situ hybridization showing bcd mRNA localization in wild-type ovaries (a, c) and in ovaries expressing TnBVank1 (b, d). a Stage 9 egg chamber showing that bcd mRNA is mostly localized at the anterior corners of the oocyte. c Stage 10B egg chamber showing that bcd transcript is distributed into the anterior cortex of the oocyte. b Stage 9 egg chamber showing that bcd mRNA appears diffuse throughout the oocyte. d Stage 10B egg chamber in which bcd mRNA is not properly localized along the anterior cortical region of the oocyte. In situ hybridization showing oskar mRNA localization in wild-type ovaries (e, g) and in ovaries expressing TnBVank1 (f, h). e–f osk mRNA distribution from the germarium until mid-oogenesis. e At stage 9 osk mRNA transitorily accumulates in the oocyte and then becomes localized at the posterior pole of the oocyte. g Stage 10B egg chamber showing osk mRNA accumulation at the posterior pole of the oocyte. f In stage 7 and 9 egg chambers osk mRNA is more concentrated within the nurse cells and in the oocyte cytoplasm, instead of being associated with the posterior cortex of the oocyte. h At stage 10B osk transcript is not found at the posterior pole of the oocyte. In situ hybridizations were performed in parallel with wild-type controls, and all samples were stained for equal lengths of time. Immunolocalization of Oskar protein as assayed by using anti-osk antibody in wild-type ovaries (i) and in ovaries expressing TnBVank1(j, k). i Stage 10B egg chamber showing Osk accumulation at the posterior pole of the oocyte. j and k Stage 10B egg chambers showing, respectively, a very weak posterior Osk signal and a cortical localization of this protein. In all panels, the egg chambers are oriented with the anterior region toward the left. Scale bars 20 μm. St stage
We next tested the distribution of oskar mRNA in egg chambers expressing the TnBVank1 gene. In wild-type ovaries, osk mRNA is produced by the nurse cells and accumulates in the oocyte during early stages of oogenesis (stage 7, Fig. 4e). During stages 8–10 of oogenesis, osk mRNA moves to the posterior pole of the oocyte (Fig. 4e, g) where it is translated (Fig. 4i). In ovaries expressing TnBVank1, osk mRNA was localized to the oocyte in early stages, but appeared more concentrated within the nurse cells (Fig. 4f). During mid-oogenesis, in 75% (n = 100) of stage 9 egg chambers osk mRNA was found as a diffuse signal throughout the oocyte cytoplasm and was less accumulated at the posterior pole (stage 9, Fig. 4f). At stage 10B osk transcript was not detected at the posterior pole of the oocyte in 78% (n = 100) of the egg chambers (Fig. 4h). In addition to the effect on osk mRNA localization, the accumulation of Osk protein is also affected. In 77% (n = 100) egg chambers expressing TnBVank1 its distribution was affected, ranging from a very weak posterior signal (Fig. 4j) to localization over the oocyte cortex (Fig. 4k).
Collectively, the effects on the localization of grk, bcd, and osk mRNAs indicate that TnBVANK1 interferes with the microtubule cytoskeleton.
Effects of the expression of TnBVank1 on microtubule-based transport in the oocyte
We evaluated the organization of the microtubule cytoskeleton in ovaries expressing TnBVank1 by analyzing the microtubule motors. Proper localization of grk and bcd mRNAs is directed by the minus-end-directed microtubule motor dynein [21]. We analyzed the distribution of dynein by using the mouse anti-dynein heavy-chain P1H4 monoclonal antibody [49]. In the early stages of oogenesis, dynein heavy chain (DHC) localizes around the oocyte nucleus (Fig. 5a), while at mid-oogenesis it gradually becomes concentrated to the oocyte posterior end (stage 9, Fig. 5c) [50], where it is presumably stored to allow repeated rounds of minus end-directed transport. In egg chambers expressing the TnBVank1 gene, DHC appeared properly localized during early stages (Fig. 5b). However, the levels of DHC were variably reduced throughout oogenesis and in 88% (n = 100) of stage 9 egg chambers DHC was not or barely detectable at the posterior end of the oocyte (Fig. 5d). This result indicates that transport to minus end of the microtubules is in some way altered in the egg chambers expressing TnBVank1.
Effects of TnBVank1 expression on microtubule-motor proteins in the oocyte. Localization of dynein heavy chain (DHC) in wild-type ovaries (a, c) and in ovaries expressing TnBVank1 (b, d). a From germarium until stage 8 DHC is localized within the oocyte around the oocyte nucleus. c Stage 9 egg chamber showing that DHC accumulates at the posterior end of the oocyte. b DHC is localized to the oocyte in early stages until stage 8. d At stage 9 DHC is not localized at the posterior end of the oocyte. Nod-β-gal distribution as assayed with antibodies against β-gal in wild-type ovaries (e) and ovaries expressing TnBVank1 (f). e Stage 9 egg chamber showing that Nod-β-gal is enriched around the oocyte nucleus. f Stage 9 egg chamber in which Nod-β-gal signal is only faintly detected at the dorsoanterior corner adjacent to oocyte nucleus. Localization of KHC-β-gal, as assayed by antibodies against β-gal, in wild-type (g) and TnBVank1 egg chambers (h). g Stage 9 egg chamber showing that KHC-β-gal accumulates at the posterior pole of the oocyte. h Stage 9 egg chamber showing that KHC-β-gal is mislocalized in the oocyte and does not concentrate at the posterior pole. In all panels, the egg chambers are oriented with the anterior region toward the left. In panels from c to h the dorsal side is up. Scale bars 20 μm. St stage
We next analyzed the distribution of Nod-β-gal, a fusion protein of the motor-like domain of Nod, a kinesin-related protein, and β-galactosidase [27]. Differently from Nod, which preferentially binds microtubule plus-ends in vivo [51, 52], Nod-β-gal localizes to the oocyte microtubule minus-ends, for which it has been reliably used as a marker [27, 53, 54]. By stage 9, the Nod-β-gal protein becomes concentrated around the oocyte nucleus (stage 9, Fig. 5e). We found that in ovaries expressing TnBVank1 Nod-β-gal was either absent or only weakly detectable around the oocyte nucleus in 83% (n = 100) of stage 9 egg chambers (Fig. 5f). This result suggests that microtubule minus-ends are not properly distributed when TnBVANK1 protein is present.
Microtubules and the plus-end-directed microtubule motor kinesin are required for the selective accumulation of osk mRNA at the posterior cortex of the Drosophila oocyte. We investigated the distribution of kinesin by following the expression of a Kinesin Heavy Chain-β-galactosidase fusion (KHC-β-lacZ) transgene. In wild-type stage 9 egg chambers, the KHC-β-gal is localized to the posterior pole of the oocyte (Fig. 5g) [26]. In 79% (n = 100) of stage 9 egg chambers expressing TnBVANK1, the KHC-β-gal was not properly accumulated at the posterior pole of the oocyte and it appeared also mislocalized in the oocyte (Fig. 5h).
Since osk mRNA particle movement is kinesin dependent, in the ovaries expressing the TnBVank1 gene, as expected the penetrance of the kinesin localization defects concords with the one observed for osk mRNA localization.
Microtubules are essential for TnBVANK1 cortical localization
The experimental data presented above strongly indicate the occurrence of a disrupting effect exerted by TnBVank1 gene on the oocyte microtubule network. To assess any direct interaction between TnBVANK1 and microtubules, we analyzed by confocal microscopy the immunolocalization of TnBVANK1 and the dependence of its profile on microtubule stability.
As shown in Fig. 6a–f, at mid-oogenesis the TnBVANK1 protein, detected by using the anti-TnBVANK1 antibody, was localized around the oocyte cortex, outlining the cortical layer of actin distribution but not colocalizing with it and displaying granular appearance (see Fig. S1 in Electronic Supplementary Material, ESM). Similarly, TnBVANK1 showed cortical distribution in the nurse cells (Fig. 6g–i). Furthermore, the TnBVANK1 protein did not appear to affect the cortical actin cytoskeleton, as compared with wild type egg chambers (data not shown). These results indicate that TnBVANK1 may be anchored to the oocyte cortex by contacting the microtubules ends. To test this hypothesis, we examined the effect of microtubule depolymerization on the localization of TnBVANK1 in the oocyte. Egg chambers expressing the TnBVank1 gene were dissected and cultured in the presence of the microtubule inhibitor nocodazole. After drug treatment, microtubules, detected by using an anti-β-tubulin antibody, appeared strongly affected (Fig. 7b), and the TnBVANK1 protein lost its cortical localization and showed a punctuated distribution in the cytoplasm (Fig. 7a, c, c′). This result corroborates the hypothesis that this protein interacts with microtubules ends. Accordingly, the egg chambers treated with paclitaxel, which stabilizes the microtubules, showed a stronger β-tubulin staining (Fig. 7e) and an increase of the TnBVANK1 cortical signal (Fig. 7d, f, f′), as compared with control egg chambers treated only with DMSO, the solvent for both nocodazole and paclitaxel drugs (Fig. 7g, i, i′). The DMSO treatment did not affect the cortical distribution of TnBVANK1, as also shown in Fig. 7j, k where stage 9 and 10B egg chambers were carefully analyzed by taking confocal optical cross sections through the center of the egg chambers.
Intracellular localization of TnBVANK1 protein. Confocal cross section of a stage 10B egg chamber stained with anti-TnBVANK1 antibody (green a) and TRITC-conjugate phalloidin (red b). Within the oocyte the TnBVANK1 protein is distributed on the surface just underneath the oocyte cortical actin cytoskeleton. c Merged image of TnBVANK1 (green), TRITC-conjugate phalloidin (red) and To-Pro-3 (cyan) signals is shown in c. d–f Higher magnification of the dotted area in a–c. g–i Confocal surface section of the same stage 10B egg chamber shown in a–f. TnBVANK1 displays a cortical localization in nurse cells and appears also distributed as bundles in the nurse cells cytoplasm. In all panels, the egg chamber is oriented with the anterior region toward the left
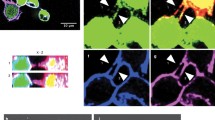
Ex vivo analysis of TnBVANK1 distribution in ovaries treated with drugs affecting the microtubule polymerization. Confocal sections of stage 9 egg chambers treated with nocodazole (a–c′), paclitaxel (d–f′) and DMSO (g–i′). These egg chambers have been stained with anti-TnBVANK1 antibody (green in a, d, g) and with anti-β-tubulin antibody (red in b, e, h). c, f, i Merged images of TnBVANK1, β-tubulin and To-Pro-3 (blue) nuclear staining signals, and the higher magnification views of the boxed regions are respectively shown in c′, f′, i′. Nocodazole treated stage 9 egg chamber showing that microtubules are dramatically altered (b), and the localization on the oocyte cortex of the TnBVANK1 protein is lost (a, c, c′). In paclitaxel treated stage 9 egg chamber microtubules are more abundant (e) and TnBVANK1 (d, f, f′) is more abundantly localized to the cortical surface of the oocyte (see arrows in f, f′). Microtubules cytoskeleton (h) and TnBVANK1 distribution on the oocyte cortex (g, i, i′) are unaffected in DMSO treated stage 9 egg chamber (see arrowhead in i, i′). j–k Confocal optical cross sections through the center of stage 9 and 10B egg chambers treated with DMSO, showing the cortical localization of TnBVANK1. In all panels, the egg chambers are oriented with the anterior region toward the left. Scale bars 20 μm
Discussion
The increasing number of studies addressing the mechanisms of host regulation mediated by PDVs clearly indicate the occurrence of immunosuppressive strategies based on virulence factors hitting multiple host targets, in order to insure a more effective immune disguise [1, 2]. The complexity of the molecular network finely tuning the host regulation process is further reinforced by the presence of gene families, which show members characterized by tissue/temporal specific profiles of expression, likely influencing different host physiological pathways [9–11, 55]. These multiple effects are well documented by recent studies on vankyrin proteins, encoded by both bracoviruses and ichnoviruses, for which a role in preventing apoptosis [56] and in immune suppression [10, 11] has been demonstrated. The pleiotropic effects of vankyrin proteins could be in part explained by the large number of pathways controlled by NF-κB transcription factors and by the conserved structural features of its interacting molecules, which may account also for the observed biological activity of these IκB-like proteins in evolutionary unrelated organisms [11].
Here we shed some light on this issue, by analyzing the impact of a TnBV-encoded vankyrin on cytoskeleton dynamics, which is a key aspect of cellular immune response. We used Drosophila oogenesis as model system to investigate the impact of TnBVank1 expression in a well-characterized cell biology context. Our data clearly show that the expression of the TnBVank1 gene interferes with proper mRNAs localization driven by the minus- and plus-end-directed microtubule motors. Remarkably, the defects of grk mRNA positioning on the dorsoanterior corner of the oocyte cause altered Grk/Egfr signaling and generate ventralized eggs.
Our analysis of the distribution of TnBVANK1 protein within the oocyte reveals that this protein is localized at the oocyte cortex, just underneath the actin cytoskeleton. We found that treatment of ovaries with nocodazole, a microtubules depolymerizing drug, abolishes TnBVANK1 cortical localization. Therefore, TnBVANK1 could affect the polarized microtubule network by anchoring to microtubules ends.
This finding suggests that in the host cells the TnBVANK1 protein could play a double function in suppressing the NF-κB activity. On one hand, it may bind irreversibly NF-κB repressing the nuclear import of this protein. On the other hand, it may also affect NF-κB nuclear translocation by altering the microtubule network. However, we must take into account that the altered microtubule network, as we have shown for the localization of maternal cues in the Drosophila oocyte, might also impair other cellular processes. Among these, of remarkable importance is the microtubule-induced cortical Rac1 activation and lamellipodium formation during polarized cell migration [57], a process largely studied in leukocyte chemotaxis and highly conserved in eukaryotic cells [58, 59]. Basically, the microtubules plus ends deliver GEFs (Guanine nucleotide Exchange Factors) at the leading edge cortex, stimulating Rac1 activation and subsequent actin polymerization [57]. Thus, a disruption of microtubules architecture may well result in a failure of actin-mediated cell behaviors, which are essential in cellular immunity.
The complex interplay among different components of the cytoskeleton in cellular immune responses is a relevant issue that deserves further research effort. Other biological roles for ankyrin proteins disrupting microtubules functioning have been described in the literature. In this regard, it is interesting to note that some intracellular bacterial pathogens disrupt the function of eukaryotic factors by introducing ankyrin proteins into the host cells [60]. This study has shown that the intracellular pathogens Legionella pneumophila and Coxiella burnetii deliver into eukaryotic cells a large number of different bacterial proteins containing ankyrin repeat homology domains. Particularly, the L. pneumophila AnkX interferes with minus end-directed transport of vesicles on microtubules.
In conclusion, the major outcome of our study is the finding that the TnBVANK1, a IκB-like protein, disrupts the microtubule network. This proposes a novel function for vankyrin proteins and sets the stage for more specific in vivo studies on the suppressive mechanisms of the host immune response against eukaryotic parasites of insects.
References
Pennacchio F, Strand MR (2006) Evolution of developmental strategies in parasitic hymenoptera. Annu Rev Entomol 51:233–258
Webb BA, Strand MR (2005) The biology and genomics of polydnaviruses. In: Gilbert LI, Iatrou K, Gill SS (eds) Comprehensive molecular insect science, vol 6. Elsevier, San Diego, pp 323–360
Webb BA, Beckage NE, Hayakawa Y, Krell PJ, Lanzrein B, Stoltz DB, Strand MR, Summers MD (2000) Family Polydnaviridae. In: van Regenmortel MHV, Fauquet CM, Bishop DHL, Carstens EB, Estes MK, Lemon SM, Maniloff J, Mayo MA, McGeoch DJ, Pringle CR, Wickner RB et al (eds) Virus taxonomy: seventh report of the international committee on taxonomy of viruses. Academic Press, San Diego, pp 253–260
Deng L, Stoltz DB, Webb BA (2000) A gene encoding a polydnavirus structural polypeptide is not encapsidated. Virology 269:440–450
Bézier A, Annaheim M, Herbinière J, Wetterwald C, Gyapay G, Bernard-Samain S, Wincker P, Roditi I, Heller M, Belghazi M, Pfister-Wilhem R, Periquet G, Dupuy C, Huguet E, Volkoff AN, Lanzrein B, Drezen JM (2009) Polydnaviruses of braconid wasps derive from an ancestral nudivirus. Science 323:926–930
Bézier A, Herbinière J, Lanzrein B, Drezen JM (2009) Polydnavirus hidden face: the genes producing virus particles of parasitic wasps. J Invertebr Pathol 101:194–203
Espagne E, Dupuy C, Huguet E, Cattolico L, Provost B, Martins N, Poirié M, Periquet G, Drezen JM (2004) Genome sequence of a polydnavirus: insights into symbiotic virus evolution. Science 306:286–289
Silverman N, Maniatis T (2001) NF-κB signaling pathways in mammalian and insect innate immunity. Genes Dev 15:2321–2342
Kroemer JA, Webb BA (2005) IκB-related vankyrin genes in the Campoletis sonorensis ichnovirus: temporal and tissue-specific patterns of expression in parasitized Heliothis virescens lepidopteran hosts. J Virol 79:7617–7628
Thoetkiattikul H, Beck MH, Strand MR (2005) Inhibitor κB-like proteins from a polydnavirus inhibit NF-κB activation and suppress the insect immune response. Proc Natl Acad Sci USA 102:11426–11431
Falabella P, Varricchio P, Provost B, Espagne E, Ferrarese R, Grimaldi A, de Eguileor M, Fimiani G, Ursini MV, Malva C, Drezen JM, Pennacchio F (2007) Characterization of the IκB-like gene family in polydnaviruses associated with wasps belonging to different Braconid subfamilies. J Gen Virol 88:92–104
Lapointe R, Tanaka K, Barney WE, Whitfield JB, Banks JC, Béliveau C, Stoltz D, Webb BA, Cusson M (2007) Genomic and morphological features of a banchine polydnavirus: comparison with bracoviruses and ichnoviruses. J Virol 81:6491–6501
Tian SP, Zhang JH, Wang CZ (2007) Cloning and characterization of two Campoletis chlorideae ichnovirus vankyrin genes expressed in parasitized host Helicoverpa armigera. J Insect Physiol 53:699–707
Shi M, Chen YF, Huang F, Liu PC, Zhou XP, Chen XX (2008) Characterization of a novel gene encoding ankyrin repeat domain from Cotesia vestalis polydnavirus (CvBV). Virology 375:374–382
Spencer W, Kwon H, Crepieux P, Leclerc N, Lin R, Hiscott J (1999) Taxol selectively blocks microtubule dependent NF-κB activation by phorbol ester via inhibition of IκBα phosphorylation and degradation. Oncogene 18:495–505
Mikenberg I, Widera D, Kaus A, Kaltschmidt B, Kaltschmidt C (2007) Transcription factor NF-κB is transported to the nucleus via cytoplasmic dynein/dynactin motor complex in hippocampal neurons. PLoS ONE 2:e589
Shrum CK, DeFrancisco D, Meffert MK (2009) Stimulated nuclear translocation of NF-κB and shuttling differentially depend on dynein and the dynactin complex. Proc Natl Acad Sci USA 106:2647–2652
Rizki TM, Rizki RM (1994) Parasitoid-induced cellular immune deficiency in Drosophila. Ann N Y Acad Sci 712:178–194
Glatz RV, Asgari S, Schmidt O (2004) Evolution of polydnaviruses as insect immune suppressors. Trends Microbiol 12:545–554
Spradling AC (1993) Developmental genetics of oogenesis. In: Bate M, Martinez-Arias A et al (eds) The development of Drosophila melanogaster. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 1–70
Steinhauer J, Kalderon D (2006) Microtubule polarity and axis formation in the Drosophila oocyte. Dev Dyn 235:1455–1468
Becalska AN, Gavis ER (2009) Lighting up mRNA localization in Drosophila oogenesis. Development 136:2493–2503
Cha BJ, Koppetsch BS, Theurkauf WE (2001) In vivo analysis of Drosophila bicoid mRNA localization reveals a novel microtubule-dependent axis specification pathway. Cell 106:35–46
MacDougall N, Clark A, MacDougall E, Davis I (2003) Drosophila gurken (TGFalpha) mRNA localizes as particles that move within the oocyte in two dynein-dependent steps. Dev Cell 4:307–319
Zimyanin VL, Belaya K, Pecreaux J, Gilchrist MJ, Clark A, Davis I, St Johnston D (2008) In vivo imaging of oskar mRNA transport reveals the mechanism of posterior localization. Cell 134:843–853
Clark I, Giniger E, Ruohola-Baker H, Jan LY, Jan YN (1994) Transient posterior localization of a kinesin fusion protein reflects anteroposterior polarity of the Drosophila oocyte. Curr Biol 4:289–300
Clark IE, Jan LY, Jan YN (1997) Reciprocal localization of Nod and kinesin fusion proteins indicates microtubule polarity in the Drosophila oocyte, epithelium, neuron and muscle. Development 124:461–470
Brand AH, Perrimon N (1993) Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118:401–415
Rorth P (1998) Gal4 in the Drosophila female germline. Mech Dev 78:113–118
Rubin GM, Spradling AC (1982) Genetic transformation of Drosophila with transposable element vectors. Science 218:348–353
Tautz D, Pfeifle C (1989) A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback. Chromosoma 98:81–85
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Andrenacci D, Cernilogar FM, Taddel C, Rotoli D, Cavaliere V, Graziani F, Gargiulo G (2001) Specific domains drive VM32E protein distribution and integration in Drosophila eggshell layers. J Cell Sci 114:2819–2829
Prasad M, Jang AC, Starz-Gaiano M, Melani M, Montell DJ (2007) A protocol for culturing Drosophila melanogaster stage 9 egg chambers for live imaging. Nat Protoc 2:2467–2473
Van Doren M, Williamson AL, Lehmann R (1998) Regulation of zygotic gene expression in Drosophila primordial germ cells. Curr Biol 8:243–246
Neuman-Silberberg FS, Schüpbach T (1993) The Drosophila dorsoventral patterning gene gurken produces a dorsally localized RNA and encodes a TGF alpha-like protein. Cell 75:165–174
Neuman-Silberberg FS, Schüpbach T (1996) The Drosophila TGF-alpha-like protein Gurken: expression and cellular localization during Drosophila oogenesis. Mech Dev 59:105–113
Schüpbach T (1987) Germ line and soma cooperate during oogenesis to establish the dorsoventral pattern of egg shell and embryo in Drosophila melanogaster. Cell 49:699–707
Wasserman JD, Freeman M (1998) An autoregulatory cascade of EGF receptor signaling patterns the Drosophila egg. Cell 95:355–364
Peri F, Bokel C, Roth S (1999) Local Gurken signaling and dynamic MAPK activation during Drosophila oogenesis. Mech Dev 81:75–88
Gonzalez-Reyes A, Elliott H, St Johnston D (1995) Polarization of both major body axes in Drosophila by gurken-torpedo signalling. Nature 375:654–658
Roth S, Neuman-Silberberg FS, Barcelo G, Schüpbach T (1995) Cornichon and the EGF receptor signaling process are necessary for both anterior-posterior and dorsal-ventral pattern formation in Drosophila. Cell 81:967–978
Hawkins NC, Van Buskirk C, Grossniklaus U, Schüpbach T (1997) Post-transcriptional regulation of gurken by encore is required for axis determination in Drosophila. Development 124:4801–4810
Saunders C, Cohen RS (1999) The role of oocyte transcription, the 5′UTR, and translation repression and derepression in Drosophila gurken mRNA and protein localization. Mol Cell 3:43–54
Norvell A, Kelley RL, Wehr K, Schüpbach T (1999) Specific isoforms of squid, a Drosophila hnRNP, perform distinct roles in Gurken localization during oogenesis. Genes Dev 13:864–876
St Johnston D, Nüsslein-Volhard C (1992) The origin of pattern and polarity in the Drosophila embryo. Cell 68:201–219
Berleth T, Burri M, Thoma G, Bopp D, Richstein S, Frigerio G, Noll M, Nüsslein-Volhard C (1988) The role of localization of bicoid RNA in organizing the anterior pattern of the Drosophila embryo. EMBO J 7:1749–1756
Ephrussi A, Dickinson LK, Lehmann R (1991) Oskar organizes the germ plasm and directs localization of the posterior determinant nanos. Cell 66:37–50
McGrail M, Hays TS (1997) The microtubule motor cytoplasmic dynein is required for spindle orientation during germline cell divisions and oocyte differentiation in Drosophila. Development 124:2409–2419
Li M, McGrail M, Serr M, Hays TS (1994) Drosophila cytoplasmic dynein, a microtubule motor that is asymmetrically localized in the oocyte. J Cell Biol 126:1475–1494
Matthies HJ, Baskin RJ, Hawley RS (2001) Orphan kinesin NOD lacks motile properties but does possess a microtubule-stimulated ATPase activity. Mol Biol Cell 12:4000–4012
Cui W, Sproul LR, Gustafson SM, Matthies HJ, Gilbert SP, Hawley RS (2005) Drosophila Nod protein binds preferentially to the plus ends of microtubules and promotes microtubule polymerization in vitro. Mol Biol Cell 16:5400–5409
Shapiro RS, Anderson KV (2006) Drosophila Iκ2, a member of the IκB kinase family, is required for mRNA localization during oogenesis. Development 133:1467–1475
Abdu U, Bar D, Schüpbach T (2006) spn-F encodes a novel protein that affects oocyte patterning and bristle morphology in Drosophila. Development 133:1477–1484
Provost B, Varricchio P, Arana E, Espagne E, Falabella P, Huguet E, La Scaleia R, Cattolico L, Poirié M, Malva C, Olszewski JA, Pennacchio F, Drezen JM (2004) Bracoviruses contain a large multigene family coding for protein tyrosine phosphatases. J Virol 78:13090–13103
Fath-Goodin A, Kroemer JA, Webb BA (2009) The Campoletis sonorensis ichnovirus vankyrin protein P-vank-1 inhibits apoptosis in insect Sf9 cells. Insect Mol Biol 18:497–506
Siegrist SE, Doe CQ (2007) Microtubule-induced cortical cell polarity. Genes Dev 21:483–496
Affolter M, Weijer CJ (2005) Signaling to cytoskeletal dynamics during chemotaxis. Dev Cell 9:19–34
Etienne-Manneville S (2006) In vitro assay of primary astrocyte migration as a tool to study Rho GTPase function in cell polarization. Methods Enzymol 406:565–578
Pan X, Lührmann A, Satoh A, Laskowski-Arce MA, Roy CR (2008) Ankyrin repeat proteins comprise a diverse family of bacterial type IV effectors. Science 320:1651–1654
Acknowledgments
Special thanks go to Carla Malva who inspired this work with her really original and open-minded vision of science. We thank Trudi Schüpbach, Anne Ephrussi, Tom Hays, and Daniel St. Johnston for kindly providing us antibodies and fly strains. We also thank the Bloomington Stock Center for fly stocks and the Developmental Studies Hybridoma Bank for antibodies. We are grateful to Angela Algeri for the careful reading of the manuscript. Our work was supported by grants from the MIUR and University of Bologna (Prin 2006/2008, RFO 2006; 2007). SD had a PhD fellowship from the University of Bologna.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
18_2010_273_MOESM1_ESM.jpg
Distribution of TnBVANK1 protein. TnBVANK1 protein is localized in the cortical region of the oocyte, as shown by analyzing the oocyte just underneath its surface (A) where the protein shows a granular appearance, which is also detected in the cross section of the same stage 10B egg chamber (B). (JPG 1329 kb)
Rights and permissions
About this article
Cite this article
Duchi, S., Cavaliere, V., Fagnocchi, L. et al. The impact on microtubule network of a bracovirus IκB-like protein. Cell. Mol. Life Sci. 67, 1699–1712 (2010). https://doi.org/10.1007/s00018-010-0273-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0273-2