Abstract
Docetaxel is a chemotherapy drug to treat breast cancer, however as with many chemotherapeutic drugs resistance to docetaxel occurs in 50% of patients, and the underlying molecular mechanisms of drug resistance are not fully understood. Gene regulation through microRNAs (miRNA) has been shown to play an important role in cancer drug resistance. By directly targeting mRNA, miRNAs are able to inhibit genes that are necessary for signalling pathways or drug induced apoptosis rendering cells drug resistant. This study investigated the role of differential miRNA expression in two in vitro breast cancer cell line models (MCF-7, MDA-MB-231) of acquired docetaxel resistance. MiRNA microarray analysis identified 299 and 226 miRNAs altered in MCF-7 and MDA-MB-231 docetaxel-resistant cells, respectively. Docetaxel resistance was associated with increased expression of miR-34a and miR-141 and decreased expression of miR-7, miR-16, miR-30a, miR-125a-5p, miR-126. Computational target prediction revealed eight candidate genes targeted by these miRNAs. Quantitative PCR and western analysis confirmed decreased expression of two genes, BCL-2 and CCND1, in docetaxel-resistant cells, which are both targeted by miR-34a. Modulation of miR-34a expression was correlated with BCL-2 and cyclin D1 protein expression changes and a direct interaction of miR-34a with BCL-2 was shown by luciferase assay. Inhibition of miR-34a enhanced response to docetaxel in MCF-7 docetaxel-resistant cells, whereas overexpression of miR-34a conferred resistance in MCF-7 docetaxel-sensitive cells. This study is the first to show differences in miRNA expression, in particular, increased expression of miR-34a in an acquired model of docetaxel resistance in breast cancer. This serves as a mechanism of acquired docetaxel resistance in these cells, possibly through direct interactions with BCL-2 and CCND1, therefore presenting a potential therapeutic target for the treatment of docetaxel-resistant breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the most common disease in women worldwide with 44,000 new cases in the UK every year. One treatment strategy is chemotherapy with the taxane docetaxel (Taxotere®), which was shown to be more effective compared to other drugs including anthracyclines [1] and paclitaxel, another taxane [2]. It leads to cell cycle arrest and apoptosis by binding to and inhibiting the depolymerisation of the β-tubulin subunit of microtubules. As with other chemotherapeutic agents, many patients are either intrinsically resistant or acquire resistance to docetaxel during the course of treatment. Chemotherapy drug resistance involves various different mechanisms including gene mutations altering the mode of action of chemotherapeutic agents [3–6]. In addition, there is evidence that alteration of gene expression through DNA methylation and histone modifications can lead to drug resistance in cancer [7, 8]. MicroRNAs (miRNAs) are small molecules that are able to control gene expression by direct interaction with an mRNA target. These 19–25 nucleotide long molecules originate in the nucleus, where they are processed and exported into the cytoplasm [9]. Here, the mature miRNA exerts its gene regulatory function by binding to the 3′UTR of the target gene leading to either degradation of the mRNA transcript or inhibition of the protein translation process [10]. Advances in miRNA profiling showed that miRNAs play an important role in biological processes like development, differentiation and proliferation, therefore also influencing cancer development and progression. In addition, current research highlights their role in drug resistance in cancer. Studies have revealed a variety of miRNAs that can be directly linked to altering gene expression, therefore causing chemotherapy drug resistance in gastric [11] and prostate cancer [12]. In breast cancer, it was shown that miR-328 is associated with mitoxantrone-resistance [13] and increased miR-221 and miR-222 expression was associated with tamoxifen-resistant cells by targeting p27/kip1 [14]. Other miRNAs that can be associated with chemotherapy drug resistance in breast cancer include miR-326 [15] and miR-21 [16]. Even though miRNAs have been intensively studied, no study investigated the role of miRNA expression in docetaxel resistance in breast cancer. This study is the first to report miR-34a-mediated BCL-2 and CCND1 regulation in docetaxel-resistant breast cancer cells. More importantly, it was shown that increased miR-34a expression was associated with acquired docetaxel resistance and changes in miR-34a expression modulated response to docetaxel in these cells.
Materials and methods
Cell culture
Human breast cancer cell lines (MCF-7, MDA-MB-231) were obtained from the European Collection of Animal Cell Culture (Centre for Applied Microbiology and Research, Salisbury, UK). Each parental cell line (termed docetaxel-sensitive cells) was made resistant to docetaxel (termed docetaxel-resistant cells) following sequential exposure to docetaxel (Sanofi-Aventis, Surrey, UK) at increasing concentrations, as described previously [17]. Cells were cultured and maintained in RPMI-1640 medium including l-glutamine, supplemented with 10% (v/v) foetal calf serum and 1% (v/v) penicillin/streptomycin (100,000 U/l penicillin, 100 mg/l streptomycin), at 37°C in a humidified atmosphere containing 5% carbon dioxide.
RNA extraction
RNA was extracted from TRIzol® reagent frozen cells, according to the manufacturer’s protocol (Invitrogen, Paisley, UK). RNA quality and quantity was determined by Nanodrop spectrophotometry (Nanodrop 1000 Spectrophotometer, Thermo Scientific, Loughborough, UK). RNA was reverse transcribed with SuperScript™ II Reverse Transcriptase (Invitrogen), according to the manufacturer’s protocol, using random hexamers (GE Healthcare, Buckinghamshire, UK).
Small RNA isolation and reverse transcription
Total RNA was extracted as described. The RT2 qPCR-Grade miRNA isolation kit (SABiosciencesTM, Frederick, MD, USA) was used to isolate small RNA from total RNA. Briefly, 0.75 μg total RNA was used to isolate small RNA according to manufacturer’s instructions. Small RNA quantity and integrity was determined using the nanodrop (Thermo Scientific) and by Agilent Bioanalyser (Agilent Technologies UK Ltd., Wokingham, UK). Small RNA (150 ng) was reverse transcribed using the RT2 miRNA first strand kit according to manufacturer’s instructions (SABiosciencesTM).
Quantitative real-time PCR
Quantitative real-time PCR was performed using SYBR green master mix (Roche Diagnostics GmbH, Mannheim, Germany), according to the following PCR conditions: initial denaturation at 95°C for 5 min followed by 45 cycles of amplification at 95°C for 10 s and 60°C for 15 s. The amplified fluorescent signal was detected by Roche LightCycler 480 (Roche Diagnostics). Relative quantification was assessed using secondary derivative maximum (Roche Diagnostics). Gene expression was normalised to GAPDH and changes in expression measured relative to the control (docetaxel-sensitive cells). Table 1 represents the PCR primer sequences used (Sigma-Genosys, Haverhill, UK). For each gene studied, all experiments were repeated in triplicate using two independent cDNA extractions with RNA isolated from three independent RNA extractions.
miRNA quantitative PCR array analysis
MiRNA expression was measured using the RT2 miRNA PCR array system (SABiosciencesTM). Expression analysis of 376 miRNA sequences was performed according to manufacturer’s instructions. The LightCycler 480 was used for all reactions (Roche Diagnostics). PCR conditions were set according to SABioscience recommendations (SABiosciencesTM). Data analysis was performed using the RT² Profiler PCR Array Data Analysis Template v3.1 (SABiosciencesTM). Data was normalised to four miRNAs (hsa-SNORD-44, hsa-SNORD47, hsa-SNORD48 and hsa-U6) and relative miRNA expression levels were calculated with 2^(−ΔCt). Duplicate experiments were performed for each cell line.
miRNA target gene identification
For miRNA target gene identification computational analysis was used. The most recent version of miRecords (http://mirecords.biolead.org/; September 2009) was used to identify candidate target genes of identified miRNAs. To predict target genes, miRecords integrates predicted targets of 11 target prediction tools: DIANA-microT, MicroInspector, miRanda, MirTarget2, miTarget, NBmiRTar, PicTar, PITA, RNA22, RNAhybrid, and TargetScan/TargertScanS.
Western blot
Protein expression analysis was performed as described previously [17]. Briefly, 20 μg protein was separated by SDS-PAGE using polyacrylamide gels (Invitrogen) and transferred to a nitrocellulose membrane (Bio-Rad Laboratories Ltd., Hertfordshire, UK). BCL-2 (clone # sc-509) and cyclin D1 (clone # sc-2044) mouse monoclonal antibodies were obtained from Santa Cruz Biotechnology Inc. and used at a dilution of 1:500 and 1:750, respectively. β-actin (1:20000; clone # mAbcam 8226, Abcam plc, Cambridge, UK) was used as an internal loading control to be able to normalise the expression patterns of each protein of interest. Secondary antibodies were either horse radish peroxidase labelled goat anti-mouse, goat anti-rabbit or donkey anti-goat conjugated antibodies (Santa Cruz). All secondary antibodies were used at a 1:5000 dilution. Proteins were detected by chemiluminescence detection using the Amersham ECL plus western blotting detection kit (GE Healthcare) and visualised with the Biorad Flour S Imager (Bio-Rad) and densitometry was performed using Quantity One 4.6.3 software (Bio-Rad). Experiments were performed from three independent protein extractions.
miRNA-34a transfection
MiRNA-34a was overexpressed in MCF-7 docetaxel-sensitive cells using a miRNA-34a precursor (Applied Biosystems, Warrington, UK). In MCF-7 docetaxel-resistant cells miRNA-34a was knocked down using a miRNA-34a inhibitor (Exiqon A/S, Vedbaek, Denmark). 24 h prior to transfection, cells were plated onto a six-well plate at a density of 2 × 105 cells/well. Either 50 nM of the precursor or inhibitor molecule was transfected into the cells using jetPrime transfection reagent (Autogen Bioclear UK Ltd,Wilts, UK) according to manufacturer’s instructions. Three independent experiments were performed. Per experiment a positive control of Cy3-labelled scrambled miRNA oligoprobe (Applied Biosystems) was used to measure transfection efficiency. Transfection efficiency was calculated in percent where total cell count of mock-transfected cells was used as a control (100%). Typical transfection efficiencies in MCF-7 docetaxel-sensitive cells and resistant cells were ~90 and 65%, respectively.
Dual luciferase assay
The pGL3-BCL-2 wild-type and pGL3-BCL-2 mutant constructs were a kind gift of Dr Guido Bommer (De Duve Institute, Brussel, Belgium) [50]. Either 400 ng pGL3-BCL-2 wild-type or pGL3-BCL-2 mutant and 8 ng pGL4.70[hRluc] (Promega Corp., Madison, WI, USA) plasmid was co-transfected with 50 nM miRNA-34 precursor (Applied Biosystems). Transfected cells were incubated for 24 h before performing the dual luciferase assay according to manufacturer’s instructions (Promega). Firefly luciferase activity of the pGL3-control plasmid was normalised to Renilla luciferase activity of pGL4.70[hRluc]. Three independent experiments were performed.
Cell viability assay
In order to measure cell viability before and after transfection, the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay was used as described previously [17]. Briefly, 24 h post-transfection, 5 × 104 cells per well were seeded onto a 96-well plate in 100 μl medium and incubated for 24 h prior to docetaxel treatment. Appropriate docetaxel concentrations made from a 40 mg/ml stock were added at 100 μl and cells were incubated with the drug for 24 h. MTT (12 mM) (Sigma-Aldrich, Dorset, UK) was added to each well and cell viability was measured as described previously. Three independent experiments with six replicates each were performed.
Statistical analysis
Unpaired Student’s t test was performed for all experiments. All data, unless stated otherwise, are expressed as mean ± standard error of mean (SEM). A P value of less than 0.05 was considered statistically significant. SPSS Software, version 17 (SPSS Inc., Chicago, IL, USA) was used for all statistical analyses.
Results
miRNA expression is altered in docetaxel-resistant human breast cancer cells
To identify changes in miRNA expression in docetaxel-resistant human breast cancer cells, miRNA expression profiling of 376 human miRNAs was performed in two different in vitro breast cancer models of acquired docetaxel resistance (MCF-7, MDA-MB-231). About 40% of miRNAs had Ct values higher than 35, which were not included in the subsequent analysis. In MCF-7 docetaxel-resistant breast cancer cells miR-141 and miR-34a were increased in expression whereas miR-16, miR-138, miR-195, miR-7, miR-190b, miR-532-5p, miR-30d, miR129-3p, miR-30a, miR-151-3p, miR-590-3p, miR-151-5p, miR-126, miR-363, miR-125-5p, miR-149, and miR-361-5p were down regulated in MCF-7 docetaxel-resistant cells (Table 2). In contrast, only one of the altered miRNA sequences in MDA-MB-231 docetaxel-resistant cells reached statistical significance (Table 2). Here, miR-429 was downregulated in the docetaxel-resistant subline.
Identification of miRNA target genes
To identify gene targets for the altered miRNAs, computational in silico analysis was performed using the miRecords database. For the miRNAs that were significantly altered in MCF-7 docetaxel-resistant breast cancer cells, a subset of eight genes (Table 3) was chosen for further validation by qRT-PCR analysis to investigate their expression levels in MCF-7 docetaxel-sensitive and docetaxel-resistant breast cancer cells. These eight genes were chosen for further downstream analysis due to their known role in apoptosis, cell cycle regulation, and drug resistance.
Validation of identified target genes in docetaxel-resistant human breast cancer cells
In order to validate expression changes of the genes that were chosen to be further investigated, quantitative real-time PCR was performed in docetaxel-sensitive and docetaxel-resistant breast cancer cells. In MCF-7 cells, BCL-2, CCND1, CDK6, ERBB3, HMGB1, RAF1, SIRT1, and VEGF mRNA expression was measured. In MCF-7 docetaxel-resistant cells, BCL-2 (2.1-fold ± 0.24; P = 0.049), CCND1 (2-fold ± 0.21; P = 0.046), RAF1 (2-fold ± 0.03; P = 0.04), and HMGB1 (2.5-fold ± 0.09; P = 0.002) mRNA expression was significantly decreased compared to MCF-7 docetaxel-sensitive cells (Fig. 1). In contrast, SIRT1, VEGF, CDK6, and ERBB3 were not significantly altered. In MDA-MB-231 docetaxel-resistant cells, TUBB2A was not significantly changed, compared to MDA-MB-231 docetaxel-sensitive cells.
mRNA expression of potential miRNA target genes in human breast cancer cells. a Quantitative PCR measured BCL-2, CCND1, HMGB1, RAF1 gene expression between MCF-7 docetaxel-sensitive cells and docetaxel-resistant cells. The results are expressed as fold change (±SEM) relative to the control. Docetaxel-sensitive cells served as control. All experiments were repeated in triplicate using two independent cDNA extractions with RNA isolated from three independent RNA extractions. *P < 0.05, **P < 0.01
To investigate whether the selected genes exert functional effects, their expression was also measured on protein level. Eight proteins were investigated in MCF-7 docetaxel-sensitive and docetaxel-resistant cells and four proteins were investigated in MDA-MB-231 docetaxel-sensitive and docetaxel-resistant cells. Only BCL-2 and cyclin D1 showed statistically significant changes in MCF-7 docetaxel-resistant cells. BCL-2 was significantly downregulated more than 10-fold (±0.02, P < 0.001), and cyclin D1 expression by 4.6-fold (±0.11, P < 0.001) compared to docetaxel-sensitive cells (Fig. 2).
BCL-2 and cyclin D1 are decreased in MCF-7 docetaxel-resistant cells. Differences in protein expression of BCL-2 and cyclin D1 between docetaxel-sensitive cells and docetaxel-resistant cells. a Representative western blots of each protein investigated. b Relative protein expression is shown as fold change (±SD) relative to the control. Docetaxel-sensitive cells served as a control. β-actin was used as an internal loading control. All experiments were repeated in triplicate. *P < 0.05, **P < 0.01
miR-34a regulates BCL-2 and cyclin D1 in MCF-7 breast cancer cells
BCL-2 as well as cyclin D1 expression was decreased in MCF-7 docetaxel-resistant cells both at mRNA and protein level. Since these genes are known to be targeted by miR-34a, which was significantly increased in the docetaxel-resistant cells, it was hypothesised that miR-34a regulates BCL-2 and cyclin D1 expression. For this purpose, miR-34a was inhibited in MCF-7 docetaxel-resistant cells and BCL-2 and cyclin D1 protein expression was measured. After transfection with the miR-34a inhibitor, BCL-2 expression was increased 92.33-fold (±11.43; P < 0.001) compared to mock-transfected cells (Fig. 3a). Cyclin D1 was also increased after miR-34a inhibition by 1.85-fold (±0.53; P < 0.05) in MCF-7 docetaxel-resistant cells, compared to mock-transfected cells (Fig. 3a).
miR-34a regulates BCL-2 and cyclin D1 in MCF-7 breast cancer cells. a Differences in protein expression of BCL-2 and cyclin D1 in MCF-7 docetaxel-resistant cells transfected with or without miR-34a inhibitor. Representative western blot of each protein investigated (above). β-actin served as internal loading control. Relative protein expression (below) is shown as fold change (±SD) relative to the control. Docetaxel-resistant cells served as control. β-actin was used as an internal loading control. All experiments were repeated in triplicate. b BCL-2 mRNA transcript structure and miR-34a target site within human BCL-2. MiR-34a seed region in 202–208 base position in the BCL-2 3′UTR with seed matches is shown in the bottom box. c Relative luciferase levels were measured in MCF-7 docetaxel-resistant cells transfected with either pGL3 + BCL-2 wild-type plasmid or pGL3 + BCL-2 mutant plasmid and miR-34a precursor. The results are expressed as percentage of control (±SD). Cells transfected with pGL3 + BCL-2 wild-type plasmid or pGL3 + BCL-2 mutant plasmid only served as control (100%). All experiments were repeated in triplicate. *P < 0.05, **P < 0.01
In order to further investigate the regulatory function of miR-34a in BCL-2 expression, a dual luciferase reporter system was used to measure direct interaction between miR-34a and BCL-2 (Fig. 3b). Upon transfection of the miR-34a precursor, luciferase activity significantly decreased by 31.3% (±19.8; P < 0.05) in docetaxel-resistant cells that contained the 3′UTR with the miR-34a seed region of BCL-2 (Fig. 3c). In contrast, addition of the miR-34a precursor did not alter luciferase activity in docetaxel-resistant cells that contained a mutated seed region in the 3′UTR of the BCL-2 sequence (100.9% ± 15.9) (Fig. 3c). By reducing luciferase activity, this shows that miR-34a directly interacts with the 3′UTR region of the BCL-2 gene.
Modulation of miR-34a expression alters response to docetaxel in MCF-7 breast cancer cells
MiR-34a was significantly upregulated in MCF-7 docetaxel-resistant cells. To investigate the role of miR-34a in acquired docetaxel resistance cell viability was measured by MTT assay after transfection with a miR-34a precursor in MCF-7 docetaxel-sensitive cells or after transfection with a miR-34a inhibitor in MCF-7 docetaxel-resistant cells. Treatment with the miR-34a precursor affected cell growth particularly at lower docetaxel concentrations (Fig. 4a, left) and significantly increased resistance to docetaxel in docetaxel-sensitive cells (IC50sensitive: 2.9 μM ± 4.9 versus IC50miR-34a precursor: 19.6 μM ± 5.8; P = 0.01) (Fig. 4a, right). Vice versa, in MCF-7 docetaxel-resistant cells, inhibition of miR-34a diminished cell viability (Fig. 4b, left) and significantly enhanced response to docetaxel (IC50resistant: 22.1 μM ± 1.3 versus IC50miR-34a inhibitor: 18.8 μM ± 0.4; P = 0.01) (Fig. 4b, right).
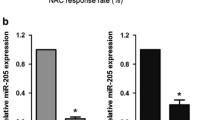
miR-34a modulates docetaxel response in MCF-7 breast cancer cells. Cellular viability was measured in MCF-7 cells either mock transfected or treated with 50 nM a miR-34a precursor (MCF-7 docetaxel-sensitive) or b miR-34a inhibitor (MCF-7 docetaxel-resistant) for 24 h. Left: Mean MTT readings of three independent experiments showing growth effects upon addition of miR-34a precursor relative to mock treatment in MCF-7 sensitive cells (a) or miR-34a inhibitor in MCF-7 resistant cells (b), respectively. Right: The results are expressed as IC50 values (±SD), which represent docetaxel concentrations that inhibit 50% cell growth. All experiments were repeated in triplicate with six replicates per experiment. *P < 0.05
Discussion
MiR-34a is a widely studied human miRNA that was associated with cancer cell growth and proliferation in a variety of cancers [26, 27] and recently, evidence emerged that miR-34a is also associated with drug resistance in cancer [28, 29]. Here, both studies identified miR-34a in prostate cancer cells that, upon overexpression sensitised the cells towards camptothecin and paclitaxel, respectively, possibly by targeting SIRT1 and BCL-2. In contrast, increased miR-34a expression was seen in the present study in MCF-7 docetaxel-resistant breast cancer cells and docetaxel response could be altered upon miR-34a modulation. It was shown that miR-34a induces G1 cell cycle arrest in cancer cells [12, 30]. Docetaxel primarily acts in G2-M phase, whereas it has diminished activity in G1 phase. Increased miR-34a expression may therefore be able to inhibit docetaxel activity by arresting cells in G1 phase. MiRNAs have not only been described to act as tumour suppressors but also as oncogenes [31, 32]. This study, therefore, suggests that miR-34a exerts oncogenic properties hence mediating docetaxel resistance in these cells.
Cyclin D1 was significantly decreased at mRNA and protein level in MCF-7 docetaxel-resistant cells and was identified as candidate target gene for miR-34a. By binding to and forming a complex with CDK4 and CDK6, cyclin D1 supports cell cycle transition from G1 to S phase. Besides its role in cell cycle regulation and dependent on the tissue, it is expressed in cyclin D can either be pro-apoptotic or anti-apoptotic [33]. Overexpression of cyclin D1 has been shown to be associated with cisplatin resistance in cancers [34–36]. In contrast, decreased cyclin D1 expression was found in MCF-7 doxorubicin-resistant and MCF-7 cisplatin-resistant breast cancer cells [37] and was associated with multi-drug resistance in human colon carcinoma cells [38]. Also, in this study decreased cyclin D1 mRNA and protein expression was observed in MCF-7 docetaxel-resistant breast cancer cells. Since cyclin D1 has different functions dependent on cell or tissue type it may be that its pro-apoptotic effects may be more prevalent in the MCF-7 cells, whereas its role as tumour suppressor and its anti-apoptotic effects may be more prevalent in other studies. Docetaxel inhibits microtubule depolymerisation in G2-M phase that subsequently causes cell cycle arrest and apoptosis. Cyclin D1 is required for G1-S transition, therefore in the case of absent or decreased cyclin D1 expression, as observed in the MCF-7 docetaxel-resistant cells, G1 arrest occurs [39–42]. Docetaxel only exerts minimal toxicity in G1 phase so that decreased cyclin D1 expression would counteract docetaxel-induced apoptosis hence leading to docetaxel resistance in MCF-7 cells.
Decreased BCL-2 expression was observed in MCF-7 docetaxel-resistant cells and BCL-2 represented another candidate gene of miR-34a that was identified through computational in silico analysis. In tamoxifen-resistant breast cancer cells, anti-sense BCL-2 treatment enhanced sensitivity to tamoxifen in HER2-positive cells [43]. In addition, BCL-2 overexpression, causes paclitaxel resistance in MCF-7 cells [44]. In this study, however, MCF-7 docetaxel-resistant cells, showed a significant decrease in BCL-2 expression both on mRNA and protein level. This is an unexpected finding, since a decrease of anti-apoptotic protein would lead to the hypothesis that apoptosis would increase with a chemotherapeutic drug such as docetaxel. Besides its anti-apoptotic function, however, increased BCL-2 expression is indicative of a favourable prognostic response in breast cancer patients [45]. Hormonal treatment in breast cancer treatment was also shown to be more beneficial in breast cancer patients, with longer disease-free survival and overall survival rates, when BCL-2 was increased [46–48]. Cardoso and colleagues reported a greater benefit of tamoxifen treatment in BCL-2-positive patients with node-positive breast cancer [49]. These studies show, that, even though BCL-2 is known as an anti-apoptotic protein, increased BCL-2 expression can be of favour in breast cancer treatment, whereas decreased BCL-2 expression is associated with drug resistance or poorer outcome. The mechanisms by which decreased BCL-2 expression can negatively influence the activity of drugs like docetaxel are currently not known. The apoptotic pathway is a complex machinery that involves a variety of apoptotic genes, therefore it may be possible that other factors influence the cell whether to enter or evade docetaxel-induced apoptosis.
In this study, a direct interaction between miR-34a and the 3′UTR of BCL-2 was shown in MCF-7 docetaxel-resistant cells by dual luciferase assay, supporting various reports of the current literature on miR-34a-BCL-2 interactions [26, 28, 50, 51]. In addition, miR-34a modulated response to docetaxel in breast cancer cells. This suggests that miR-34a-mediated docetaxel resistance occurs through regulation of BCL-2. However, BCL-2 is known to be anti-apoptotic, therefore the biological role of decreased BCL-2 in docetaxel resistance remains unclear and can only be speculated upon. The BCL-2 family contains a variety of pro-apoptotic and anti-apoptotic genes which may also be a target of miR-34a. Therefore, alterations of miR-34a may trigger a whole cascade of apoptotic gene changes, whereby BCL-2 expression is predominated by increased expression of other anti-apoptotic genes, which in turn may lead to resistance to docetaxel.
Whereas miR-34a expression could be associated with docetaxel resistance in MCF-7 cells, no such association was identified in MDA-MB-231 cells. This may have several reasons and possible implications for our understanding about cell line specific miRNA-mediated gene regulation. MDA-MB-231 cells are of a more aggressive phenotype and compared to the luminal-like MCF-7 cells represent the basal type of breast cancer in vitro. It was shown that miRNA regulatory networks depend on oestrogen receptor alpha which is expressed in luminal-like breast cancer cells [52, 53]. Particularly miR-34a could be a miRNA that is at least in part dependent on factors like oestrogen receptor alpha, and therefore was not identified in the basal type MDA-MB-231 cells. Additionally, the miRNAs investigated in the present study may not have included other miRNAs that could have played an important role in acquired docetaxel resistance in MDA-MB-231 cells which needs to be further investigated in the future. Besides showing that miR-34a is important in docetaxel resistance in MCF-7 cells, this study also presents some data on cell line-specific effects of miRNAs which is an important finding considering future therapeutical applications where miRNA profiling may be applied to identify responders as well as non-responders to chemotherapeutic treatment in breast cancer.
References
Smith IC, Heys SD, Hutcheon AW, Miller ID, Payne S, Gilbert FJ, Ah-See AK, Eremin O, Walker LG, Sarkar TK, Eggleton SP, Ogston KN (2002) Neoadjuvant chemotherapy in breast cancer: significantly enhanced response with docetaxel. J Clin Oncol 20:1456–1466
Jones SE, Erban J, Overmoyer B, Budd GT, Hutchins L, Lower E, Laufman L, Sundaram S, Urba WJ, Pritchard KI, Mennel R, Richards D, Olsen S, Meyers ML, Ravdin PM (2005) Randomized phase III study of docetaxel compared with paclitaxel in metastatic breast cancer. J Clin Oncol 23:5542–5551
Modok S, Mellor HR, Callaghan R (2006) Modulation of multidrug resistance efflux pump activity to overcome chemoresistance in cancer. Curr Opin Pharmacol 6:350–354
Miyoshi Y, Taguchi T, Kim SJ, Tamaki Y, Noguchi S (2005) Prediction of response to docetaxel by immunohistochemical analysis of CYP3A4 expression in human breast cancers. Breast Cancer 12:11–15
Noguchi S (2006) Predictive factors for response to docetaxel in human breast cancers. Cancer Sci 97:813–820
Shalli K, Brown I, Heys SD, Schofield AC (2005) Alterations of beta-tubulin isotypes in breast cancer cells resistant to docetaxel. FASEB J 19:1299–1301
Kastl L, Brown I, Schofield AC (2010) Altered DNA methylation is associated with docetaxel resistance in human breast cancer cells. Int J Oncol 36:1235–1241
Zhang XT, Yashiro M, Ren J, Hirakawa K (2006) Histone deacetylase inhibitor, trichostatin A, increases the chemosensitivity of anticancer drugs in gastric cancer cell lines. Oncol Rep 16:563–568
Miska EA (2005) How microRNAs control cell division, differentiation and death. Curr Opin Genet Dev 15:563–568
Ying SY, Chang DC, Lin S (2008) The microRNA (miRNA): overview of the RNA genes that modulate gene function. Mol Biotechnol 38:257–268
Xia L, Zhang D, Du R, Pan Y, Zhao L, Sun S, Hong L, Liu J, Fan D (2008) miR-15b and miR-16 modulate multidrug resistance by targeting BCL-2 in human gastric cancer cells. Int J Cancer 123:372–379
Sun F, Fu H, Liu Q, Tie Y, Zhu J, Xing R, Sun Z, Zheng X (2008) Downregulation of CCND1 and CDK6 by miR-34a induces cell cycle arrest. FEBS Lett 582:1564–1568
Pan YZ, Morris ME, Yu AM (2009) MicroRNA-328 negatively regulates the expression of breast cancer resistance protein (BCRP/ABCG2) in human cancer cells. Mol Pharmacol 75:1374–1379
Miller TE, Ghoshal K, Ramaswamy B, Roy S, Datta J, Shapiro CL, Jacob S, Majumder S (2008) MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J Biol Chem 283:29897–29903
Liang Z, Wu H, Xia J, Li Y, Zhang Y, Huang K, Wagar N, Yoon Y, Cho HT, Scala S, Shim H (2010) Involvement of miR-326 in chemotherapy resistance of breast cancer through modulating expression of multidrug resistance-associated protein 1. Biochem Pharmacol 79:817–824
Mei M, Ren Y, Zhou X, Yuan XB, Han L, Wang GX, Jia Z, Pu PY, Kang CS, Yao Z (2010) Downregulation of miR-21 enhances chemotherapeutic effect of taxol in breast carcinoma cells. Technol Cancer Res Treat 9:77–86
Brown I, Shalli K, McDonald SL, Moir SE, Hutcheon AW, Heys SD, Schofield AC (2004) Reduced expression of p27 is a novel mechanism of docetaxel resistance in breast cancer cells. Breast Cancer Res 6:601–607
Webster RJ, Giles KM, Price KJ, Zhang PM, Mattick JS, Leedman PJ (2009) Regulation of epidermal growth factor receptor signaling in human cancer cells by microRNA-7. J Biol Chem 284:5731–5741
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M, Wojcik SE, Aqeilan RI, Zupo S, Dono M, Rassenti L, Alder H, Volinia S, Liu CG, Kipps TJ, Negrini M, Croce CM (2005) miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 102:13944–13949
Liu Q, Fu H, Sun F, Zhang H, Tie Y, Zhu J, Xing R, Sun Z, Zheng X (2008) miR-16 family induces cell cycle arrest by regulating multiple cell cycle genes. Nucleic Acids Res 36:5391–5404
Nakamoto M, Jin P, O’Donnell WT, Warren ST (2005) Physiological identification of human transcripts translationally regulated by a specific microRNA. Hum Mol Genet 14:3813–3821
Yamakuchi M, Ferlito M, Lowenstein CJ (2008) miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci USA 105:13421–13426
Scott GK, Goga A, Bhaumik D, Berger CE, Sullivan CS, Benz CC (2007) Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b. J Biol Chem 282:1479–1486
Liu B, Peng XC, Zheng XL, Wang J, Qin YW (2009) MiR-126 restoration down-regulate VEGF and inhibit the growth of lung cancer cell lines in vitro and in vivo. Lung Cancer 66:169–175
Beitzinger M, Peters L, Zhu JY, Kremmer E, Meister G (2007) Identification of human microRNA targets from isolated argonaute protein complexes. RNA Biol 4:76–84
Cole KA, Attiyeh EF, Mosse YP, Laquaglia MJ, Diskin SJ, Brodeur GM, Maris JM (2008) A functional screen identifies miR-34a as a candidate neuroblastoma tumor suppressor gene. Mol Cancer Res 6:735–742
Li N, Fu H, Tie Y, Hu Z, Kong W, Wu Y, Zheng X (2009) miR-34a inhibits migration and invasion by down-regulation of c-Met expression in human hepatocellular carcinoma cells. Cancer Lett 275:44–53
Fujita Y, Kojima K, Hamada N, Ohhashi R, Akao Y, Nozawa Y, Deguchi T, Ito M (2008) Effects of miR-34a on cell growth and chemoresistance in prostate cancer PC3 cells. Biochem Biophys Res Commun 377:114–119
Kojima K, Fujita Y, Nozawa Y, Deguchi T, Ito M (2010) MiR-34a attenuates paclitaxel-resistance of hormone-refractory prostate cancer PC3 cells through direct and indirect mechanisms. Prostate 70:1501–1512
Tarasov V, Jung P, Verdoodt B, Lodygin D, Epanchintsev A, Menssen A, Meister G, Hermeking H (2007) Differential regulation of microRNAs by p53 revealed by massively parallel sequencing: miR-34a is a p53 target that induces apoptosis and G1-arrest. Cell Cycle 6:1586–1593
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Kent OA, Mendell JT (2006) A small piece in the cancer puzzle: microRNAs as tumor suppressors and oncogenes. Oncogene 25:6188–6196
Han EK, Ng SC, Arber N, Begemann M, Weinstein IB (1999) Roles of cyclin D1 and related genes in growth inhibition, senescence and apoptosis. Apoptosis 4:213–219
Biliran H Jr, Wang Y, Banerjee S, Xu H, Heng H, Thakur A, Bollig A, Sarkar FH, Liao JD (2005) Overexpression of cyclin D1 promotes tumor cell growth and confers resistance to cisplatin-mediated apoptosis in an elastase-myc transgene-expressing pancreatic tumor cell line. Clin Cancer Res 11:6075–6086
Henriksson E, Baldetorp B, Borg A, Kjellen E, Akervall J, Wennerberg J, Wahlberg P (2006) p53 mutation and cyclin D1 amplification correlate with cisplatin sensitivity in xenografted human squamous cell carcinomas from head and neck. Acta Oncol 45:300–305
Zhang P, Zhang Z, Zhou X, Qiu W, Chen F, Chen W (2006) Identification of genes associated with cisplatin resistance in human oral squamous cell carcinoma cell line. BMC Cancer 6:224
Lukyanova NY, Rusetskya NV, Tregubova NA, Chekhun VF (2009) Molecular profile and cell cycle in MCF-7 cells resistant to cisplatin and doxorubicin. Exp Oncol 31:87–91
Fan CW, Chan CC, Chao CC, Fan HA, Sheu DL, Chan EC (2004) Expression patterns of cell cycle and apoptosis-related genes in a multidrug-resistant human colon carcinoma cell line. Scand J Gastroenterol 39:464–469
Resnitzky D, Gossen M, Bujard H, Reed SI (1994) Acceleration of the G1/S phase transition by expression of cyclins D1 and E with an inducible system. Mol Cell Biol 14:1669–1679
Resnitzky D, Reed SI (1995) Different roles for cyclins D1 and E in regulation of the G1-to-S transition. Mol Cell Biol 15:3463–3469
Lukas J, Aagaard L, Strauss M, Bartek J (1995) Oncogenic aberrations of p16INK4/CDKN2 and cyclin D1 cooperate to deregulate G1 control. Cancer Res 55:4818–4823
Lukas J, Pagano M, Staskova Z, Draetta G, Bartek J (1994) Cyclin D1 protein oscillates and is essential for cell cycle progression in human tumour cell lines. Oncogene 9:707–718
Kim R, Tanabe K, Emi M, Uchida Y, Toge T (2005) Modulation of tamoxifen sensitivity by antisense Bcl-2 and trastuzumab in breast carcinoma cells. Cancer 103:2199–2207
Huang Y, Ray S, Reed JC, Ibrado AM, Tang C, Nawabi A, Bhalla K (1997) Estrogen increases intracellular p26Bcl-2 to p21Bax ratios and inhibits taxol-induced apoptosis of human breast cancer MCF-7 cells. Breast Cancer Res Treat 42:73–81
Hori M, Nogami T, Itabashi M, Yoshimi F, Ono H, Koizumi S (1997) Expression of Bcl-2 in human breast cancer: correlation between hormone receptor status, p53 protein accumulation and DNA strand breaks associated with apoptosis. Pathol Int 47:757–762
Elledge RM, Green S, Howes L, Clark GM, Berardo M, Allred DC, Pugh R, Ciocca D, Ravdin P, O’Sullivan J, Rivkin S, Martino S, Osborne CK (1997) bcl-2, p53, and response to tamoxifen in estrogen receptor-positive metastatic breast cancer: a Southwest Oncology Group Study. J Clin Oncol 15:1916–1922
el-Ahmady O, El-Salahy E, Mahmoud M, Wahab MA, Eissa S, Khalifa A (2002) Multivariate analysis of bcl-2, apoptosis, P53 and HER-2/neu in breast cancer: a short-term follow-up. Anticancer Res 22:2493–2499
Bilalovic N, Vranic S, Hasanagic S, Basić H, Tatarević A, Beslija S, Selak I (2004) The Bcl-2 protein: a prognostic indicator strongly related to ER and PR in breast cancer. Bosn J Basic Med Sci 4:5–12
Cardoso F, Paesmans M, Larsimont D, Durbecq V, Bernard-Marty C, Rouas G, Dolci S, Sotiriou C, Piccart MJ, Di Leo A (2004) Potential predictive value of Bcl-2 for response to tamoxifen in the adjuvant setting of node-positive breast cancer. Clin Breast Cancer 5:364–369
Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, Zhai Y, Giordano TJ, Qin ZS, Moore BB, MacDougald OA, Cho KR, Fearon ER (2007) p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 17:1298–1307
Wang X, Liu P, Zhu H, Xu Y, Ma C, Dai X, Huang L, Liu Y, Zhang L, Qin C (2009) miR-34a, a microRNA up-regulated in a double transgenic mouse model of Alzheimer’s disease, inhibits BCL-2 translation. Brain Res Bull 80:268–273
Cicatiello L, Mutarelli M, Grober OM, Paris O, Ferraro L, Ravo M, Tarallo R, Luo S, Schroth GP, Seifert M, Zinser C, Chiusano ML, Traini A, De Bortoli M, Weisz A (2010) Estrogen receptor alpha controls a gene network in luminal-like breast cancer cells comprising multiple transcription factors and microRNAs. Am J Pathol 176:2113–2130
Kondo N, Toyama T, Sugiura H, Fujii Y, Yamashita H (2008) miR-206 Expression is down-regulated in estrogen receptor alpha-positive human breast cancer. Cancer Res 68:5004–5008
Acknowledgments
The authors especially thank Dr Alun Hughes, Dr Scott Davidson and Dr Sandra Stoppelkamp for their help with performing the quantitative PCR and the dual luciferase studies. Special acknowledgements to Dr Guido Bommer for providing the pGL3-BCL2 wild type and mutant plasmids. This work was supported by TENOVUS Scotland, the Dr James Alexander Mearns, PhD studentship and the Fraserburgh Moonlight Prowl Fund.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kastl, L., Brown, I. & Schofield, A.C. miRNA-34a is associated with docetaxel resistance in human breast cancer cells. Breast Cancer Res Treat 131, 445–454 (2012). https://doi.org/10.1007/s10549-011-1424-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-011-1424-3