Abstract
The CMV (cucumber mosaic virus) is the most frequently occurring virus in chili pepper farms. A variety of peppers that are resistant to CMVP0 were developed in the middle of 1990s through a breeding program, and commercial cultivars have since been able to control the spread of CMVP0. However, a new pathotype (CMVP1) that breaks the resistance of CMVP0-resistant peppers has recently appeared and caused a heavy loss in productivity. Since no genetic source of this new pathotype was available, a traditional breeding method cannot be used to generate a CMVP1-resistant pepper variety. Therefore, we set up a transformation system of pepper using Agrobacterium that had been transfected with the coat protein gene, CMVP0-CP, with the aim of developing a new CMVP1-resistant pepper line. A large number of transgenic peppers (T1, T2 and T3) were screened for CMVP1 tolerance using CMVP1 inoculation. Transgenic peppers tolerant to CMVP1 were selected in a plastic house as well as in the field. Three independent T3 pepper lines highly tolerant to the CMVP1 pathogen were found to also be tolerant to the CMVP0 pathogen. These selected T3 pepper lines were phenotypically identical or close to the non-transformed lines. However, after CMVP1 infection, the height and fruit size of the non-transformed lines became shorter and smaller, respectively, while the T3 pepper lines maintained a normal phenotype.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant virus infection is a major factor in crop yield and has been responsible for causing severe losses in pepper crop production. The cucumber mosaic virus (CMV), which is in the cucumovirus genus, has been the most detrimental in regard to pepper cultivation. Since the early 1980s the FNY-CMV strain (known as CMVP0 or Ca-P0, Lee et al. 2006) has spread over most of the Korean pepper farms. In the middle of the 1990s, CMVP0-resistant pepper strains were developed by pepper breeders in Korea and have been commercially available since then. However, very recently, the CMVP0-resistant strains have become susceptible to infection by a new CMVP0 resistance-breaking virus. This new CMV strain was identified as CMVP1 (called as Ca-P1, Lee et al. 2006) and the CMVP1 outbreak has damaged a large portion of pepper cultivation and production. Recently, this virus has been detected in almost all Asian countries and since only a couple of recessively inherited domestic varieties are available, it has been very difficult to develop a resistant cultivar to this new CMV strain (personal conversation with Dr. Moon Hwan Lee, Nongwoo Bio Co.).
However, a genetic transformation technique could overcome the problems that are typically associated with an ordinary breeding program. A CP (coat protein) gene has been widely used to enhance the tolerance levels of viral disease in different plant species, including tobacco (Powell-Abel et al. 1986; Cuozzo et al. 1988; Nida et al. 1992; Linbo and Dougherty 1992); tomato (Nelson et al. 1988; Zrachya et al. 2007); cantaloupe (Clough and Hamm 1995), melon (Fuchs et al. 1998a), grapevine (Krastanova et al. 1995; Gölles et al. 2000; Mauro et al. 1995; Maghuly et al. 2006; Ling et al. 2008), papaya (Fitch et al. 1992; Tennant et al. 1994; Bau et al. 2003; Davis and Ying 2004; Krubphachaya et al. 2007), orange (Iwanami and Shimizu 2004), sweet orange (Zanek et al. 2008), soybean (Di et al. 1996; Reddy et al. 2001; Tougou et al. 2006), squash (Clough and Hamm 1995; Fuchs and Gonsalves 1995; Tricoli et al. 1995; Fuchs et al. 1998a, b; Pang et al. 2000; Klas et al. 2006), sugarcane (Jin et al. 2007) and watermelon rootstock (Park et al. 2005). Among these genetically modified plants, tolerant strains against the papaya ring virus and the watermelon mosaic virus squash have been successfully commercialized (James 2008).
A similar method that used the CP gene has also been successfully demonstrated in the chili pepper plant (CMV and ToMV, Shin et al. 2002; CMV and TMV, Cai et al. 2003; Lee et al. 2004) and sweet peppers (Zhu et al. 1996). Since the previous transformation methods were still recalcitrant in chili pepper, GM chili peppers are not yet commercially available. However, very recently, virus resistant GM sweet peppers have been cultivated in China (James 2008). There are three major steps that must be overcome before virus tolerant GM peppers can become commercially available. The first of these steps is to use a reliable transformation method. The second is to have a CP gene that can cover a broad spectrum of the viruses’ defenses through DNA sequence identity. The third is to develop a transgenic pepper line with multi-resistant traits through breeding (transgenic breeding).
Here we present transgenic pepper lines developed using a CP gene cloned from CMVP0, a previously dominant virus. The transgenic peppers were highly tolerant to CMVP1, the new pathotype, as well as CMVP0.
Materials and methods
Genetic transformation of pepper
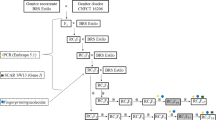
Seeds of three pepper inbred lines (P915, P2377 and Ph240; properties of Nongwoo Bio Co.) were surface-disinfected in 95% EtOH for 30 s and 25% bleach (Yuhanrox) for 30 min, and then rinsed three times with sterilized water. The sterilized seeds were placed in 1/2 MS medium (Murashige and Skoog 1962) and allowed to germinate in the dark at 25°C. Cotyledons from 3-day-old seedlings were excised and used as explants for regeneration and transformation. For the Agrobacterium-mediated transformation of peppers, explants were transferred to a pre-culture medium that consisted of MS medium supplemented with zeatin 2.0 mg l−1 and IAA 0.1 mg l−1. The explants were then placed in a lighted room at 25°C for 36 h. Agrobacterium EHA105, which contained a binary vector with a 35S CaMV promoter and the NPTII gene for kanamycin selection along with the CMVP0-CP gene of the FNY-CMV strain, was grown to the log phase in YEP liquid media (OD600: 0.3–0.5) and used for the co-culture. The pepper transformation method used in this study was modified from the one described by Lee et al. (2004).
PCR analysis
To detect the CMVP0-CP gene in transformed pepper plants by PCR, total DNA was isolated using a DNA extraction kit (iNtRON Biotechnology, http://www.intronbio.com). The PCR primer sequences used for detecting the CMVP0-CP gene insertion were: 5′-ATGACGCACAATCCCACTAT-3′ (sense: 35S promoter region at 3,185-3,204 bp of accession number X84105) and 5′-GGGGTACCTCAGACTGGGAGCACTCC-3′ (antisense: CMVP0-CP gene at 639–657 bp of accession number D10538). PCR analysis was carried out using these primers in a reaction solution that contained 0.65 μM, 299 μM dNTP, 1 U/μM of Taq DNA polymerase (BioLabs, http://www.neb.com) in 50 mM KCl, 1.5 mM MgCl2, and 10 mM Tris–HCl pH 8.3. The PCR program consisted of 35 amplification cycles of 94, 55 and 72°C, each for 1 min.
Southern blot analysis
For the genomic Southern blot analysis, DNA from T0 plants was isolated using the method described by Sambrook et al. (1989). 30 μg of DNA was then digested with DraI and XbaI and fractionated on 0.8% agarose gel. Southern blotting was performed as previously described (Church and Gilbert 1984; Sambrook et al. 1989) using Hybond N membranes (Amersham Biosciences, http://www.amershambiosciences.com) and hybridization with a 32P-labeled probe containing the CMVP0-CP gene (657 bp; D10538) as instructed by the manufacturer (Amersham Biosciences, http://www.amershambiosciences.com).
Test for disease tolerance to CMVP1 and selection for tolerant T generation
In 2004, CMVP1 was isolated from the Manidda, which is one of the hot pepper varieties grown in Korea. CMVP1 was propagated by inoculating it in tobacco and applying the crude sap of the tobacco leaves to the pepper (as described in Lee et al. 2006). This method was also used for CMVP0 inoculation. Approximately 60–120 seeds from seven independent T1 pepper lines were planted in a small multi-plug (4 × 4 × 4 cm). When the small seedlings were at the two-leaf stage, they were exposed to CMVP1 by scraping with carborundum that had been dipped in the crude sap. A month later, a leaf disk was taken from each transgenic pepper and subjected to an ELISA assay using the indirect ELISA kit (Bioreba, http://www.bioreba.ch). Readings were taken at A405 nm using an ELISA Thermo Max Microplate Reader (Molecular Devices, http://www.moleculardevices.com). The CMVP1-tolerant peppers from the T1 population were selected and self-crossed. Seedlings from the T2 pepper lines were exposed to CMVP1, and the CMVP1 tolerant T2 peppers were selected and self-crossed. The same experiment was performed as described earlier with the T3 peppers and CMVP1 tolerant T3 peppers were selected. These T3 peppers were exposed to CMVP0 using the same method as described earlier for the CMVP1 treatment.
Results and discussion
Genetic transformation of pepper
Four years ago, A CP (coat protein) gene was cloned from the CMVP0 (FNY-CMV) strain and subcloned into a pCambia vector 2300 that had been modified for genetic transformation with Agrobacterium (Fig. 1). A total of 1,932 explants from inbred lines (P915, P2377 and Ph240) were used for co-culture and 9 T0 peppers were obtained with a transformation ratio of 0.43–0.66% (Table 1). The transformation method used here was the callus-induced transformation (CIT) method, which was established by modifying the callus-mediated shoot formation method (Lee et al. 2004).
The insert gene for genetic transformation was FNY CMVP0-CP, which shares 93% DNA identity (Fig. 2) and 96% AA identity with the newly identified CMVP1-CP gene (data not shown). This difference is due to the subgroup specification of the cucumovirus. By sequence comparison of RNA3 (encoding movement protein and coat protein) with known representative strains of CMV, the phylogenetic tree analysis showed that the Ca-P1-CMV belongs to a typical member of the CMV subgroup IB while the FNY-CMVP0 strain belongs to the subgroup 1A (Lee et al. 2006). All of the Korean pepper lines and varieties that were resistant to CMVP0 (Ca-P0-R gene included) were susceptible to CMVP1, indicating that the pathogenicity of CMVP1 was much greater than that of CMVP0. However, the phenotypes of the mosaic patterns on the leaves were not distinguishable when the peppers were exposed to either CMVP0 or CMVP1.
Nucleotide sequence comparison between CMVP0-CP (657 bp) and CMVP1-CP (657 bp). The nucleotide sequence identity was 93% and the amino acid sequence identity was 96%. The start and stop codons were colored blue and red, respectively. The different base pairs were in bold. By sequence comparison of the RNA3 (encoding movement protein and coat protein) of CMV strains, the phylogenetic tree analysis indicates the Ca-P1-CMV belongs to a typical member of the CMV subgroup IB while the FNY-CMVP0 strain belongs to the subgroup 1A
PCR and Southern blot analysis
The genomic DNA from nine putative T0 peppers was isolated and subjected to PCR. The 740-bp band in lanes 1, 2, 5, 6, 7, 8 and 9 in Fig. 3 was the PCR product, which includes the 3′ end region of the 35S promoter (88 bp) and the CMVP0-CP gene. However, two T0 pepper lines (lane 3 and 4) did not display this band. Any band that appeared on the gel with the 35S promoter region was believed to be the real PCR product containing the insert gene.
Genomic DNA was then isolated from six T0 peppers samples (1, 2, 5, 6, 7 and 9) and 30 μg of the genomic DNA was digested with DraI and XbaI (enzymes that do not cut the insert) and fractionated on 0.8% agarose gel. The resulting restriction bands shown in the Southern blot confirmed the presence of the CMVP0-CP gene in the T0 peppers (Fig. 4). Lanes 1, 2, 7 and 9 showed a different single band when digested with each restriction enzyme, DraI or XbaI, suggesting that these peppers contain a single copy gene of CMVP0-CP. Sample 5 and 6 had the same band at 3.6 kb after DraI digestion and treatment with XbaI resulted in two weak bands at 9 and 1.5 kb and one strong band at 7 kb in both sample 5 and 6. These results indicate that samples 5 and 6 were of the same origin and must have probably been obtained from different shoots of the same callus origin.
CMVP1 tolerance test of transgenic peppers
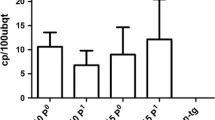
Independent T0 peppers with a single copy of CMVP0-CP were self-crossed and T1 peppers from each T0 plant were obtained. A total of 595 T1 pepper seedlings, at the two-leaf stage, were exposed to the CMVP1 pathogen and 3 weeks after exposure, an initial screen was conducted by checking for the existence of mosaic symptoms in the leaves (eye-judgment) (Table 2). 208 T1 peppers did not show any CMV symptoms while all of the non-transformed peppers showed mosaic symptoms. PCR analysis was performed to determine the presence of the CMVP0-CP insert in the 208 T1 peppers that did not show any CMV symptoms. From this analysis, 97% of the T1 peppers were shown to contain the insert gene (data not shown). The first ELISA test was performed 1 month after exposure and the second ELISA test was conducted 2 months after exposure. The two consecutive ELISA analyses revealed that 19 peppers were highly tolerant to CMVP1 infection. These 19 plants that possessed the insert and were tolerant to CMVP1 for up to 120 days after inoculation in the plastic house (data not shown), were self-crossed and T2 peppers were obtained. A total of 357 T2 pepper seedlings were exposed to the CMVP1 pathogen and 112 of the T2 peppers were found to be highly tolerant to CMVP1 (Table 3). These 112 plants possessed the insert and were tolerant to CMVP1 for up to 90 days after inoculation (stopped at 90 days). These selected T2 peppers were self-crossed to obtain T3 seeds. We planted 153 T3 peppers that were generated from the E7 line containing the insert and exposed them to CMVP1. Of the 153 T3 peppers, 142 T3 peppers or 92% were tolerant to CMVP1 for up to 85 days after inoculation (stopped at 85 days) (Fig. 5). From these peppers, the ones that displayed good phenotypes (breeder’s judgment) were selected and used for self-crossing to the next generation and for crossing to the elite lines for transgenic breeding.
There have been lots of studies that have examined the gene silencing mechanism for gene regulation (Gura 2000; Benedito et al. 2004). In plants, the typical mechanism for gene regulation is post-transcriptional gene silencing (PTGS). In this mechanism, when the DNA sequence identity between the two comparable genes is higher, gene suppression is higher. The first clue in identifying the molecular mechanisms underlying PTGS was the observation that a class of small RNAs of about 25 nucleotides, which degraded from the double-stranded RNA that was generated from the transgene, triggered the signal for gene silencing (Hamilton and Baulcombe 1999). In this mechanism, over-expression of a certain gene induces the co-suppression of the endogenous gene that it shares overall sequence homology with, for example, 74% with PFG (PETUNIA FLOWERING GENE) or 88% with MADS box (Immink et al. 1999; Ratcliff et al. 2001). In addition, it is not necessary to use the whole coding gene to induce gene silencing; only a fragment with high homology is required (Ruiz et al. 1998). As a result, we used this same strategy to develop virus-resistant chili pepper transformed with CMVP0-CP. Here, the DNA sequence identity between CMVP1-CP of the virus and CMVP0-CP inserted in the transgenic pepper was 93%, which is high enough to co-suppress gene activity.
Phenotypical differences between tolerant and susceptible peppers
Several major differences between the T1 peppers and non-transformed peppers were observed when these peppers were exposed to CMVP1. First, the mosaic occurrence on the leaves of non-transgenic peppers was severe and it was distributed to all the leaves of matured peppers while no mosaic occurrence was seen on the leaves of the transgenic peppers during growth (Fig. 6a). Second, because the presence of the mosaic virus causes the leaf surface to wrinkle, the development and growth of the non-transformed peppers was hindered resulting in peppers with much smaller heights, indicative of a stunt phenotype (Fig. 6b). Third, the transgenic green fruits and red fruits were phenotypically normal while the non-transformed peppers after CMVP1 infection generated much shorter fruits with a small number of seeds (Fig. 6c). In addition, leaf length, leaf width, and fruit width, 90 days after inoculation, were much smaller in the P2377 and P915 lines (non-transformed) than the CMVP1-tolerant transgenic peppers (Table 4). Similar phenotypes were observed in the T2 peppers (Fig. 7).
Generally, when the peppers were cultivated in the plastic house, virus infections rarely occurred naturally and therefore the infection did not appreciably affect productivity. However, virus infection did lower pepper productivity when the peppers were cultivated in the open field and this loss was dependent on the level of infection. If CMVP1 infection was not severe in the field, peppers did not have shorter heights nor did they produce smaller fruit sizes (data not shown). Phenotype changes in peppers were related to the specificity of CMVP1 pathogenicity. We did not observe a dramatic difference in height and fruit size between the non-transformed peppers and the peppers transformed with TMV-CP (Lee et al. 2004), and PepMoV-CP (data not shown) when they were cultured in the plastic house after exposure to TMV and PepMoV, respectively.
The phenotype and growth pattern of CMVP1 tolerant transgenic peppers (T3) were identical or close to non-transformed peppers when the transgenic peppers and non-transformed peppers were grown for 120 days in the plastic house without exposure to CMVP1 (Table 5). In addition, no significant difference between the phenotype of the transgenic peppers and non-transformed peppers were observed (P > 0.05; data not shown). Three different transgenic peppers selected for CMVP1 tolerance (T3), E7, B20 and H15, were chosen for subsequent experiments. E7 and B20 were transgenic lines from the P915 inbred line while the H15 transgenic line was from the P2377 inbred line. The Ph240 inbred line produced no tolerant peppers.
Tolerance levels to CMVP1 in the field
In order to examine the CMVP1 tolerance of transgenic peppers in the open field, 150 peppers from T3 homozygotes (E7, B20 and H15; 50 peppers for each) and non-transformed inbred lines (P915 and P2377; 50 peppers for each) were planted and grown for 143 days after planting (Fig. 8). The aphids did not appear in the field even up to 50 days after planting and the non-transformed inbred lines were still intact during this period. After 106 days in the field, the P915 inbred pepper line was only 15% tolerant, while the E7 and B20 transgenic peppers had a tolerance slightly over 50 and 80%, respectively (Fig. 8a). After 120 days when most of the fruit harvest ends, approximately 30% of the transgenic peppers were still intact after natural exposure to CMV and most of the non-transformed lines were infected.
In the field, the tolerance efficiency of the H15 transgenic peppers was approximately 80, 70, 50 and 35% at 106, 120, 130 and 143 days after planting, respectively (Fig. 8b), while the tolerance efficiency of the non-transformed inbred line P2377 was about 50, 20, 5 and 0% during the same period. The tolerance levels of the three transgenic peppers were similar 106 days after planting. However, at increased cultivation times the tolerance of the H15 transgenic peppers to CMV was apparently higher than the E7 and B20 transgenic peppers.
There was a clear difference in the tolerance levels of transgenic peppers cultivated in the plastic house with artificial inoculation and in the field with natural infection. There was a decrease in tolerance efficiency in the field at longer pepper cultivation periods because other infections aside from CMV can occur during cultivation. A similar study was conducted at the field with T3 peppers by a group of scientists at Korea Research Institute of Bioscience and Biotechnology (KRIBB) for an environmental risk assessment. In this study, the tolerance against the CMV was maintained over a long time period with a similar efficiency pattern, as observed here (data not shown).
One interesting observation was that some of the CMV tolerant transgenic peppers after 143 days in the field were indeed still intact and phenotypically normal. Several periods of rain and high temperatures during the 5 months did not affect the growth and development of these transgenic peppers. The CMVP0 tolerant and commercially available varieties did not stay intact for a long period in the field under natural infection (data not shown). These results here, therefore, are very promising in terms of using these transgenic lines in a breeding program. A similar study of the field performance of a CMV-CP transgenic chili pepper was conducted by Cai et al. (2003). They showed that the transgenic peppers displayed delayed symptom development with milder disease severity in the field. However, 10 weeks post transplantation, a high disease incidence was observed for both transgenic and non-transgenic peppers. In this report, after 143 days in the field we selected peppers that showed no symptoms and appeared almost completely resistant (3 from 50 peppers of B20, 5 from 50 peppers of E7, 18 from 50 peppers of H15). The discrepancy in the field trial between this report and Cai et al.’s report is probably due to the facts that, first, the transgenic lines are different. The transgene location in the genome of each transgenic line may affect different levels of tolerance. Second, the field environment may differ and the field of Cai et al. could have more spread of CMVP1. In addition, during long-term cultivation in the field, they found that other diseases were accompanied by infections by different viruses and other pathogen. Those could make peppers more vulnerable to CMV infection.
CMVP0 tolerance test of T3 peppers
One question raised was whether the T3 peppers that were homozygote and tolerant to CMVP1 would also display tolerance to CMVP0. In order to test this, a total of 49 T3 peppers generated from the E7 line were exposed to CMVP0. From these experiments, 36 transgenic peppers were determined to be tolerant while all the non-transformed peppers were susceptible to infection (Table 6). The tolerance rate was approximately 74% and the tolerant peppers did not show any symptom during 90 days of cultivation in the field. However, 26% of the T3 peppers were susceptible to CMVP0 although the symptoms were relatively weak during the same period. These results were somewhat unexpected because the same badge of T3 peppers showed only 8% susceptible to CMVP1 (Fig. 5). The insert gene, CMVP0-CP, was isolated from CMVP0 and therefore shares 100% DNA sequence identity with the one inserted in the transgenic peppers. When gene silencing by the viral gene occurs, higher DNA sequence identity is typically better (Morroni et al. 2008); however, this was not the case here. The differences between the pathogenicity of CMVP0 and CMVP1 in pepper plants are not known except for the fact that CMVP1 (Ca-P1) is a CMVP0 (Ca-P0) resistant breaking virus and the Ca-P0-R line becomes susceptible by Ca-P1 (Lee et al. 2006). Although several possible mechanisms involved in conferring the resistance to CMV were suggested (Lin et al. 2007), such as protein-mediated resistance, RNA-mediated resistance and cross-protection, the interaction between different CMV strain pathogenicity and the resistance mechanism of peppers must be further studied. One thing that is clear from our study is that the transgenic peppers selected for their high tolerance to CMVP1 and CMVP0 are more likely resistant lines. The selected peppers are currently being used in a breeding program.
Here we present CMVP0-CP transgenic peppers that are tolerant to two CMV pathotypes, suggesting that these transgenic peppers would be tolerant to any CMVs, even new CMV strains that have not yet occurred, as long as the DNA sequence identity of the CP genes is very close.
References
Bau HJ, Cheng YH, Yu TA, Yang JS, Yeh SD (2003) Broad spectrum resistance to different geographic strains of papaya ringspot virus in coat protein gene transgenic papaya. Phytopathology 93:112–120
Benedito VA, Visser PB, Angenent G, Krens FA (2004) The potential of virus-induced gene silencing for speeding up functional characterization of plant genes. Genet Mol Res 3:323–341
Cai WQ, Fang RX, Shang HS, Wang X, Zhang FL, Li YR, Zhang JC, Cheng XY, Wang GL, Mang KQ (2003) Development of CMV- and TMV-resistant transgenic chili pepper: field performance and biosafety assessment. Mol Breed 11:25–35
Church GM, Gilbert W (1984) Genomic sequencing. Proc Natl Acad Sci USA 81:1991–1995
Clough GH, Hamm PB (1995) Coat protein transgenic resistance to watermelon mosaic virus and zucchini yellow mosaic virus in squash and cantaloupe. Plant Dis 79:1107–1109
Cuozzo M, O’Connell KM, Kaniewski WK, Fang RX, Chua NH, Tumer NE (1988) Viral protection in transgenic tobacco plants expressing the cucumber mosaic virus coat protein or its antisense RNA. Biotechnology 6:549–557
Davis MJ, Ying Z (2004) Development of papaya breeding lines with transgenic resistance to papaya ringspot virus. Plant Dis 88:352–358
Di R, Purcell V, Collins GB, Ghabrial SA (1996) Production of transgenic soybean lines expressing the bean pod mottle virus coat protein precursor gene. Plant Cell Rep 15:746–750
Fitch MMM, Manshardt RM, Gonsalves D, Slightom JL, Sanford C (1992) Virus resistant papaya derived from tissues bombarded with the coat protein gene of papaya. Biotechnology 10:1466–1472
Fuchs M, Gonsalves D (1995) Resistance of transgenic hybrid squash ZW-20 expressing the coat protein genes of zucchini yellow mosaic virus and watermelon mosaic virus 2 to mixed infections of both potyviruses. Biotechnology 13:1466–1473
Fuchs M, Klas FE, McFerson JR, Gonsalves D (1998a) Transgenic melon and squash expressing coat protein genes of aphid-borne viruses do not assist the spread of an aphid non-transmissible strain of cucumber mosaic virus. Transgenic Res 7:449–462
Fuchs M, Tricoli DM, Carney KJ, Schesser M, McFerson JR, Gonsalves D (1998b) Comparative virus resistance and fruit yield of transgenic squash with single and multiple coat protein genes. Plant Dis 82:1350–1356
Gölles R, Moser R, Puhringer H, Katinger H, Leimer da Camara Machado M, Minafra A, Savino V, Saldarelli P, Martelli GP, Camara MA (2000) Transgenic grapevines expressing coat protein gene sequences of Grapevine fanleaf virus, Arabis mosaic virus, Grapevine virus A and Grapevine virus B. Acta Hortic 528:305–311
Gura T (2000) A silence that speaks volumes. Nature 404:804–808
Hamilton AJ, Baulcombe DC (1999) A species of small antisense RNA in post-transcriptional gene silencing in plants. Science 286:950–952
Immink RGH, Hannapel DJ, Ferrario S, Busscher M, Franken J, Lookeren Campagne MM, Angenent GC (1999) A petunia MADS box gene involved in the transition from vegetative to reproductive development. Development 126:5117–5126
Iwanami T, Shimizu T (2004) Tolerance to citrus mosaic virus in transgenic trifoliate orange lines harboring capsid polyprotein gene. Plant Dis 88:865–868
James C (2008) Global status of commercialized biotech/GM crops. Report on global status of biotech/GM Crops by ISAAA (International Service for the Acquisition of Agri-biotech Applications)
Jin M, Chengalrayan K, Abouzid A, Gallo M (2007) Prospecting the utility of a PMI/mannose selection system for the recovery of transgenic sugarcane (Saccharum spp. hybrid) plants. Plant Cell Rep 26:581–591
Klas FE, Fuchs M, Gonsalves D (2006) Comparative spatial spread overtime of zucchini yellow mosaic virus (ZYMV) and watermelon mosaic virus (WMV) in fields of transgenic squash expressing the coat protein genes of ZYMV and WMV, and fields of nontransgenic squash. Trasngenic Res 15:527–541
Krastanova S, Perrin M, Barbier P, Demangeat G, Cornuet P, Bardonnet N, Otten L, Pinck L, Walter B (1995) Transformation of grapevine rootstocks with the coat protein gene of Grapevine fanleaf nepovirus. Plant Cell Rep 14:50–554
Krubphachaya P, Juricek M, Kerbundit S (2007) Induction of RNA-mediated resistance to papaya ringspot virus type W. J Biochem Mol Biol 40:404–411
Lee YH, Kim HS, Kim JY, Jung M, Park YS, Lee JL, Choi SH, Her NH, Lee JH, Hyung NI, Lee CH, Yang SG, Harn CH (2004) A new selection method for pepper transformation: callus-mediated shoot formation. Plant Cell Rep 23:50–58
Lee MY, Lee JH, Ahn HI, Kim MJ, Her NH, Choi JK, Harn CH, Ryu KH (2006) Identification and sequence analysis of RNA3 of a resistance-breaking isolate of cucumber mosaic virus from Capsicum annuum. Plant Pathol J 22:265–270
Lin SS, Henriques R, Wu HW, Niu QW, Yeh SD, Chua NH (2007) Strategies and mechanism of plant virus resistance. Plant Biotechnol Rep 1:125–134
Linbo JA, Dougherty WG (1992) Pathogen-derived resistance to a potyvirus: Immune and resistance phenotypes in transgenic tobacco expressing altered forms of a potyvirus coat protein nucleotide sequence. Mol Plant Microbe Interact 5:144–153
Ling KS, Zhu HY, Gonsalves D (2008) Resistance to grapevine leafroll associated virus-2 is conferred by post-transcriptional gene silencing in transgenic Nicotiana benthamiana. Transgenic Res 17:733–740
Maghuly F, Leopold S, Machado AC, Fernadez EB, Khan MA, Gambino G, Gribaudo I, Schartl A, Laimer MA (2006) Molecular characterization of grapevine plants transformed with GFLV resistance genes. Plant Cell Rep 25:546–553
Mauro MC, Tortain S, Walter B, Pinck L, Otten L, Coutos-Thevento P, Deloire A, Barbier P (1995) High efficiency regeneration of grapevine plants transformed with the GFLV coat protein gene. Plant Sci 112:97–106
Morroni M, Thompson JR, Tepfer M (2008) Twenty years of transgenic plants resistant to cucumber mosaic virus. Mol Plant Microbe Interact 21:675–684
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol Plant 5:473–479
Nelson RS, McCormick SM, Delannay X, Dube P, Layton J, Andoson EJ, Kaniewska M, Proksch RK, Horsch RB, Rogers SG, Fraley RT, Beachy RN (1988) Virus tolerance, plant growth, and field performance of transgenic tomato plants express coat protein from tobacco mosaic virus. Biotechnology 6:403–409
Nida DL, Anjos JR, Lomonossoff GP, Ghabrial SA (1992) Expression of cow-pea mosaic virus coat protein precursor in transgenic tobacco plants. J Gen Virol 73:157–163
Pang SZ, Jan FJ, Tricoli DM, Russell PF, Carney KJ, Hu JS, Fuchs M, Quemada HD, Gonsalves D (2000) Resistance to squash mosaic comovirus in transgenic squash plants expressing its coat protein genes. Mol Breed 6:87–93
Park SM, Lee JS, Jegal S, Jeon BY, Jung M, Park YS, Ryu KH, Han SL, Shin YS, Her NH, Lee JH, Yang SG, Harn CH (2005) Transgenic watermelon rootstock resistant to CGMMV (cucumber green mottle mosaic virus) infection. Plant Cell Rep 24:350–356
Powell-Abel P, Nelson RS, De B, Hoffman N, Rogers SG, Fraley RT, Beachy R (1986) Delay of disease development in transgenic plants that express the Tobacco Mosaic Virus coat protein gene. Science 232:738–743
Ratcliff F, Martin-Hernandez AM, Baulcombe D (2001) Tobacco rattle virus as a vector for analysis of gene function by silencing. Plant J 25:237–245
Reddy MSS, Ghabrial SA, Redmond CT, Dinkins RD, Collins GB (2001) Resistance to bean pod mottle virus in transgenic soybean lines expressing the capsid polyprotein. Phytopathology 91:831–838
Ruiz MT, Voinnet O, Baulcombe D (1998) Initiation and maintenance of virus-induced gene silencing. Plant Cell 10:937–946
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual (2nd edn). Cold Spring Harbor Laboratory, Cold Spring Harbor
Shin R, Han JH, Lee GJ, Paek KH (2002) The potential use of a viral coat protein gene as a transgene screening marker and multiple virus resistance of pepper plants coexpressing coat proteins of cucumber mosaic virus and tomato mosaic virus. Transgenic Res 11:215–219
Tennant PF, Gonsalves C, Ling KS, Fitch M, Manshardt R, Slightom JL, Gonslaves D (1994) Different protection against papaya ringspot virus isolates in coat protein gene transgenic papaya and classically cross-protected papaya. Phytopathology 84:1359–1366
Tougou M, Furutani N, Yamagishi N, Shizukawa Y, Takahata Y, Hidaka S (2006) Development of resistant transgenic soybeans with inverted repeat-coat protein genes of soybean dwarf virus. Plant Cell Rep 25:1213–1218
Tricoli DM, Carney KJ, Russell PF, McMaster JR, Groff DW, Hadden KC, Himmel PT, Hubbard JP, Boeshore ML, Quemada HD (1995) Field evaluation of transgenic squash containing single or multiple virus coat protein gene constructs for resistance to cucumber mosaic virus. Biotechnology 13:1458–1465
Zanek MC, Reyes CA, Cervera M, Pena EJ, Velazquez K, Costa N, Plata MI, Grau O, Pena L, Garcia ML (2008) Genetic transformation of sweet orange with the coat protein gene of Citrus psorosis virus and evaluation of resistance against the virus. Plant Cell Rep 27:57–66
Zhu YX, Qu Y, Wen J, Zhang Y, Chen ZL (1996) Transgenic sweet pepper plants from Agrobacterium-mediated transformation. Plant Cell Rep 16:71–75
Zrachya A, Kumar PP, Ramakrishnan U, Levy Y, Loyter A, Arazi T, Lapidot M, Gafni Y (2007) Production of siRNA targeted against TYLCV coat protein transcripts leads to silencing of its expression and resistance to the virus. Transgenic Res 16:385–398
Acknowledgments
This research was supported mainly from a grant of the Crop Functional Genomics Center of the 21st Century Frontier Research Program funded by the Ministry of Education, Science and Technology and partially from a grant of the BioGreen 21 research fund of the Rural Development of Administration from the Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. R. Liu.
Rights and permissions
About this article
Cite this article
Lee, Y.H., Jung, M., Shin, S.H. et al. Transgenic peppers that are highly tolerant to a new CMV pathotype. Plant Cell Rep 28, 223–232 (2009). https://doi.org/10.1007/s00299-008-0637-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-008-0637-3