Abstract
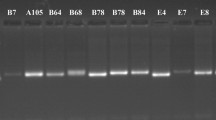
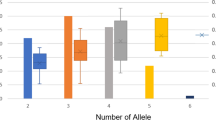
During the past five decades, a large number of tobacco varieties have been developed for different end uses in India through pure line selection from local land races, mutation breeding, and hybridization involving local selections and exotic introductions followed by pedigree selection. No systematic effort has been made to understand the existing diversity pattern in these varieties, which is crucial to define future breeding strategy in this important commercial crop. We characterized 46 varieties belonging to 10 different manufacturing tobacco types cultivated under different agro-climatic conditions in India along with two wild species of Nicotiana using 40 arbitrary primers in RAPID. The level of polymorphism among the varieties of N. tabacum was 59.4%, which was more than double the level observed in the other cultivated species N. rustica (25.2%). A broader range (0.64 to 0.94) of pair wise similarity measures in N. tabacum than in N. rustica (0.83 to 0.92) reflected the more diversified breeding efforts in the major cultivated species. The two wild species namely, N. glutinosa and N. gossei clustered separately from the two cultivated species. Molecular classification of the varieties corresponded largely with their manufacturing trait and parentage. RAPID markers provided sufficient resolution to distinguish among closely related tobacco types. Nine RAPID markers were found conserved across all the varieties and species. The markers found specific to the varieties can be used in correct identification of the carrier genotypes in trade and commerce. This is the first report on the molecular diversity analysis of Indian tobacco.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Narayan RK, Plant Syst Evol, 157 (1987) 161.
Anonymous, (2003) Information bulletin on crop improvement, Central Tobacco Research Institute, Rajahmundry, Andhra Pradesh, India.
Singh KD, Narishmha Rao CV, Prabhu SR & Subba Rao R, CORESTA Congress, New Orleans, USA (2002)
Clegg MT, In Plant population genetics, breeding and genetic resource (AHD Brown et al, Editors), Sinauer Associate Inc, Sunderland, MA, USA (1990) p 98.
Bai D, Reedler R & Brandle JE, Theor Appl Genet, 91 (1995) 1184.
Rufty R, Yi YH & Wernsman EA, CORESTA Congress, Montréal, Canada (1997) p 50.
Yi HY, Rufty RC & Wernsman EA, Plant Dis, 82 (1998) 1319.
Del Piano L, Abet M, Sorrenlino C, Acanfora F, Cozzolino E, Acanfora F, Cozzolino E & DiMuro A, Beitrege Zur Tab Int, 19 (2000) 1.
Doyle JJ & Doyle JL, Focus, 12 (1990) 13.
Williams JGK, Kubelik AR, Livak KJ, Rafalski JA & Tingey SV, Nucleic Acids Res, 18 (1990) 6531.
Rohlf FJ, NTSYS-pc: Numerical taxonomy and multivariate analysis system. Version 2.02. Exter Software, Setauket, New York (1998).
Jaccard P, Bull Soc Vaeed Sci Nat, 44 (1908) 223.
Kelm P, Beavis W, Schupp J & Freestone R, Theor Appl Genet, 85 (1992) 205.
Yap I & Nelson RJ, IRRI Discussion Paper Series No. 14, International Rice Research Institute, Manila, Philippines (1996).
Ren N & Timko MP, Genome, 44 (2002) 559.
Yang BC, Xiao BG, Chen XJ & Shi CH, Ann Appl Biol, 150 (2007) 393.
Ford R & Taylor PWJ, Australian J Agril Res, 48 (1997) 1213.
Koundal M, Sharma DR, Mohapatra T & Koundal KR, J Plant Biochern Biotech, 15 (2006) 15.
Goodspeed TH, The genus Nicotiana, Chronica Bot Wallham, Mass, USA (1954).
Cao W, Hucl P, Scoles G, Chibbar RN, Fox PN & Skovmand B, Wheat Inf Ser, 95 (2002) 29.
Fu YB, Rowland GG, Duguid SD & Richards KW, Crop Sci, 43 (2003) 1510.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
SivaRaju, K., Madhav, M.S., Sharma, R.K. et al. Molecular Diversity in Indian Tobacco Types as Revealed by Randomly Amplified DNA Polymorphism. J. Plant Biochem. Biotechnol. 17, 51–56 (2008). https://doi.org/10.1007/BF03263259
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03263259