Abstract
Background and Objective: It has been shown that somatic missense mutations in codon 132 of the NADP+ dependent isocitrate dehydrogenase 1 (IDH1) gene occur frequently in primary brain tumors including highly malignant glioblastoma (GBM). The aim of this study was to evaluate a PCR-restriction fragment length polymorphism (RFLP)-based method for missense mutation detection and to estimate the prognostic value of the two most frequent IDH1 codon 132 mutations, R132H and R132C, in patients with newly diagnosed GBM treated with radiation combined with temozolomide.
Methods: DNA was extracted from formalin-fixed, paraffin-embedded tissue. The PCR-RFLP method was adapted to IDH1 codon 132 mutation screening. The mutation status was determined in a group of 58 patients.
Results: We found R132H mutations in 14% of patients. No R132C mutation was found in this study. Median follow-up for living patients was 31 (range 17–51) months. Median progression-free survival in the group of patients with IDH1 mutation was 29 months compared with 10 months in the IDH1 wild-type group (p = 0.004; hazard ratio [HR] 3.09, 95% CI 1.25, 4.78). Median overall survival in the group with IDH1 mutation has not been reached, whereas in the group with wild-type IDH1 it was 19.5 months (p < 0.001; HR 4.76, 95% CI 1.22, 6.30). Three-year overall survival was 60% in the group with IDH1 mutation while in the wild-type IDH1 group it dropped to 29%. IDH1 mutations significantly correlated with younger age(p = 0.02).
Conclusions: Our results indicate that the IDH1 R132H mutation is a powerful prognostic marker in GBM treated with chemoradiation. The PCR-RFLP method allows for a fast, inexpensive, and sensitive mutation screening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastoma (GBM), corresponding to WHO grade IV, is the most frequent primary brain tumor in adults. It represents 54% of all tumors originating from glial cells (Central Brain Tumor Registry of the United States, http://www.cbtrus.org), and is characterized by poor prognosis, with a median overall survival of 12 months.[1] Most GBMs (approximately 90%) develop rapidly de novo as a so-called primary GBM without histological evidence of transition from more differentiated astrocytic precursor lesions of lower grade stage. In contrast, secondary GBMs develop through progression from diffuse astrocytoma (WHO grade II) or anaplastic astrocytoma (WHO grade III). Primary and secondary GBMs, although difficult to discriminate in standard diagnostics, differ in terms of age of diagnosis (median 62 years for primary vs 45 years for secondary GBM), median survival (shorter for primary GBM), and genetic pathways.[2] In most Western countries, the current standard treatment of newly diagnosed GBM involves surgery and combined chemoradiotherapy with the use of temozolomide followed by adjuvant temozolomide in 21-day cycles.[3]
Located in the chromosome region 2q33, isocitrate dehydro-genase 1 gene (IDH1) belongs to a group of five genes encoding human isocitrate dehydrogenases, enzymes which catalyze the reaction of transition of isocitrate to α-ketoglutarate. Four of the human isocitrate dehydrogenase enzymes are mitochondrial; only IDH1 is present mainly in cytoplasm and peroxisomes.[4] The physiological function of IDH1 remains unclear. Presumably, it plays a role in cytoplasmic nicotinamide adenine dinucleotide phosphate (NADPH) generation as well as fatty acid oxidation and degradation of phytanic acid in peroxisomes.
Previous studies showed that, in contrast to the majority of solid tumors, mutations in the IDH1 gene occur frequently in brain cancers.[5–7] They are also evident in some patients with acute myeloid leukemia.[8] These mutations are limited to the small hot-spot region in exon 4, codon 132, which encodes arginine located in the active site of the enzyme. Substitution of this amino acid impairs interaction with isocitrate and dramatically reduces enzymatic activity of IDH1 against its natural substrate. Normally, IDH1 functions as a homodimer. When a mutant molecule binds to a wild-type protein it results in formation of inactive heterodimer. Therefore, mutations in codon 132 have dominant effects and their presence in at least one of the alleles leads to inactivation of the enzyme, increased levels of isocitrate, and decreased levels of α-ketoglutarate.[9] Recent findings demonstrated accumulation of isocitrate metabolite R-(−)-2-hexoglutarate in cells with an IDH1 mutation.[10] At the cellular level, the consequences of these dramatic changes are not clear. Based on functional studies and the finding of increased levels of hypoxia inducible factor (HIF1α) in mutant cells, Zhao et al. proposed that IDH1 may act in the hypoxic pathway as a tumor suppressor.[11] These results are inconsistent with clinical observations in patients with glioma. Despite the conclusion from functional studies, GBM patients harboring IDH1 mutations have better outcomes than those with the wild-type gene.[12,13] In previous studies, patients carrying the mutant allele had a median overall survival (OS) of 31 months, compared with 15 months for patients with wild-type IDH1. IDH1 mutations occur mainly in secondary GBMs (approximately 85% of these patients) and lower grade tumors. They are observed much less frequently in primary GBMs (5% of cases).[14–16] This means that IDH1 mutation might be considered as the best known single genetic marker for diagnosis of secondary GBM.[17]
In previously published studies, IDH1 mutations have been detected by genomic sequencing.[13–19] The fact that these mutations occur in one codon creates an opportunity for application of a simple test based on restriction digestion. In a study by Balss et al.,[19] 92.7% of all codon 132 mutations were the R132H substitution and the second most frequent mutation was R132C (3.6%). All other mutations accounted for only 3.7%.
In this study we developed a restriction fragment length polymorphism (RFLP) method suitable for detecting the most frequent IDH1 mutations, R132H and R132C. We screened a group of patients with newly diagnosed GBM treated with standard radiotherapy with temozolomide and performed survival analysis of IDH1 wild-type versus mutated patients.
Materials and Methods
Patients
This study involved 58 patients with newly diagnosed GBM, WHO grade IV, who were treated between 2002 and 2007. All patients had good performance status at the beginning of the treatment and underwent surgical tumor resection. Patients were treated with temozolomide according to the following scheme: 75mg/m2 of body surface area of temozolomide during radiotherapy (total fraction 60 Gy) and adjuvant temozolomide therapy for 5 days in 21-day cycles in a dose of 150–200 mg/m2. In cases of tumor progression, second-line chemotherapy with lomustine (CCNU) was administered. In this group of patients the best known predictive molecular factor, MGMT promoter hypermethylation, was determined previously (manuscript in preparation) with the use of methylation-specific PCR.[20] Patients’ characteristics are listed in table I.
The study protocol was approved by the Independent Ethics Committee of the Cancer Centre and Institute of Oncology, Warsaw, Poland.
DNA Samples Preparation
All DNA samples were obtained from formalin-fixed, paraffin-embedded (FFPE) tissue, using the Scherlock AX kit (A&A Biotechnology, Gdynia, Poland). The quality and integrity of DNA samples from formalin-fixed tissue were determined by multiplex PCR reaction designed for the BIOMED-2 study,[21] with a set of primers that amplify PCR products of various sizes (range 100–400 bp). All DNA samples included in the study allowed for efficient amplification of 200 bp PCR product.
Mutational Analysis by PCR-RFLP
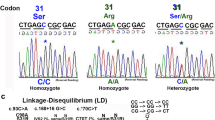
To determine IDH1 mutation status in nucleotide positions 395G>A (R132H) and 394C>T (R132C), we developed a new nested PCR-RFLP assay (figure 1). In the first step a 166 bp fragment of IDH1 exon 4 was amplified. Twenty cycles of PCR reaction were performed in a total volume of 15 µL, consisting of 1x PCR buffer, 2mmol/L MgCl2, 250µmol/L of each deoxynucleotide, 0.1µmol/L of each PCR primer, 0.5 U of FastStart Taq DNA Polymerase (Roche, Mannheim, Germany), and 50 ng of DNA template. The PCR product was diluted ten times and reamplified in two parallel second-step reactions with two sets of primers designed for R132H and R132C mutation discrimination. One of the primers in each primer pair included a mismatch that created the restriction site for Hsp92II or Psp1405I enzymes for R132H and R132C mutation detection, respectively. The second primer in each pair included a mismatch introducing a control restriction site for the same enzyme that was used for mutation detection (i.e. Hsp92II or Psp1405I). Digestion of the control site in every PCR product excluded false negative results that might have occurred due to incomplete digestion by the specific enzyme used. The second PCR step was carried out for 30 cycles in a total volume of 15 µL, consisting of 1x PCR buffer, 3 mmol/L MgCl2, 250µmol/L of each deoxynucleotide, 0.15µmol/L of each PCR primer, 0.5 U of FastStart Taq DNA Polymerase, and 1 µL of diluted first-step nested PCR product. All PCR reactions were performed in GeneAmp® system 2700 (Applied Biosystems) thermocycler. Primer sequences and PCR conditions are listed in table II. In this step, the size of the amplified products was 109 bp in the reaction to detect the R132H mutation, and 123 bp in the reaction to detect the R132C mutation. Incorporation of the restriction sites was confirmed by DNA sequencing of inner products of the selected DNA samples.
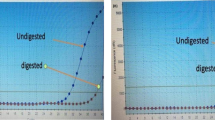
Half of the final volume of the inner PCR product was digested with the restriction enzyme Hsp92II (Promega) or Psp1406I (Fermentas) for R132H and R132C mutation detection, respectively. Digestion was performed for 4 hours in a total volume of 20 µL, containing 1x digestion buffer and 10 U of restriction enzyme. Digestion products were electrophoresed in 8% polyacrylamide gels (acrylamide/Bis 19:1) and visualized with ethidium bromide. Digestion of PCR products from wild-type template DNA occurred only in the control site. In the completely digested wild-type sample only one band on the gel was visible: 97 bp in R132H and 112 bp in R132C mutation test. In the mutated IDH1 allele an additional restriction site was introduced in the PCR step and the mutation was confirmed by the presence of an additional 76 bp or 88 bp fragment that indicated R132H or R132C substitutions, respectively (figure 2).
An example of the PCR-RFLP result for IDH1 R132H mutation detection, showing the polyacrylamide gel electrophoresis of the second-step PCR product (109bp) after restriction digestion. Lanes 1–7 contain patients’ samples, with samples 1 and 4 showing the IDH1 R132H mutation. H 2 O = no template control; K= undigested PCR control; M = 10 bp DNA ladder (Fermentas).
All positive and negative RFLP results were confirmed by DNA sequencing of first-step PCR products with BigDye® Terminator v3.1 on an ABI PRISM 3100 Genetic Analyzer (Applied Biosystems).
Statistical Analysis
The follow-up period was terminated in May 2009. OS was defined as the time from surgery to a patient’s death or last follow-up contact before the moment of completion of data collection. Progression-free survival (PFS) was defined as the time from surgery to radiographic evidence of tumor progression or the date of recraniotomy. OS and PFS curves were estimated by the Kaplan-Meier method and compared by the use of two-sided log-rank test. The statistical relationship between various parameters was analyzed using the Fisher’s exact test.
Results and Discussion
Using the RFLP-based approach we identified IDH1 R132H mutations in 14% (8/58) of the patients. No R132C mutation was found in the studied group of GBM patients. The results of PCR-RFLP analysis were validated by genomic sequencing. All eight positive and 50 negative DNA samples underwent verification. We obtained consistent results confirming the usefulness of the PCR-RFLP approach, as well as its sensitivity and specificity compared with the DNA sequencing method. Sequencing analysis showed that all identified mutations were heterozygous.
Median PFS in the group of patients with mutated IDH1 was 29 months compared with 9 months in the group with wild-type IDH1(p = 0.004; HR 3.09, 95% CI 1.25, 4.78) [figure 3]. Three-year OS was 60% in the group of patients with mutated IDH1 and 29% in the group with wild-type IDH1. Median OS in the group with mutated IDH1 has not been reached in the follow-up period. Median OS in the wild-type group was 19.5 months (p < 0.001; HR4.76,95% CI 1.22,6.30) [figure 3]. Seven of eight patients with IDH1 somatic mutations were younger than 49 years (median age of the group) and this difference is statistically significant (p = 0.023) at significance level of 0.05.
The frequency of IDH1 mutations was also higher in the group of patients with MGMT promoter hypermethylation, although this difference is not statistically significant. In six of eight cases with mutated IDH1, the detailed histopathological examination provided evidence of secondary GBM. However, these patients did not have a history of prior diagnosis of a lower-grade glioma, which is one of the possible criteria of secondary GBM.
Combined chemoradiotherapy with temozolomide with adjuvant temozolomide has become a standard treatment of patients with newly diagnosed GBM. To our best knowledge, there are no reports on the mutation analysis of the IDH1 gene and its role in clinical outcomes of patients treated with this regimen. In this study we performed analysis of IDH1 in a demographically diverse group of GBM patients treated with this combined therapy.
IDH1 mutations are limited to the small single hot-spot in codon 132 of exon 4. From the diagnostic point of view this fact is very important because well defined single nucleotide genomic variations are easy to evaluate with the use of restriction enzymes.[22] In this study we proposed the PCR-RFLP-based approach for IDH1 mutation detection. This technique allows for a fast and simple analysis, which can be performed in 1 day from DNA isolation to the final result.
The RFLP approach has several important advantages over other methods including genomic sequencing or real-time PCR instrument-based techniques such as high resolution melting point analysis or allelic discrimination. It is sensitive, cost effective, does not require any sophisticated equipment, and is at least as fast as other approaches. All of these features make this method very suitable for patient screening and diagnostics. In our experience, this approach was comparably sensitive and specific to genomic sequencing, the method that was previously shown to be useful for IDH1 mutation detection in FFPE tissue.[18]
During the data collection period, a similar method was published by Meyer et al.[23] These authors proposed PCR-RFLP for five known IDH1 mutations including the least frequent changes. Our approach differs in technical details. We designed PCR primers with the control restriction site. In our system, digestion of the PCR product results in formation of a specific control DNA fragment. The respective band on the gel indicates complete PCR product digestion. This internal control prevents eventual false-negative results caused by incomplete DNA restriction cleavage. We also designed shorter PCR products that may be more useful in analysis of low quality DNA, such as that isolated from FFPE tissue.
Recently, a monoclonal antibody that specifically recognizes the IDH1 R132H mutant protein has been described.[24,25] This antibody was successfully used for mutation detection on the protein level either by Western blot assay or immunohistochemistry. Although Western blot analysis seems not to be useful in clinical diagnostics, a specific immunohistochemical staining may find application in a standard histopathological evaluation. This approach was previously shown to be at least as useful as DNA sequencing.[26] Although comparison of the PCR-RFLP assay needs further methodological study, based on reported data we can expect that it should show similar sensitivity.
In our opinion, PCR-based methods of mutation detection provide at least one significant advantage. The results are generated as simple binary data, such as presence or lack of an additional band on the gel in the PCR-RFLP. Consequently, these methods are not susceptible to errors resulting from subjective microscopic observation. On the other hand, the PCR-based methods are prone to false negative results resulting from contamination of analyzed tumor samples with normal cells: either normal glial cells, blood microvessels or infiltrating lymphocytes. In our study, in all cases of mutated samples, the bands on the gel corresponding to the mutant allele were less intense than the wild-type allele bands. In our interpretation this results from the fact that all diagnosed mutations were heterozygous and all tissue samples contained a combination of neoplastic and other cells. The problem of contamination of the tumor tissue with normal cells may be resolved with the use of micro- or macrodissection of the analyzed tissue sample. Such an approach has already been shown to improve detection of IDH1 mutations with the use of genomic sequencing.[18]
Previous reports indicate that IDH1 mutations may be an important factor in the pathogenesis of GBM.[12,14] They also seem to be the best known genetic marker for secondary GBM tumors which, contrary to primary GBM, develop from better-differentiated astrocytic tumors (WHO grade II and III) and are diagnosed in younger patients. We observed the IDH1 R132H mutation in 14% of patients, and significantly more frequently in younger patients. In most of these cases histopathological evaluation documented, at least focally, better-differentiated neoplastic astrocytes, indicating secondary GBM. We also observed the previously reported association between IDH1 mutations and MGMT promoter methylation.[16] However, in our study this relationship was not statistically significant, probably because of the small number of patients.
Similarly to the previous reports, our results showed that IDH1 mutations are associated with better clinical outcome in terms of PFS and OS. It is not clear whether IDH1 mutations are a prognostic factor per se or a predictive factor in the context of the specific therapy regimen. Our analysis includes only patients with GBM treated with the current standard approach, i.e. concomitant radiotherapy and temozolomide with adjuvant temozolomide. In most of the previously reported studies that indicate the role of IDH1 mutation for patient survival the treatment regimen of patients was not specified. In this study we observed better prognosis for GBM patients compared with previously published results by Sanson et al.[16] and Yan et al.[14] This difference may result from younger age of the included patients, but also may suggest that the observed better clinical outcomes in our study is caused by the interaction of temozolomide treatment and IDH1 function. Systematic prospective studies as well as more detailed retrospective analysis are needed to explain this relationship.
Conclusions
Our study showed usefulness of the PCR-RFLP technique for detection of point mutations in codon 132 of IDH1 gene. This technique provides sensitivity and specificity similar to the genomic sequencing method. IDH1 mutations occur more frequently in younger patients and correlate with better progression-free survival and overall survival in glioblastoma patients treated with current standard regimen of concomitant radiotherapy and temozolomide followed by adjuvant temozolomide. Therefore these point mutations appear to be important prognostic factors.
References
Gorlia T, van den Bent MJ, Hegi ME, et al. Nomograms for predicting survival of patients with newly diagnosed glioblastoma: prognostic factor analysis of EORTC and NCIC trial 26981-22981/CE.3. Lancet Oncol 2008 Jan; 9(1): 29–38
Ohgaki H, Kleihues P. Genetic pathways to primary and secondary glioblastoma. Am J Pathol 2007 May; 170(5): 1445–53
Stupp R, Mason WP, van den Bent MJ, et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 2005 Mar 10; 352(10): 987–96
Geisbrecht BV, Gould SJ. The human PICD gene encodes a cytoplasmic and peroxisomal NADP(+)-dependent isocitrate dehydrogenase. J Biol Chem 1999 Oct 22; 274(43): 30527–33
Bleeker FE, Lamba S, Leenstra S, et al. IDH1 mutations at residue p.R132 (IDH1(R132)) occur frequently in high-grade gliomas but not in other solid tumors. Hum Mutat 2009 Jan; 30(1): 7–11
Kang MR, Kim MS, Oh JE, et al. Mutational analysis of IDH1 codon 132 in glioblastomas and other common cancers. Int J Cancer 2009 Jul 15; 125(2): 353–5
Hartmann C, Meyer J, Balss J, et al. Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol 2009 Oct; 118(4): 469–74
Mardis ER, Ding L, Dooling DJ, et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med 2009 Sep 10; 361(11): 1058–66
Jennings GT, Minard KI, McAlister-Henn L. Expression and mutagenesis of mammalian cytosolic NADP+-specific isocitrate dehydrogenase. Biochemistry 1997 Nov 4; 36(44): 13743–7
Dang L, White DW, Gross S, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 2009 Dec 10; 462(7274): 739–44
Zhao S, Lin Y, Xu W, et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1 alpha. Science (NY) 2009 Apr 10; 324(5924): 261–5
Parsons DW, Jones S, Zhang X, et al. An integrated genomic analysis of human glioblastoma multiforme. Science (NY) 2008 Sep 26; 321(5897): 1807–12
Weller M, Felsberg J, Hartmann C, et al. Molecular predictors of progression-free and overall survival in patients with newly diagnosed glioblastoma: a prospective translational study of the German Glioma Network. J Clin Oncol 2009 Dec 1; 27(34): 5743–50
Yan H, Parsons DW, Jin G, et al. IDH1 and IDH2 mutations in gliomas. N Engl J Med 2009 Feb 19; 360(8): 765–73
Watanabe T, Nobusawa S, Kleihues P, et al. IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. Am J Pathol 2009 Apr; 174(4): 1149–53
Sanson M, Marie Y, Paris S, et al. Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J Clin Oncol 2009 Sep 1; 27(25): 4150–4
Nobusawa S, Watanabe T, Kleihues P, et al. IDH1 mutations as molecular signature and predictive factor of secondary glioblastomas. Clin Cancer Res 2009 Oct 1; 15(19): 6002–7
Horbinski C, Kofler J, Kelly LM, et al. Diagnostic use of IDH1/2 mutation analysis in routine clinical testing of formalin-fixed, paraffin-embedded glioma tissues. J Neuropathol Exp Neurol 2009 Dec; 68(12): 1319–25
Balss J, Meyer J, Mueller W, et al. Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol 2008; 116: 597–602
Esteller M, Hamilton SR, Burger PC, et al. Inactivation of the DNA repair gene O6-methylguanine-DNA methyltransferase by promoter hypermethylation is a common event in primary human neoplasia. Cancer Res 1999 Feb 15; 59(4): 793–7
van Dongen JJ, Langerak AW, Bruggemann M, et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 2003 Dec; 17(12): 2257–317
Parsons BL, Heflich RH. Genotypic selection methods for the direct analysis of point mutations. Mutat Res 1997 Oct; 387(2): 97–121
Meyer J, Pusch S, Balss J, et al. PCR- and restriction endonuclease-based detection of IDH1 mutations. Brain Pathol 2010 Mar; 20(2): 298–300
Kato Y, Jin G, Kuan CT, et al. A monoclonal antibody IMab-1 specifically recognizes IDH1R132H, the most common glioma-derived mutation. Biochem Biophys Res Commun 2009 Dec 18; 390(3): 547–51
Capper D, Zentgraf H, Balss J, et al. Monoclonal antibody specific for IDH1 R132H mutation. Acta Neuropathol 2009 Nov; 118(5): 599–601
Capper D, Weissert S, Balss J, et al. Characterization of R132H mutation-specific IDH1 antibody binding in brain tumors. Brain Pathol 2010 Jan; 20(1): 245–54
Acknowledgments
No sources of funding were used to assist in the preparation of this article. The authors have no conflicts of interest that are directly relevant to the content of the study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bujko, M., Kober, P., Matyja, E. et al. Prognostic Value of IDH1 Mutations Identified with PCR-RFLP Assay in Glioblastoma Patients. Mol Diag Ther 14, 163–169 (2010). https://doi.org/10.1007/BF03256369
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03256369