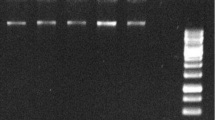
Abstract
As opposed to standard polymerase chain reaction (PCR) using specific primers, genome analysis involving short random primers, for example RAPD, may yield inconsistent results if crude plant DNA preparations are used as the template. When RNase A, a thermostable enzyme, was added to such reactions, highly repeatable banding patterns were obtained from crude plant DNA, thus speeding up analyses substantially.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Edwards, K., Johnstone, C., and Thompson, C. (1991) A simple and rapid method for the preparation of plant genomic DNA for PCR analysis.Nucleic Acid Res. 19, 1349.
Williams, J. G. K., Kubelik, A. R., Livak, K. J., Rafalski, J. A., and Tingey, S. V. (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers.Nucleic Acid Res. 18, 6531–6535.
Welsh, J. and McClelland, M. (1990) Fingerprinting genomes using PCR with arbitrary primers.Nucleic Acid Res. 18, 7213–7218.
Cardineau, G. A. and Filner, P. (1992) Compositions and methods for analyzing genomic variation. WO patent 92/07948, 14/5/92.
Caetano-Anollés, G., Bassam, B. J., and Gresshoff, P. M. (1991) DNA amplification fingerprinting using very short arbitrary oligonucleotide primers.Biotechnology 9, 553–557.
Muralidharan, K. and Wakeland, E. K. (1993) Concentration of primer and template qualitatively affects products in random amplified polymorphic DNA PCR.Biotechniques 14, 362.
Ellsworth, D. E., Rittenhouse, K. D., and Honeycutt, R. L. (1993) Artificial variation in randomly amplified polymorphic DNA banding patterns.Biotechniques 14, 214–217.
Büscher, N., Zyprian, E., and Blaich, R. (1993) Identification of grapevine cultivars by DNA analyses—pitfalls of random amplified polymorphic DNA techniques using 10mer primers.Vitis 32, 187–188.
Pikaart, M. J. and Villeponteau, B. (1993) Suppression of PCR amplification by high levels of RNA.Biotechniques 14, 24–27.
Yoon, C.-S. and Glawe, D. A. (1993) Pretreatment with RNase to improve PCR amplification of DNA using 10-mer primers.Biotechniques 14, 908–910.
{au{fnHeinze}, {gnB.}} ({dy1993}) {btGenetic stability in Norway spruce plants and embryos derived from somatic embryogenesis as analysed by randomly amplified DNA, PhD thesis}, {pmUniversität für Bodenkultur}, {plVienna, Austria}.
Sambrook, J., Fritsch, E. F., and Maniatis, T. (1989)Molecular Cloning. A Laboratory Manual. 2nd ed., Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Dhillon, S. S. (1987) DNA in tree species, inCell and Tissue Culture in Forestry, vol. 1: General Principles and Biotechnology (Bonga, J. M. and Durzan, D. J., eds.), Martinus Nijhof, Dordrecht, The Netherlands, pp. 298–313.
Walder, R. Y., Hayes, J. R., and Walder, J. A. (1993) Use of PCR primers containing a 3′-terminal ribose residue to prevent cross-contamination of amplified sequences.Nucleic Acids Res. 21, 4339–4343.
Jones, M. D. (1993) Reverse transcription of mRNA byThermus aquaticus DNA polymerase followed by polymerase chain reaction.Methods Enzymol. 218, 413–419.