Abstract
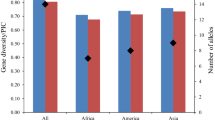
There are abundant soybean germplasm in China. In order to assess genetic diversity of Chinese summer soybean germplasm, 158 Chinese summer soybean accessions from the primary core collection ofG. max were used to analyze genetic variation at 67 SSR loci. A total of 460 alleles were detected, in which 414 and 419 alleles occurred in the 80 Huanghuai and the 78 Southern summer accessions, respectively. The average number of alleles per locus was 6.9 for all the summer accessions, and 6.2 for both Huanghuai and Southern summer accessions. Marker diversity (D) per locus ranged from 0.414 to 0.905 with an average of 0.735 for all the summer accessions, from 0.387 to 0.886 with an average of 0.708 for the Huanghuai summer accessions, and from 0.189 to 0.884 with an average of 0.687 for the Southern summer accessions. The Huanghuai and Southern summer germplasm were different in the specific alleles, allelic-frequencies and pairwise genetic similarities. UPGMA cluster analysis based on the similarity data clearly separated the Huanghuai from Southern summer soybean accessions, suggesting that they were different gene pools. The results indicate that Chinese Huanghuai and Southern summer soybean germplasm can be used to enlarge genetic basis for developing elite summer soybean cultivars by exchanging their germplasm.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Gai, J. Y., Zhao, T. I., Cui, Z. L. et al., The Genetic base for 651 soybean cultivars released during 1923–1995 in China, Scientia Agriculture Sinica, 1998, 31(5): 35–43.
Akkaya, M. S., Bhagwat, A. A., Cregan, P. B., Length polymorphism of simple sequence repeats DNA in soybean, Genetics, 1992, 132: 1131–1139.
Maughan, P. J., Saghai Maroof, M. A., Buss, G. R., Microsatellite and amplified sequence length polymorphism in cultivated and wild soybean, Genome, 1995, 38: 715–723.
Rongwen, J., Akkaya, M. S., Bhahwat, A. A. et al., The use of microsatellite DNA markers for soybean genotype identification, Theor. Appl. Genet, 1995, 90: 43–48.
Diwan, N., Cregan, P. B., Automated sizing of fluorescent-labeled simple sequence repeat (SSR) markers to assay genetic variation in soybean, Theor. Appl. Genet., 1997, 95: 723–733.
Narvel, J. M., Fehr, W. R., Chu, W. C. et al., Simple sequence repeat diversity among soybean plant introductions and elite genotypes, Crop Sci., 2000, 40: 1452–1458.
Brown-Guedira, G. L., Thompson, J. A., Nelson, R. L. et al., Evaluation of genetic diversity of soybean introductions and North American ancestors using RAPD and SSR markers, Crop Sci., 2000, 40: 815–823.
Abe, J., Xu, D. H., Suzuki, Y. et al., Soybean germplasm pools in Asia revealed by nuclear SSRs, Theor. Appl. Genet., 2003, 106: 445–453.
Institute of Oil Crops, Chinese Academy of Agricultural Sciences. A catalog of soybean germplasm resources from China, Beijing: Agricultural Press, 1982.
Institute of Crop Germplasm Resources, Chinese Academy of Agricultural Sciences, A catalog of soybean germplasm resources from China (Continuation 1), Beijing: Agricultural Press, 1991.
Institute of Crop Germplasm Resources, Chinese Academy of Agricultural Sciences, A catalog of soybean germplasm resources from China (Continuation 2), Beijing: Agricultural Press, 1996.
Zhou, X. A., Peng, Y.H., Wang, G. X. et al., Study on classifying and researching of soybean varieties in China, Crop Variety Germplasm, 1998, 1: 1–4.
Keim, P., Olson, T. C., Shoemaker, R. C., A rapid protocol for isolating soybean DNA, Soybean Genet. Neswsl., 1988, 15: 150–152.
Cregan, P. B., Jarvik, T., Bush, A. L. et al., An integrated genetic linkage map of the soybean genome, Crop Sci., 1999, 39: 1464–1490.
Qiu, L. J., Nelson, R. L., Vodkin, L. O., Evaluation of soybean germplasm with random amplification polymorphic DNA (RAPD) markers, Acta Agronomica Sinica, 1997, 23(4): 408–417.
Li, Z. L., Qiu, L. J., Thompaon, J. A. et al., Molecular genetic analysis of the U.S. and Chinese soybean ancestral lines, Crop Sci., 2001, 41(5): 1330–1336.
Li, Z. L., Nelson, R. L., Genetic diversity among soybean accessions from three countries measured by RAPDs, Crop Sci., 2001, 41: 1337–1347.
Scott, K. D., Eggler, P., Seaton, G. et al., Analysis of SSRs derived from grape ESTs, Theor. Appl. Genet., 2000, 100: 723–726.
Cho, Y.G., Ishil, T., Temnykh, S. et al., Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.), Theor. Appl. Genet., 2000, 100: 713–722.
Eujayl, I., Sorrells, M. E., Baum, M. et al., Isolation of EST-derived microsatellite markers for genetyping the A and B genomes of wheat, Theor. Appl. Genet., 2002, 104: 399–407.
Hackauf, B., Wehling, P., Identification of microsatellite polymorphisms in an expressed portion of the rye genome, Plant Breeding, 2002, 121: 17–25.
Thiel, T., Michalek, W., Varsheny, K. et al., Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.), Theor. Appl. Genet., 2003, 106: 411–422.
Gao, L. F., Jing, R. L., Huo, N. X. et al., One hundred and one new microsatellite loci derived from ESTs (EST-SSRs) in bread wheat, Theor. Appl. Genet, 2004, 108(7): 1392–1400.
Xu, Y., Ma, R. C., Xie, H.et al., Development of SSR markers for the phylogenetic analysis of almond trees from China and the Mediterranean Region, Genome, 2004, 47: 1091–1104.
Tian, A. G., Wang, J., Cui, P. et al., Characterization of soybean genomic features by analysis of its expressed sequence tags, Theor. Appl. Genet, 2004, 108: 903–913.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Xie, H., Guan, R., Chang, R. et al. Genetic diversity of Chinese summer soybean germplasm revealed by SSR markers. Chin.Sci.Bull. 50, 526–535 (2005). https://doi.org/10.1007/BF02897476
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02897476