Abstract
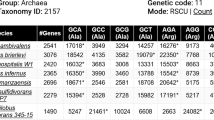
A computer program (PCBI) was developed to quickly calculate codon bias index (CBI). PCBI can analyze a gene containing introns. The 22 preferred codons defined fromSaccharomyces cerevisiae were used in PCBI as the standard to measure the CBI values. However, users can modify the preferred codons to suit each organism. The data PCBI provides include DNA sequence of open reading frame without introns, amino acid sequence of gene product, a table of amino acid composition, a table of codon usage and (G+C) content, parameters for calculating CBI, and the value of CBI. PCBI runs on DOS or Windows environment, but results can be saved in ASCII text format.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Grantham, R., Gautier, C., Gouy, M., Mercier, R., and Pave, A. (1980) Codon catalog usage and the genome hypothesis.Nucleic Acids Res. 8, 49–63.
Grantham, R., Gautier, C., Gouy, M., Jacobzone, M., and Mercier, R. (1981) Codon catalog usage is a genome strategy modulated for gene expressivity.Nucleic Acids Res. 9, 43–74.
Ikemura, T. (1981) Correlation between the abundance ofEscherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes.J. Mol. Biol. 146, 1–21.
Ikemura, T. (1981) Correlation between the abundance ofEscherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: A proposal for a synonymous codon choice that is optimal for theE. coli translational system.J. Mol. Biol. 151, 389–409.
Ikemura, T. (1982) Correlation between the abundance of yeast transfer RNAs and the occurrence of the respective codons in its protein genes.J. Mol. Biol. 158, 573–597.
Bennetzen, J. L., and Hall, B. D. (1982) Codon selection in yeast.J. Biol. Chem. 257, 3026–3031.
Kinnaird, J. H., Burns, P. A., and Fincham, J. R. (1991) An apparent rare-codon effect on the rate of translation of aNeurospora gene.J. Mol. Biol. 221, 733–736.
Solomovici, J., Lesnik, T., and Reiss, C. (1997) DoesEscherichia coli optimize the economics of the translation process?J. Theor. Biol. 185, 511–521.
Sharp, P. M. and Li, W. H. (1987) The codon Adaptation Index—a measure of directional synonymous codon usage bias, and its potential applications.Nucleic Acids Res. 15, 1281–1295.
Freire-Picos, M. A., Gonzalez-Siso, M. I., Rodriguez-Belmonte, E., Rodriguez-Torres, A. M., Ramil, E., and Cerdan, M. E. (1994) Codon usage inKluyveromyces lactis and in yeast cytochrome c-encoding genes.Gene 139, 43–49.
Wright, F. (1990) The effective number of codons used in a gene.Gene 87, 23–29.
Nagasu, T. and Hall, B. D. (1985) Nucleotide sequence of theGDH gene coding for the NADP-specific glutamate dehydrogenase ofSaccharomyces cerevisiae.Gene 37, 247–253.
Wang, T. T., Lee, C. F., and Lee, B. H. (1998) The molecular biology ofSchwanniomyces occidentalis Klocker.CRC Crit. Rev. Biotechnol. (Submitted).
Sharp, P. M. and Cowe, E. (1991) Synonymous codon usage inSaccharomyces cerevisiae.Yeast 7, 657–678.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, TT., Cheng, WC. & Lee, B.H. A simple program to calculate codon bias index. Mol Biotechnol 10, 103–106 (1998). https://doi.org/10.1007/BF02760858
Issue Date:
DOI: https://doi.org/10.1007/BF02760858