Abstract
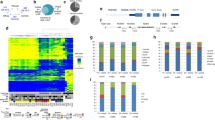
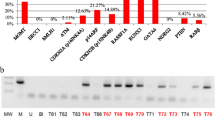
The underlying basis of the malignant progression of astrocytomas is a specific and cumulative series of genetic alterations, most of which are confined to high-grade tumors. In contrast, a proportion of low-grade astrocytomas have a relatively normal-appearing genome when examined with standard genetic screening methods. These methods do not detect epigenetic events such as aberrant methylation of CpG island, which result in transcriptional silencing of important cancer genes. To determine if aberrant methylation is involved in the early stages of astrocytoma development, we assessed the methylation status of 1184 genes in each of 14 low-grade astrocytomas using restriction landmark genome scanning (RLGS). The results showed nonrandom and astrocytoma-specific patterns of aberrantly methylated genes. We estimate that an average of 1544 CpG island-associated genes (range, 38 to 3731) of the approximately 45,000 in the genome are aberrantly methylated in each tumor. Expression of a significant proportion of the genes could be reactivated by 5-aza-2-deoxycytidine-induced demethylation in cultured glioma cell lines. The data suggest that aberrant methylation of genes is more prevalent than genetic alterations and may have consequences for the development of low-grade astrocytomas.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Kleihues P, Ohgaki H (1999) Primary and secondary glioblastoma: from concept to clinical diagnosis. Neuro-Oncology 1:44–51
Cavenee WK, Furnari FB, Nagane M et al. (2000) Diffusely infiltrating astrocytomas. In: Cavenee WK, Kleihues P (eds) Tumours of the nervous system. IARC Press, Lyon, pp 9–52
Baylin SB, Herman JG, Graff JR, et al. (1998) Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res 72:141–196
Jones PA, Laird PW (1999) Cancer epigenetics comes of age. Nature Gene 21:163–167
Gama-Sosa MA, Slagel VA, Trewyn RW, et al. (1983) The 5-methylcytosine content of DNA from human tumors. Nucl Acids Res 11:6883–6894
Chen RZ, Pettersson U, Beard C et al. (1998) DNA hypomethylation leads to elevated mutation rates. Nature 395:89–93
Stirzaker C, Millar DS, Paul CL et al. (1997) Extensive DNA methylation spanning the Rb promoter in retinoblastoma tumors. Cancer Res 57:2229–2237
Herman JG, Latif F, Weng, Y, et al. (1994) Silencing of the VHL tumor-suppressor gene by DNA methylation in renal carcinoma. Proc Nat Acad Sci USA 91:9700–9704
Kane MF, Loda M, Gaida GM, et al. (1997) Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch repair-defective human tumor cell lines. Cancer Res 57:808–811
Costello JF, Futscher BW, Kroes RA, et al. (1994) Methylation-related chromatin structure is associated with exclusion of transcription factors from and suppressed expression of the O-6-methylguanine DNA methyltransferase gene in human glioma cell lines. Mol Cell Bio 14:6515–6521
Li Q, Ahuja N, Burger PC, et al. (1999) Methylation and silencing of the thrombospondin-1 promoter in human cancer. Oncogene 18:3284–3289
Graff JR, Herman JG, Lapidus RG, et al. (1995) E-cadherin expression is silenced by DNA hypermethylation in human breast and prostate carcinomas. Cancer Res 55:5195–5199
Antequera F, Bird A (1993) Number of CpG islands and genes in human and mouse. Proc Nat Acad USA 90:11995–11999
Hatada I, Hayashizaki Y, Hirotsune S, et al. (1991) A genomic scanning method for higher organisms using restriction sites as landmarks. Proc Nat Acad Sci USA 88:9523–9527
Lindsay S, Bird AP (1987) Use of restriction enzymes to detect potential gene sequences in mammalian DNA. Nature 327:336–338
Plass C, Shibata H, Kalcheva I, et al. (1996) Identification of Grf1 on mouse chromosome 9 as an imprinted gene by RLGS-M. Nature Genetics 14:106–109
Hayashizaki Y, Shibata, H, Hirotsune S, et al. (1994) Identification of an imprinted U2af binding protein related sequence on mouse chromosome 11 using the RLGS method. Nature Genet 6:33–40
Costello JF, Plass C, Arap W, et al. (1997) Cyclin-dependent kinase 6 (CDK6) amplification in human gliomas identified using two-dimensional separation of genomic DNA. Cancer Res 57:1250–1254
Frühwald MC, O'Dorisio MS, Rush L, et al. (2000) Gene amplification in PNET/medulloblastoma: mapping of a novel amplified gene within the MYCN amplicon. J Med Genet 37:501–509
Costello JF, Fruhwald MC, Smiraglia DJ, et al. (2000) Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nature Genet 24:132–138
Plass C, Yu F, Yu L, et al. (1999) Restriction landmark genome scanning for aberrant methylation in primary refractory and relapsed acute myeloid leukemia; involvement of the WIT-1 gene. Oncogene 18:3159–3165
Akama TO, Okazaki Y, Ito M, et al. (1997) Restriction landmark genomic scanning (RLGS-M)-based genome-wide scanning of mouse liver tumors for alterations in DNA methylation status. Cancer Res 57:3294–3299
Yoshikawa H, de la Monte S, Nagai H, et al. (1996) Chromosomal assignment of human genomicNotI restriction fragments in a two-dimensional electrophoresis profile. Genomics 31:28–35
Graff JR, Herman JG, Myöhänen S, et al. (1997) Mapping patterns of CpG island methylation in normal and neoplastic cells implicates both upstream and downstream regions in de novo methylation. J Biol Chem 272:22322–22329
Graff JR, Greenberg VE, Herman JG, et al. (1998) Distinct patterns of E-cadherin CpG island methylation in papillary, follicular, Hurthle's cell, and poorly differentiated human thyroid carcinoma. Cancer Res 58:2063–2066
Plass C, Weichenhan D, Catanese J, et al. (1997) An arrayed humanNotI-EcoRV boundary library as a tool for RLGS spot analysis. DNA Res 4:253–255
Smiraglia DJ, Frühwald MC, Costello JF, et al. (1999) A new tool for the rapid cloning of amplified and hypermethylated human DNA sequences from restriction landmark genomic scanning gels. Genomics 58:254–262
Bachman KE, Herman JG, Corn PG, et al. (1999) Methylation-associated silencing of the tissue inhibitor of metalloproteinase-3 gene suggests a suppressor role in kidney, brain, and other human cancers. Cancer Res 59:798–802
Merlo A, Herman JG, Mao L et al. (1995) 5′ CpG island methylation is associated with transcriptional silencing of the tumour suppressor p16/CDKN2/MTS1 in human cancers. Nature Med 1:686–692
Myöhânen SK, Baylin SB, Herman JG (1998) Hypermethylation can selectively silence individual p16ink4A alleles in neoplasia. Cancer Res 58:591–593
Nan X, Tate P, Li E, et al. (1996) DNA methylation specifies chromosomal localization of MeCP2. Mol Cell Biol 16:414–421
Cameron EE, Bachman KE, Myöhänen S, et al. (1999) Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nature Genet 21:103–107
Bermingham JR Jr, Scherer SS, O'Connell S, et al. (1996) Tst-1/Oct-6/SCIP regulates a unique step in peripheral myelination and is required for normal respiration. Genes Dev 10:1751–1762
Nambu JR, Franks RG, Hu S, et al. (1990) The single-minded gene of Drosophila is required for the expression of genes important for the development of CNS midline cells. Cell 63:63–75
Offermanns S, Simon MI (1996) Organization of transmembrane signalling by heterotrimeric G proteins. Cancer Surv 27:177–198
Kallioniemi A, Kallioniemi OP, Sudar D, et al. (1992) comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 258:818–821
Cavenee WK, Dryja TP, Phillips RA, et al. (1983) Expression of recessive alleles by chromosomal mechanisms in retinoblastoma. Nature 305:779–784
Mertens F, Johansson B, Höglund M, et al. (1997) Chromosomal imbalance maps of malignant solid tumors: a cytogenetic survey of 3185 neoplasms. Cancer Res 57:2765–2780
Gardiner-Garden M, Frommer M (1987) CpG islands in vertebrate genomes. J Mol Biol 196:261–282
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Costello, J.F., Plass, C. & Cavenee, W.K. Aberrant methylation of genes in low-grade astrocytomas. Brain Tumor Pathol 17, 49–56 (2000). https://doi.org/10.1007/BF02482735
Issue Date:
DOI: https://doi.org/10.1007/BF02482735