Summary
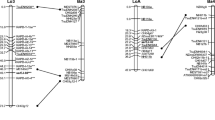
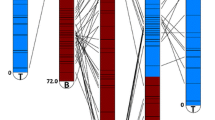
A detailed linkage map ofB. rapa (syn.campestris) was constructed based on segregation of 280 restriction fragment length polymorphism loci, detected by using 188 genomic DNA clones as probes on DNAs from a F2 population of Chinese cabbage ‘MichihilF’בSpring broccoli.’ These genetic markers covered 1,850 centiMorgans (cM) and defined ten linkage groups, which equals the haploid chromosome number of this species. Extensive sequence duplication was evident by the detection of two or more segregating loci with each of 69 clones (36.7% of the total). Although some duplicated loci were randomly distributed throughout the genome, many had linkage arrangements that were conserved on different linkage groups, suggesting that large chromosome fragments were present in multiple copies. However, conservation in the linkage arrangement of duplicate loci throughout entire pairs of linkage groups was not observed. Single-copy loci were often found to be located within conserved duplicated regions, and linkage distances between some loci having conserved duplicated arrangements were substantially different between the duplicated regions. Structural rearrangements, such as insertions, deletions, and inversions or combinations of these events, seemed to be related to the alternations of map distances between duplicated loci and to the dispersal of duplicated chromosome fragments. These results suggest thatB. rapa has evolved in part by duplication of chromosomes or large chromosome fragments with subsequent structural rearrangements.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Armstrong KC, Keller WA (1981) Chromosome pairing in haploids ofBrassica campestris. Theor Appl Genet 59:49–52
Attia T, Robbelen G (1986) Cytogenetic relationship within cultivatedBrassica analyzed in amphidiploids from the three diploid ancestors. Can J Genet Cytol 28:323–329
Beckmann JS, Soller M (1986) Restriction fragment length polymorphisms and genetic improvement of agricultural species. Euphytica 35:111–124
Bernatzky R, Tanksley SD (1986) Toward a saturated linkage map in tomato based on isozymes and random cDNA sequences. Genetics 112:887–898
Bonierbale MW, Plaisted RL, Tanksley SD (1988) RFLP maps based on a common set of clones reveal modes of chromosomal evolution in potato and tomato. Genetics 120:1095–1103
Burr BS, Evola V, Burr FA, Beckmann JS (1986) The application of restriction fragment length polymorphism to plant breeding. In: Setlow J, Hollaender A (eds) Genetic engineering, vol 5. Plenum, New York, pp 45–59
Catcheside DG (1937) Secondary pairing inBrassica oleracea.
Helentjaris T, King G, Slocum M, Siedenstrang C, Wegman S (1985) Restriction fragment polymorphism as probes for plant diversity and their development as tools for applied plant breeding. Plant Mol Biol 5:109–118
Helentjaris T, Slocum M, Wright S, Schaefter A, Nienhuis J (1986) Construction of genetic linkage maps in maize and tomato using restriction fragment length polymorphisms. Theor Appl Genet 72:761–769
Helentjaris T, Weber D, Wright S (1988) Identification of the genomic locations of duplicated sequences in maize by analysis of restriction fragment length polymorphisms. Genetics 118:353–363
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg I. (1987) Mapmaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Landry BS, Kesseli RV, Farrara B, Michelmore RW (1987) A genetic map of lettuce (Lactuca sativa L.) with restriction fragment length polymorphism, isozyme, disease resistance, and morphological markers. Genetics 116:331–337
McCouch SR, Kocher G, Yu ZG, Wang ZY, Khush GS, Coffman WR, Tanksley SD (1988) Molecular mapping of rice chromosomes. Theor Appl Genet 76:815–829
Mizushima U (1980) Genome analysis inBrassica and allied genera. In: Tsunoda S, Hinata K, Gomez-Campo C (eds)Brassica crops and wild allies. Japanese Scientific Society Press, Tokyo, pp 89–105
Osborn TC, Alexander DC, Fobes JF (1987) Identification of restriction fragment length polymorphisms linked to genes controlling soluble solid content in tomato fruit. Theor Appl Genet 73:350–356
Prakash S, Hinata K (1980) Taxonomy, cytogenetics, and origin of cropBrassica, a review. Opera Bot 55:1–59
Robbelen G (1960) Beiträge zur Analyse desBrassica genomes. Chromosoma II:205–228
Slocum MK, Figdore SS, Kennard WC, Suzuki JY, Osborn TC (1990) Linkage arrangement of restriction fragment length polymorphism loci inBrassica oleracea. Theor Appl Genet 80:57–64
Song KM, Osborn TC, Williams PH (1988a)Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 1. Genome evolution of diploid and amphidiploid species. Theor Appl Genet 75:784–794
Song KM, Osborn TC, Williams PH (1988b)Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 2. Preliminary analysis of subspecies withinB. rapa (syn.campestris) andB. oleracea. Theor Appl Genet 76:593–600
Song KM, Osborn TC, Williams PH (1990)Brassica taxonomy based on nuclear RFLPs. 3. Genome relationships inBrassica and related genera and the oriein ofB. oleracea and
Author information
Authors and Affiliations
Additional information
Communicated by J. Beckmann
Rights and permissions
About this article
Cite this article
Song, K.M., Suzuki, J.Y., Slocum, M.K. et al. A linkage map ofBrassica rapa (syn.campestris) based on restriction fragment length polymorphism loci. Theoret. Appl. Genetics 82, 296–304 (1991). https://doi.org/10.1007/BF02190615
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02190615