Summary
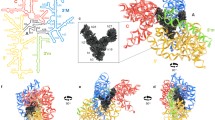
We present the sequence of the nuclearencoded ribosomal small-subunit RNA from soybean. The soybean 18S rRNA sequence of 1807 nucleotides (nt) is contained in a gene family of approximately 800 closely related members per haploid genome. This sequence is compared with the ribosomal small-subunit RNAs of maize (1805 nt), yeast (1789 nt),Xenopus (1825 nt), rat (1869 nt), andEscherichia coli (1541 nt). Significant sequence homology is observed among the eukaryotic small-subunit rRNAs examined, and some sequence homology is observed between eukaryotic and prokaryotic small-subunit rRNAs. Conserved regions are found to be interspersed among highly diverged sequences. The significance of these comparisons is evaluated using computer simulation of a random sequence model. A tentative model of the secondary structure of soybean 18S rRNA is presented and discussed in the context of the functions of the various conserved regions within the sequence. On the basis of this model, the short basepaired sequences defining the four structural and functional domains of all 18S rRNAs are seen to be well conserved. The potential roles of other conserved soybean 18S rRNA sequences in protein synthesis are discussed.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bolivar F, Rodriguez RL, Greene PJ, Betlach MC, Heyneker HL, Boyer HW (1977) Construction and characterization of new cloning vehicles: II. A multipurpose cloning system. Gene 2:95–113
Bolivar F, Backman K (1979) Plasmids ofE. coli as cloning vectors. Methods Enzymol 68:245–267
Brimacombe R (1982) The secondary structure of ribosomal RNA, and its organization within the ribosomal subunits. Biochem Soc Symp 47:49–60
Brown AHD, Clegg MT (1983) Analysis of variation in related DNA sequences. In: Weir BS (ed) Statistical analysis of DNA sequence data. Marcel Dekker, New York, pp 107–132
Carreira LH, Carlton BC, Bobbio SM, Nagao RT, Meagher RB (1980) Construction and application of a modified “Gene Machine”: A circular concentrating preparative gel electrophoresis device employing discontinuous elution. Anal Biochem 106:455–468
Chao S, Sederoff RR, Levings CS III (1984) Nucleotide sequence and evolution of the 18S ribosomal RNA gene in maize mitochondria. Nucleic Acids Res 12:6629–6644
Connaughton JF, Rairkar A, Lockard RE, Kumar A (1984) Primary structure of rabbit 18S ribosomal RNA determined by direct RNA sequence analysis. Nucleic Acids Res 12:4731–4745
Denhardt D (1966) A membrane-filter technique for the detection of complementary DNA. Biochem Biophys Res Commun 23:641–646
Eckenrode VK (1983) Sequence analysis of an 18S rRNA gene and physical mapping of an rDNA repeat unit from soybean. Ph.D. dissertation, University of Georgia, Athens, Georgia
Elleman TC (1978) A method for detecting distant evolutionary relationships between protein or nucleic acid sequences in the presence of deletions and insertions. J Mol Evol 11:143–161
Eperon IC, Anderson S, Nierlich DP (1980) Distinctive sequence of human mitochondrial ribosomal RNA genes. Nature 286:460–467
Erdmann VA, Huysmens E, Vandenberghe A, De Wachter R (1983) Collection of published 5S and 5.8S ribosomal RNA sequences. Nucleic Acids Res 11:r105-r133
Erickson BW, Sellers PH (1983) Recognition of patterns in genetics sequences. In: Sankoff D, Kruskal JB (eds) Time warps, string edits, and macromolecules. Addison-Wesley, Reading, Massachusetts, pp 55–91
Friedrich H, Hemleben V, Meagher RB, Key JL (1979) Purification and restriction endonuclease mapping of soybean 18S and 25S ribosomal RNA genes. Planta 146:467–473
Hartigan JA (1975) Clustering algorithms. John Wiley & Sons, New York, pp 191–215
Herr W, Chapman NM, Noller HF (1979) Mechanism of ribosomal subunit association: discrimination of specific sites in 16S RNA essential for association activity. J Mol Biol 130:433–449
Jackson PJ, Lark KG (1982) Ribosomal RNA synthesis in soybean suspension cultures growing in different media. Plant Physiol 69:234–239
Knuth D (1981) The art of computer programming. In: Seminumerical algorithms, 2nd ed vol 2. Addison-Wesley, Reading, Massachusetts, pp 1–77
Kozak M (1983) Comparison of initiation of protein synthesis in procaryotes, eucaryotes, and organelles. Microbiol Rev 47:1–45
Krayev AS, Kramerov DA, Skryabin KG, Ryskov AP, Bayev AA, Georgiev GP (1980) The nucleotide sequence of the ubiquitous repetitive DNA sequence B1 complementary to the most abundant class of mouse foldback RNA. Nucleic Acids Res 8:1201–1215
Kruskal JB (1983) An overview of sequence comparison. In: Sankoff D, Kruskal JB (eds) Time warps, string edits, and macromolecules. Addison-Wesley, Reading, Massachusetts, pp 1–40
Küntzel H, Köchel HG (1981) Evolution of rRNA and origin of mitochondria. Nature 293:751–755
Kushner SR (1978) An improved method for transformation ofEscherichia coli with colE1 derived plasmids. In: Boyer HW, Nicosia S (eds) Genetic engineering. Elsevier/North Holland, New York, pp 17–23
Lake JA (1983) Evolving ribosome structure: domains in archaebacteria, eubacteria, and eucaryotes. Cell 33:318–319
Liljas A (1982) Structural studies of ribosomes. Prog Biophys Mol Biol 40:161–228
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Maxam AM, Gilbert W (1980) Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol 65:499–560
McCarroll R, Olfen GJ, Stahl YD, Woese CR, Sogin ML (1983) Nucleotide sequence of theDictyostelium discoideum small-subunit ribosomal ribonucleic acid inferred from the gene sequence: evolutionary implications. Biochemistry 22:5858–5868
Meagher RB, Tait RC, Betlach M, Boyer HW (1977a) Protein expression inE. coli minicells by recombinant plasmids. Cell 10:521–536
Meagher RB, Shepherd RJ, Boyer HW (1977b) The structure of cauliflower moisaic virus: I. A restriction endonuclease map of cauliflower mosaic virus DNA. Virology 80:362–375
Messing J, Carlson J, Hagen G, Rubenstein I, Oleson A (1984) Cloning and sequencing of the ribosomal RNA genes in maize: the 17S region. DNA 3:31–40
Needleman SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino-acid sequence of two proteins. J Mol Biol 48:443–453
Noller HF (1980) Structure and topography of ribosomal RNA. In: Chambliss G, Craven GR, Davies J, Davis K, Kahan L, Nomura M (eds) Ribosomes: structure, function and genetics. University Park Press, Baltimore, Maryland, pp 3–22
Noller HF, Woese CR (1981) Secondary structure of 16S ribosomal RNA. Science 212:403–411
Ofengand J, Gornicki P, Chakraburtty K, Nurse K (1982) Functional conservation near the 3′ end of eukaryotic small subunit RNA: photochemical crosslinking of P site-bound acetylvalyl-tRNA to 18S RNA of yeast ribosomes. Proc Natl Acad Sci USA 79:2817–2821
Rubstov PM, Musakhonov MM, Zakharyev VM, Krayev AS, Skryabin KG, Bayev AA (1980) The structure of the yeast ribosomal RNA genes: I. The complete nucleotide sequence of the 18S ribosomal RNA gene fromSaccharomyces cerevisiae. Nucleic Acids Res 8:5779–5794
Salim M, Maden BEH (1981) Nucleotide sequence ofXenopus laevis 18S ribosomal RNA inferred from the gene sequence. Nature 291:205–208
Sankoff D, Cedergren RJ (1973) A test for nucleotide sequence homology. J Mol Biol 77:159–164
Sankoff D, Cedergren RJ, Lapalme G (1976) Frequency of insertion-deletion, transversion, and transition in the evolution of 5S ribosomal RNA. J Mol Evol 7:133–149
Shah DM, Hightower RC, Meagher RB (1982) Complete nucleotide sequence of a soybean actin gene. Proc Natl Acad Sci USA 79:1022–1026
Shine J, Dalgarno L (1974) The 3′-terminal sequence ofEscherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci USA 71:1342–1346
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
Steitz JA, Jakes K (1975) How ribosomes select initiator regions in mRNA: base pair formation between the 3′ terminus of 16S rRNA and the mRNA during initiation of protein synthesis inEscherichia coli. Proc Natl Acad Sci USA 72:4734–4738
Stiegler P, Carbon P, Ebel J-P, Ehresmann C (1981) A general secondary-structure model for prokaryotic and eucaryotic RNAs of the small ribosomal subunits. Eur J Biochem 120: 487–495
Thompson JF, Hearst JE (1983) Structure-function relations inE. coli 16S RNA. Cell 33:19–24
Torczynski R, Bollon AP, Fuke M (1983) The complete nucleotide sequence of the rat 18S ribosomal RNA gene and its comparison with the respective yeast and frog genes. Nucleic Acids Res 11:4879–4890
Varsanyi-Breiner A, Gusella JF, Keys C, Honsman DE, Sullivan D, Brisson N, Verma DPS (1979) The organization of a nuclear DNA sequence from a higher plant: molecular cloning and characterization of soybean ribosomal DNA. Gene 7:317–334
Wool IG (1980) The structure and function of eucaryotic ribosomes. In: Chambliss G, Craven GR, Davies J, Davis K, Kahan L, Nomura M (eds) Ribosomes: structure, function and genetics. University Park Press, Baltimore, Maryland, pp 797–824
Zwieb C, Glotz C, Brimacombe R (1981) Secondary structure comparisons between small subunit ribosomal RNA molecules from six different species. Nucleic Acids Res 9:3621–3640
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Eckenrode, V.K., Arnold, J. & Meagher, R.B. Comparison of the nucleotide sequence of soybean 18S rRNA with the sequences of other small-subunit rRNAs. J Mol Evol 21, 259–269 (1985). https://doi.org/10.1007/BF02102358
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02102358