Summary
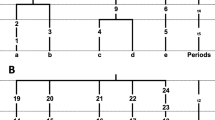
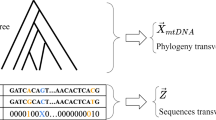
A molecular clock analysis was carried out on the nucleotide sequences of parts of the major noncoding region of mitochondrial DNA (mtDNA) from the major geographic populations of humans. Dates of branchings in the mtDNA tree among humans were estimated with an improved maximum likelihood method. Two species of chimpanzees were used as an outgroup, and the mtDNA clock was calibrated by assuming that the chimpanzee/human split occurred 4 million years ago, following our earlier works. A model of homogeneous evolution among sites does not fit well with the data even within hypervariable segments, and hence an additional parameter that represents a proportion of variable sites was introduced. Taking account of this heterogeneity among sites, the date for the deepest root of the mtDNA tree among humans was estimated to be 280,000±50,000 years old (±1 SE), although there remains uncertainty about the constancy of the evolutionary rate among lineages. The evolutionary rate of the most rapidly evolving sites in mtDNA was estimated to be more than 100 times greater than that of a nuclear pseudogene.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Akaike H (1974) A new look at the statistical model identification. Inst Electrical Electronics Engineers Trans Autom Contr AC-19:716–723
Anderson S, Bankier AT, Barrell BG, de Bruijn MHL, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith ALH, Staden R, Young IG (1981) Sequence and organization of the human mitochondrial genome. Nature 290:457–464
Brown WM (1980) Polymorphism in mitochondrial DNA of humans as revealed by restriction endonuclease analysis. Proc Natl Acad Sci USA 77:3605–3609
Brown WM, Prager EM, Wang A, Wilson AC (1982) Mitochondrial DNA sequences of primates: tempo and mode of evolution. J Mol Evol 18:225–239
Cann RL, Stoneking M, Wilson AC (1987) Mitochondrial DNA and human evolution. Nature 325:31–36
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1986) An estimation of the number of invariable sites is necessary for the accurate estimation of the number of nucleotide substitutions since a common ancestor. In: Gershowitz H, Rucknagel DL, Tashian RE (eds) Evolutionary perspectives and the new genetics. Liss, New York
Foran DR, Hixson JE, Brown WM (1988) Comparisons of ape and human sequences that regulate mitochondrial DNA transcription and D-loop DNA synthesis. Nucleic Acids Res 16: 5841–5861
Greenberg BD, Newbold JE, Sugino A (1983) Intraspecific nucleotide sequence variability surrounding the origin of replication in human mitochondrial DNA. Gene 21:33–49
Hasegawa M, Kishino H (1989) Confidence limits on the maximum-likelihood estimate of the hominoid tree from mitochondrial-DNA sequences. Evolution 43:672–677
Hasegawa M, Kishino H (1990) DNA sequence analysis and evolution of Homoinoidea. In: Kimura M, Takahata N (eds) New aspects of the genetics of molecular evolution. Springer-Verlag (in press)
Hasegawa M, Kishino H, Yano T (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Hasegawa M, Kishino H, Yano T (1989) Estimation of branching dates among primates by molecular clocks of nuclear DNA which slowed down in Hominoidea. J Hum Evol 18:461–476
Hasegawa M, Kishino H, Hayasaka K, Horai S (1990) Mitochondrial DNA evolution in primates: transition rate has been extremely low in lemur. J Mol Evol 31:113–121
Hixson JE, Brown WM (1986) A comparison of the small ribosomal RNA genes from the mitochondrial DNA of the great apes and humans: sequence, structure, evolution, and phylogenetic implications. Mol Biol Evol 3:1–18
Horai S, Matsunaga E (1986) Mitochondrial DNA polymorphism in Japanese. II. Analysis with restriction enzymes of four or five base pair recognition. Hum Genet 72:105–117
Horai S, Hayasaka K (1990) Intraspecific nucleotide sequence differences in the major noncoding region of human mitochondrial DNA. Am J Hum Genet 46:828–842
Horai S, Gojobori T, Matsunaga E (1986) Distinct clustering of mitochondrial DNA types among Japanese, Caucasians and Negroes. Jpn J Genet 61:271–275
Johnson MJ, Wallace DC, Ferris SD, Rattazzi MC, Cavalli-Sforza LL (1983) Radiation of human mitochondrial DNA types analyzed by restriction endonuclease cleavage patterns. J Mol Evol 19:255–271
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Kishino H, Hasegawa M (1990) Converting distance to time: an application to human evolution. Methods Enzymol 183: 550–570
Kocher TD, Thomas WK, Meyer A, Edwards SV, Pääbo S, Villablanca FX, Wilson AC (1989) Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci USA 86:6196–6200
Lanave C, Preparata G, Saccone C, Serio G (1984) A new method for calculating evolutionary substitution rates. J Mol Evol 20:86–93
Vigilant L, Pennington R, Harpending H, Kocher TD, Wilson AC (1989) Mitochondrial DNA sequences in single hairs from a southern African population. Proc Natl Acad Sci USA 86:9350–9354
Wilson AC, Cann RL, Carr SM, George M, Gyllensten UB, Helm-Bychowski KM, Higuchi RG, Palumbi SR, Prager EM, Sage RD, Stoneking M (1985) Mitochondrial DNA and two perspectives on evolutionary genetics. Biol J Linn Soc Lond 26: 375–400
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hasegawa, M., Horai, S. Time of the deepest root for polymorphism in human mitochondrial DNA. J Mol Evol 32, 37–42 (1991). https://doi.org/10.1007/BF02099927
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02099927