Abstract
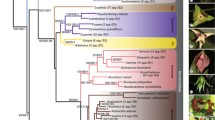
In contrast to animals, the slowly evolving mitochondrial nucleotide sequences of plants appear well suited to investigate phylogenetic relations between old taxonomic groups. Analysis ofnad5 gene sequences in 47 bryophytes, the living representatives of very early land plants, confirm this assessment. Statistically reliable phylogenetic trees are obtained with different mathematical approaches. A group I intron sequence conserved in thenad5 gene of all 30 mosses and 15 liverworts investigated supports a sister group relationship of the two classes. The intron sequence adds phylogenetic information for fine resolution on top of the conserved exon sequences down to the level of classically defined orders or families, respectively. This intron is not present in the hornwortsAnthoceros husnotii andA. punctatus. The results allow statements on diverging taxonomic interpretations and support the monophyly of the liverworts, mosses, Jungermanniidae, Marchantiidae and Bryidae, and allow recognition of subclasses like Hypnanae and Dicrananae. Among the mosses, the derived orders (subclass Bryidae) are confidently set apart from the Sphagnales, Andreaeales, Polytrichales and Tetraphidales with Buxbaumiales occupying a mediating position. Among the liverworts, full support is found for the classic separation of simple (jungermanniid) and complex thalloid (marchantiid) species with a strikingly low mitochondrial sequence divergence among the latter.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bopp M., Capesius I. (1996) New aspects of bryophyte taxonomy provided by a molecular approach. Bot. Acta 109: 368–372.
Bowe L. M., dePamphilis C. W. (1996) Effects of RNA editing and gene processing on phylogenetic reconstruction. Mol. Biol. Evol. 13: 1159–1166.
Capesius I. (1995) A molecular phylogeny of bryophytes based on the nuclear encoded 18S rRNA genes. J. Plant Physiol. 146: 59–63.
Capesius I., Stech M. (1997) Molecular relationships within mosses based on 18S rRNA gene sequences. Nova Hedwigia 64: 525–533.
Capesius I., Bopp M. (1997) New classification of liverworts based on molecular and morphological data. Plant Syst. Evol. 207: 87–97.
DeBoer S. H., Ward L. J., Li X., Chittarajan S. (1995) Attenuation of PCR inhibition in the presence of plant compounds by addition of BLOTTO. Nucl. Acids Res. 23: 2567–2568.
Devereux R., Haeberli P., Smithies O. (1984) A comprehensive set of sequence analysis programs for the VAX. Nucl. Acids. Res. 12: 387–395.
Doyle J. J., Doyle J. L. (1990) Isolation of plant DNA from fresh tissue. Focus 12: 13–15.
Felsenstein J. (1995) PHYLIP version 3.57c manual. University of Washington, Seattle.
Frahm J.-P., Frey W. (1992) Moosflora. UTB, Stuttgart.
Fukarek F., Schultze-Motel J., Siegel M. (1992) Moose, Farne, Nacktsamer. Urania, Leipzig.
Hedderson T. A., Chapman R. L., Rootes W. L. (1996) Phylogenetic relationships of bryophytes inferred from nuclear-encoded rRNA gene sequences. Plant Syst. Evol. 200: 213–224.
Hiesel R., von Haeseler A., Brennicke A. (1994) Plant mitochondrial nucleic acid sequences as a tool for phylogenetic analysis. Proc. Natl. Acad. Sci. USA 91: 634–638.
Knoop B. (1984) Culture methods for bryophytes. In: Vasil I. (ed.) Cell culture and somatic cell genetics of plants. Academic Press, New York, pp. 96–106.
Knoop V., Schuster W., Wissinger B., Brennicke A. (1991) Trans splicing integrates an exon of 22 nucleotides into thenad5 mRNA in higher plant mitochondria. EMBO J. 10: 3483–3493.
Lewis L. A., Mishler B. D., Vilgalys R. (1997) Phylogenetic relationships of the liverworts (Hepaticae), a basal embryophyte lineage, inferred from nucleotide sequence data of the chloroplast generbcL. Mol. Phyl. Evol. 7: 377–393.
Malek O., Lättig K., Hiesel R., Brennicke A., Knoop V. (1996) RNA editing in bryophytes and a molecular phylogeny of land plants. EMBO J. 15: 1403–1411.
Mishler B. D., Churchill S. P. (1984) A cladistic approach to the phylogeny of the “bryophytes”. Brittonia 36: 406–424.
Mishler B. D. (1986) A Hennigian approach to bryophyte phylogeny. J. Bryol. 14: 71–81.
Oda K., Yamato K., Ohta E., Nakamura Y., Takemura M., Nozato N., Akashi K., Kanegae T., Ogura Y., Kohchi T., Ohyama K. (1992) Gene organization deduced from the complete sequence of liverwortMarchantia polymorpha mitochondrial DNA. A primitive form of plant mitochondrial genome. J. Mol. Biol. 223: 1–7.
Samigullin T. H., Valiejo-Roman K. M., Troitsky A. V., Bobrova V. K., Filin V. R., Martin W., Antonov A. S. (1998) Sequences of rDNA internal transcribed spacers from the chloroplast DNA of 26 bryophytes: properties and phylogenetic utility. FEBS Lett. 422: 47–51.
Schuster R. M. (ed.) (1984) New manual of bryology. Hattori Botanical Laboratory, Nichinan.
Sitte P., Ziegler H., Ehrendorfer F., Bresinsky A. (1998) Strasburger — Lehrbuch der Botanik für Hochschulen, 34th ed. Gustav Fischer, Stuttgart Jena New York.
Vitt D. H. (1984) Classification of the Bryopsida. In: Schuster R. M. (ed.) New manual of bryology. Hattori Botanical Laboratory, Nichinan, pp. 696–759.
Walther K. (1983) Kapitel V,2, Brophytina. Laubmoose. In: Gerloff J., Poelt J. (eds.) Syllabus der Pflanzenfamilien. Gebrüder Borntraeger, Berlin.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Beckert, S., Steinhauser, S., Muhle, H. et al. A molecular phylogeny of bryophytes based on nucleotide sequences of the mitochondrialnad5 gene. Pl Syst Evol 218, 179–192 (1999). https://doi.org/10.1007/BF01089226
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01089226