Abstract
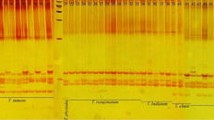
The recent development of molecular marker technology is revolutionising the study of plant populations, providing opportunities to address questions requiring a precise knowledge of pedigrees. We applied Inter-Simple Sequence Repeat (ISSR) PCR to severalBrassica oleracea accessions and toBrassica napus. Four microsatellite-primers were screened and, among the 136 reproducible fragments recorded, 25 (18.4%) fragments were common for allBrassica, 27 (19.9%) were unique and 84 (61.7%) were phylogenetically informative. Each individual test sample exhibited a unique molecular genotype. ISSR markers provided a rapid approach to analyse genetic diversity and reflected the known genetic relationships among selected entries. ISSR markers appeared of great value in gene bank management and the establishment of genetic similarity, and can be applied to allogamous (autumn and winter cauliflower) crops.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Akkaya M. A., Bhagwat A. A., Cregan P. B. (1992) Length polymorphisms of simple sequence repeat DNA in soybean. Genetics 132: 1131–1139.
Blair M. W., Panaud O., McCouch S. R. (1999) Inter-Simple sequence repeat (ISSR) amplification for analysis of microsatellite motif frequency and fingerprinting in rice (Oryza sativa L.). Theor. Appl. Genet. 99: 780–792.
Charters Y. M., Robertson A., Wilkinson M. J., Ramsay G. (1996) PCR analysis of oilseed rape cultivars (Brassica napus L. ssp.oleifera) using 5′-anchored simple sequence repeat (SSR) primers. Theor. Appl. Genet. 92: 442–447.
Dias J. S. (1995) The Portuguese tronchuda cabbage and galega kales landraces: a historical review. Genet. Res. Crop Evol. 42: 179–184.
Fang D. Q., Roose M. L. (1997) Identification of closely related citrus cultivars with inter-simple sequence repeat markers. Theor. Appl. Genet. 95: 408–417.
Felsenstein J. (1993) PHYLIP — PHYLogeny Inference Package (Version 3.5c) Department of Genetics, University of Washington, Seattle/WA.
Fisher P. J., Gardner R. C., Richardson T. E. (1996) Single locus microsatellites isolated using 5′ anchored PCR. Nucl. Acids Res. 24: 4369–4371.
Gootlieb M., Chavko M. (1987) Silver staining of native and denatured eukaryotic DNA in agarose gels. Anal. Bioch. 165: 33–37.
Kantety R. V., Zeng X. P., Bennetzen J. L., Zehr B. E. (1995) Assessment of genetic diversity in dent and popcorn (Zea mays L.) inbred lines using inter-simple sequence repeat (ISSR) amplification. Mol. Breed. 1: 365–373.
Kojima T., Nagaoka T., Noda K., Ogihara Y. (1998) Genetic linkage map of ISSR and RAPD markers in Einkorn wheat in relation to that of RFLP marker. Theor. Appl. Genet. 96: 37–45.
Litt M., Luty J. A. (1989) A hypervariable microsatellite revealed by in vitro amplification of a dinucleotide repeat within the cardiac muscle actin gene. A. J. Hum. Genet. 44: 388–396.
Margalé E., Hervé Y., Hu J., Quiros C. F. (1995) Determination of genetic variability by RAPD markers in cauliflower, cabbage and kale local cultivars from France. Gen. Res. Crop Evol. 42: 281–289.
Nagaoka T., Ogihara Y. (1997) Applicability of inter-simple sequence repeat polymorphisms in wheat for use as DNA markers in comparison to RFLP and RAPD markers. Theor. Appl. Genet. 94: 597–602.
Nei M., Li W. H. (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA 76: 5269–5273.
Saghai-Maroof M. A., Soliman K. M., Jorgensen R. A., Allard R. W. (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc. Natl. Acad. Sci. USA 81: 8014–8018.
Song K. M., Osborn T. C., Williams P. H. (1988)Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 2. Preliminary analysis of subspecies withinB. rapa (syn.campestris) andB. oleracea. Theor. Appl. Genet. 76: 593–600.
Song K. M., Osborn T. C., Williams P. H. (1990)Brassica taxonomy based on restriction fragment length polymorphisms (RFLPs). 3. Genome relationships inBrassica and related genera and the origin ofB. oleracea andB. rapa (syn.campestris). Theor. Appl. Genet. 79: 497–506.
Weber J. L., May P. E. (1989) Abundant class of human DNA polymorphism which can be typed using the polymerase chain reaction. A. J. Hum. Genet. 44: 388–396.
Wolff K., Zietkiewicz E., Hofstra H. (1995) Identification ofChrysanthemum cultivars and stability of DNA fingerprint patterns. Theor. Appl. Genet. 91: 439–447.
Zietkiewicz E., Rafalski A., Labuda D. (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20: 176–183.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Leroy, X.J., Leon, K. & Branchard, M. Characterisation ofBrassica oleracea L. by microsatellite primers. Pl Syst Evol 225, 235–240 (2000). https://doi.org/10.1007/BF00985470
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00985470