Abstract
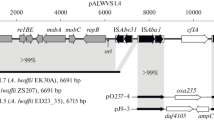
The genetic organization of the DNA region encoding the phenol degradation pathway ofPseudomonas putida H has been investigated. This strain can utilize phenol or some of its methylated derivatives as its sole source of carbon and energy. The first step in this process is the conversion of phenol into catechol. Catechol is then further metabolized via themeta-cleavage pathway into TCA cycle intermediates. Genes encoding these enzymes are clustered on the plasmid pPGH1. A region of contiguous DNA spanning about 16 kb contains all of the genetic information necessary for inducible phenol degradation. The analysis of mutants generated by insertion of transposons and cassettes indicates that all of the catabolic genes are contained in a single operon. This codes for a multicomponent phenol hydroxylase andmeta-cleavage pathway enzymes. Catabolic genes are subject to positive control by the gene product(s) of a second locus.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bagdasarian M, Timmis KN (1981) Host:vector systems for gene cloning inPseudomonas. In: Goebel W, Hofschneider PG (eds) Gene cloning in organisms other thanE. coli. Springer-Verlag, Berlin Heidelberg New York, pp 47–67
Bartilson M, Nordlund I, Shingler V (1990) Location and organization of the dimethylphenol catabolic genes ofPseudomonas CF600. Mol Gen Genet 220: 294–300
Bartilson M, Shingler V (1989) Nucleotide sequence and expression of the catechol 2,3-dioxygenase encoding gene of phenolcatabolizingPseudomonas CF600. Gene 85: 233–238
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7: 1513–1523
Boyer HW, Roulland-Dussoix D (1969) A complementation analysis of the restriction and modification of DNA inEscherichia coli. J Mol Biol 41: 459–472
Cane PA, Williams PA (1986) A restriction map of naphthalene catabolic plasmid pWW60-1 and the location of some of its catabolic genes. J Gen Microbiol 132: 2919–2929
Chakrabarty AM, Chou G, Gunsalus IC (1973) Genetic regulation of octane dissimilation plasmid inPseudomonas. Proc Natl Acad Sci USA 70: 1137–1140
Franklin FCH, Bagdasarian M, Bagdasarian MM, Timmis KN (1981) Molecular and functional analysis of the TOL plasmid pWWO from putida and cloning of genes for the entire regulated aromatic ringmeta cleavage pathway. Proc Natl Acad Sci USA 78: 7458–7462
Fürste JP, Pansegrau W, Frank R, Blöcker H, Scholz P, Bagdasarian M, Lanka E (1986) Molecular cloning of the RP4 DNA primase region in multirangetacP expression vector. Gene 48: 119–131
Gribskov M, Devereux J, Burges RR (1984) The codnn preference plot: graphic analysis of protein coding sequences and prediction of gene expression. Nucleic Acids Res 12: 539–549
Haas D, Holloway BW (1976) R factor variants with enhanced sex factor activity inPseudomonas aeruginosa. Mol Gen Genet 144: 243–251
Hanahan D (1985) Techniques for transformation ofEscherichia coli. In: Glover DM (ed) DNA cloning. A practical approach, vol 1. IRL Press, Oxford Washington, pp 109–135
Harayama S, Lehrbach PR, Timmis KN (1984) Transposon mutagenesis analysis of meta-cleavage pathway operon genes of the TOL plasmid ofPseudomonas putida mt-2. J Bacteriol 160: 251–255
Harayama S, Rekik M, Wasserfallen A, Bairoch A (1987) Evolutionary relationships between catabolic pathways for aromatics: conservation of gene order and nucleotide sequences of catechol oxidation genes of pWWO and NAH7 plasmids. Mol Gen Genet 210: 241–247
Harayama S, Polissi A, Rekik M (1991) Divergent evolution of chloroplast-type ferredoxins. FEBS Lett 285: 85–88
Herrmann H, Klopotowski T, Günther E (1986) The Hfr status ofPseudomonas aeruginosa is stabilized by integrative suppression. Mol Gen Genet 204: 519–523
Herrmann H, Janke D, Krejsa S, Kunze I (1987) Involvement of the plasmid pPGHI in the phenol degradation ofPseudomonas putida strain H. FEMS Microbiol Lett 43: 133–137
Herrmann H, Janke D, Krejsa S, Roy M (1988)In vivo generation of R68.45-pPGHI hybrid plasmids conferring a Phl+ (meta pathway) phenotype. Mol Gen Genet 214: 173–176
Janke D (1987) Use of salicylate to estimate the “threshold” inducer level for de novo synthesis of phenol degrading enzymes inPseudomonas putida strain H. J Basic Mikrobiol 27: 83–89
Kaminski U, Janke D, Prauser H, Fritsche W (1983) Degradation of aniline and monochloroaniline byRhodococcus sp. An117 and a pseudomonad: a comparative study. Z Allg Mikrobiol 23: 235–246
Keil H, Lebens MR, Williams PA (1985) TOL plasmid pWW15 contains two nonhomologous independent regulated catechol 2,3-oxygenase genes. J Bacteriol 163: 248–255
Kokotek W, Lotz W (1989) Construction of aLacZ-kanamycin-resistance cassette, useful for site-directed mutagenesis and as a promoter probe. Gene 84:467–471
Kukor JJ, Olsen RH (1990) Molecular cloning, characterization, and regulation of aPseudomonas pickettii PKO1 gene encoding phenol hydroxylase and expression of the gene inPseudomonas aeruginosa PAO1c. J Bacteriol 172: 4624–4630
Lehrbach PR, McGregor I, Ward JM, Broda P (1983) Molecular relationships betweenPseudomonas Inc P-9 degradative plasmids TOL, NAH, SAL. Plasmid 10: 164–174
Mead DA, Szczesna-Skorupka E, Kemper B (1986) Single stranded DNA “blue” T7 promoter plasmids: a versatile tandem promoter system for cloning and protein engineering. Protein Eng 1: 67–74
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Nakazawa T, Inouye S, Nakazawa A (1980) Physical and functional mapping of RP4-TOL plasmid recombinants: analysis of insertions and deletion mutants. J Bacteriol 144: 222–231
Nies D, Mergeay M, Friedrich B, Schlegel HG (1987)Cloning of plasmid genes encoding resistance to cadmium, zinc and cobalt inAlcaligenes eutrophus CH34. J Bacteriol 169:4865–4868
Nordlund I, Powlowski J, Shingler V (1990) Complete nucleotide sequence and polypeptide analysis of multicomponent phenol hydroxylase fromPseudomonas sp. strain CF600. J Bacteriol 172: 6826–6833
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74: 5463–5467
Shingler V, Powlowski J, Marklund U (1992) Nucleotide sequence and functional analysis of the complete phenol/3,4-dimethylphenol catabolic pathway ofPseudomonas sp. strain CF600. J Bacteriol 174: 711–724
Simon R, Priefer U, Pühler A (1983) A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in Gram negative bacteria. Biotechnology 1: 784–790
Simon R, Quant J, Klipp W (1989) New derivatives of transposon Tn5 suitable for mobilization of replicons, generation of operon fusions and induction of genes in Gram-negative bacteria. Gene 80: 161–169
Southern EH (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98: 503–517
Tietze E, Brevet J, Tschäpe H (1987) Relationships among the streptothricin resistance transposons Tn1825 and Tn1826 and the trimethoprim resistance transposon Tn7. Plasmid 18: 246–249
Vogel HJ, Bonner DM (1956) Acetylornithinase ofEscherichia coli: partial purification and some properties. J Biol Chem 218: 97–106
Yen K-M, Gunsalus IC (1982) Plasmid gene organization: naphthalene/salicylate oxidation. Proc Natl Acad Sci USA 79: 874–878
Author information
Authors and Affiliations
Additional information
Communicated by W. Goebel
Rights and permissions
About this article
Cite this article
Herrmann, H., Müller, C., Schmidt, I. et al. Localization and organization of phenol degradation genes ofPseudomonas putida strain H. Molec. Gen. Genet. 247, 240–246 (1995). https://doi.org/10.1007/BF00705655
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00705655